Abstract
Introduction
Hepatocellular carcinoma (HCC) screening in patients with hepatic cirrhosis (HC) relies on ultrasound and alpha-fetoprotein (US + AFP), which has limitations in sensitivity, particularly for early-stage HCC detection. This study aims to evaluate the performance of a novel multi-omics blood test, HCCscreen, with its individual components (methylation, AFP, Des-γ-Carboxy Prothrombin (DCP), mutations) and the standard US + AFP for HCC screening in a hepatic cirrhotic population.
Methods
A total of 5078 patients with known high-risk for HCC were recruited. A prospective screening study was conducted on 650 patients with hepatic cirrhosis identified by ultrasound. Blood samples were collected from all patients before the confirmation of diagnosis by imaging and/or pathological examinations. The performance of HCCscreen, individual markers and US + AFP were calculated and compared. Statistics was performed with Graphpad Prism 5.0.
Results
HCCscreen exhibited a sensitivity of 86.3% at a specificity of 81.3%, with a positive predictive value (PPV) of 28.2% and a negative predictive value (NPV) of 98.6%. The positive likelihood ratio (LR+) was 4.61 and the negative LR (LR-) was 0.17. The positive detection rate (PDR) for all markers increased with more advanced HCC stages, whether Barcelona Clinic Liver Cancer (BCLC) or clinical staging. Among the single-omics, methylation showed the highest PDR, followed by AFP, DCP and mutations. HCCscreen demonstrated superior overall performance with an AUC of 0.87, outperforming individual markers like methylation (AUC = 0.76), AFP (AUC = 0.83), and DCP (AUC = 0.77). Crucially, HCCscreen's PDR was significantly higher than US + AFP in early-stage HCC (BCLC-0 and clinical stage I). Furthermore, while AFP's PDR varied significantly by sex, HCCscreen's performance remained consistent across all demographics. Correlation analysis revealed a significant association only between the HCCscreen score and the methylation score.
Conclusions
The multi-omics approach of HCCscreen significantly enhances early HCC detection in patients with hepatic cirrhosis compared to both its individual components and the current standard of US + AFP. Its robust and consistent performance across patient demographics underscores its potential as a superior tool for population-wide early HCC screening.
Introduction
Hepatocellular carcinoma (HCC) is the fifth most common and the third most deadly cancer, with a 5-year survival rate of only 12.1%.1,2 The most effective way to improve the survival rate of HCC is to detect the lesion at an early stage and provide timely radical treatment. 3 Therefore, the application of effective screening methods and early intervention is crucial for the prevention and treatment of HCC.
Cirrhosis is a crucial step in the progression from various types of hepatitis and liver diseases to HCC. Taking hepatitis B infection as an example, it is generally believed that patients will go through a pathological development process of hepatitis B virus infection - chronic hepatitis B - hepatitis B cirrhosis - liver nodules - HCC. At the cirrhosis stage, stem cells have already suffered irreversible damage. If the pathogenic factors such as hepatitis B persist, cirrhosis usually continues to progress to liver nodules, precancerous lesions, and even HCC.4,5 Therefore, cirrhosis is a high-risk factor for HCC. Current epidemiological data show that the annual incidence of HCC in cirrhosis patients (3%-6%) is significantly higher than that in patients with chronic hepatitis B (∼1%). 6 HCC screening in cirrhosis patients can help improve the early detection rate and early intervention rate of HCC, and help reduce the incidence of HCC.
Among the current screening methods, ultrasound (US) combined with alpha-fetoprotein (AFP) remains the standard screening method recommended by guidelines and consensus, but its sensitivity for HCC screening is limited and its compliance is not ideal. 7 In recent years, liquid biopsy based on cell-free circulating DNA (cfDNA) has emerged as a promising noninvasive screening method in clinical applications. 8 In the field of HCC screening, several studies have confirmed the feasibility of liquid biopsy for detecting mutations and methylation.9–11 However, due to the low content of cfDNA in plasma, detecting low-frequency mutations is challenging, which may lead to false-negative results, especially common in early-stage cancer patients. 12 Compared with mutations, cfDNA methylation detection has been widely used in cancer screening, auxiliary diagnosis, prognosis prediction, and monitoring. 13 Currently, bisulfite conversion followed by sequencing is considered the standard method for DNA methylation analysis, but this approach can cause DNA damage, reducing the detectability of cfDNA and the sensitivity of the assay. 14 Given these issues, the combination of multiple biomarkers from different omics has the potential to improve the performance of cfDNA-based cancer detection. 15 Omics-based detection refers to the use of multiple omics rather than a single omics for testing, including but not limited to genomics, transcriptomics, proteomics, metabolomics, lipidomics, and glycomics. Omics-based detection examines multiple dimensions of sample indicators at an overall level, typically integrating two or more single omics for comprehensive analysis. Currently, the amount of cfDNA obtained from blood draws is insufficient to support parallel multiple tests, and most previous tests have focused on a single type of biomarker, making it difficult to achieve parallel omics-based detection.
To address the aforementioned issues, we have selected a technology that supports parallel analysis of mutations and methylation, namely the Mutation Capsule Plus (MCP) technology, along with the corresponding multi-omics test (HCCscreen, a multi-omics combined early screening test for HCC that integrates multiple gene methylations, TP53/TERT/CTNNB1 mutations, AFP, and Des-γ-Carboxy Prothrombin (DCP)). This test has been extensively validated in a large-scale prospective study among hepatitis B surface antigen (HBsAg)-positive individuals.9,10 In this study, we aim to verify the performance of HCCscreen in screening for HCC in patients with liver cirrhosis in a real-world hospital setting through screening, hoping to provide strong evidence for the application of HCCscreen in screening high-risk populations for HCC in real-world scenarios.
Methods and Materials
Ethics and Informed Consent
This study was approved by the Ethics Committee of The First People's Hospital of Yunnan Province, The Affiliated Hospital of Kunming University of Science and Technology before its commencement and was conducted in accordance with the hospital's guidelines for the management of clinical research (approval number: KHLL2021-KY108). All patients signed an informed consent form before participating in this study. Patients were informed of the purpose of blood collection and the content of the tests, and were also informed of the test results. All patients were informed that the test results might be used for statistical purposes in scientific research and were aware that no personal information would be disclosed in this study.
Study Design
The objective of this study is to verify the effectiveness of HCCscreen testing for screening of HCC high-risk populations (cirrhosis) in a hospital setting. A prospective study design was adopted. According to the definition of cirrhosis in the “Guidelines for Stratified Screening and Monitoring of Primary Liver Cancer (2020 Edition)”, all patients diagnosed cirrhosis by ultrasound were prospectively enrolled and blood samples were collected. 6 HCCscreen testing was conducted, and ultrasound and AFP testing were performed as comparisons. Patients were screened according to the inclusion and exclusion criteria. All patients underwent imaging and/or pathological examinations after blood collection for disease confirmation.
Patients, Sample Collection and Clinical Diagnosis
The enrollment period for patients in this study was from February, 2023 to July, 2025. The inclusion criteria for patients in this study were as follows: aged 18 to 80 years, regardless of gender, patients with liver cirrhosis caused by HBV, HCV and other etiologies as defined by guidelines, with or without liver nodules. 6 The exclusion criteria included: patients with a history of malignant tumors, patients who had received anti-tumor treatment before this visit, patients who did not agree to participate in this study, patients with unqualified sample collection or quality control, and patients whose final diagnosis information could not be traced. Subsequently, patients were retrospectively grouped and analyzed based on their disease types. As a result, 5078 patients with known high-risk for HCC were recruited, and all patients underwent US + AFP screening, and those with confirmed diagnosis of hepatic cirrhosis by US were involved in subsequent HCCscreen screening and diagnosis with CT/MRI/Pathology examinations. As a result, 51 patients were diagnosed HCC. The detailed procedure of patient recruitment and examination is illustrated in Figure 1, and patient categories are listed in Table 1.
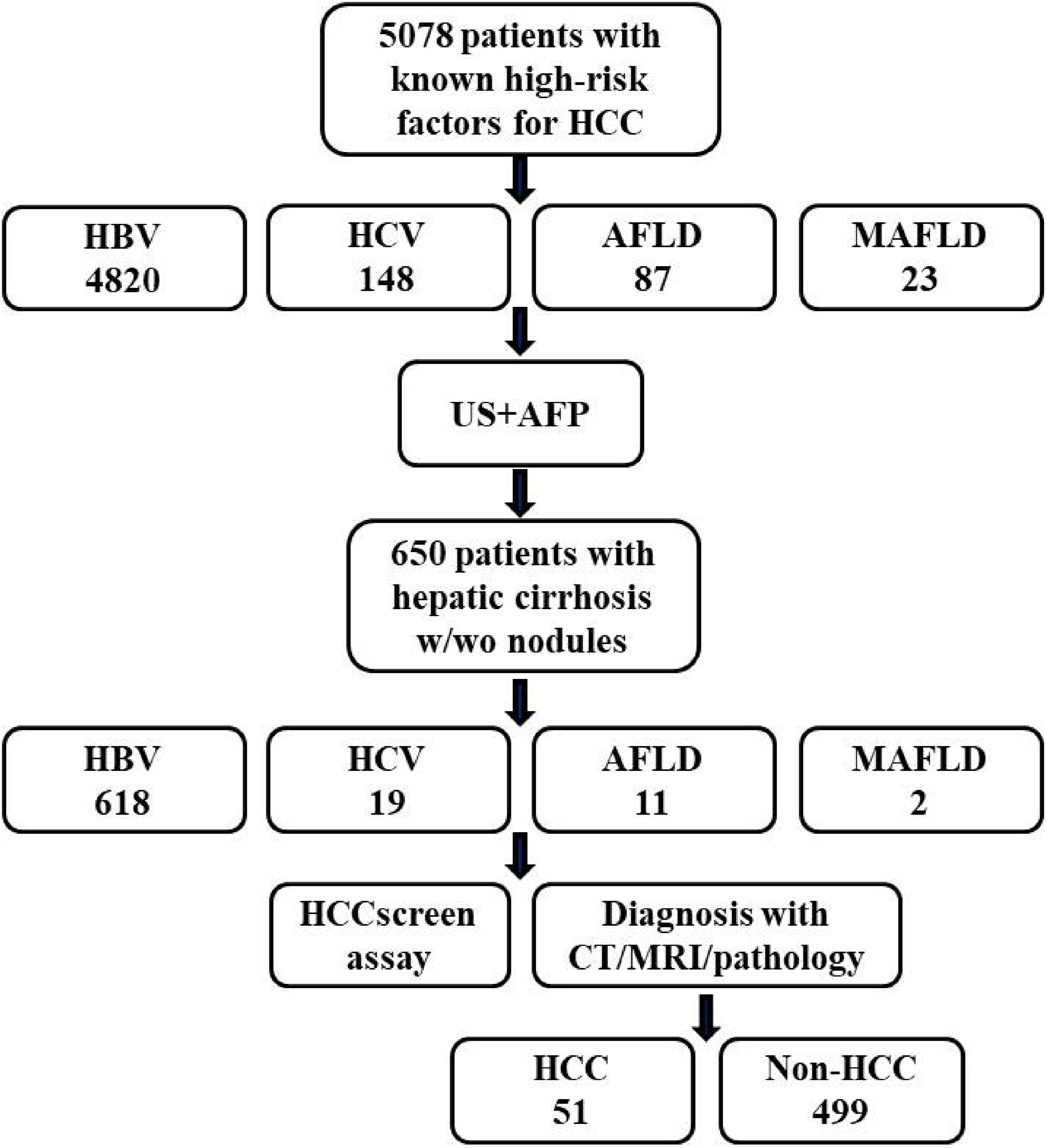
Flowchart for study design and implementation. 5078 patients with known high-risk for HCC were recruited, in which 4820 were HBV- infected, 148 were HCV infected, 87 with alcoholic fatty liver disease (AFLD) and 23 with metabolic associated fatty liver disease (MAFLD). All patients underwent US + AFP screening, and those with confirmed diagnosis of hepatic cirrhosis by US were involved in subsequent HCCscreen screening and diagnosis with CT/MRI/Pathology examinations. As a result, 51 patients were diagnosed HCC. HCC, hepatocellular carcinoma; HC, hepatic cirrhosis; HBV, hepatitis B virus; HCV, hepatitis C virus; AFLD, alcoholic fatty liver disease; MAFLD, metabolic-associated fatty liver disease; US, ultrasound; AFP, alpha fetoprotein; w/wo, with or without.
Clinical Characteristics of Patients Involved in This Study.
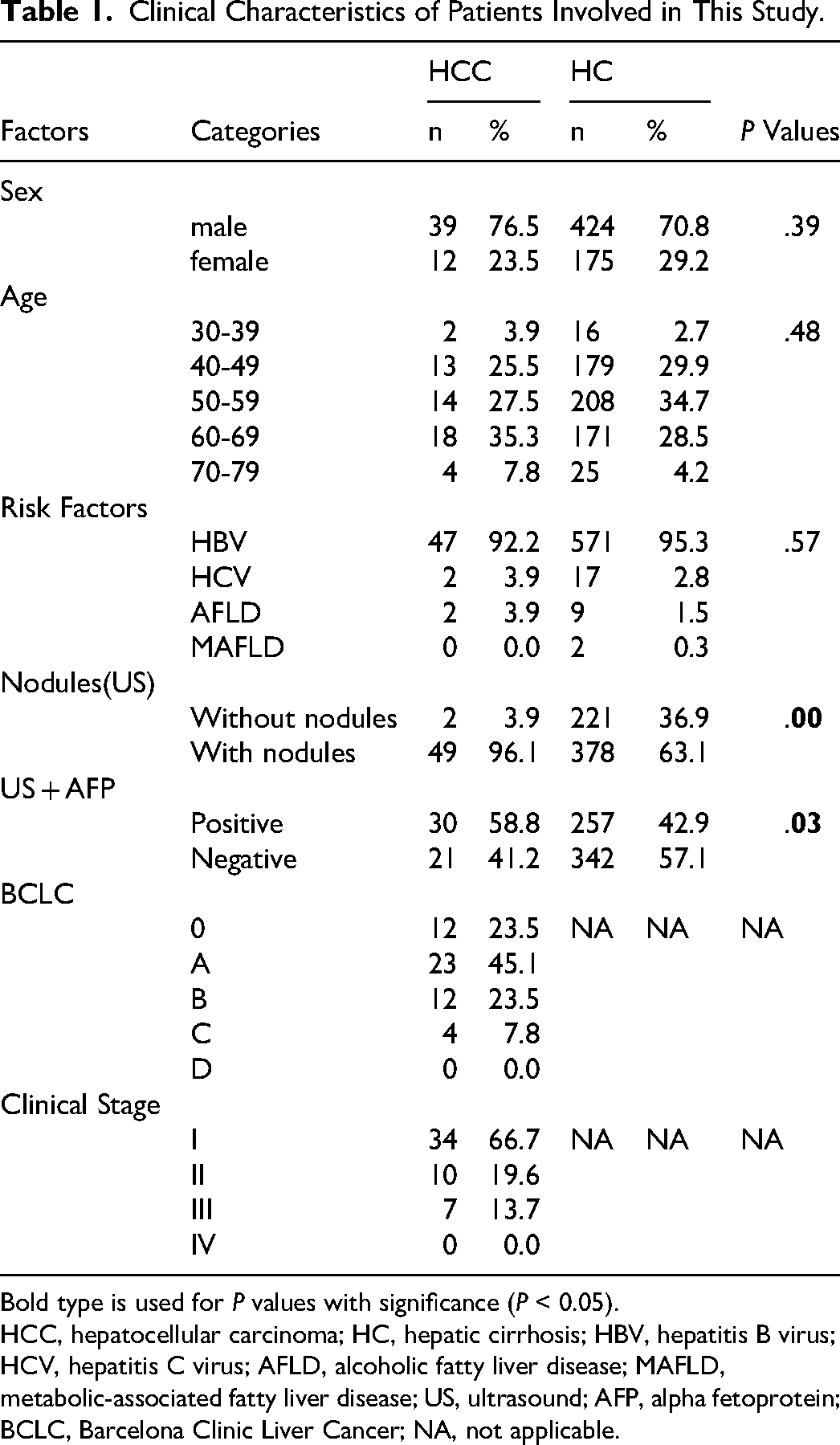
Bold type is used for P values with significance (P < 0.05).
HCC, hepatocellular carcinoma; HC, hepatic cirrhosis; HBV, hepatitis B virus; HCV, hepatitis C virus; AFLD, alcoholic fatty liver disease; MAFLD, metabolic-associated fatty liver disease; US, ultrasound; AFP, alpha fetoprotein; BCLC, Barcelona Clinic Liver Cancer; NA, not applicable.
The Performance of HCCscreen and a Comparison with US + AFP.

Bold type is used for P values with significance (P < 0.05).
US, ultrasound; AFP, alpha-fetoprotein; DCP, Des-γ-Carboxy Prothrombi; BCLC, Barcelona Clinic Liver Cancer.
All patients had blood drawn before the diagnosis in this study, using 10 mL K2EDTA anticoagulant tubes to collect 10 mL of venous whole blood. Patients underwent ultrasound and AFP tests before blood drawing. The diagnosis methods in this study included dynamic contrast-enhanced computed tomography (CT), dynamic contrast-enhanced MRI, gadoxetic acid disodium-enhanced MRI and/or pathological examination. The diagnostic criteria for HCC were in accordance with the standards of the guidelines (Diagnosis and Treatment Guidelines for Primary Liver Cancer (2024 Edition)). 16 According to the diagnosis results, patients were retrospectively classified and statistically analyzed (Table 1). The determination of US + AFP positivity was based on the definition from previous literatures. 9 In brief, ‘AFP/US-positive’individuals had any of the following: 1) serum AFP levels of >400 ng/mL regardless of US-detected nodule; 2) US-detected nodule of ≧2 cm in size regardless of serum AFP concentration; 3) US detected nodule of ≧1 cm in size with serum AFP≧200 ng/mL. The ‘AFP/US-negative’ individuals were defined as having serum AFP levels of <20 ng/mL and/or without a US-detected liver nodule.
DNA Extraction, NGS Library Construction, Targeted Sequencing, AFP and DCP Detection
The methods for DNA extraction, library construction, and targeted sequencing were consistent with those previously reported. 10 Firstly, cfDNA was extracted from plasma samples using the Apostle MiniMax cfDNA Isolation Kit (C43468, Apostle), and white blood cell DNA was extracted using the QIAamp DNA Mini Kit (51306, Qiagen). DNA concentration was determined using the Qubit dsDNA HS Assay Kit (Q32854, Thermo Fisher Scientific). Then, the samples were digested with methylation-sensitive restriction endonuclease Hha I (R0139L, New England BioLabs) at 37 °C for 30 min and at 65 °C for 20 min. Next, the UltraII DNA Library Prep Kit (E7370L, New England BioLabs) was used to prepare the library (pre-mutation capsule plus library, MCP pre-library) according to the manufacturer's instructions. Subsequently, the MCP library was directionally amplified and prepared as described in previous literature. 10 Finally, the polymerase chain reaction (PCR) library products were purified and sequenced on the Illumina NovaSeq 6000 platform (Illumina). Serum AFP and DCP levels were determined using commercial kits compatible with the Abbott ARCHITECT i2000SR chemiluminescence immunoassay analyzer (Abbott Laboratories; Chicago, IL, USA) according to the manufacturer's instructions.
Algorithm and Threshold Determination
The HCCscreen algorithm and interpretation method were consistent with previous reports. 10 The final HCCscreen detection model included mutation, methylation scores, and the detection values of AFP and DCP, which were comprehensively integrated to establish the model and form the HCCscreen score. In the preliminary study, random forest algorithm was used to screen out 10 methylation markers with diagnostic value from the training cohort containing HCC and non-HCCsamples, 10 and based on this, a methylation score model was constructed. Meanwhile, the detection of multiple reliable HCC-related mutations was defined as a binary mutation score. Additionally, the quantitative detection results of the commonly used clinical protein markers AFP and DCP were integrated with the methylation score, mutation score, and a multi-dimensional HCC early screening integrated model was constructed using the random forest method. After obtaining the initial MCP score by maximizing the methylation score and mutation score, we further included AFP and DCP in the final model training. The model performance was evaluated by AUC in both the training set and the test set, and the sensitivity and specificity were determined based on the cutoff value of 0.70 optimized by the Youden index in the leave-one-out cross-validation of the training set. According to the corresponding kit instructions, the positive interpretation threshold for AFP was 20 ng/mL, and that for DCP was 40 mAU/mL.
Statistical Analysis and Graphing
GraphPad Prism (version 5.0) software was used to construct bar charts, establish receiver operating characteristic (ROC) models and perform curve analysis, correlation analysis and regression analysis. Chi-square test, corrected chi-square test or Fisher's exact probability test was used to compare the differences in rates. When the data distribution was non-normal, non-parametric tests (Mann-Whitney) were used to compare the differences between two groups of data. Hosmer-Lemeshow test was performed using the corresponding R package to evaluate the goodness of fit of Logistic regression models.
Results
HCCscreen Exhibited Optimal Performance in HCC Screening in HC Patients
We first conducted a systematic study on the performance of HCCscreen and the included single-omics tests in screening for HCC in patients with liver cirrhosis. A total of 5078 patients with known high-risk for HCC were recruited, and screened with US for hepatic cirrhosis. As a result, 650 patients with hepatic cirrhosis with or without nodules were identified, in which fifty-one out of 650 patients were diagnosed HCC, and the rest 599 patients were diagnosed hepatic cirrhosis (HC) by CT/MR and/or pathological examinations (Figure 1, Table 1). Figure 2 shows the scatter plots and positive detection rate (PDR) of HCCscreen and methylation, AFP, DCP, and mutations in patients at various stages of HCC, and makes a comparison. The overall sensitivity was 86.3% (44/51) at a specificity of 81.3% (487/599). The positive predictive value (PPV) was 28.2% (44/157), and the negative predictive value (NPV) was 98.6% (487/494) (Table 1, Figure 2). The corresponding positive likelihood ratio (LR+) was 4.61 and the negative LR(LR−) was 0.17. Figure 2A presents the scatter plots and PDR of HCCscreen and each single-omics in patients at different Barcelona Clinic Liver Cancer (BCLC) stages, while Figure 2B shows the scatter plots and PDR of patients at different clinical stages. It can be seen that, regardless of whether it is the BCLC stage or the clinical stage, the HCCscreen score shows an increasing trend with the advancement of the stage. The same trend can be observed in the methylation score, and similar trends can also be seen in AFP, DCP, and mutations. Correspondingly, the PDR also shows an increasing trend with the advancement of the stage, which is consistent with the changing trends of the above markers. Thus, it can be suggested that the level of PDR is determined by the marker level, and PDR shows stage dependance. Additionally, we observed that in early-stage patients, such as those at BCLC-0 and clinical stage I, the marker levels were lower compared to patients at other stages, and the corresponding PDRs were also lower.

The stage-dependent performance of HCCscreen and its components in HCC screening in HC patients. Panel A: scatter plot of individual biomarker levels and HCCscreen scores in BCLC stages. Panels of scatter plots and comparison of PDR are shown for HCCscreen score, methylation score, AFP, DCP and mutation, as indicated. Panel B: scatter plot of HCCscreen scores and biomarkers levels in clinical stages. Panels of scatter plots and comparison of PDR are shown for HCCscreen score, methylation score, AFP, DCP and mutation, as indicated.
We further compared the detection performance of different markers. Figure 3 shows the comparison of detection performance (PDR) of different markers in each stage of BCLC or clinical staging. It can be found that, regardless of BCLC staging or each clinical stage, the order of PDR is mutation < DCP < AFP < methylation. The same is true in the overall statistics including all stages. Thus, among the markers included in HCCscreen, the PDR of methylation is higher than that of the two proteins, while the performance of mutation is relatively weak. Their combined detection can help improve the overall detection performance. The ROC curve further validates this conclusion. As can be seen from Figure 4, the overall AUC of HCCscreen reaches 0.87 (95% CI: 0.81∼0.92), which is superior to methylation (AUC = 0.76, 95% CI: 0.69∼0.84), AFP (AUC = 0.83, 95% CI: 0.77∼0.89) and DCP (AUC = 0.77, 95% CI: 0.67∼0.86). Statistics shows that the AUC between HCCscreen and methylation was significant (P = .030), while it was not significant between HCCscreen and AFP (P = .420), or between HCCscreen and DCP (P = .076). Hosmer-Lemeshow test was performed to evaluate the goodness of fit of the Logistic regression model (overall, Figure 4), and the P value was calculated based on the chi-square distribution and degrees of freedom. P value of .264 was obtained, indicating that the model fits well in the validation.

Comparisons of performance among mutation, DCP, AFP, methylation and HCCscreen at each BCLC or clinical stage. PDR for individual biomarkers is compared for each stage, as indicated.

The ROC curves for HCCscreen, methylation, AFP and DCP. The AUC is labeled in each panel.
We further investigated the relationship between performance and gender or age. Figure 5 compares the differences in various markers in terms of gender and age, including the existing screening method, US + AFP. The results showed that the PDR of AFP in female HCC patients was significantly higher than that in male patients (P < .05). However, there was no significant difference in the PDR of other markers between male and female (Figure 5A). Additionally, the PDR of US + AFP in patients aged 60 and above was significantly higher than that in patients aged 50-59, while there was no significant difference in the PDR of the other markers among the three age groups (Figure 5B). Overall, sex and age did not influence the performance of HCCscreen.

The influence of sex and age in the performance of HCCscreen and individual markers. Panal A: A comparison of biomarker performance between two sexes. Female showed significantly higher PDR than male in AFP. Panel B: A comparison of biomarker performance among various age groups. Patients over 60 years old exhibited significantly higher PDR in US + AFP than patients between 50 and 59 years old.
HCCscreen Exhibit Better Performance Than US ± AFP in HCC Screening in HC Patients
US + AFP is currently the main method for HCC screening. To understand the performance differences between HCCscreen and US + AFP, we directly compared their performance and studied the performance of their combined use. As shown in Figure 6A and B and Table 2, the overall performance of HCCscreen was significantly better than that of US + AFP (P < .01). In the BCLC staging (Figure 6A), the performance of HCCscreen in the BCLC-A stage was significantly better than that of US + AFP (P < .01) (Table 2). Similar trends were also observed in BCLC-0 and BCLC-B stages, although they were not statistically significant. In the BCLC-C stage, the PDR of both was 100%. Similarly, in the clinical staging (Figure 6B), the PDR of HCCscreen in the clinical stage I was significantly better than that of US + AFP (P < .01) (Table 2). In stage II, a similar trend was also observed, although it was not statistically significant. In stage III, the PDR of both was 100%. The above observations indicate that the performance difference between HCCscreen and US + AFP mainly comes from the fact that the former has a significantly higher detection rate of early-stage HCC (BCLC-A, stage I) than the latter. Therefore, HCCscreen is more suitable for early screening of HCC than US + AFP.
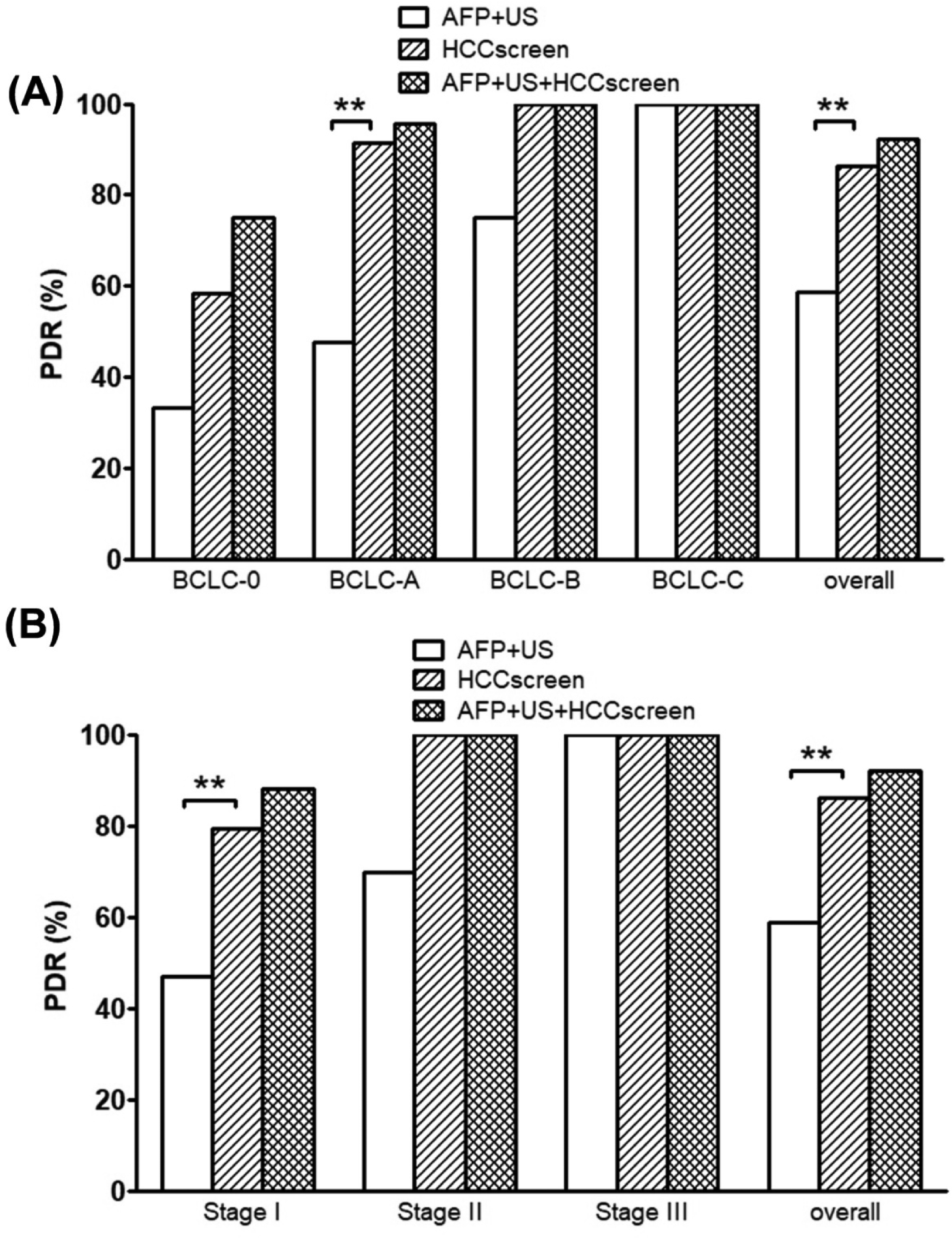
Comparison of performance and combination of HCCscreen and AFP + US. Panel A: A comparison of performance in AFP + US, HCCscreen and their combination based on BCLC staging. HCCscreen exhibited significantly better performance than AFP + US in BCLC-A and overall groups. Panel B: A comparison of performance in AFP + US, HCCscreen and their combination based on clinical staging. HCCscreen exhibited significantly better performance than AFP + US in stage I and overall groups.
Finally, we investigated the correlations among the various markers. As shown in Figure 7, only a significant linear correlation was found between the HCCscreen score and the methylation score (r2 = 0.27, P = .0001). No significant correlations were observed among the other markers, including between the HCCscreen score and AFP, the HCCscreen score and DCP, the methylation score and AFP, the methylation score and DCP, and AFP and DCP.

Correlation between individual biomarkers. The correlation between HCCscreen score and methylation score, between HCCscreen score and AFP, between HCCscreen score and DCP, between methylation score and AFP, between methylation score and DCP, between AFP and DCP is shown as indicated.
Discussion
Currently, the recommended methods for HCC screening in guidelines and consensuses are AFP combined with ultrasound.6,16 This method is convenient, noninvasive, inexpensive and accessible, making it the first choice for both general and high-risk populations. However, the sensitivity of AFP combined with ultrasound for HCC screening is not high. According to the results of a meta-analysis, its sensitivity for early HCC (tumor diameter <3 cm) is only 63%, 17 which means that more than one-third of HCC patients may be missed. Besides the issue of sensitivity, the awareness and frequency of screening in the population are also inadequate, leading to low compliance with screening. According to a retrospective study, the single screening compliance rate of AFP combined with ultrasound in high-risk liver cancer populations in China was only 46.87% (range 62.50% to 38.59%), 7 and in another study, only 7.3% of HBsAg-positive patients were able to complete six consecutive ultrasound screenings at six-month intervals. 18 Therefore, the overall screening performance of AFP combined with ultrasound is not ideal. Blood tests have higher compliance in cancer screening than invasive screening methods and other sample sources. 19 Traditional tumor blood tests mainly target various proteins, including Carcinoembryonic Antigen (CEA), AFP and carbohydrate antigen series, etc Among them, AFP is a relatively specific marker for HCC, but its sensitivity for HCC, especially for early stage HCC, is low, therefore, guidelines do not recommend AFP alone as a screening indicator. 6 DCP is also a relatively specific tumor marker for HCC, and its detection performance is comparable to that of AFP.20,21 Therefore, AFP and DCP are the most commonly used HCC-specific blood protein markers in clinical practice and are also the targets included in the HCCscreen test. In recent years, research on new blood markers for tumor screening has grown explosively, mainly including markers represented by methylation, hydroxymethylation, mutations, copy number variation (CNV), microRNA (miRNA), etc.22,23 These markers generally use plasma as the detection object and require the extraction of cfDNA from plasma. Due to the low content and easy degradation of cfDNA in plasma, developing highly sensitive markers for cfDNA and corresponding detection methods is the main challenge in tumor screening based on new blood markers. In addition, if multi-dimensional information in cfDNA can be detected and used in combination as markers for early tumor screening, the sensitivity of early screening can be improved. However, due to the generally low amount of cfDNA in a fixed volume of blood and large fluctuations among different individuals, as well as the limitations of detection methods, it is technically difficult to perform parallel detection of multiple markers.
In this study, the HCCscreen test has addressed the technical issue of parallel detection of multi-dimensional markers with a limited sample size. By using the HCCscreen test based on the MCP technology, we can perform multiple detections on a single cfDNA sample without sacrificing sensitivity, that is, simultaneously detecting methylation, mutations, CNVs, HBV integration, and proteins in one sample. This technology can amplify the information of genomic and methylation variations, supporting various downstream detections, including whole-genome level variations and methylation analysis, which can be used to develop new markers. The multiple parallel detections not only preserve the detection sensitivity of each omics but also improve the sensitivity due to the combined detection of multiple omics. 10 From the performance validation of the HCCscreen test in high-risk cirrhosis populations for HCC in this study, the sensitivity of the HCCscreen test is significantly higher than the existing screening method of AFP combined with ultrasound. HCCscreen appears to provide an effective detection method and strong evidence for the early screening and intervention of HCC. HCCscreen may also provide a new path and monitoring method for the early screening of HCC. Patients do not need to frequently undergo imaging monitoring but can achieve effective monitoring at a higher frequency through convenient blood sampling, meeting the monitoring frequency recommended by the guidelines. If positive changes are detected during monitoring, subsequent diagnosis and intervention can be carried out, greatly promoting the implementation of screening, the continuity of monitoring, and the targeting of diagnosis. It has the potential to become a new tool to replace AFP combined with ultrasound.24,25
Methylation analysis at the single-gene or combination and whole-genome levels has been widely applied in many scenarios such as cancer screening and therapeutic efficacy evaluation. Methylation detection based on PCR and next-generation sequencing (NGS) technologies has been used for cancer screening and early detection. Large-scale parallel methylation sequencing based on NGS technology for biomarker screening and early screening test development has been a research hotspot in recent years. Targeted sequencing of multiple genes or whole-genome epigenetic feature identification and subsequent modeling is the main strategy for detecting various types of cancers. The prototype method of targeted methylation sequencing was reported in 2009, and a method for specifically capturing genomic targets for single-molecule bisulfite sequencing was used for single-nucleotide resolution DNA methylation quantification detection. 26 In the field of early screening for HCC, using the above method, a study constructed an HCC diagnostic prediction model with a sensitivity of 85.7% and a specificity of 94.3% in the training dataset, and a sensitivity of 83.3% and a specificity of 90.5% in the validation dataset, 27 demonstrating the promising prospects of methylation sequencing in early screening for HCC. Subsequently, many studies have confirmed that detection based on multiple-gene methylation markers can be used for HCC screening.28–33 In this study, the methylation detection method we adopted is different from the above bisulfite sequencing-based methods. We used enzymatic digestion without altering the DNA sequence, effectively preserving methylation information, which can be used to simultaneously detect methylation and mutations, improving sample utilization efficiency and sequencing detection efficiency, and facilitating the development of multi-biomarker blood tests and continuous method optimization.
For an individual cancer patient, the higher the level of the target methylation markers in their plasma (ie, the higher the methylation fraction), the greater the likelihood of a positive test result. This is because a higher tumor DNA signal is more easily distinguishable from the background noise of normal cfDNA. In cancer patients, the average methylation fraction is usually positively correlated with the stage of cancer. Therefore, for a patient group of the same stage (such as stage I and II for early screening), the overall sensitivity (positive detection rate) of the detection method is directly related to the lowest methylation fraction that the method can stably detect (ie, the detection limit). Additionally, any diagnostic test requires setting a “threshold” to determine whether a result is positive or negative. This threshold typically corresponds to a specific methylation fraction. Lowering the threshold means that even weaker methylation signals are judged as positive, which usually increases the sensitivity (positive detection rate), but raising the threshold requires a stronger methylation signal to be judged as positive, which improves specificity and reduces false positives, but lowers sensitivity and may miss some early-stage patients. A single marker may have insufficient signal in early-stage cancer. By combining multiple markers and calculating a comprehensive methylation score or integrated interpretation score, even if the signal at a single site is weak, the accumulation of weak signals from multiple sites can improve the overall detection rate, thereby enhancing the positive detection rate at low methylation fractions. In summary, in tumor early screening, the methylation fraction in plasma is a key direct variable determining the positive detection rate (sensitivity). To achieve a high detection rate for low methylation fractions (corresponding to early-stage cancer) while maintaining high specificity, current research and product development mainly focus on screening high-specificity marker panels, developing ultra-sensitive detection technologies, and using machine learning algorithms to integrate multi-omics dimension information to optimize models, in order to break through the technical bottleneck of low ctDNA abundance in early-stage cancer.
In the field of epigenetics, in addition to methylation detection, hydroxymethylation and miRNA detection methods also show promising application prospects in HCC screening. Researchers have detected the whole-genome 5-hydroxymethylcytosine (5hmC) in cfDNA samples from a large cohort including 2554 Chinese subjects and established a diagnostic model for early HCC. A diagnostic model containing 32 genes could accurately distinguish early HCC (0/A stage) from non-HCC in both the training set and validation set (training set: AUC = 92.3%; sensitivity = 89.6%; specificity = 78.9%; validation set: AUC = 88.4%; sensitivity = 82.7%; specificity = 76.4%). 34 Similarly, detection methods based on multiple miRNA markers also show potential for early HCC screening.35–37
Compared with methylation-based blood tests, the detection of mutations in plasma has higher specificity. This is because methylation gradually occurs in a sequential manner during hepatitis, liver fibrosis, cirrhosis, precancerous diseases, and cancer stages, and DNA itself has a certain methylation background. Additionally, the methylation level is relatively high in the elderly.38,39 Therefore, changes in methylation marker levels are relatively sensitive to tumor occurrence, but the above reasons can lead to a higher false positive rate in non-cancerous benign diseases. Compared with methylation, the occurrence of mutations is a relatively late event in the process of tumor development, and mutations can generally be detected in the precancerous and early cancer stages. 40 Therefore, although the overall positive detection rate of mutations in blood is not as high as that of methylation, as a more specific marker, it can complement the insufficiency of methylation specificity and play a certain synergistic role in terms of sensitivity. In HCC, the mutation frequencies of TP53, TERT, and CTNNB1 genes are the highest, among which TERT has relatively liver specificity. If a mutation in the TERT promoter region is detected, it can accurately locate the liver.41,42 Therefore, in this study, HCCscreen detects genes with high mutation frequencies in HCC. Its algorithm takes into account both high sensitivity and high specificity, ensuring good positive and negative predictive values in population screening. Due to the very high negative predictive value of HCCscreen, it can be used to rule out potential HCC patients, playing an important role in alleviating patient anxiety and providing an additional reference dimension for patients who are difficult to diagnose with existing clinical methods.
Considering the distinct pathophysiological backgrounds of AFLD/MAFLD and viral hepatitis (HBV/HCV), differences in DNA methylation patterns, genetic mutations and protein markers may be expected among these population groups. From the perspective of methylation detection, the integration of HBV DNA into the host genome is a core feature for HBV-related liver cancer, which often involves genes related to the Wnt/β-catenin pathway and cell cycle regulation, such as RASSF1A, GSTP1, CDKN2A, etc In contrast, HCV usually does not involve viral DNA integration, and its methylation pattern is more driven by the background of chronic inflammation and liver cirrhosis, including hypermethylation of genes such as RASSF1A, APC, SFRP1, etc In AFLD and metabolic associated fatty liver disease (MAFLD), methylation changes of specific markers related to metabolic and insulin signaling pathways may be more prominent, such as PPARγ, SOCS1, etc In early screening tests, due to the particularly low sensitivity of AFP in NAFLD/AFLD-HCC, methylation markers are expected to become key supplementary or alternative tools. Methylation panels designed for metabolic pathways may be more effective for this type of HCC.43–45
From the perspective of gene mutations, gene mutations in ctDNA (such as TP53, CTNNB1, TERT promoter, etc) can also be used for early screening. In HBV-related liver cancer, the incidence of TERT promoter mutations and TP53 mutations is very high (especially in Chinese patients). This is the most prominent feature. In HCV-related liver cancer, CTNNB1 (β-catenin) mutations are relatively more common. The mutation rate of TP53 is usually lower than that of HBV-HCC. In AFLD/NAFLD-related liver cancer, CTNNB1 mutations and TERT promoter mutations are also common, but the mutation rate of TP53 is relatively low. In addition, mutations in genes related to metabolism and chromatin remodeling (such as ARID1A, ARID2) may be more common. In summary, regardless of the cause of liver cancer, the mutation spectrum is scattered, and the sensitivity of a single mutation marker may be limited, requiring a broader gene panel.46–48
From the perspective of proteins such as AFP and DCP, AFP has moderate sensitivity (about 60%-70%) in HBV-HCC, but its specificity is acceptable. In early screening, approximately 30%-40% of early-stage HCC patients do not show elevated AFP levels. HCV-HCC is similar to HBV-HCC in terms of sensitivity, which is also moderate. AFLD/NAFLD-HCC has the lowest sensitivity (possibly as low as 40%-50% or less), especially in early-stage HCC without a background of liver cirrhosis. Therefore, using AFP alone for early screening in the NAFLD/AFLD population is not very effective. The production of DCP is related to abnormal vitamin K metabolism and tumor angiogenesis. In HBV/HCV-HCC, DCP is more sensitive to advanced HCC and has limited sensitivity to early-stage HCC, but it is generally better than AFP, and has a complementary role to AFP. In AFLD/NAFLD-HCC, there is relatively less research data, but it is generally believed that its diagnostic performance in NAFLD-HCC is better than that of AFP. In the early screening of HCC, the combination of DCP and AFP can significantly improve the diagnostic performance of viral HCC. In the NAFLD population, incorporating DCP into models (such as the HES algorithm) also shows better results than AFP alone.44,49
Since most patients with HBV/HCV infection receive anti-viral therapy during HCC screening, the influence of anti-viral therapy on screening performance should be considered. The comprehensive impact of antiviral therapy on the performance of early screening mainly includes the following aspects. First, it can enhance specificity (reduce false positive rate), which is the most significant positive effect. Methylation, mutation, and protein markers all decrease non-cancer-related elevations/positives due to the reduction of the liver inflammation background caused by antiviral therapy, thereby more accurately pointing early screening positive results to true tumor risks. This is crucial for large-scale screening in high-risk populations and can avoid a large number of unnecessary panics and excessive examinations. Second, the impact of antiviral therapy on sensitivity is more complex. Its positive effect lies in the reduction of background noise, which helps detect weak true signals (especially for methylation and mutations). The potential challenge is that tumors may grow more slowly and be more “inert” (due to the anti-cancer effect of antiviral therapy), releasing fewer markers; the warning threshold of traditional protein markers (AFP) may need to be adjusted. Therefore, more sensitive detection techniques and multi-dimensional joint modeling are needed to ensure and even improve sensitivity. Third, antiviral therapy may change the PPV and NPV. PPV may increase: under the same sensitivity, the increase in specificity directly leads to an increase in PPV. This means that a positive result from a liquid biopsy is more likely to indicate the actual presence of liver cancer, and the clinical action value is greater. NPV may remain or slightly increase: the reliability of excluding liver cancer based on a negative result is very high, which is very important for alleviating patient anxiety.
This study still has some limitations and areas for improvement. Firstly, the development of the HCCscreen test was based on HBsAg-positive individuals, including low-, medium-, and high-risk groups for liver cancer. However, in this study, only high-risk and extremely high-risk patients were involved. Therefore, the HCCscreen test model still needs to be continuously optimized for this population. Secondly, the number of patients in this study, especially those with HCC, is relatively small. Further tracking of the test performance is needed in real-world studies with larger sample sizes. Thirdly, the performance of the HCCscreen test in non-HBV-related high-risk groups for HCC, such as those with hepatitis C, alcoholic liver disease, and nonalcoholic fatty liver disease, has not been systematically verified. Subsequent research needs to optimize the model to enable screening in these populations.
Conclusions
This study indicates that the novel multi-omics blood test technology, HCCscreen, demonstrates significant advantages over the current standard protocol (US + AFP) and single-omics biomarkers in screening for HCC in the liver cirrhosis population. It exhibits excellent overall detection performance, particularly with a significantly higher detection rate of early HCC compared to US + AFP. Additionally, HCCscreen shows stable performance across different demographic characteristics and has a high sensitivity and negative predictive value, which helps to reduce the risk of missed diagnosis. Therefore, as a noninvasive and efficient plasma multi-omics detection tool, HCCscreen is expected to become a better choice for early HCC screening in liver cirrhosis patients and has the potential to be widely applied in clinical practice.
Footnotes
Abbreviations
Acknowledgements
Not applicable
Ethics Approval and Consent to Participate
This study was approved by the Ethics Committee of The First People's Hospital of Yunnan Province, The Affiliated Hospital of Kunming University of Science and Technology (KHLL2021-KY108) before its commencement and was conducted in accordance with the hospital's guidelines for the management of clinical research. All patients signed an informed consent form before participating in this study.
Consent for Publication
All patients signed informed consent, which included the consent for publication.
Author's Contributions
All authors designed the study and were responsible for project management and implementation. All authors collected the data and performed the data analysis, initial interpretation, and the final statistics and made the figures and tables. All authors wrote the manuscript and proof read the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the General Surgical Key Clinical Specialty Department (Department of Hepatopancreatobiliary Surgery) of The First People's Hospital of Yunnan Province Platform Opening Project (No.2024GDKFKT-02).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of Data and Materials
The datasets generated and/or analyzed during the current study are included in the current manuscript.
