Abstract
Introduction:
Heat stabilization treatment preserves the in vivo state of biological samples by rapidly inactivating enzymes that cause degradation of proteins and nucleic acids. Historically, proteomics studies used this technique as an alternative to chemical fixation. More recently, microbiologists discovered that heat stabilization treatment rapidly inactivates pathogens present in tissue samples and preserves deoxyribonucleic acid (DNA) in the tissue. However, these recent studies did not investigate the inactivation of high-density bacterial suspensions and the quality of bacterial DNA.
Methods and Results:
High-density suspensions of Escherichia coli (>109 cfu/mL) were completely inactivated by heat stabilization treatment using the Denator Stabilizor T1 instrument at 72°C and 95°C for 45 seconds. Using the heat stabilization instrument, a panel of 30 species, 20 Gram-negative and 10 non-endospore-forming Gram-positive species, were fully inactivated by treatment (95°C for 45 seconds). DNA was isolated from bacterial suspensions of Gram-negative bacteria, including E. albertii, E. coli, Shigella dysenteriae, and S. flexneri, following inactivation via heat stabilization treatment and without treatment. DNA isolated following heat stabilization treatment was fully compatible with all downstream molecular applications tested, including next-generation sequencing, pulsed-field gel electrophoresis, multiplex polymerase chain reaction (PCR), and real-time PCR.
Conclusions and Discussion:
Heat stabilization treatment of Gram-negative and non-endospore-forming Gram-positive pathogens completely inactivates high-density bacterial suspensions. This treatment is compatible with downstream DNA molecular assays, including next-generation sequencing, pulsed-field gel electrophoresis, and PCR. Inactivation by heat stabilization is a rapid process that may increase safety by decreasing risks for laboratory-associated infections and risks associated with transportation of infectious materials.
Keywords
Laboratory work with bacterial pathogens and potentially infectious materials presents a hazard to laboratory employees. Despite strict adherence to biological safety practices and the implementation of appropriate engineering controls, the risks cannot be completely eliminated when working with infectious materials and laboratory-acquired infections may still occur.1,2 The development of techniques for the inactivation of bacterial and viral pathogens for downstream molecular analyses is a recent topic of interest, as expensive instruments are often not present in high-containment laboratories, biosafety level (BSL) 3 or 4.3,4 Following verification of inactivation of potentially infectious samples, the biological samples may be further analyzed in laboratories with lower levels of biocontainment using instruments that may generate aerosols without the risk for laboratory-acquired infection. 1 The heat stabilization technology Denator Stabilizor T1 instrument (Denator, Gothenburg, Sweden) rapidly inactivates pathogens present in tissue samples, which facilitated proteomic analysis using matrix-assisted laser desorption/ionization mass spectrometry imaging by eliminating the risk for aerosolized infectious materials. 4
Preservation of biological samples is essential to reduce signaling cascades in cells that results in measurable changes of cellular biomarkers. 5 Developed for various proteomic applications, the Denator Stabilizor T1 instrument preserves the proteome of biological tissue samples without the introduction of chemicals that may interfere with downstream analysis. 6 The combination of heat and vacuum pressure causes a rapid increase in the sample temperature, which thermally denatures proteins within the sample. The inactivation of enzymes by heat stabilization treatment is compatible with subsequent proteomic analysis using matrix-assisted laser desorption/ionization mass spectrometry imaging.6,7 Heat stabilization of research and clinical samples yields significantly higher amounts of phosphorylated proteins and decreased proteolytic cleavages.6 -10 These studies show that elimination of ex vivo post-translational modifications during tissue sample processing prevented changes and degradation of biomarkers due to processing delays inherent in the process. Elimination of these post-translational modifications allows a better understanding of the in vivo state of protein biomarkers.
Although the benefits of heat stabilization of biological samples for proteomic studies have been extensively studied,6 -10 a limited number of studies have investigated the effects of heat stabilization on nucleic acids.8,11 These studies show, when using a single-step ribonucleic acid (RNA) isolation reagent, that snap-freezing tissues yields a higher concentration of RNA than heat stabilization of tissues,8,11 but snap freezing and heat stabilization perform equally for tissue presolubilized with urea. 11 One study showed heat stabilization treatment of brain tissue samples increases the yield of deoxyribonucleic acid (DNA), 11 which suggested that heat stabilization technology may be incorporated into workflows for molecular analysis of nucleic acid including next-generation sequencing.
The aims of this study were to determine whether the Denator Stabilizor T1 instrument reliably inactivates high-density bacterial suspensions and to determine if heat inactivation of bacteria using the heat stabilization technology is compatible with downstream molecular analyses for DNA, including next-generation sequencing on PacBio and Illumina platforms.
Methods
Bacterial Strains, Media, and Growth Conditions
Bacterial strains used in this study included 6 Escherichia strains, E. albertii 2014C-4356, E. coli 2011C-4315, E. coli 2012C-4431, E. coli D181, Shigella dysenteriae BU53M1, S. flexneri 94-3007, S. flexneri 71-2783, 12 Clostridium cadaveris isolated from the spleen of a cow, and 26 additional species listed in Table 1. Work was performed in a BSL-2 laboratory within a Class II biological safety cabinet. All strains were routinely grown on commercial or in-house preparations of trypticase soy agar II containing 5% sheep’s blood, MacConkey agar, or Mueller-Hinton agar, as appropriate for the organism studied. Plate cultures were grown at 37°C for 14 to 18 hours or longer on the basis of growth requirements for the organism. Bacterial suspensions were made by transferring fresh colonies from overnight cultures in sterile molecular grade water, saline (0.85% NaCl), or 33% sucrose. Turbidity of suspensions was adjusted as appropriate for downstream molecular applications. Bacterial concentrations were determined by serial dilution in saline and subsequent plating onto a sufficient agar medium for determination of colony-forming unit concentrations reported as colony-forming units per milliliter (cfu/mL).
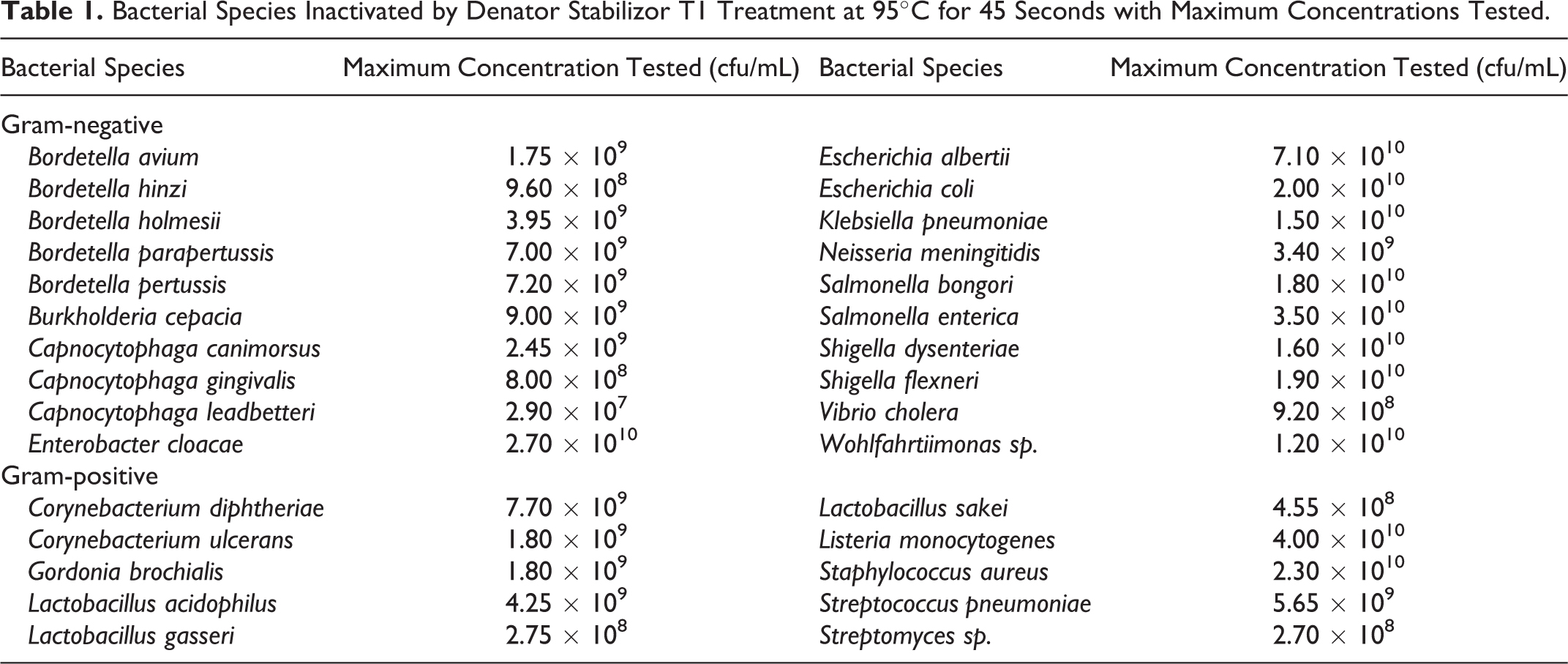
Bacterial Species Inactivated by Denator Stabilizor T1 Treatment at 95°C for 45 Seconds with Maximum Concentrations Tested.

Heat Stabilization for Bacterial Inactivation and Verification of Inactivation
Bacteria were heat inactivated using the Stabilizor T1 instrument. Bacterial suspensions (450-800 μL) were transferred to a Denator Maintainor Liquid card and processed in accordance with the manufacturer’s protocol using a custom setting of 95°C, heater position 2.1 mm, upper heater scale, cavity vacuum off at 0 mbar, and a treatment time of 45 seconds. Temperature and treatment times were varied during inactivation validation for E. coli 2012C-4431 and E. coli 2011C-4315; each combination was performed in at least triplicate. E. albertii 2014C-4356, S. dysenteriae BU53M1, S. flexneri CDC 94-3007, S. flexneri NCDC 71-2783, an additional 16 Gram-negative species, 10 Gram-positive species, and 1 endospore-forming species (C. cadaveris) were tested for inactivation through treatment at 95°C for 45 seconds (Table 1). Heat inactivation of bacterial suspensions was verified by inoculation of 100 μL of the treated bacterial suspensions onto agar plates, which were then incubated in conditions appropriate for the organism for 18 hours to 5 days. The remaining volumes of inactivated bacterial suspensions were used for downstream molecular analyses as described below.
DNA Sequencing
The MasterPure Gram Positive DNA Purification Kit (Epicentre, Chicago, IL) was used to isolate high molecular weight DNA from heat stabilization treated and control (untreated) bacterial suspensions for whole-genome shotgun sequencing using the PacBio RSII (Pacific Biosciences, Menlo Park, CA) and Illumina MiSeq (Illumina, San Diego, CA) platforms. DNA templates, libraries, and sequence reactions, data analysis, and genomic assemblies were performed as described previously. 12
Multiplex Polymerase Chain Reaction and Real-Time Polymerase Chain Reaction
DNA purified with the Epicentre kit was diluted and used as templates for polymerase chain reaction (PCR). Multiplex PCR was performed using HotStarTaq polymerase (Qiagen, Hilden, Germany) and primers (EA_F and EA_R, EF_F and EF_R, EC_F and EF_R). 13 Real-time PCR was performed for E. coli 2011C-4315 and E. coli 2012C-4431 using SsoAdvanced Universal SYBR Green Supermix (Bio-Rad, Hercules, CA) and uidA primers (uidA-784F gtgtgatatctacccgcttcgc and uidA-866R agaacggtttgtggttaatcagga) 14 on a CFX96 instrument (Bio-Rad) following the commercial protocol with melt-curve analysis.
Pulsed-Field Gel Electrophoresis
A routine method of pulsed-field gel electrophoresis (PFGE) for enteric bacteria was implemented for six bacterial strains treated with heat stabilization and untreated control suspensions. Overnight bacterial cultures were suspended in 2 mL of cell suspension buffer, and an 800-μL aliquot was treated using the Denator Stabilizer T1 instrument. SeaKem Gold agarose plugs were prepared from 400 μL of the untreated bacterial suspension and 400 μL of the treated suspensions. Plugs were processed according to standard conditions including digestion with XbaI. 15
Results
To determine whether heat stabilization treatment would effectively inactivate E. coli, high-density suspensions of E. coli 2011C-4315 and E. coli 2012C-4431 (1.6 × 109 to 8.0 × 109 cfu/mL) were heated for 45 seconds with a temperature gradient (50°C, 60°C, 65°C, 72°C, and 95°C) using the Denator Stabilizor T1 instrument, all performed at least in triplicate. The lower temperatures (50°C, 60°C, and 65°C) failed to inactivate the suspension: at 50°C, approximately 90% of E. coli survived, at 60°C approximately 70% of E. coli survived, and at 65°C ≤1% of E. coli survived (Figure 1). Heat stabilization treatment at 72°C and 95°C for 45 seconds fully inactivated the suspensions, as no growth was observed on the plates inoculated with treated suspensions. The effect of shorter treatment times was tested, and the suspensions were also inactivated when treated for 15 and 30 seconds at 95°C (data not shown).
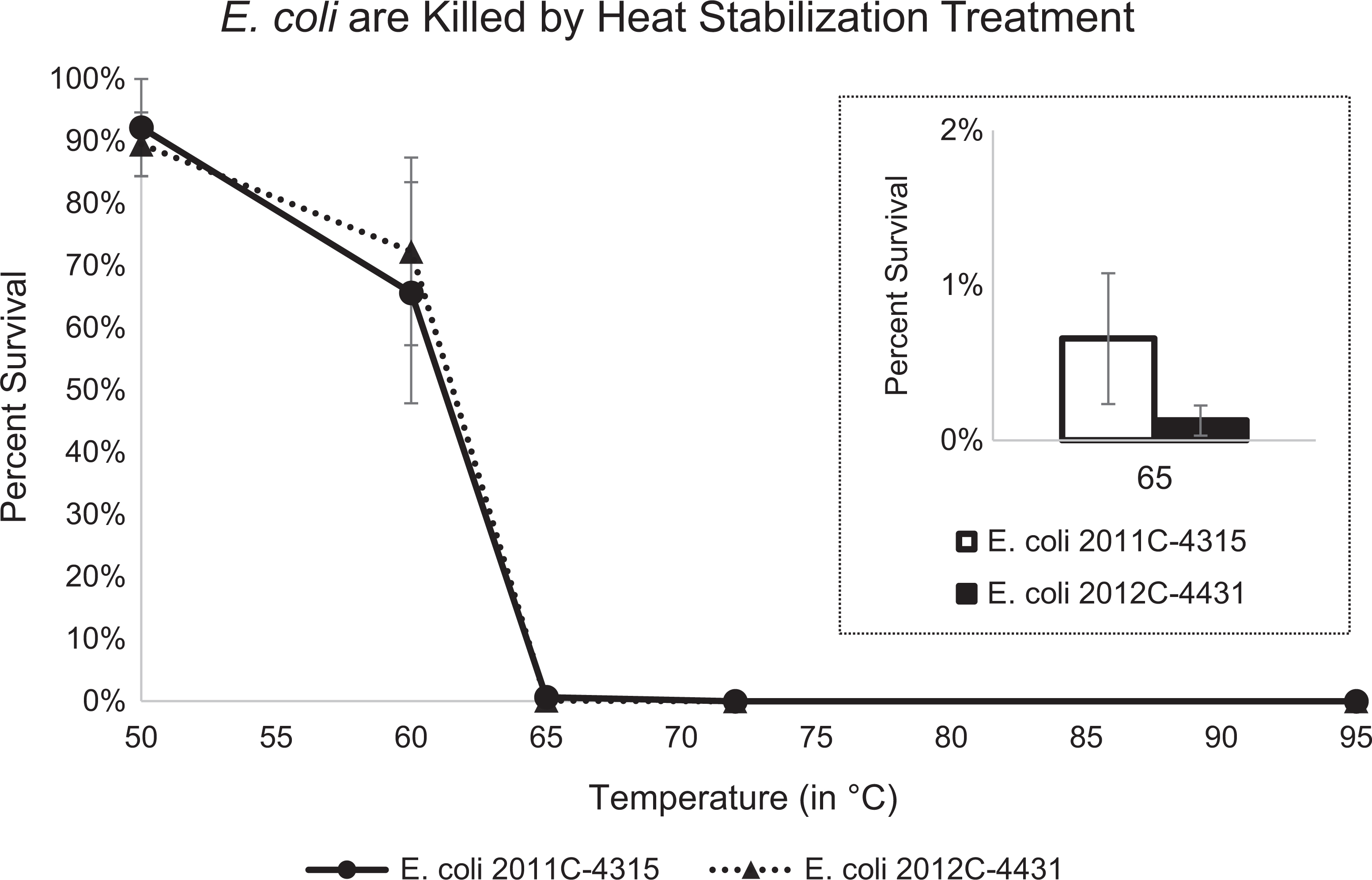
The survival of Escherichia coli strains 2011C-4315 (circles and solid line) and 2012C-4431 (triangles and dotted line) following heat stabilization treatment using the Denator Stabilizer T1 instrument for 45 seconds at 50°C to 95°C. Error bars denote standard error of the mean.
To more fully determine the spectrum of organisms that are inactivated by heat stabilization when treated at 95°C for 45 seconds, high-density bacterial cultures of E. albertii 2014C-4356, S. dysenteriae BU53M1, S. flexneri CDC 94-3007, and S. flexneri NCDC 71-2783 were tested in at least triplicate (concentrations ranged from 5.7 × 108 to 7.1 × 1010 cfu/mL). Each time the heat stabilization treatment fully inactivated the bacterial suspensions. An additional 16 Gram-negative and 10 Gram-positive species were also fully inactivated by heat stabilization treatment (Table 1). Heat stabilization treatment at 95°C for 45 seconds failed to inactivate C. cadaveris, the only endospore-forming bacterium tested. To date, all Gram-negative and non-endospore-forming Gram-positive bacteria tested have been inactivated with these treatment conditions.
High molecular weight DNA isolated using the Epicentre kit from heat stabilization treated and control untreated bacterial suspensions were purified to similar concentrations and qualities (Supplemental File 1) and were equally compatible with downstream DNA molecular analyses. Heat stabilization treatment prior to DNA isolation did not appear to affect sequence reads obtained from next-generation sequencing from the PacBio (Supplemental File 2) or Illumina (Supplemental File 3) platforms, and combined data were used to complete genomes. 12 DNA isolated after heat stabilization treatment was also compatible with multiplex PCR (Supplemental Figure S1) and real-time PCR (Supplemental Figure S2). The PFGE banding patterns were virtually identical when comparing patterns obtained for untreated control and heat stabilization treated bacterial suspensions (Supplemental Figure S3).
Discussion
Inactivation of infectious materials is a common laboratory practice, and the Biosafety in Microbiological and Biomedical Laboratories Manual requires inactivation of all infectious materials before removal from BSL-3 and BSL-4 laboratories. 1 A recent study showed that heat stabilization treatment using the Denator Stabilizor T1 instrument completely inactivates viral and bacterial pathogens in tissue samples from infected mice. 4 The authors then used these inactivated tissue samples for downstream proteomic analysis via matrix-assisted laser desorption ionization mass spectrometry imaging. 4 Here, we sought to determine whether high-density suspensions of bacterial pathogens are completely inactivated by heat stabilization treatment using the Denator Stabilizor T1 instrument and whether these inactivated suspensions are compatible with downstream DNA molecular assays.
Our data indicate heat stabilization completely inactivates high-density suspensions of E. coli in 45 seconds when treated at 72°C and 95°C (Figure 1) and that 95°C inactivation is achieved in as little as 15 seconds. A panel of 20 Gram-negative species and 10 Gram-positive species were inactivated by heat stabilization treatment for 45 seconds at 95°C (Table 1). Following heat stabilization treatment, the C. cadaveris suspension grew on the agar medium, which indicated a failure to inactivate endospore-forming bacteria with the conditions tested. To date, all Gram-negative and non-endospore-forming Gram-positive bacteria tested by our laboratory have been completely inactivated with this heat stabilization setting. This is the first demonstration, to our knowledge, that the heat stabilization technology (Denator Stabilizor T1) inactivates high-density (106 to 1010 cfu/mL) suspensions of Gram-negative and non-endospore-forming Gram-positive pathogens. Although our data suggest that heat stabilization using the Denator Stabilizor T1 instrument fully inactivates all Gram-negative and non-endospore-forming Gram-positive bacteria, verification of inactivation should be performed each time prior to transfer of potential infectious materials from BSL-3 and BSL-4 laboratories and when first added to a BSL-1 or BSL-2 laboratory workflow to ensure proper function of the instrument. We recommend verification of inactivation include inoculating at least 10% volume of the bacterial suspension onto agar media and incubation for a duration sufficient for the organism to produce colonies.
The endospore-forming bacterium C. cadaveris survived heat stabilization treatment (95°C for 45 seconds), which is consistent with a previous study suggesting that longer treatment times are needed to inactivate endospores by heat alone. 16 Recent research into the inactivation of Bacillus and Clostridium endospores focused on high temperatures combined with high pressure (eg, autoclave),17 -19 vaporized hydrogen peroxide, 20 and electric arcs. 21 Because we investigated only one endospore-forming species, additional species should be studied. Further studies into the inactivation of endospore-forming bacteria through heat stabilization treatment should also consider longer durations of treatment, higher temperatures, and/or combined chemical inactivation.
We show that DNA prepared from bacteria inactivated by heat stabilization treatment is compatible with next-generation sequencing with the PacBio and Illumina platforms, multiplex PCR, real-time PCR, and PFGE. This method of heat stabilization addresses the need to improve inactivation of bacterial pathogens, which may reduce the risk for downstream laboratory-associated infections. Heat stabilization treatment as a method of pathogen inactivation is easily integrated into existing workflows required by research, public health, and diagnostic laboratories. Here, we demonstrated that rapid inactivation by of Gram-negative and non-endospore-forming Gram-positive bacteria by heat stabilization is compatible with downstream DNA molecular applications, including next-generation sequencing.
Conclusions
Heat stabilization treatment using the Denator Stabilizor T1 instrument (95°C for 45 seconds) effectively inactivates high-density bacterial suspensions (the highest tested was 7.1 × 1010 cfu/mL) of Gram-negative and non-endospore-forming Gram-positive pathogens. Inactivation using this heat stabilization technology is compatible with downstream DNA analysis including next-generation sequencing (PacBio and Illumina), PFGE, multiplex PCR, and real-time PCR. Adoption of this technique may decrease the risk for laboratory-acquired infections and the risk associated with transferring and shipping infectious samples, and it is simple to integrate into existing molecular workflows.
Footnotes
Acknowledgments
We would like to thank Lisley Garcia-Toledo and Rebecca Lindsey for performance of multiplex PCR, Joseph Lavin for assistance with performing PFGE, and Dhwani Batra and Kristen Knipe for assistance with genomic sequence analysis. The various laboratories at the Centers for Disease Control and Prevention provided the bacterial strains used in this study.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The authors declare that there is no conflict of interest. The findings and conclusions of this article are those of the authors and do not necessarily represent the views of or an endorsement by the Centers for Disease Control and Prevention.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by U.S. government funds, the Centers for Disease Control and Prevention Laboratory Safety Science and Innovation Intramural Research Fund, and an Evaluation Fellowship provided by Oak Ridge Institute for Science and Education.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
