Abstract
Pathogenic variants in genes encoding ion channels are frequently discovered in monogenic disorders associated with epilepsy and neurodevelopmental disorders. This review covers advances in the use of automated patch clamp recording for determining the functional consequences of epilepsy-associated ion channel variants and the use of induced pluripotent stem cell (iPSC) derived neurons for in-depth investigations of the physiological consequences of such variants. The combination of these advanced technologies was a focus of the recently completed NINDS-funded Channelopathy-associated Epilepsy Research Center without Walls.
The accelerated use of genetic testing as an essential component of the clinical evaluation of early-onset epilepsy has led to an explosive growth in the number and diversity of identified ion channel variants. These discoveries represent a rich data source valuable for understanding genetic etiologies, genotype-phenotype relationships, and for stratifying therapeutic approaches in individuals with epilepsy. However, a large fraction of newly discovered variants are classified as variants of uncertain significance (VUS) because there is insufficient knowledge to confidently assess the likelihood of pathogenicity.
One strategy to gain a better understanding of pathogenicity in channelopathies is to experimentally determine the functional consequences of individual variants—particularly those that alter the protein-coding sequence. While determining ion channel variant function can be accomplished using well-established methods including voltage-clamp recording, the typical approach (eg, manual patch clamp recording) is low throughput and not sufficient to meet the demands of an ever-expanding variant landscape. Fortunately, newer technologies enabled by automated patch clamp recording are more scalable and capable of at least an order of magnitude greater throughput.
In parallel with the need to scale throughput for determining the functional consequences of epilepsy-associated ion channel variants, there is a need to investigate the physiological consequences of variants in relevant cell models. The emergence of iPSC technology enabled the generation and in vitro investigation of human, patient-specific neurons with known pathogenic variants. 1 The remarkable advances in iPSC technology applied to genetic epilepsies provide a novel cellular platform with which to investigate disease mechanisms and trial therapeutic approaches.
Automated Patch Clamp in Channelopathy-Associated Epilepsy
In the ion channel field, most consider voltage-clamp recording using the patch clamp technique to be the “gold” standard approach for determining the functional consequences of a genetic variant. In the early era of channelopathies, conventional patch clamp recording techniques were sufficient for studying the limited number of known ion channel variants. However, with advances and widespread use of DNA sequencing in research and clinical medicine, the number of variants exploded and quickly overwhelmed the collective capacity of conventional electrophysiology laboratories.
Interestingly, the evolution of ion channel electrophysiology parallels that of DNA sequencing technology, albeit with a significant lag time. In the early days, DNA sequencing was a difficult, low-throughput method performed only in elite molecular biology laboratories with specialized equipment and expertise. The development of automated sequencing instruments enabled more users and larger-scale projects. The evolution of patch clamp electrophysiology followed a similar trajectory. Initially, only a few advanced neuroscience laboratories performed patch clamp recording. Standardization and commercialization of equipment led to a democratization of the method, but throughput remained meager due to the one-cell-at-a-time format. The conceptual advance of planar patch clamp enabled the development of automated platforms that could perform multiple recordings in parallel. Rather than navigating a micropipette electrode to an individual cell as in conventional patch clamp recording, a planar patch flips this around such that cells are dissociated and applied to a glass-bottomed multiwell plate in which a tiny hole is engraved in each well (Figure 1). Capturing a single cell in the hole followed by establishing a high (gigaohm) resistance electrical seal with the cell membrane enables electrophysiological recording.
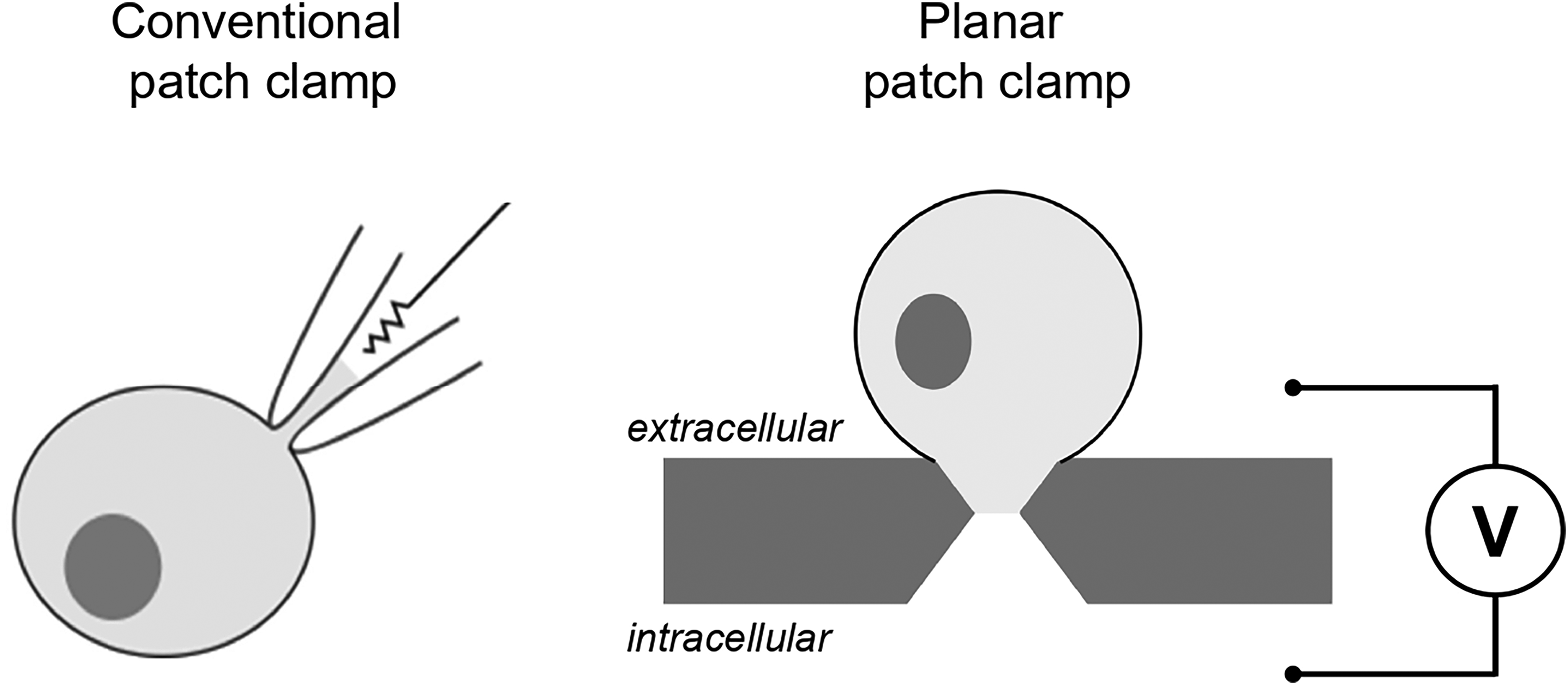
Cartoons illustrating conventional (left) versus planar (right) patch clamp configurations.
Originally developed for drug screening in the pharmaceutical industry, automated patch clamp platforms have been successfully repurposed to study epilepsy-associated ion channel gene variants.2-4 In addition to higher throughput, these approaches offer other advantages that improve experimental rigor including more robust sample sizes for greater statistical power, standardization of conditions for large numbers of variants, and lack of operator bias in selecting cells for recording. The lack of cell selection bias is particularly important for avoiding potential skewing of results based on operator-dependent preferences for choosing cells, and additionally allows for blinded measurements. High throughput capabilities also enable the parallel recording of more cells with multiple voltage-clamp protocols that can reveal complex patterns of channel dysfunction.5,6 These platforms also offer the capability to perform pharmacological studies on variants.3 This ability to assess pharmacology experimentally may be valuable for determining clinical trial eligibility for investigational new drugs that target specific functional subsets of variants (eg, gain-of-function sodium channel variants).
By design, this experimental approach typically uses nonneuronal heterologous cell models (eg, HEK293 cells) that offer a rapid turnaround, robust expression of recombinant ion channels, and enables recording in the absence of confounding endogenous ion currents that exist in native neurons. The inherent limitations of using heterologous cells are balanced by the high throughput and low background noise. Scaling the generation of recombinant plasmids encoding ion channel variants and efficient, high throughput methods to transduce cells are additional requirements. High-efficiency electroporation has proven to be particularly valuable for automated patch clamp recording studies and obviates the need to make stable cell lines, which are time and cost intensive. 7 A potential disadvantage of automated patch clamp relates to the large initial cost burden (∼ US$700,000) of purchasing such systems. Challenges also exist in the management and analysis of large datasets that dwarf those obtained by conventional patch clamp recording.
Automated patch clamp recording has been applied to determine the functional consequences of variants in multiple voltage-gated ion channel genes including genes encoding potassium channels (eg, KCNB1 and KCNQ2),2,3 sodium channels (eg, SCN2A and SCN8A),5,6 and calcium channels (CACNA1I). 4 Large-scale functional studies using more conventional methods have also been reported for variants in NMDA-type glutamate receptor genes (eg, GRIN2A and GRIN2B). 8
Studying large numbers of variants on a single platform provides opportunities for discerning more granular genotype-phenotype relationships with less interlaboratory variability. For example, recent studies of SCN2A-related disorders found that the relationships among distinct clinical presentations and variant function were more nuanced than previously recognized. This observation was made possible by examining detailed functional properties of several dozen missense variants based on data representing more than 12,500 individual cell recordings.5,6 Furthermore, the high throughput capabilities of automated patch clamp recording enabled studies of several of these variants in different alternatively spliced isoforms of the channel. Accomplishing this feat with traditional manual patch clamp recording would have required an army of electrophysiologists, which would be cost prohibitive. A similar high throughput study provided evidence that a developmentally regulated splice isoform influences the functional properties of SCN8A variants. 9
A large-scale study of epilepsy-associated KCNQ2 variants performed using automated patch clamp recording reported both functional and pharmacological properties of 80 distinct alleles.3 This study demonstrated the capability to couple biophysical analyses with drug responses in the same experiment, which would not be feasible using conventional recording methods. Similar studies have been conducted on variants in the voltage-gated potassium channel gene KCNB1 and a voltage-gated calcium channel gene (CACNA1I), revealing extensive genotype-phenotype correlations that were not previously apparent.2,4
Investigating Physiological Impact of Genetic Variants in Human Neurons
While automated patch clamp recording in heterologous cells generates high throughput data on the molecular defects conferred by genetic variants, there is a parallel need for investigating the physiological impact of variants on human neurons. This need is satisfied by the use of iPSC technologies, which enable the production of human neurons for in vitro investigations.1,10,11 The Use of iPSC-derived neurons provides high information content generated by electrophysiological and cell biological methods that can inform about cellular and network mechanisms of epilepsy. Human neuron models of genetic epilepsy are also valuable for testing therapeutic approaches, including small molecules, antisense oligonucleotides (ASOs), and gene therapies. The advent of CRISPR/Cas9 gene editing technologies has allowed for the generation of isogenic control iPSC lines, where ion channel mutations can be “corrected” in patient iPSCs, or “introduced” in the context of a healthy genetic background.
There have been multiple informative investigations of channelopathy-associated epilepsy using iPSC-neuron technology including studies of Dravet syndrome,12-15 KCNQ2-developmental and epileptic encephalopathy (DEE), 16 KCNT1-associated epilepsy, 17 SCN2A-related disorders,18-22 SCN3A encephalopathy, 23 and SCN8A-related disorders. 24
Conventional patch clamp recording has been applied to iPSC-neurons to learn about the physiological consequences of pathogenic ion channel variants. Recording action potentials in current clamp mode or individual ion currents in voltage-clamp mode has been informative for several genetic epilepsies. For example, in a study of iPSCs from individuals with Dravet syndrome associated with a missense SCN1A variant, whole-cell patch clamp recording of medial ganglionic eminence-like inhibitory neurons demonstrated smaller sodium currents and impaired action potential firing consistent with the accepted pathophysiology of this condition. 14 In an unrelated study, patch-clamp recordings performed on iPSC-derived cortical excitatory neurons heterozygous for pathogenic SCN8A variants demonstrated abnormal levels of persistent and resurgent sodium current accompanied by early afterdepolarizations and slower action potential repolarization. 24 Potassium channelopathies modeled in iPSC-neurons have also been investigated with patch clamp recording, including our investigation of KCNQ2 DEE that revealed abnormal neuronal excitability compounded by a dyshomeostatic enhancement of Ca2+-activated potassium current. 16
While patch clamp recording of single neurons can be highly informative, this method is low throughput. Attempts to use automated patch clamps to record from iPSC-neurons have been described,25,26 but these methods have not gained wide usage. Challenges to using automated patch clamps for induced neurons relate to the need to acutely dissociate cells and transfer them to the planar patch apparatus, which can disrupt morphological features and damage fragile membranes.
Another commonly used approach to interrogate the activity of iPSC-neurons noninvasively involves multielectrode array (MEA) recording. Culturing neurons on a grid of tiny electrodes enables extended live cell recording of complex neural activities (“spikes”) represented by extracellular field potentials. Spikes recorded using an MEA likely represent ensembles of action potentials, presynaptic, and postsynaptic events. MEA recording can be performed repeatedly over time in culture allowing temporal trends to be investigated without damaging or manipulating neurons externally. Furthermore, recording simultaneously from multiple electrodes provides information about neural network activity. MEA recording can be performed in multiwell plates affording greater throughput. Multiwell formats enable side-by-side comparisons with control cells and testing of therapeutic molecules.
MEA recording has proven useful for interrogating network activities in two-dimensional iPSC-neurons cultures and for testing the effects of pharmacological agents. Recent studies of two pathogenic SCN2A variants with divergent effects on channel function demonstrated that induced cortical excitatory neurons heterozygous for a gain-of-function variant (p.R1882Q) exhibited higher mean firing rate and higher burst firing frequency by MEA recording relative to a largely loss-of-function variant (p.R853Q) that showed lower mean firing rate.21,22 Additionally, the MEA platform was used to demonstrate the efficacy of phenytoin in correcting abnormal firing patterns exhibited by the gain-of-function variant neurons. MEA recording also revealed sustained phasic bursting activity in mutant KCNQ2 DEE excitatory neurons that developed over several weeks in culture. 16 While MEA recording provides high information content about neuronal networks, more detail about individual cell behaviors can be obtained from optical recording approaches with chemical or genetically encoded Ca2+ or voltage reporters including all-optical electrophysiology.27,28
Important considerations in using iPSC technology to model the cellular physiology of channelopathy-associated epilepsy include the subtype of neurons to study, age of cultures, availability of genetically matched control cells, and biological replicates. Limitations of using iPSC-neurons relate to the heterogeneity of the cultures and the immature developmental stage of the neurons, the latter resulting from the slow development of human cells and an inability to drive some neurons to full maturity. Neuronal immaturity, however, may be an advantage when modeling early-onset neurodevelopmental disorders. Another limitation is that two-dimensional iPSC-neuron cultures do not represent native brain circuitry and extrapolating results from recordings of such models to native brain should be done cautiously. Next-generation iPSC-derived neuronal models such as spheroids, organoids, assembloids of regionally patterned brain organoids, and transplantation of cells into rodent brain may offer advantages to studying disease pathogenesis, but these systems are inherently slow and low throughput.10,29,30
In summary, breakthroughs in electrophysiological methods have been applied to address the expanding number of ion channel variants discovered in genetic epilepsies. Combining large surveys of variant function with in-depth studies of channelopathy-associated epilepsy in human neurons advances knowledge regarding genotype-phenotype correlations and the pathophysiology of these important and pervasive neurogenetic disorders.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by NIH grant U54NS108874.
