Abstract
Diabetic foot ulcers, with worldwide prevalence ranging from 12%-25%, are an important cause of nontraumatic lower limb amputation. Evidence-based assessment of early infection can help the clinician provide the right first line treatment thus helping improve the wound closure rate. Illuminate®, a novel point of care device working on multispectral autofluorescence imaging, helps in the rapid identification and classification of bacteria. This study was aimed to evaluate the diagnostic accuracy of the device in detecting bacterial gram type against standard culture methods. A total of 178 patients from a tertiary care center for diabetes was recruited and 203 tissue samples were obtained from the wound base by the plastic surgeon. The device was handled by the trained investigator to take wound images. The tissue samples were taken from the color-coded infected region as indicated by the device's Artificial Intelligence algorithm and sent for microbial assessment. The results were compared against the Gram type inferred by the device and the device was found to have an accuracy of 89.54%, a positive predictive value of 86.27% for detecting Gram-positive bacteria, 80.77% for Gram-negative bacteria, and 91.67% for no infection. The negative predictive value corresponded to 87.25% for Gram-positive, 92% for Gram-negative, and 96.12% for no infection. The Results exhibited the accuracy of this novel autofluorescence device in identifying and classifying the gram type of bacteria and its potential in significantly aiding clinicians towards early infection assessment and treatment.
Keywords
Introduction
The International Diabetes Federation estimates that the expected prevalence of diabetes mellitus will rise to more than 570 million in 2030. 1 Diabetic foot ulcers (DFU), and diabetic foot infections are major contributors of worldwide morbidity and mortality as they lead to limb amputation in 12%-25% of individuals with diabetes. 2 DFU initially begins with peripheral neuropathy and finally lead to ulcerations. Secondary infections leads to chronically infected wounds over a period of time. 3 Prompt diagnosis of bacteria in these wounds and timely treatment help speed up the wound healing process. However, it is clinically difficult to diagnose the causative pathogens causing secondary infections. Routine microbial-based culture methods, the current standard for identifying and classifying bacteria infecting the wounds, requires taking a swab or biopsy from the infected region and take 2 to 3 days for diagnosis, causing a delay in providing first-line treatment and/or inappropriate usage of antibiotics leading to the development of bacterial resistance. 4
Autofluorescence-based multispectral imaging has recently emerged as a useful technique for the identification of bacteria and involves shining multiple wavelengths of light on wounds and collecting their emission responses thus helping understand the pathophysiology of the wounds. Fluorescence methods have been found to be one of the most straightforward, noncontact, and sensitive methods for the detection of biomolecules (intrinsic fluorophores) such as tryptophan, nicotinamide adenine dinucleotide hydride (NADH), flavins, etc. 5 Different bacteria contain varying amounts of these fluorophores in differing microenvironments which help in distinguishing different types of bacteria. 6 This study was carried out to evaluate the accuracy of Illuminate®, one such autofluorescence-based point of a care imaging device that uses a multispectral imaging technique to shine multiple excitation sources on the region of the wound and collects spectral images. An Artificial Intelligence (AI) algorithm then runs on the collected spectral images to compare the biomolecule fluorophore intensity toward Gram-type bacterial classification. The results obtained from this device were compared against the current gold standard method of bacteriological culture.
Methods
Study Design
An interventional single arm comparative study was conducted at a tertiary care center for diabetes after obtaining institutional ethics committee approval (IEC/N-009/10/2017) registered with the Clinical Trial Registry of India (Reg. No. CTRI/2018/10/016147). Both acute and chronic wounds were considered for imaging and the total sample size required for detection of infection using the device was estimated as 140 (for 99% confidence level, assumed prevalence of infection as 70% and a precision of 10%. 7 Consecutive patients attending the outpatient department were recruited after screening for inclusion criteria and obtaining both informed and written consent forms. The persons with type 2 diabetes (T2DM) and with new and/or chronic DFU with nonintact skin (stasis ulcers both arterial and venous) were included in the study. The persons with an existing autoimmune disease and skin diseases such as eczema, psoriasis in the area close to the wounds were excluded. We excluded patients with osteomyelitis, as the bone region is high autofluorescent in nature and needs additional algorithm training. A total of 178 persons with T2DM with acute and/or chronic DFU were recruited into the study. Patients’ peri lesion noninfected site served as their own control.
Illuminate device and Imaging Procedure with Illuminate®
Intrinsic fluorescence exhibited by bacteria and fungi are attractive methods for the rapid identification of infection on wounds without any additional steps. Previous studies have reported that bacteria have characteristic emission fluorescence when excited in the UV and blue regions of light contributed mainly by metabolic and infectious markers such as NADH, Flavin and Porphyrin. 8 Shelly et al, 9 reported fluorescence fingerprinting by characterizing each bacteria’s excitation and characteristic emission wavelengths for rapid and accurate identification of infection especially Pseudomonas species and Staphylococcus aureus. On the other hand, Ammor et al 5 developed a method towards species level bacterial detection leveraging the intrinsic fluorescence of aromatic amino acids and nucleic acids, suggesting that autofluorescence spectral analysis can be a rapid and inexpensive technique. Based on the above principles of using autofluorescence imaging of bacteria, a novel imaging device named Illuminate® (Adiuvo Diagnostics Private Limited, Chennai, India) (Figure 1) was developed, combining multiple light sources (370, 395, and 415 nm) to shine on the wound region and collects multispectral images in approximately 30 sec to detect infection on wounds. An image processing algorithm then analyses the spectral images, and a pretrained machine learning algorithm compares the relative intensity between different autofluorescence biomarkers 10 thus enabling classification of bacteria into Gram-positive and Gram-negative in 2 min.10,11 The device is noncontact, does not require the addition of any reagents and removes the burden of interpreting spectral images. The AI Engine runs on the device and can even be used in remote places with no internet access. The device provides a comprehensive report along with automated wound measurement options including a progressive report, which can be synced with the hospital’s central database. The device is easy to use and requires training to image the wound regions consistently without shaking.

Illuminate® imaging device.
Illuminate device was used to collect Multispectral wound images either in a dark room or using a black hood covering the wound region. The wound imaging was done by the trained investigator. Before imaging, wounds were first cleaned with normal saline solution. The study participants’ details were entered in the Illuminate Imaging Application of the device. The user was guided to image the wound at a distance of 10 to 12 cm from the area of interest. A series of 16 multispectral images were automatically captured by the device in ∼20 sec. The user was then prompted to trace the region of the wound post image capture. In under 2 min, an algorithm was processed in the background and displayed a color-coded wound image along with a Gram type of bacteria (if infected) along with the length and breadth of the wound.
Specimen Collection
Tissue samples were taken with Alley’s tissue holding forceps and scissors by the attending plastic surgeon. These tissues were collected before and after debridement from the color-coded region as indicated by the device. Specimens were collected from all the study participants and placed in a sterile transport container and sent to the microbiology laboratory in the hospital for routine standard culture and biochemical methods for identifying and classification of bacteria.
Culture
All tissue samples were cultured in the microbiology lab. All tissue samples were analyzed initially by Gram-staining and bacterial identification using conventional aerobic phenotypic methods by plating into various enriched and differential growth mediums and biochemical assays. 12 Bacterial load was estimated by plate count semiquantitative analysis method and then graded as scanty, light, moderate, or heavy (1+, 2+, 3+ or 4+) of which moderate and heavy growth indicated a significant bacterial load (ie, greater than 100 000 per gram of tissue). The antibiotics susceptibility profile was then obtained using the disk diffusion method as per Clinical Laboratory Standards Institute (CLSI) guidelines. 13 The results of gram-staining, bacterial identification, and its drug susceptibility profile were obtained from the microbiology lab in a report format for each patient sample.
Statistical Analysis
The estimates of diagnostic tests such as true positives, true negatives, false positives, false negatives were first evaluated and used for the calculation of sensitivity, specificity, accuracy, positive predictive value, and negative predictive value. The results obtained from the imaging device were compared with the results of the tissue culture to calculate accuracy, sensitivity, specificity, positive predictive value, and negative predictive value. The statistical analysis was performed using Python software.
Results
Clinical Characteristics of the Study Participants
The imaging of the wound was done for a total of 178 persons with DFU. The tissue sample was taken from 203 sites and analyzed. Out of 178 study participants, 144 were male and 34 were female with a median age of 59 years. 13 Patients were on prior antibiotics usage and were included in the study (Table 1). About 60% of the participants were found to have third-grade ulcers based on the Wagner’s wound classification score, while 34.3% had grade 2 and 6.2% had grade 1 ulcers respectively. A total of 131 patients underwent debridement, while 20 participants were directed towards cleaning and dressing, 17 towards sequestrectomy, 3 underwent exostectomy, 1 escharotomy, 1 bursa excision, and 16 patients had to undergo an amputation. 39 wounds had exudates. Around 63% of the study participants were found to have an HbA1c level >8%.
Basic and Clinical Profile of the Study Participants.
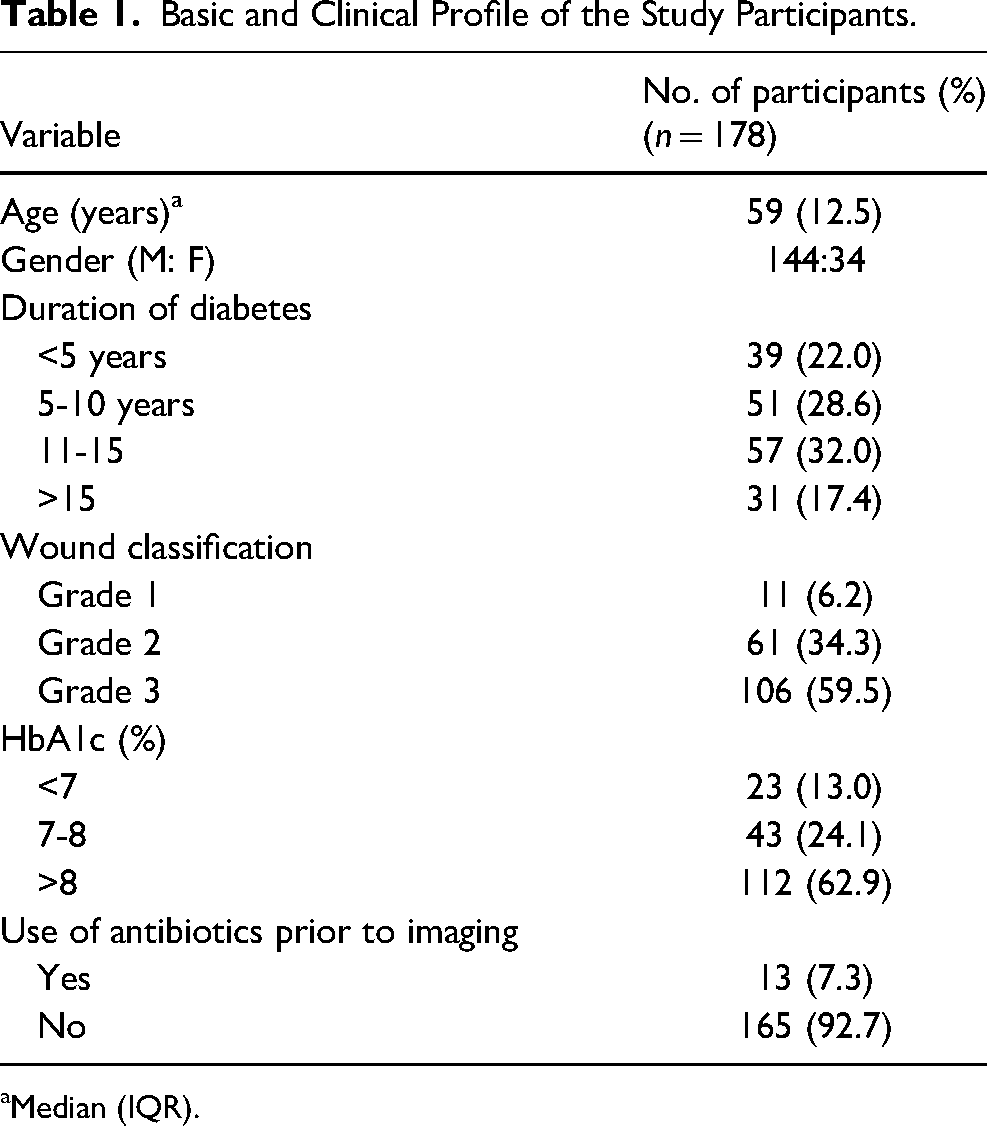
Median (IQR).
The percentage of treatment procedures is shown in Figure 2.
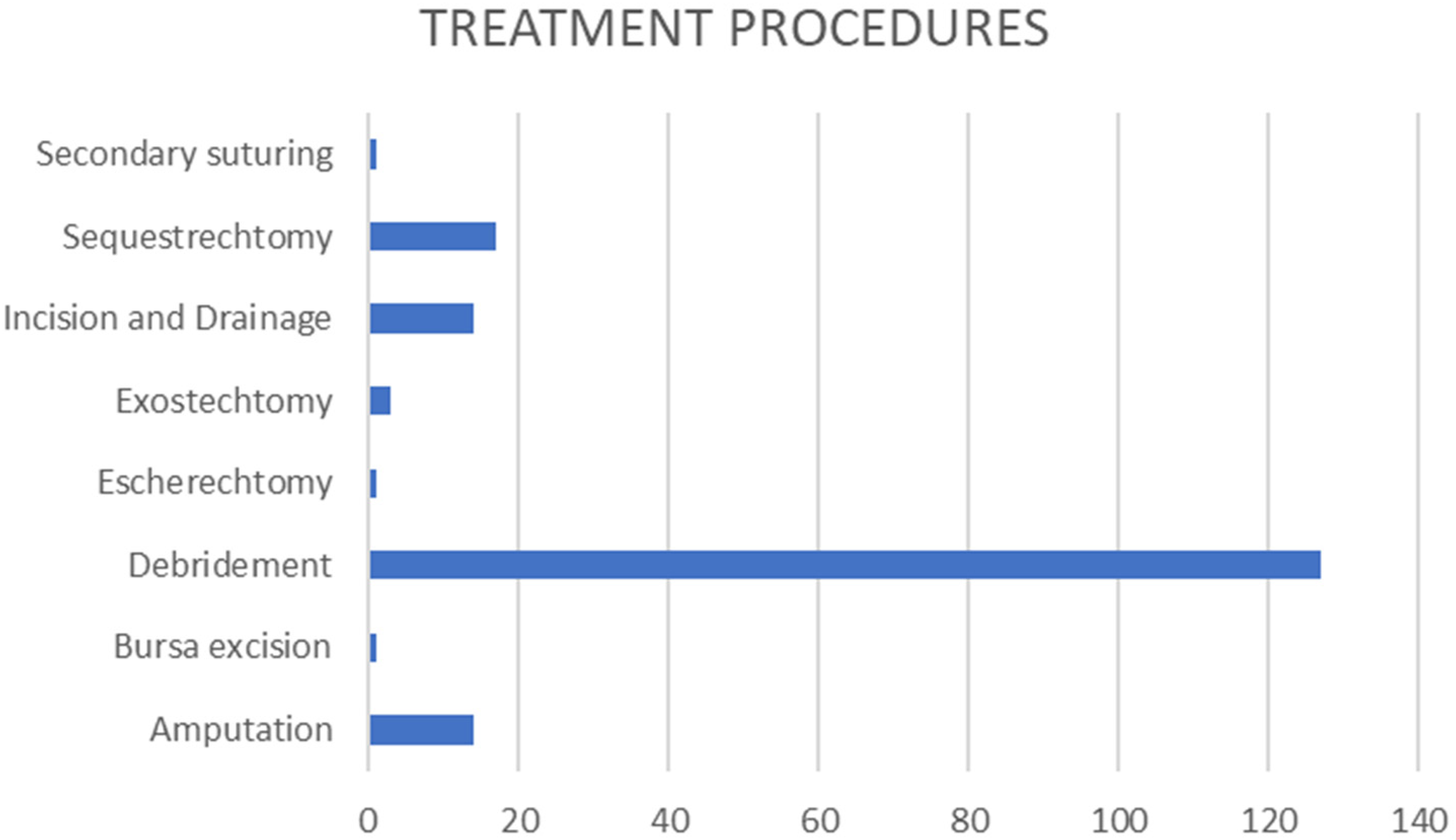
Percentage of treatment procedures.
Organisms Isolated in Culture
From the 203 tissue samples collected, 14 clinically relevant pathogens were isolated (Table 2) while 28 cases showed no growth which included reference sites as well. 63 samples revealed one Gram-negative bacteria, 53 had one Gram-positive bacteria, while 6 tissues had 2-gram-negatives, 4 samples had 2-gram-positives and 49 patients had both gram-positive and gram-negative bacteria (Table 3).
Details of Pathogens Isolated From Wound Samples by Culture Methods.
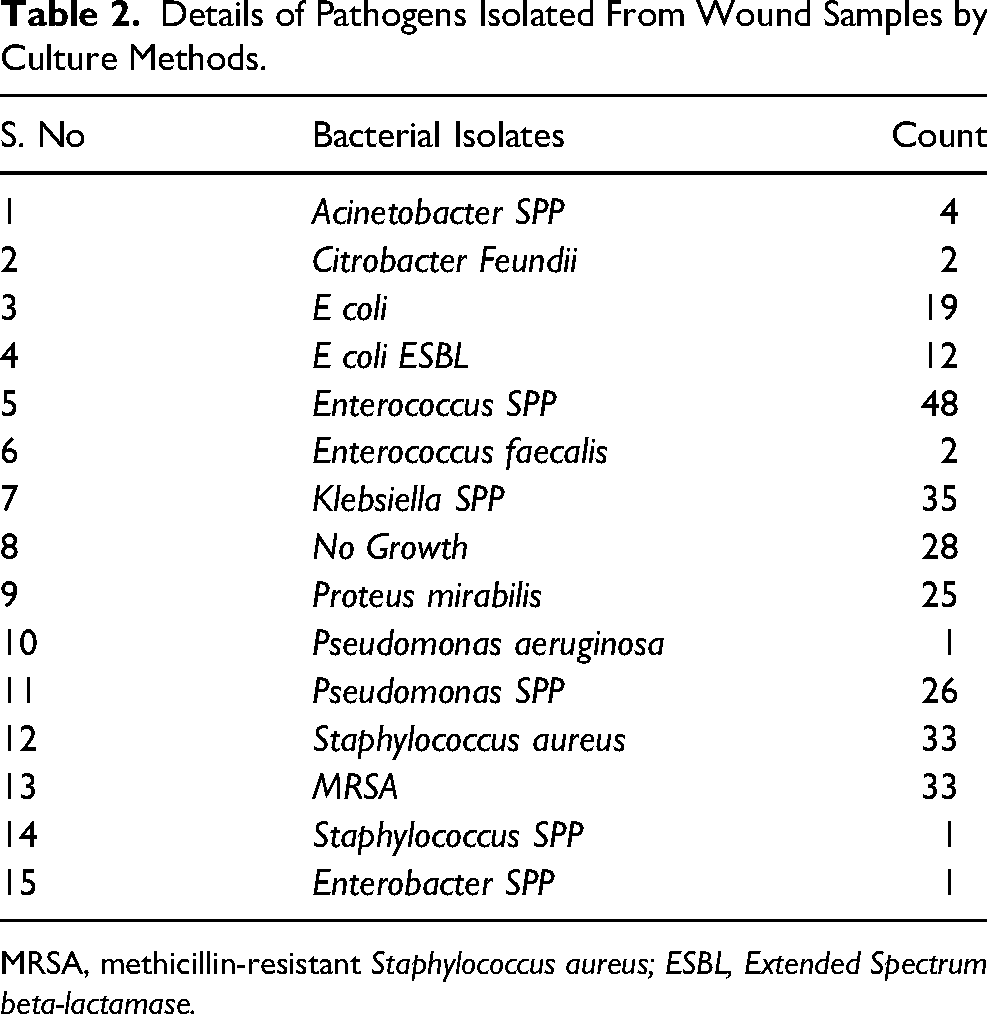

MRSA, methicillin-resistant Staphylococcus aureus; ESBL, Extended Spectrum beta-lactamase.
Identification of Gram-Positive and Gram-Negative Organisms in Wound Samples by Culture Methods vs Illuminate Device.
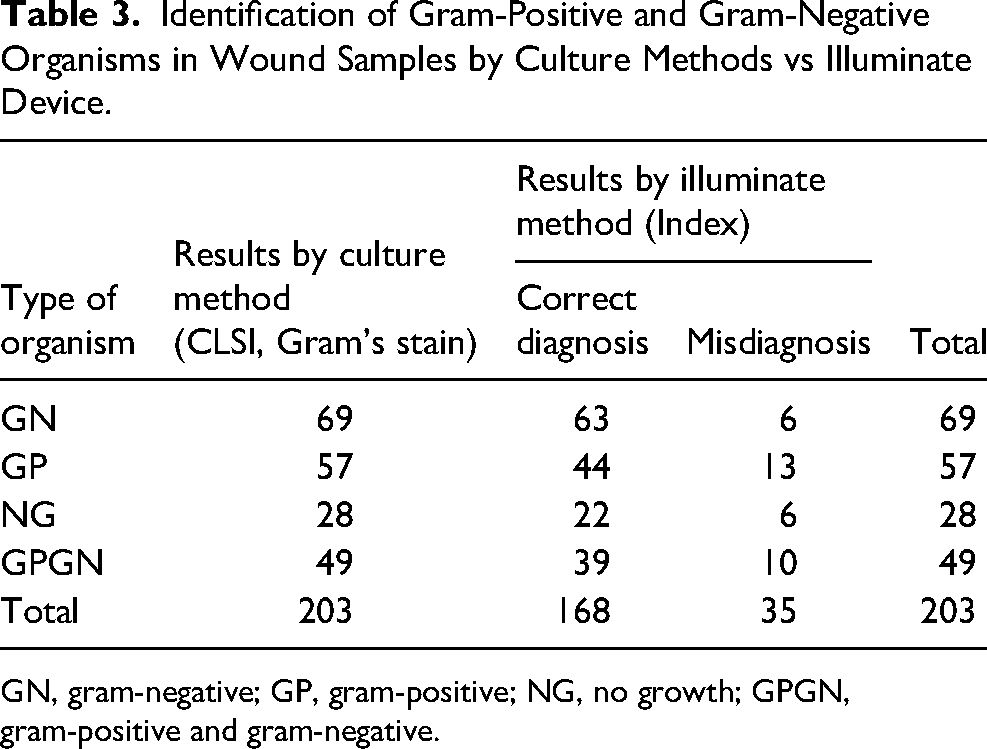

GN, gram-negative; GP, gram-positive; NG, no growth; GPGN, gram-positive and gram-negative.
Illuminate Inference
The device color codes Gram-positive bacteria with red (Figure 3b) and gram-negative with green (Figure 3a, c). Tissue culture gram type results were compared against the device gram type results. Illuminate device displayed 63-gram-negative bacteria, 44 gram-positive, 22 No growths. 49 wounds were found to have a polymicrobial infection containing both gram-negative and gram-positive bacteria (Figureure 3d) and the device identified the same in 39 patients. 6 ulcers were found to have only gram-negative colonization and 4, gram-positive colonization.
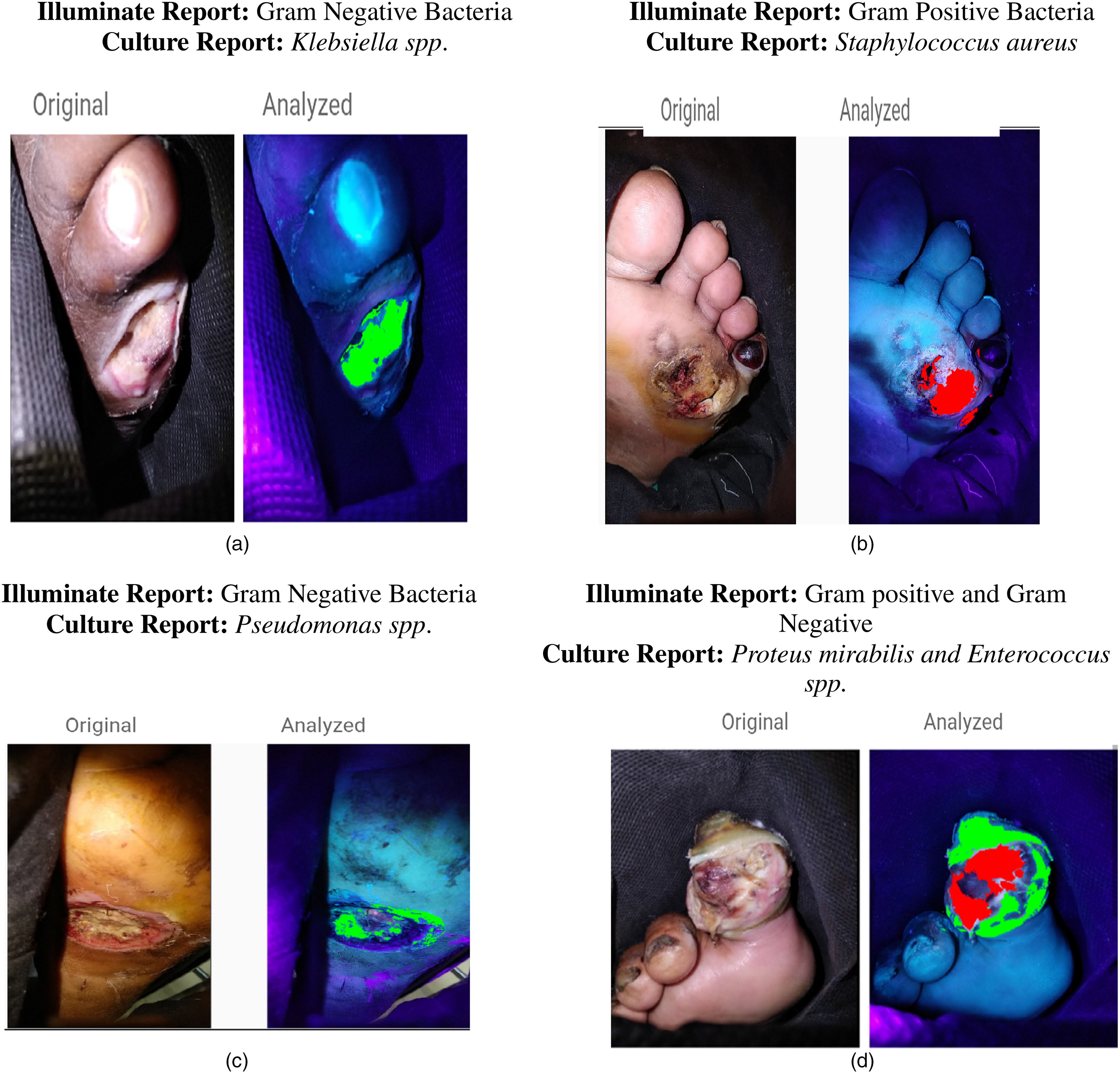
(a) Gram-negative bacteria-infected wound. (b) Gram-positive bacteria-infected wound. (c) Gram-negative bacteria-infected wound. (d) Gram-positive and gram-negative bacteria-infected wound. Figures 3a-3d are examples of clinical image and Illuminate® predicted image of Patient’s wound region. Figure 3a and 3c show a Green overlay indicating Gram-Negative infection and the corresponding culture reports indicated Klebsiella Spp. and Pseudomonas spp. Figure 3b indicated red over overlay predicting Gram-positive bacteria and the corresponding culture report was Staphylococcus aureus, 3d is an example of a wound infected by both Gram-positive and negative bacteria.
Further statistical inference was made by comparing the culture results with Illuminate device results (Table 3).
The results of the machine learning algorithm are summarized in the confusion matrix provided in Table 4. The accuracy of the device was found to be 89.54% with a positive predictive value of 80.77% for detecting gram-negative bacteria, 86.3% for gram-positive bacteria, and 91.67% for no growth. The negative predictive value corresponded to 92.0% for gram-negative bacteria, 87.2% for gram-positive bacteria, and 96.12% for no infection. The sensitivity for gram-negative bacteria was 91.30%, 77.2% for Gram-positive, and 81.5% for No growth. Specificity for Gram-negative was 82.14%, 92.71% for gram-positive, and 98.4% for No Growth. Multiclass area under the curve (AUC) score calculated using the One-versus-One method was 0.86.
Confusion Matrix and Results for the Gram Type Classification Using Illuminate® Device With the Standard Culture-Based Test.
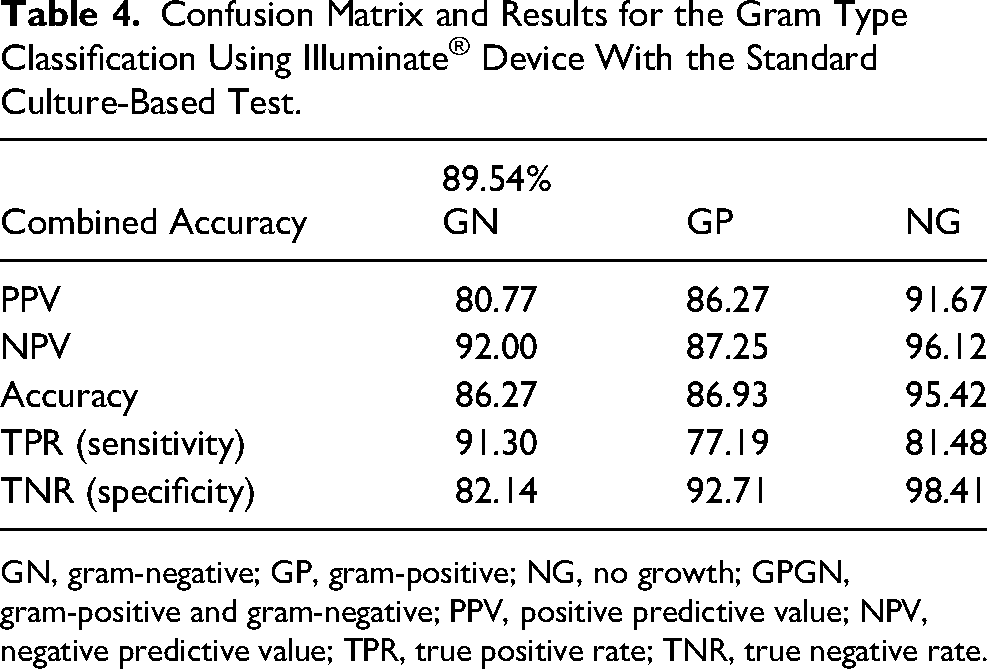
GN, gram-negative; GP, gram-positive; NG, no growth; GPGN, gram-positive and gram-negative; PPV, positive predictive value; NPV, negative predictive value; TPR, true positive rate; TNR, true negative rate.
Discussion
Understanding the pathophysiology of DFU is essential for effective treatment and prevention of its downstream complications. Clinical Signs and Symptoms Score continue to be the recommended way of detecting wound infection. 14 However, this assessment can be very subjective as demonstrated by various studies. An accurate, objective method would be a boon to the doctors to better understand the infection levels and to provide tailored antibiotic treatments.
Our study findings showed that the sensitivity for gram-negative and gram-positive bacteria was 91.3% and 77.2% respectively. A sensitivity of 81.5% was found for no growth of any bacteria. The specificity of the test of the device was 82.1% for gram-negative organisms, 92.7% for gram-positive, and 98.4% for no growth of the organisms. AUC score of 0.86 also indicates a significant discriminating ability of the device to the presence of pathogens.
A pilot study conducted in a wound care clinic of Ireland on the efficacy of a bacterial fluorescence imaging device (Moleculight) showed a sensitivity of 100% and specificity of 78%. 15 The same study demonstrated that the device has sensitivity and specificity of 100% in identifying the Pseudomonas spp. by visually interpreting the cyan fluorescence from the autofluorescence images. This study also showed the effect of appropriate selection of antibiotics and further recovery of the wound within a 2-week period. It is to be noted that these images need to be manually interpreted. 16 and cannot automatically distinguish Gram-positive and Gram-Negative bacteria.
Our study of 178 patients using the Illuminate® imaging device can detect infections and differentiate the gram type of bacteria using multispectral image information and AI. The device has an accuracy of 89.54%, a positive predictive value of 86.27% for detecting Gram-positive bacteria, 80.77% for Gram-negative bacteria, and 91.67% for no infection. The negative predictive value corresponded to 87.25% for Gram-positive, 92% for Gram-negative, and 96.12% for no infection.
The study conducted by DaCosta RS., et al showed that only 52.5% of the bacterial load was accurately identified by clinical assessment whereas 74.6% of the occurrences of wound infection was correctly identified by fluorescent imaging-based technique. 17 Another study has re-confirmed the high accuracy in the detection of the pathogenic infection of the wound by an autofluorescence imaging device compared to the clinical evaluation of signs and symptoms in detecting the wounds. It also demonstrated an improved sensitivity of the combined use of bacterial fluorescence imaging with clinical signs and symptoms (72%) compared to the use of only clinical signs and symptoms (22%) in the identification of wounds with moderate to heavy bacterial loads (≥104CFU/g). 18
Predominantly Gram-positive bacteria such as methicillin-resistant Staphylococcus aureus 19 , methicillin sensitive S. aureus, Streptococcus species are typically identified in the acute stage of wounds while gram-negative bacteria such as Escherichia coli, Pseudomonas aeruginosa, Klebsiella spp., are detected in chronic wounds.20–22 Our study showed the presence of Enterococcus spp. (17.8%) majorly followed with the presence of Klebsiella spp. (13%) and Staphylococcus aureus (12.2%). A bacterial concentration >104 CFU/g of tissue is necessary to cause infection in complex extremity wounds.23,24 Infections may then lead to a delay in wound healing, thus early detection of bacteria above 104 CFU/g of tissue becomes imperative to prevent complications.
The high sensitivity and specificity observed in the present study, along with an exploratory study done earlier 10 reiterates that this novel device can be used as a screening device for early infection detection and understanding the gram type of bacteria colonizing the wound. Definitive tests such as culture, polymerase chain reaction take time are reagent intensive, and are error prone due to reasons such as error during sample transport, proximity to a diagnostic lab, etc. This device can thus be used as a screening tool to overcome these shortcomings. The device can also help monitor the treatment efficacy over a period of wound treatment and identify regions to take a tissue sample. From our results, we could see a marked decrease in bacterial fluorescence spatially, before and post debridement and there was a 93% organism correlation from the culture report obtained. Hence it can be used to assess wounds qualitatively post debridement. 11
Currently, wounds are assessed by visual inspection, and it is difficult to track wound infection and healing post debridement. More importantly, an instant decision on the gram type of bacteria infecting the wound can aid the doctors towards the first line of antibiotic prescription. This device will reduce the time and aid the clinicians ineffective treatment by understanding the microbial burden on the wound region in real time. Manual interpretation of images is very tough and requires trained personnel and possesses some challenges in ensuring the accuracy of decisions. Incorporation of machine learning algorithm aids in easier understanding of infection regions. In future, with more images, genus and species level machine learning training can also be performed which can guide towards effective antibiotic treatment.
Limitations
Bone, tendon, and fat along with betadine and other cleaning solutions are autofluorescent in nature. Hence wounds are to be cleaned with normal saline before imaging. Also, the device cannot be used to interpret infections in the bone/fat region. The light sources used here have a penetration depth from 0.5 mm to ∼1 mm and hence the device cannot understand the status of infection in closed wounds. In the case of polymicrobial infection, the device displays the gram type that is more in number in that particular spatial region.
Conclusion
Illuminate®, a novel hand device developed based on multispectral imaging of autofluorescence exhibited by bacterium has proven to be an early screening device in the identification and classification of bacteria. The device looks at both metabolic and infectious markers of the underlying pathogen exhibiting a good overall accuracy and thus does not detect commensals. The sensitivity to detect no infection wounds is high and hence the prescription of antibiotics can be provided judiciously. The device’s high sensitivity to detect gram-negative bacteria may aid physicians toward the administration of agents with activity against gram-negative bacteria. The high specificity to detect Gram-positive bacteria can eliminate any burden faced by clinicians to combat gram-positive infections. In addition, infections in the peri-wound region and away from the wound region can be detected by the device, thus aiding clinicians in the better debridement process and improved understanding of the wound infection levels qualitatively. These results observed are closer to the gold standard culture methods and thus aids the clinicians towards the first line treatment. This device will also be helpful in convincing the patients easily to proceed with treatment by showing the infection status of the wound empirically in the outpatient clinical setting. The noninvasive and less time-consuming procedure are the added advantages of the device.
Footnotes
We thank the support from Ms Malathi, for her patient assistance, the microbiology team for providing insights from culturing perspective, Mr Anand Kumar, Dr Satyavani, and Ms Suganya for their support during the trial.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Ethical Approval
Institutional Ethics Committee approval (IEC/N-009/10/2017) was obtained before the initiation of the study.
