Abstract
Introduction
Owing to the poor clinical outcomes associated with the unlimited proliferation of tumor cells, the incidence of lung cancer and the mortality associated with the disease is on the rise globally.1,2 Studies on tumor suppression have been on the rise. Research on the induction of cell senescence and the molecular mechanisms underlying senescence in cancer cells has been gaining attention worldwide.3 -5 The Chinese medicine, Jinfukang (JFK), has been traditionally used in China for the treatment of lung cancer. It improves immunity, prevents metastasis, overcomes targeted drug resistance, alleviates chemotherapy-related adverse effects, and prolongs survival in patients with lung cancer.6 -8
Previous studies indicate that JFK can induce cellular apoptosis through the activation of DR4 and FAS and it exerts synergistic effects on apoptosis in lung cancer cells when used in combination with chemotherapy.9,10 Recently, we analyzed the composition of JFK and optimized its ingredients into the Qidongning Formula (QDF) by using computational algorithms. 11 We have further analyzed components of QDF and conducted comprehensive in vivo and in vitro studies to clarify the formula’s capacity to induce apoptosis in cancer cells. Previous research has confirmed the material basis for efficacy and tissue distribution of QDF administered orally in rats, and a novel HPLC-QQQ-MS/MS quantitative analytical method was established to determine five markers in QDF simultaneously. 12 Compared to JFK, QDF has a stronger inhibitory effect on the proliferation of lung cancer cells and a smaller impact on normal lung cells. 13 In research on the anti-lung cancer mechanism of QDF, it was found that it can upregulate the expression of p53 and p21 proteins, promoting cellular aging in lung cancer. Our previous research indicates that the transcription factor early growth response 1 (EGR1) is a crucial target for QDF’s anti-lung cancer effect. 14 EGR1 can exert anti-cancer effects by regulating the expression of p53 and p21.15,16 EGR1 plays an important role in Ras-mediated cellular aging. 17 In this study, we examined the mechanism underlying the antitumor ability of QDF and demonstrated that QDF induces senescence in lung cancer cells by activating the transcriptional regulator EGR1.
Materials and Methods
Preparation of QDF
QDF was prepared using a combination of Astragalus, Paris polyphylla, Glossy Privet Fruit, Ophiopogonis Radix, and Fiveleaf Gynostemma. All the crude herbs were provided by the Department of Pharmacy, Yueyang Hospital. The herbs were ground and mixed to obtain a weight ratio of 3:2:1:1:1. Subsequently, five times the volume of 70% alcohol and 30% pure water were added, and extracted by performing ultrasonication thrice (60 minutes each time). Subsequently, the supernatant was collected, the alcohol was evaporated by rotary evaporation, and the mixture was freeze-dried to form a powder. For in vitro experiments, the QDF powder was dissolved in the culture medium before use. The culture medium without QDF was used as the control.
Cell Culture
The human non-small cell lung cancer (NSCLC) cell lines A549 and NCI-H1975 were procured from the Shanghai Cell Bank, Chinese Academy of Sciences. The packaging plasmid for overexpression was sourced from Shanghai Jirui Biological Engineering Co. Ltd. Fetal bovine serum was acquired from Biological Industries (Catalog No. 04-001-1ACS). The EGR1 lentiviral expression vector, pLVX-IRES-ZsGreen1, was obtained from Shanghai Siding Biotechnology Co. The reverse transcription kit (RR420A) and PCR kit (R045) were purchased from TAKARA Bio Inc. TRIZOL reagent was supplied by Thermo Fisher Scientific (Catalog No. 15596018). Anti-EGR1 antibody was from Proteintech (Catalog No. 22008-1-AP), Lipofectamine 3000 from Thermo Fisher Scientific (Catalog No. L3000008), and anti-rabbit IgG from Proteintech (Catalog No. SA00001-2). RPMI 1640 medium was provided by Gibco (Catalog No. 11875093).
Construction of an EGR1-Overexpressing Lentivirus
EGR1 was cloned into the FLAG-tagged lentiviral vector pLVX-IRES-ZsGreen1, and the recombinant vector was validated using double digestion, protein expression studies, and DNA sequencing. HEK-293T cells were evenly spread in a 10 cm dish and transfected after 12-24 hours when the cell density reached approximately 60%-70% confluence. Transfection was performed by co-transferring the lentiviral vector EGR1 (5 μg) along with the packaging plasmids PMD2G (2.5 μg) and PSPAX2 (5 μg), respectively, into HEK-293T cells. Subsequently, the cells were transferred to 15 mL of fresh DMEM high-glucose medium and placed in the incubator for 24 hours. Transfection was deemed successful if observation under a fluorescence microscope at 24 hours post-transfection revealed green fluorescence. At 48 hours post-transfection, the medium was transferred into a 15 mL centrifuge tube, and the supernatant containing the virus particles was filtered through a 0.45 μm filter to remove the cell debris and impurities from medium. The resulting virus particles were held at 4°C for the subsequent infection of the cells. Lung cancer cells A549 and NCI-H1975 were evenly spread in three wells of a 6-well plate at a density of about 5 × 105 cells per well (2 mL per well), and incubated for 12-24 hours before being used in infection studies. After 12 hours, the virus solution was aspirated and transferred to a DMEM high glucose medium containing 10% FBS. The infection was deemed successful if green fluorescence was detected when observed under a fluorescence microscope after about 48 hours, since the lentiviral vector used in the study, pLVX-IRES-ZsGreen1, carried a green fluorescent protein screening marker. The approximate infection efficiency was calculated preliminarily. The infected cells were digested and passaged for amplification after about 72 hours.
β-Galactosidase (SA-β-GAL) Cytochemical Stain
Lung cancer cells in the logarithmic growth phase were seeded in 6-well plates at a density of 1 × 105 cells/well, incubated for 24 hours, washed once with PBS, fixed with 2% paraformaldehyde at room temperature (18~25°C) for 15 minutes, and washed twice with PBS. Subsequently, 1 mL SA-β-gal solution was appended to each well, and cells were dried in a drying oven at temperature of 37°C overnight. On the following day, the cells were observed under a microscope to study the cell morphology and color. Three replicates per group were performed.
Western Bolt Assay
Lung cancer cells were cultured using the same method as described above. Subsequently, the supernatant was discarded, and the pellet was washed twice with pre-cooled PBS, lysed on ice for 30 mintues, mixed every 10 minutes, and centrifuged at 12,000 rpm for 15 minutes at 4°C. The supernatant was collected, and the protein concentration was measured using BCA. Subsequently, the proteins were separated by gel electrophoresis, transferred to a membrane, and blocked using 5% skim milk at room temperature for 2 hours. Primary antibodies were then added and the membrane was incubated overnight at 4°C. Post-incubation, the membrane was washed thrice with TBST for 5 minutes. Secondary antibodies were added, and the membrane was again incubated at room temperature for 2 hours. Subsequently, the membrane was washed thrice with TBST for 5 minutes, and the protein bands were visualized and analyzed densitometrically using Quantity One software.
RT-qPCR
RNA extraction and reverse transcription were carried out as described in our previous study. 14 The mRNA levels of the individual genes were detected by quantitative real-time PCR (RT-qPCR) on a StepOne Plus Real-Time PCR system. The primer sequences are shown in Table 1.
The Primer Sequences of RT-qPCR.

RNA-seq and ChIP-seq Data Analysis
The transcriptome data and ChIP seq data were taken from previous publications. 14 We analyzed the EGR1-associated transcription network by intersecting ChIP-seq data with the transcriptome data. To obtain downstream senescence-related genes, we compared these overlap genes with the senescence-related genes in the Cellage database (https://genomics.senescence.info/cells/) to obtain the senescence genes downstream of EGR1. We applied Heml 1.0.3.7 to construct gene heat maps. The protein interaction network diagram were obtained from the GENEMANIA website (http://genemania.org/). We uploaded the 12 immune-related genes obtained from the analysis to the GENEMANIA website (http://genemania.org/) for protein interaction analysis, and we obtained the protein interaction network diagram.
Statistical Analyses
Differences in the mean values between groups were analyzed using a non-paired t-test. All data were analyzed using GraphPad Prism 9 (GraphPad Software Inc., La Jolla, CA, United States). P-values were calculated using two-tailed Student’s t-test. Values of P < 0.05, were considered to indicate a statistically significant difference.
Results
QDF Inhibits Proliferation and Induces Senescence in Lung Cancer Cells
To study the effect of QDF on lung cancer cell proliferation, A549 and NCI-H1975 were treated with different concentrations of QDF for 24, 48, and 72 hours, respectively. Results indicate that the proliferation of lung cells was significantly inhibited in a time- and dose-dependent manner (Figure 1A, Table S1). This indicates that the inhibitory effect of QDF on lung cancer cell growth intensifies with extended exposure and higher concentrations of the drug.
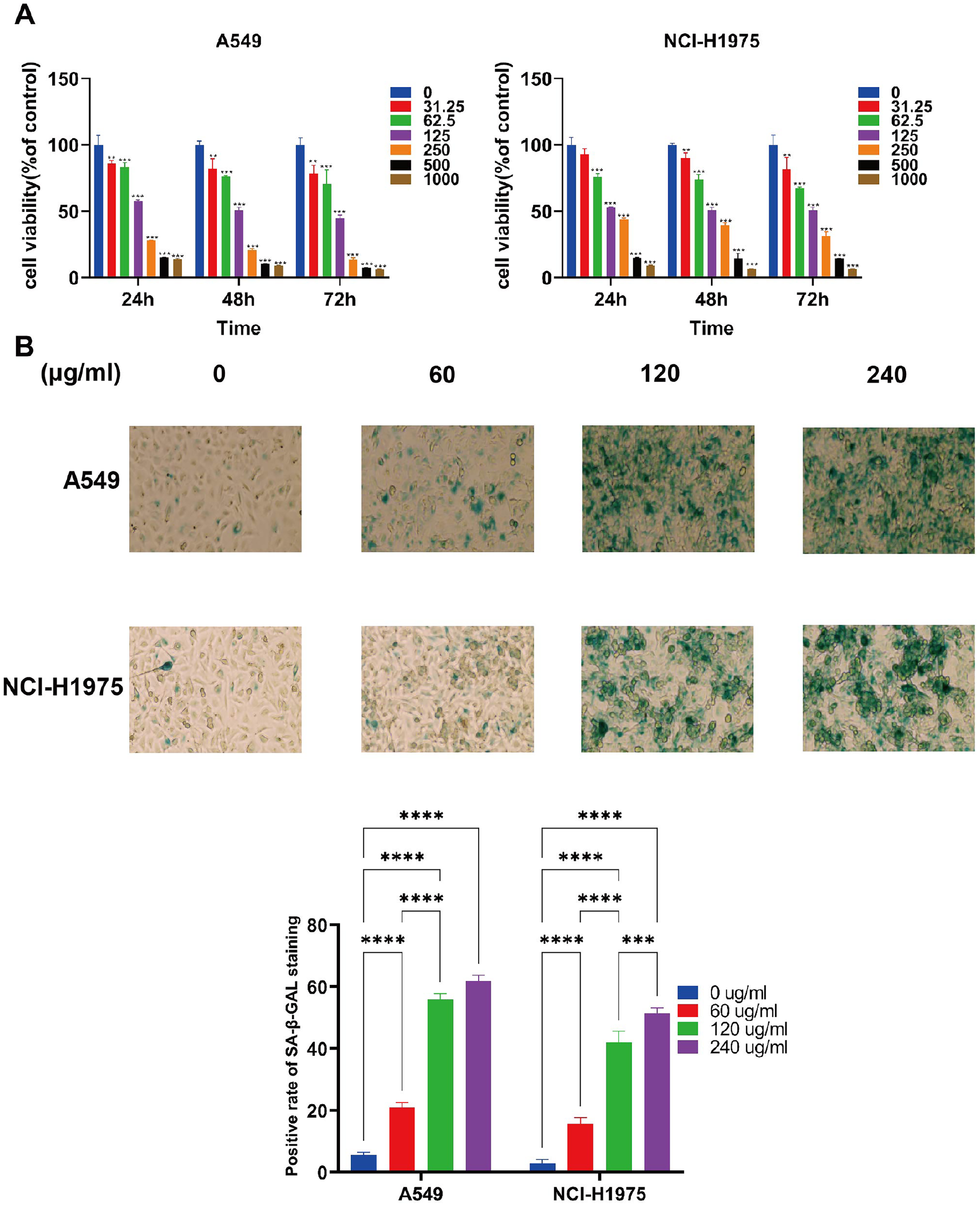
The effects of QDF on A549 and NCI-H1975 cells proliferation and senescence-associated β-galactosidase (SA-β-gal) activity. (A) The effects of various concentrations of YYJD on A549 and NCI-H1975 cells viability after 24, 48, and 72 hours treatment. *P < 0.05, **P < 0.01, ***P < 0.001. (B) SA-β-gal analysis of senescence in lung cancer cells (200X).
Senescence is associated with permanent cell proliferation. To examine the effect of QDF on cellular senescence, A549 and NCI-H1975 cells were stained with SA-β-GAL, a marker of senescent cells. After the QDF intervention, the number of SA-β-gal-positive cells increased significantly in a concentration-dependent manner compared with the control. After the addition of 60 μg/mL QDF to A549 cells, the β-galactosidase staining positivity rate was 21.10 ± 1.93%, and this rate increased to 61.79 ± 2.36% when the QDF concentration reached 240 μg/mL. Similar trends were observed in NCI-H1975 cells. (Figure 1B, Table S2).
QDF can Induce the Expression of Senescence-Associated Proteins in Lung Cancer Cells
Our previous study showed that the tumor suppressor early growth response 1 (EGR1) is the most activated after QDF treatment. 14 Activation of p21 and p53 is essential for induction of senescence and was also regulated by EGR1.18,19 Therefore, we examined the expression of EGR1, p53, p21 proteins in lung cancer cells after QDF intervention. The results showed that treatment with QDF led to a significant increase in the expression levels of EGR1 and the senescence-related gene p21, in both the A549 and NCI-H1975 lung cancer cells, compared with the control group (***P <0 .001). The expression level of the p53 gene increased significantly in A549 cells, but no difference was observed in NCI-H1975 cells (P > 0.05) (Figure 2A).

Effect of QDF on the expression of senescence-regulated gene and proteins EGR1, p53, p21. (A) Influence of different QDF concentrations on senescence-regulated gene in lung cancer cells. (B) Influence of different QDF concentrations on senescence-regulated proteins in lung cancer cells. (C) Western blot detection of the expression in A549 and NCI-H1975 lung cancer cells.
After the intervention with QDF, the expression levels of EGR1, p53, and p21 proteins were significantly increased in both A549 and NCI-H1975 cells compared with the control (**P <0 .001). Protein expression increased with an increasing concentration gradient (Figure 2B and C). These results suggest that QDF can induce senescence in lung cancer cells via EGR1 and its downstream proteins p21 and p53.
EGR1 is Implicated in QDF-Induced Senescence in Lung Cancer Cells
To further evaluate the role of EGR1 activation in QDF-induced senescence in lung cancer cells, we performed a rescue assay using an EGR1-overexpressing vector. Following the stable transfection of the EGR1-overexpressing lentiviral vector in A549 and H1975 cells, RT-qPCR and Western blot analysis revealed that the expression levels of EGR1 protein, and that of the EGR1 gene and protein in the A549-EGR1 and H1975-EGR1 groups increased significantly compared with that in the EGR1-overexpressing negative control group (Table S3, Figure 3A). This indicates the successful fabrication of EGR1-overexpressing cell lines using lentiviral transfection.

Effect on QDF-induced cellular senescence after overexpression of EGR1. (A) Western blot detection of the expression level of EGR1 protein. (B) Senescence-specific β galactosidase in lung cancer cells (200X).
On microscopic observation, the lung cancer cells that stained positive for senescence-specific β-galactosidase appeared blue. The staining results obtained at high power fields showed that the number of cells that stained positive was significantly higher in the EGR1-overexpressed A549 and EGR1-overexpressed NCI-H1975 groups compared with the EGR1-overexpressed negative control model, reaching 46.67 ± 3.58 and 48.71 ± 1.99, respectively.
In the EGR1-overexpressing lung cancer cell lines, We observed that transfection of A549 and H1975 lung cancer cells with EGR1 overexpression, followed by QDF treatment, resulted in a marked increase in SA-β-gal-positive cells compared to the untreated A549 and H1975 lung cancer cells. These findings indicate that EGR1 overexpression potentiates the senescence-inducing effect of QDF (Figure 3B).
EGR1 Overexpression can Enhance the Expression of QDF-Induced Senescence-Related Proteins in Lung Cancer Cells
In EGR1-overexpressing A549 and H1975, p21 gene and protein were highly expressed compared with the EGR1-overexpressed negative control group. P53 genes were highly expressed in EGR1-overexpressing H1975 cell compared with the EGR1-overexpressed negative control model. However, no difference was observed in the expression levels of the senescence-related genes p53, in A549 lung cancer cells, compared with the EGR1-overexpressed negative control model (P > 0.05) (Figure 4A). In EGR1-overexpressing A549 and H1975, p53 protein was highly expressed compared with the negative control group. These results indicated that EGR1 is implicated in the process of senescence in lung cancer cells and that the overexpression of EGR1 can induce cellular senescence in these cells.

Effect of overexpression of EGR1 on the expression of QDF-induced cellular senescence-related proteins. (A) The expression of gene EGR1, p53 and p21. (B) Western blot detection of the expression level of EGR1, p53 and p21 protein.
Post QDF intervention, we found that in A549 and H1975 lung cancer cells transfected with an EGR1-overexpressing lentiviral vector, the expression genes and proteins levels of p53 and p21 were significantly different from that of QDF group (*P < 0.05). These results indicated that overexpression of EGR1 enhanced the expression of QDF-induced senescence-related genes p53 and p21 genes and proteins (Figure 4A and B, Figure S1).
QDF Activates the EGR1 Regulatory Network During Senescence
To determine the mechanism by which EGR1 affects the expression status of the downstream senescence-related genes in QDF-treated lung cancer cells, we analyzed the EGR1 transcription network by combining the ChIP-seq data with the transcriptome data.
Among the genes in the EGR1 network, we identified 275 genes bound by EGR1, out of which 193 were upregulated and 82 were downregulated. Then, we compared these 275 genes with the senescence-related genes in the Cellage database (https://genomics.senescence.info/cells/) to obtain the senescence genes downstream of EGR1, when induced by QDF. After comparison, we obtained a total of 21 genes as shown in Figure 5A, including, SNAI1, PLD2, SIX1, TBX2, ZFP36, RBM38, MCM7, CBX7, CKB, ORAI1, FOXD1, IGFBP1, HSPB1, DUSP6, ING2, CDCA4, SIN3B, BRAF, HIVEP1, IL1R1, GLI1. We uploaded the 21 senescence-related genes obtained from the analysis to the GENEMANIA website (http://genemania.org/) for protein interaction analysis, and we obtained the following protein interaction network diagram as shown in Figure 5B. This enabled us to verify the results obtained in the phenotypic experiments and also to construct a protein interaction network diagram for QDF-induced cellular senescence in lung cancer cells (Table S4).

Regulatory network of senescence genes downstream of EGR1 by QDF. (A) Heatmap showed the downstream senescence genes of EGR1 under the interference of QDF. (B) The 21 senescence-related following protein interactions network diagram.
Discussion
Cellular senescence is a stress-responsive cell-cycle arrest program characterized by stable cell cycle withdrawal and major changes in cell morphology and physiology. 20 It is a relatively stable state characterized by the irreversible loss of proliferation capacity. 21 Key signaling components of the senescence machinery, such as p16INK4a, p21, CIP1, and p53, as well as epigenetic modifications, such as trimethylation of lysine 9 of histone H3 (H3K9me3), also operate as critical regulators of stem cell functions. 22 Characterized by a permanent proliferation arrest, cellular senescence occurs in response to endogenous and exogenous stresses, including telomere dysfunction, oncogene activation, and persistent DNA damage. 23 For tumor cells with unlimited proliferation ability, the occurrence of cellular senescence indicates inhibition of this continuous proliferation ability of these cells. However, it has a dual effect. For these cells, cellular senescence proceeds through the tumor suppressor pathway, which must be inactivated to impair the establishment of senescence and promote cancer. 24 Therefore, by overcoming or bypassing cellular senescence, cancer cells evade cell cycle checkpoint control, leading to genomic instability and uncontrolled proliferation. 25 Previous studies found that senescence can be induced in tumor cells under certain conditions. It has been found that the tumor suppressor gene WWOX (ROS-dependent senescence induction gene) can inhibit the growth of bladder cancer. 26 Ou’s study found that piRNA-17560, secreted by senescent neutrophils, prompted breast cancer cell proliferation through the FTO pathway. 27 Huang’s study found that senescent hepatocytes secrete cytokines that promote the occurrence of HCC, and that knocking out BCL3 dramatically inhibited the secretion of hepatocyte SASP factors and macrophage activation, thus inhibiting the occurrence of liver cancer. 28 Moreover, previous studies have shown that the rupture of the nuclear envelope can cause DNA damage, which in turn induces senescence in non-transformed cells and induces an aggressive phenotype in breast carcinoma cells. 29 Therefore, cell senescence is a pristine therapeutic target pathway in tumors, next in line to cell apoptosis and differentiation. Extensive research is currently underway to discover novel drugs that induce senescence in tumor cells. 30
Studies have shown that traditional Chinese medicine (TCM) has been widely used for thousands of years in China. In clinical practice, TCM has proved to be efficacious in the treatment of different kinds of cancers. The “infinite growth” characteristic of tumor cells and the cycle arrest manifested by cell senescence bring the cells to a steady state, which is known as the “Yin and Yang in equilibrium” state in TCM. In recent years, clinical and basic research on the effect of TCM in lung cancer has focused on combining the disease with syndrome theory. 31 The theory of Chinese medicine states that the lung is a tender organ that prefers “moisture” to “dryness.” When exogenous pathogenic factors invade the lungs, the “Qi and Yin” fluids get consumed. Therefore, TCM considers the “deficiency of Qi and Yin” as the key factor mediating the formation of lung cancer. The QDF recipe consisted of Astragalus, Paris polyphylla, Glossy Privet Fruit, Ophiopogonis Radix, and Fiveleaf Gynostemma. The primary role of QDF is to nourish Qi and Yin. In our previous studies, we reported that QDF exerts antitumor activity by inducing apoptosis and senescence in lung cancer cells. Moreover, we demonstrated that EGR1 contributes to QDF-induced apoptosis in A549 lung cancer cells.11,14 However, it was unclear whether QDF could induce cellular senescence by regulating EGR1.
Consistent with the previous research results, in this study, we found that QDF inhibits cell proliferation and induces cell senescence in lung cancer cells (Figure 1A and B). We also found that QDF induces EGR1 expression in lung cancer cells and promotes the expression of the senescence-associated proteins p21 and p53 (Figure 2A and B). To further elucidate the key role of EGR1 in the induction of senescence post-QDF treatment, A549 and H1975 were stably transfected with an EGR1-overexpressing lentiviral vector (Figure 3A). However, after the transfection, both p53 and p21 proteins were expressed at high levels in the EGR1-overexpressing A549 and EGR1-overexpressing H1975 groups (Figure 4A and B). These results confirmed that EGR1 is implicated in the senescence mechanism in lung cancer cells and that the overexpression of EGR1 induces cellular senescence. We further delineated the EGR1 target genes in QDF-treated A549 cells, including certain senescence-related genes, such as FOXD1, IGFBP1, DUSP6, IL1R132 -35 (Figure 5A and B). These findings suggest that the senescence induced in lung cancer cells treated with QDF is probably mediated by the EGR1-bound target genes.
In brief, we demonstrated that QDF activates the transcription factor EGR1 and that this activation is implicated in the process of senescence induced by QDF in lung cancer cells. Thus, our study provides insight into the antitumor mechanism of QDF. However, the mechanism underlying the regulation of senescence by QDF requires further study.
Supplemental Material
sj-docx-1-ict-10.1177_15347354241307007 – Supplemental material for Induction of Senescence in Lung Cancer Cells by Qidongning Formula via the Transcription Factor EGR1
Supplemental material, sj-docx-1-ict-10.1177_15347354241307007 for Induction of Senescence in Lung Cancer Cells by Qidongning Formula via the Transcription Factor EGR1 by Di Zhou, Wen-Xiao Yang, Cheng-Yan Wang, Cheng-Xin Qian, Ling Xu, Chang-Sheng Dong, Jie Chen and Ya-Bin Gong in Integrative Cancer Therapies
Footnotes
Abbreviations
EGR1, early growth response gene 1; QDF, Qidongning Formula; TCM, traditional Chinese medicine; SA-β-GAL, senescence-associated β-galactosidase; RT-qPCR, Real-time quantitative PCR; ROC, Receiver Operating Characteristic Curve; AUC, area under the curve; A549-EGR1, EGR1-overexpressing A549; NCI-H1975-EGR1, EGR1-overexpressing NCI-H1975.
Author Contributions
Jie Chen and Ya-bin Gong conceived the study design. Di Zhou and Wen-xiao Yang prepared the manuscript draft and figures. Changsheng Dong performed SA-β-GAL, Western blot and PCR. Wenxiao Yang operated Network pharmacology. Cheng-xin Qian and Cheng-yan Wang helped in the modification of the script draft. The final script has been reviewed and approved by all of the writers.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Natural Science Foundation of China (No. 82405497 and No. 82205213)
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
