Abstract
Lung cancer is the most prevalent and lethal malignant tumor in China, primarily categorized into small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC). NSCLC accounts for more than 80% of all lung cancer cases, with current treatments primarily consisting of surgery, chemotherapy, and targeted therapy. However, these treatments often come with various adverse effects and drug resistance issues, highlighting the urgent need for new NSCLC therapies. Traditional Chinese medicine serves as a natural treasury of medicinal compounds and an important avenue for discovering novel active compounds. Platycodin D (PD) is a triterpenoid saponin isolated from the roots of Platycodon, possessing various pharmacological properties. Nevertheless, the exact mechanism of PD’s anti-lung cancer activity remains unclear. In this study, 3 lung cancer cell models, A549, NCI-H1299, and PC-9, were employed. After intervention with Platycodin-D, tumor cell proliferation and migration were assessed. Cell migration ability was assessed through transwell assays, while transcriptomics was employed to explore the mechanism of PD’s anticancer activity. Bioinformatic analysis revealed significant enrichment of apoptosis and the TGFβ pathway following PD intervention, as shown in gene expression heatmaps, where genes associated with cancer were significantly downregulated by PD intervention. Subsequently, we used immunofluorescent labeling of KI-67 to evaluate cell proliferation, flow cytometry to assess apoptosis, and Western blot to detect protein expression of TGFβ and P-SMAD3. Immunofluorescence was also employed to investigate E-cadherin, vimentin, and N-cadherin. Finally, molecular docking and dynamic simulations were utilized to study the interaction between PD and TGFβ proteins. The results of this study indicate that PD exhibits robust anti-lung cancer pharmacological activity, with its primary target being TGFβ. PD may serve as a potential TGFβ inhibitor and a candidate drug for NSCLC treatment.
Keywords
Introduction
Lung cancer is a leading category of malignant tumors in China, characterized by the highest incidence and mortality rates. It can be mainly categorized into small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC), with the latter accounting for approximately 80% of cases and being the primary cause of death among lung cancer patients. 1 NSCLC often presents with non-specific symptoms in the early stages, and most patients are diagnosed in advanced stages, where clinical treatments primarily involve surgery, radiotherapy, chemotherapy, and targeted therapies. Even in cases amenable to surgical resection, adjuvant chemotherapy or targeted therapy is required to extend the patients’ survival. 2 Despite advancements in various therapeutic modalities such as surgery, radiation therapy, and chemotherapy, the 5-year survival rate for NSCLC patients remains suboptimal, with early-stage (stage IB) patients having a survival rate of only 68%, and stage IV patients having survival rates below 10%. 3 With the advent of precision medicine, breakthroughs in NSCLC treatment have been achieved through immunotherapies targeting programed cell death receptor 1 (PD-1) or programed cell death ligand 1 (PD-L1). However, such treatment approaches are not universally applicable to all patients, and issues such as drug resistance and adverse reactions often hinder the expected therapeutic outcomes.4 -6 Consequently, the quest for safe and effective new drugs to treat NSCLC is imperative.
Platycodin D (PD), a triterpenoid saponin component extracted from the traditional Chinese herb Platycodon grandiflorus (PG), is abundant in PG and serves as its principal bioactive constituent. PD has demonstrated a wide range of pharmacological properties, including anti-inflammatory, anti-allergic, anti-tumor, and immune-enhancing effects.7,8 In the context of diabetic nephropathy research, PD treatment has been shown to downregulate the expression of ACSL4 and TFR1 while upregulating FTH1 and SLC7A11, thereby inhibiting cellular iron accumulation induced by high glucose. 9 PD can also suppress airway inflammation by inhibiting the production of eosinophils and mucin in bronchial mucosa and intervening in Th2 cell cytokines. 10 Furthermore, PD has a significant impact on oxidative stress, apoptosis, and inflammation induced by cisplatin, mitigating cisplatin-induced renal toxicity. 11 PD has also been demonstrated to ameliorate liver fibrosis by promoting the activation of JNK and c-Jun, further altering autophosphorylation of hepatic stellate cells. 12 Natural products play a pivotal role in drug discovery and development, warranting further in-depth investigation.
The TGFβ signaling pathway plays a crucial role in tumor initiation and progression. Transforming growth factor β1 (TGF-β1) is a key regulator of cell growth and differentiation, which, through its interaction with its receptor, mediates downstream signaling molecules such as Smad proteins to participate in tumor cell differentiation and proliferation. 13 Epithelial to mesenchymal transition (EMT) induced by the TGFβ pathway plays a central driving role in tumor invasion and metastasis. Inducing EMT can increase the invasive capability of tumor cells, promote tumor angiogenesis, and facilitate tumor growth and dissemination. 14 TGFβ serves as a key regulatory factor in EMT, and extracellular TGFβ signaling activates EMT transcription factors via the Smad complex, resulting in increased expression of mesenchymal markers (e.g., N-cadherin and vimentin) and decreased expression of epithelial markers (eg, E-cadherin), ultimately promoting tumor cell migration and invasion. 15
In this study, we utilized the lung cancer cell model to investigate the effects of PD on tumor migration, proliferation, and apoptosis. We employed transcriptomics to explore changes in gene expression following PD intervention and conducted relevant target validation. Our aim is to elucidate the pharmacological mechanisms underlying PD’s anti-lung cancer properties and contribute to the development of drugs for the prevention and treatment of this disease.
Materials and Methods
Reagents
Platycodin D was purchased from Chenguang Biology, M.F:C57H92O27, HPLC purity > 98%, Catalog number: HS204174B1 (Baoji, China). A cell apoptosis detection kit was purchased from Keygen Biotech, Catalog number: F6012L (Nanjing, China). Cell Counting Kit-8was purchased from APExBIO, Catalog number: K1018 (Shanghai, China). Cell Proliferation EdU Image Kit, was purchased from Beyotime, Catalog number: C0071s (Shanghai, China). Transwell Staining Kit was purchased from Jiancheng Bioengineering Institute, Catalog number: 353097 (Nanjing, China). P-smad3 (Catalog number: AF3362) and N-cadherin (Catalog number: TF5311) antibody was purchased from Affinity (Changzhou, China). E-cad (Catalog number: 60335-1-Ig), Ki-67 (Catalog number: 27309-1-AP), vimentin (Catalog number: BM0135), and TGFβ (Catalog number: 21898-1-AP) antibodies were purchased from Sanying Biotechnology (Wuhan, China).
Cell Culture
The lung cancer cell models were purchased from Keycell Biotechnology (Wuhan, China). A549 Catalog number: QS-H001, RRID: CVCL 0023; PC9 Catalog number: QS-H004, RRID:CVCL 0060; NCI-H1299 Catalog number: QS-H039, RRID:CVCL B260. A549 cell culture medium: Ham’s F-12K + 10%FBS + 1% (Penicillin-Streptomycin Solution). PC9 and NCI-H1299 cell culture medium: RPMI-1640 + 10%FBS + 1% (Penicillin-Streptomycin Solution). Once cell density reached 80%, the cells were passaged at a ratio of 1:2 and cultured in 5% CO2-saturated humidity conditions at 37°C. When the cells grew well and were in the logarithmic growth phase, the single cell suspension was obtained by trypsin digestion and centrifugation, and then the follow-up experiments were carried out according to the needs of the experiment.
CCK8 Cell Viability Assay
Lung cancer cells growing well in the logarithmic phase were inoculated in 96-well plates at 3 × 103 cells/well and treated with different concentrations of PD (2.5, 5, 10, 20, 30, 40 µM) for 24 hours. After the time needed for cell culture, 10 µl of CCK8 solution was added to each well and cultured at 37°C for 4 hours. The absorbance of each well was determined using an enzyme-labeled instrument.
Detection of Apoptosis by Flow Cytometry
The lung cancer cells were placed in a medium containing 90% F-12K + 10% FBS for cell recovery. Once cell density reached 80%, the cells were passaged at a ratio of 1:2 and cultured in 5% CO2-saturated humidity conditions at 37°C. The cells that were in good condition in the logarithmic growth phase were taken and cultured overnight at 37°C and 5% CO2 in an incubator. Each group was treated with PD of corresponding concentration. After 24 hours, apoptosis was detected using an Annexin V-FITC/PI Apoptosis Detection Kit. The tests were performed with BECKMAN flow cytometry.
Transwell
The cells of each group were resuscitated in serum-free medium to prepare single cell suspension. The cell suspension with a density of 1 × 10 ~ 5 cells/ml was inoculated into the upper chamber of Transwell, and then SH was added according to the concentration gradient. The culture medium containing 20%FBS was added to the lower 24-well plate. After incubation at 37°C according to time gradient, the non-metastatic cells in the upper chamber were erased. The inferior ependyma was fixed with paraformaldehyde, stained with crystal violet staining, observed under inverted microscope and photographed and counted.
Immunofluorescence
In the culture plate, the slides of the climbed cells were washed with PBS and fixed with 4% paraformaldehyde. 0.5%TritonX-100 (prepared by PBS) was permeated at room temperature for 20 minutes, and goat serum was dripped to seal 30 minutes at room temperature. Enough diluted first antibody (Mouse monoclonal antibody) was added to the slide and incubated overnight in a wet box at 4°C. After that, fluorescent second antibody (Cy3-labeled Goat Anti-Mouse IgG) was added and incubated at 37°C for 60 minutes in a wet box, and was incubated 5 minutes without light by dripping DAPI. The film was sealed with the sealing solution containing anti-fluorescence quenching agent, and finally the image was observed and collected under the Olympus fluorescence microscope.
Transcriptome Analysis
Total RNA was extracted from the samples using the Trizol kit. RNA quality was assessed using an Agilent 2100 Bioanalyzer and detected using ribonuclease-free agarose gel electrophoresis. After the extraction of total RNA, mRNAs from eukaryotic cells were enriched with oligo (DT) pellets, cut into short fragments with fragmentation buffer, and reverse transcribed into cDNA using a NEBNext Super RNA Library Preparation Kit. The purified double-stranded DNA fragments were end-repaired, coupled with A bases, and ligated to Illumina sequencing adapters. The ligation reaction was purified with AMPure XP beads (1.0×). Size selection of the ligated fragments was performed using agarose gel electrophoresis and PCR, and transcriptomic analysis was performed with Illumina Novaseq6000 sequencing.
Western Blot Assay
The total protein of monolayer adherent cells was extracted, the protein concentration was determined, the protein was cooled to room temperature after denaturation and then stored at −20°C, and the electrophoresis gel was prepared. After the electrophoretic separation, the membrane was transferred. After the membrane transfer was completed, the gel was taken out and added to the sealing solution, the corresponding first and second antibodies were diluted with the sealing solution. Mouse monoclonal antibody β-actin (1:20000), Rabbit polyclonal antibody TGFβ1 (1:2500), Rabbit polyclonal antibody p-smad3 (1:1000). The excess second antibodies were washed, and the enhancement solution in ECL reagent was mixed with the stable peroxidase solution at 1:1. The working solution was added to the PVDF film to develop the film, scan the film, and analyze the gray value of the film by ipp.
Molecular Docking and Molecular Dynamics Simulation
The compound PD data used for molecular docking were obtained from PubChem (https://pubchem.ncbi.nlm.nih.gov/) and then imported into Chem3D software for optimization and energy minimization using the MM2 module. The data were saved in a sdf file as ligand molecules for molecular docking. The TGFB (PDB ID: 1KLD) protein structures were then obtained from the RCSB database (https://www.rcsb.org/). Finally, molecular docking and screening were performed using the standard precision (SP) method. Molecular dynamics simulations of the screened receptor protein–small molecule complexes were conducted using the Gromacs 2020 software package. The binding free energy of the ligands and proteins was calculated according to the MMPBSA method using the Gromacs 2020 program.
Statistical Analysis
Experimental data were expressed as mean ± standard deviation (SD) values. The statistical analysis in this study was performed using GraphPad Prism version 8 (GraphPad Software Inc., San Diego, CA, USA). Differences between the groups were assessed using a one-way analysis of variance (ANOVA). For all analyses, P < .05 was considered statistically significant.
Results
PD Alleviates Lung Cancer Cells Migration and Proliferation
To investigate the impact of PD (Figure 1A) on lung cancer cells, 3 lung cancer cell models, A549, NCI-H1299, and PC-9, were employed. After intervention with Platycodin-D, tumor cell proliferation and migration were assessed. The CCK8 results revealed a dose-dependent inhibition of cell proliferation by PD (Figure 1B). The optimal intervention dose for PD was chosen as 5, 10, and 20 µM, as shown under the microscope (Figure 1C), where lung cancer cell growth was inhibited following PD intervention. Cell migration was assessed using the Transwell assay, and PD intervention reduced cell migration, with the most significant effect observed at a concentration of 20 µM (Figure 1D). Subsequently, EDU labeling was used to detect tumor cell proliferation, and the results showed that PD intervention effectively reduced cell proliferation (Figure 2A).

PD alleviates lung cancer cell migration. (A) The chemical structure of platycodin D. (B) The changes of cell proliferation rate were detected by CCK8. (C) Cell proliferation rate image. (D) Cell migration was detected by transwell assay. Compared with platycodin D 0 μM group.

PD alleviates lung cancer cell proliferation. (A) Detection of cell proliferation by EDU fluorescence staining. Compared with platycodin D 0 μM group.
Transcriptomics Reveals the Mechanism of PD Intervention in Lung Cancer Cells
To explore the mechanism of PD in lung cancer cells lung cancer cells, transcriptomics was employed. PCA results revealed 2 clusters, 1 for lung cancer cells and the other for lung cancer cells after PD intervention (Figure 3A). Using a cutoff of P < .05 and a fold change >1.5, transcriptomic analysis identified 2159 upregulated genes and 2992 downregulated genes, as illustrated in the volcano plot (Figure 3B). GO enrichment analysis of cells after PD intervention revealed the top 20 differentially regulated GO Terms (Figure 3C) Cellular Component changes were mainly associated with the cell membrane, Biological Process mainly related to cell metabolism, and Molecular Function related to protein binding (Figure 3D). Subsequent KEGG enrichment analysis showed alterations in molecular pathways after PD intervention, primarily related to apoptosis and the TGFβ signaling pathway (Figure 3E). Disease Ontology enrichment analysis demonstrated that differentially expressed genes after PD intervention were mostly related to cancer (Figure 3F).
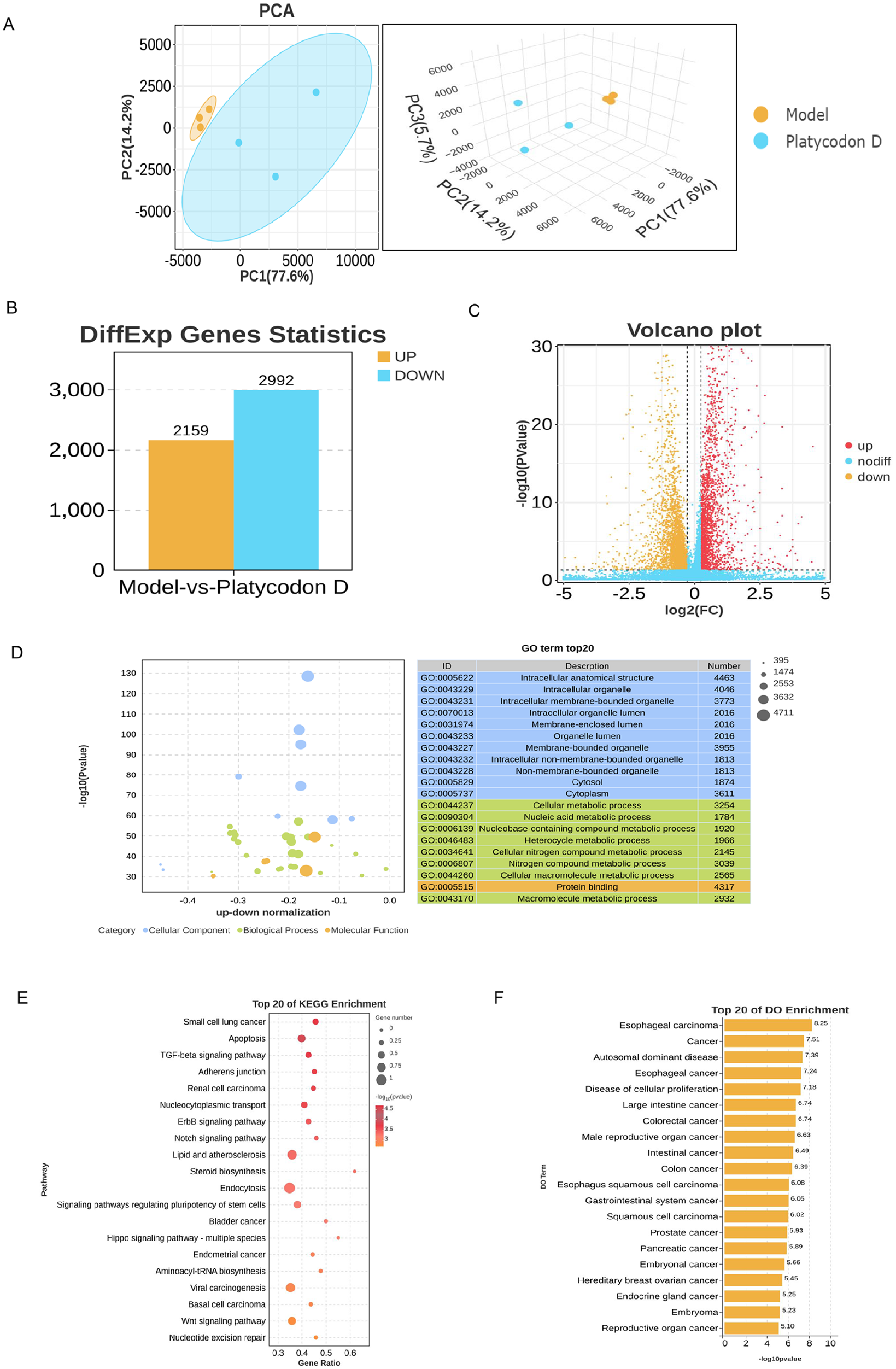
Transcriptomics reveals the mechanism of PD intervention in A549 cells. (A) PCA of the RNA-seq data from the Model and platycodin D. (B) Differential gene expression histogram. (C) Differential gene expression volcano map. (D) DO enrichment analysis. (E) KEGG enrichment analysis. (F) DO enrichment analysis.
PD’s Effects on Tumor Cell Proliferation and Apoptosis
A heatmap of differentially expressed genes related to tumor proliferation was generated, showing a significant downregulation of gene expression after PD intervention (Figure 4A). Immunofluorescent staining with KI-67 revealed a significant inhibition of KI-67 expression in tumor cells after PD intervention (Figure 4B). Flow cytometry was used to assess cell apoptosis after PD intervention, and the results showed a significant promotion of tumor cell apoptosis, with the most pronounced effect at a concentration of 20 µM (Figure 4C).

PD’s effects on tumor cell proliferation and apoptosis. (A) Heat map of gene expression associated with tumor proliferation. (B) Ki-67 was detected by immunofluorescence. (C) Flow cytometry was performed to determine apoptosis. Compared with platycodin D 0 μM group.
PD Inhibits the TGFβ Signaling Pathway
Based on bioinformatics analysis, the TGFβ signaling pathway was significantly enriched. GSEA analysis demonstrated changes in the TGF pathway, and a heatmap of differentially expressed genes on the TGFβ pathway showed a significant downregulation of TGFβ-related genes after PD intervention (Figure 5A and B). Western blot analysis confirmed a significant decrease in the expression of TGFβ and P-SMAD3 proteins after PD intervention (Figure 5C). Further immunohistochemistry showed a significant reduction in TGFβ expression after PD intervention (Figure 5D).

PD inhibits the TGFβ signaling pathway. (A) GSEA pathway enrichment analysis of TGF β signaling pathway. (B) Heat map of expression of genes related to TGF β signaling pathway. (C) The protein expression levels of TGFβ and P-SMAD3 were detected using Western blot analysis. (D) The expression level of TGFβ was detected by immunohistochemistry. Compared with platycodin D 0 μM group.
PD Modulates Tumor EMT Changes
As EMT changes play a crucial role in tumor progression, immunofluorescent staining was used to label E-cadherin, N-cadherin, and Vimentin. The results showed that N-cadherin and Vimentin expression decreased, while E-cadherin expression significantly increased after PD intervention, with the most pronounced effect observed at a concentration of 20 µM, indicating that PD can mitigate tumor cell EMT transformation (Figure 6A-C).

PD modulates tumor EMT changes. (A-C) The expression level of E-cadherin, N-cadherin, Vinmentin was detected by immunofluorescence. Compared with platycodin D 0 μM group.
Molecular Docking and Molecular Dynamics Simulation
To investigate whether PD interacts with TGFβ protein, molecular docking was performed between PD and the TGFβ target protein. The results indicated a strong binding interaction between PD and the TGFβ target protein, with a binding energy of −10.08 kcal/mol. PD could form multiple hydrogen bond interactions with amino acids at the protein’s active site, including GLU-12, GLN-67, ASN-66, GLY-71, and LYS-13, demonstrating a strong binding affinity and an essential role in anchoring small molecules in the protein pocket (Figure 7A). To further study the interaction between the small molecule and the protein, molecular dynamics simulations were conducted on the protein-small molecule complex for 100 ns. The complex displayed strong stability, and the small molecule formed a stable complex with the protein. Finally, the binding free energy between the molecule and the protein was calculated, indicating that Platycodin D exhibited a strong affinity with the TGFβ protein. These affinities promoted the formation of a stable complex, thus exerting its active function (Figure 7B-G).

Molecular docking and molecular dynamics simulation. (A) The binding mode of TGFβ protein with platycodin D. (B-E) The RMSF/RMSD/SASA/Rg analysis of platycodin D with TGFβ. (F) The hydrogen bond number of platycodin D with TGFβ. (G) The binding energy by MMGBSA (kJ/mol).
Discussion
Lung cancer is the most common malignant tumor globally, with an incidence and mortality rate that accounts for nearly 20% of all cancer-related deaths. It is projected that by the end of 2030, the mortality rate of lung cancer in China will reach 40%, imposing immeasurable burdens on society. 16 Non-small cell lung cancer (NSCLC) is the most prevalent type, representing over 80% of lung cancer cases. 17 While standard treatments, including surgery, radiation therapy, chemotherapy, and immunotherapy, have extended the overall survival of patients, real-world challenges such as chemotherapy-related side effects, drug resistance, and limited benefits from immunotherapy have created clinical bottlenecks. Therefore, the quest for safer, more effective, and reliable drugs is of paramount importance. 18 Traditional Chinese medicine (TCM) represents a natural reservoir of medicinal compounds and an important avenue for discovering novel active compounds, warranting in-depth research. Platycodin D (PD), a triterpenoid monomer isolated from Platycodon grandiflorum, serves as a primary active ingredient and has exhibited promising pharmacological activities in anti-inflammation and anti-tumor effects. 19
In this study, we employed the lung cancer cell model to investigate the effects of PD intervention. The results, as determined by CCK8 assays, indicate that PD dose-dependently inhibits cell proliferation. Transwell assays further suggest that PD intervention reduces cell migration ability, and EDU fluorescence assays validate PD’s anti-proliferative capacity. To explore the anti-lung cancer mechanisms of PD, we utilized transcriptomics in combination with bioinformatics analysis to examine differential gene expression following PD intervention. Enrichment results reveal that PD primarily pertains to apoptosis and the TGFβ signaling pathway. Heatmaps of differentially expressed genes illustrate genes associated with tumor proliferation. Immunofluorescent labeling with KI-67 and flow cytometry for apoptotic cells were employed to further assess proliferation and apoptosis changes. The results indicate that PD intervention significantly promotes tumor cell apoptosis while suppressing cell proliferation.
TGF-β belongs to the transforming growth factor family, playing a vital role in regulating cell growth and differentiation processes. It can interact with its receptor to mediate downstream signaling molecules, and Smads proteins, involved in tumor cell differentiation and proliferation. Additionally, TGF-β is a key regulator of epithelial-mesenchymal transition (EMT), which stimulates tumor angiogenesis, assists tumors in evading immune surveillance, creates an appropriate extracellular matrix microenvironment, and promotes tumor invasion and metastasis. 20 To investigate changes in TGFβ following PD intervention, we employed Western blotting to assess TGFβ and P-SMAD3 protein levels and utilized immunohistochemistry to further observe TGFβ expression changes. The results demonstrate that PD intervention significantly inhibits the expression of TGFβ and P-SMAD3 proteins.
EMT is the fundamental morphological process through which epithelial cells acquire mesenchymal characteristics, manifested by the loss of E-cadherin expression and an increase in Vimentin and N-cadherin expression. In recent years, the role of EMT in tumors has received increasing attention. The EMT process weakens the adhesive properties of tumor cells while enhancing their migration, allowing them to infiltrate lymphatic and blood vessels and spread throughout the body, a crucial mechanism for tumors to acquire enhanced migratory and invasive capabilities. 21 Research has confirmed the critical role of Smad3 in the EMT process, as Smad3 can directly or indirectly bind to various transcription factors related to EMT, influencing the expression of E-cadherin and promoting EMT.22,23 The TGF-β/Smad signaling pathway is a key regulator of EMT. To detect changes in EMT, we employed immunofluorescence to examine the expression of E-cadherin, Vimentin, and N-cadherin. The results show that PD intervention significantly increases E-cadherin expression while suppressing Vimentin and N-cadherin expression, indicating that PD modulates EMT changes in lung cancer cells.
To observe whether Platycodin-D acts through the TGFβ protein, molecular docking and molecular dynamics simulations were performed between Platycodin-D and the target protein of TGFβ.The visualizations of the complexes formed by the docking compounds and proteins revealed a clear pattern of binding between the compound and the protein pocket. Additionally, the compound fit well into the protein cavity, facilitating the formation of stable complexes between the 2. Subsequently, molecular dynamics simulations were conducted and analyzed accordingly. The RMSD plots indicated that the average RMSD of the complexes was consistently below 2.5 Å, and the complexes reached dynamic equilibrium around 40 ns, suggesting a good match between the small molecule and the target protein, leading to the formation of stable complexes. The RMSF plots showed minimal conformational changes in most amino acids, reflecting the stability of the interaction between the small molecule and the protein in forming stable complexes. According to the Rg plots, the Rg of the TGFB protein decreased significantly in the first 40 ns and then gradually declined, possibly due to the binding of the protein with the compound after molecular dynamics, which promoted the protein to maintain more hydrophobic contacts and form more effective interactions internally. Additionally, the protein cavity expanded slightly, facilitating better matching with the compound and promoting the stability of the complexes. To gain further insight into the binding of the protein and the compound, the number of hydrogen bonds between the protein and the small molecule was analyzed throughout the simulation process. Based on the hydrogen bond network diagrams for each protein-compound pair, it was evident that the compound could form 6 or more hydrogen bonds with the amino acids in the protein pocket, indicating the crucial role of these hydrogen bonds in stabilizing the interaction between the small molecule and the protein. In summary, Platycodin-D exhibited strong affinity for the TGFβ protein, enabling the formation of stable complexes between the small molecule and the protein, thereby exerting its active effects.
In this study, 3 lung cancer cell models, A549, NCI-H1299, and PC-9, were employed. After intervention with Platycodin-D, tumor cell proliferation and migration were assessed. We have discovered that PD possesses potent anti-lung cancer pharmacological effects. The safe drug concentrations of Platycodin-D against tumors were determined to be 5, 10, and 20 µM. Subsequently, A549 cells were selected for transcriptomic analysis. Transcriptomics, in conjunction with bioinformatics analysis, elucidated the mechanisms through which PD intervenes in lung cancer. This study validated PD’s anti-proliferative capacity and its ability to promote tumor cell apoptosis. The primary target of its action to be the TGFβ pathway. Thus, PD may serve as a potential TGFβ inhibitor and a candidate drug for treating NSCLC, the regulatory mechanism of PD is shown in Figure 8.

The potential role of PD in non-small cell lung cancer. PD alleviates lung cancer cell migration and proliferation, when the transcriptomics approach was used to explore its mechanism of action, PD was found to block the TGFβ signaling pathway, attenuate the EMT Changes. Molecular docking and molecular dynamics simulation was performed between PD and the TGFβ target protein. The results indicated a strong binding interaction between PD and the TGFβ target protein.
Conclusion
The results of this study indicate that PD exhibits robust anti-lung cancer pharmacological activity, Transcriptomics, in conjunction with bioinformatics analysis, elucidated the mechanisms through which PD intervenes in lung cancer, with its primary target being TGFβ. PD may serve as a potential TGFβ inhibitor and a candidate drug for NSCLC treatment.
Footnotes
Data Availability
Data will be made available on request.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Scientific and Technological Innovation Project of Information on Traditional Chinese Medicine, China Academy of Chinese Medical Sciences (No. CI2021B002).
