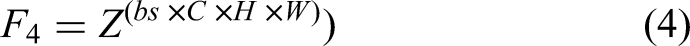Abstract
Background
By integrating Digitally Reconstructed Radiograph (DRR) images of pulmonary tumors with Electronic Portal Imaging Device (EPID) images to assist in target segmentation, and subsequently comparing morphological changes in segmented targets across different radiotherapy stages, this approach enables precise quantification of dynamic variations in target volume and shape. This methodological integration provides objective evidence for treatment response evaluation and dynamic optimization of treatment plans, thereby significantly enhancing the precision of radiotherapy delivery.
Methods
The proposed multimodal segmentation framework, named EPIDSeg-Net, comprises an encoder, a multi-scale feature layer, and a decoder. The encoder utilizes a dual-branch architecture: a CNN branch for extracting local texture features and a Swin-Transformer branch for capturing global semantic features. The model first calibrates multimodal input features through a Dual Attention Mechanism (DAM) to adaptively adjust modality-specific weights, thereby enhancing tolerance to missing image information in multi-sequence segmentation. Subsequently, two key modules are implemented within the multi-scale feature layer: a Large-Kernel Grouped Attention Gating (LKG-Gate) module to strengthen local contextual awareness, and a Multi-Path Feature Extraction (MPFE) module to improve feature robustness via a parallel structure. These designs enable the model to effectively focus on lung tumor target regions, optimize segmentation accuracy, and achieve high-performance reconstruction.
Results
The framework effectively integrates multimodal features, enabling high-precision localization and sharp boundary delineation while preserving anatomical details. Quantitative evaluations demonstrate superior performance: DICE = 93.2 (92.4∼93.9), CE = 0.352, HD95 = 9.42 (6.03∼12.8), IOU = 86.0 (84.1∼87.9), and SENCE = 0.828. Overall, the model excels at preserving gradient information, regional integrity, and fine details; effectively suppresses feature loss; and reduces missed segmentation rates, leading to improvements in both subjective and objective performance metrics.
Conclusion
The proposed segmentation method effectively integrates information from EPID and DRR images, enabling more precise localization and segmentation of lesion regions within EPID images while enhancing segmentation accuracy.
Keywords
Introduction
Primary lung cancer is a highly prevalent malignant tumor of the respiratory system, with typical symptoms including persistent cough and hemoptysis. 1 Radiotherapy serves as a cornerstone of its treatment, and post-radiotherapy imaging evaluation is crucial: quantifying morphological changes in the tumor (such as regression rate and residual volume) can assess therapeutic efficacy and predict survival, while early identification of resistance or recurrence can guide treatment modifications. Rapid imaging feedback is of significant value for dynamically adapting radiotherapy plans.
In precision radiotherapy practice, although EPID (Electronic Portal Imaging Device) imaging enables dynamic monitoring during treatment, its inherent low-contrast characteristics result in significantly limited soft tissue resolution—directly leading to core clinical issues such as blurring of tumor target boundaries and unclear organ-at-risk (OAR) contours. When anatomical displacement or morphological changes occur in the tumor during fractionated radiotherapy, single-modality EPID images often fail to provide reliable references for target delineation. This lack of reliable data undermines multi-phase treatment response assessment and adaptive radiotherapy planning, severely constraining personalized treatment decision-making. 2
To overcome this technical limitation, multimodal image fusion strategies demonstrate transformative potential: by integrating DRR (Digitally Reconstructed Radiograph)—a high-resolution “planning-level image”—a cross-modal anatomical structure mapping system can be constructed. DRR not only verifies setup errors and target coverage in the spatial domain but also, with its superior soft-tissue contrast, reveals biological tumor boundaries (such as infiltration fronts and micrometastases), providing essential anatomical references for EPID. 3 The deeper clinical value lies in Dynamic adaptation to treatment evolution: integrating DRR prior knowledge with real-time EPID images allows quantitative assessment of tumor regression throughout treatment fractions, facilitating a closed-loop adaptive radiotherapy process;Overcoming dose deposition blind spots: multimodal feature fusion can correct scatter-induced noise in EPID, reconstructing more accurate dose distribution maps and avoiding target under-dosage or OAR overdose;Enabling AI-driven decisions: constructing a dual-channel deep learning framework (“EPID-DRR”) enhances automatic segmentation robustness through cross-modal feature distillation, offering a new paradigm for precise “image–dose–response” triad analysis, ultimately advancing radiotherapy from a geometry-guided to a biology-guided era.
The diversity and complexity of medical imaging make multimodal fusion prone to issues such as data loss and nonlinear tissue contrast disparities. 4 Deep learning has partially alleviated these challenges. Mainstream multimodal fusion segmentation methods are based on CNNs and GANs. While CNNs exhibit strong feature extraction capabilities, they often lack global semantic understanding and long-range dependency capture. 5 GANs can optimize weighting through adversarial segmentation loss to improve segmentation quality, but training frequently encounters convergence difficulties and game imbalance issues.
To address these limitations, this study proposes a multimodal segmentation framework named EPIDSeg-Net, based on an early fusion strategy that combines CNN, Swin-Transformer,
6
and multi-scale feature extraction.
7
EPIDSeg-Net adopts an encoder–multi-scale feature layer–decoder architecture and takes EPID and DRR images as joint inputs, aiming to achieve accurate target segmentation on EPID images. Both the encoder and decoder integrate CNN (for multi-level feature extraction) and Swin-Transformer (to enhance global semantic modeling); the multi-scale feature layer incorporates cross-level integration to simultaneously capture fine details and high-level semantics. The core contributions of this paper are as follows:
A multimodal image segmentation framework named EPIDSeg-Net was developed based on an early-fusion strategy. It employs a CNN branch to extract local image features and incorporates Swin-Transformer to model long-range dependencies, achieving effective fusion of multimodal features. To address the issue of information attenuation during downsampling, two multi-scale feature modules—Large-Kernel Grouped Attention Gating (LKG-Gate) and Multi-Path Feature Extraction (MPFE)—were proposed. These modules enhance the joint representation of spatial details and high-level semantic information through cross-level feature integration, thereby improving segmentation accuracy. A segmentation enhancement strategy based on loss feedback was designed, which utilizes gradient information during training to strengthen the recognition of edge and detail features, further optimizing segmentation quality. For the first time, a deep learning-based method was introduced to achieve automated tumor target segmentation on full-field EPID images by incorporating DRR images. This approach provides critical data support for dynamic analysis of tumor morphological changes in patients.
Related Work
Multimodal Medical Image Processing
Multimodal medical image segmentation techniques can be broadly categorized into traditional methods and deep learning-based approaches. Among traditional methods, sparse representation-based techniques extract complementary information across modalities via linear combinations of a small set of basis vectors. These include joint sparse representation, 8 multi-task sparse representation, 9 dictionary learning, 10 and convolutional sparse representation.11,12 Transform-domain methods, on the other hand, extract salient features by projecting images into sparse domains, with representative techniques such as wavelet transform13,14 and pyramid transform.15,16. For instance, Liu G et al 17 proposed a brain tumor segmentation algorithm based on Dual-Tree Complex Wavelet Transform, which employed binary concepts to distinguish noisy regions. Singh V K et al 18 extracted frequency-domain features using Discrete Wavelet Transform and integrated a self-attention mechanism to enhance feature discrimination. Although effective in specific scenarios, traditional methods often suffer from limitations such as low computational efficiency, limited generalizability, and high complexity, which restrict their practical applicability.
Deep learning methods have demonstrated remarkable advantages due to their powerful feature extraction capabilities. Guo et al 19 proposed a cross-modal fusion supervision architecture integrating information at the feature, classifier, and decision levels, though complementary information fusion remained insufficient. Li et al 20 introduced a variational framework for PET/CT segmentation, combining a fuzzy variational model with the Split Bregman algorithm to achieve automated multimodal integration. Iqbal 21 enhanced UNet by incorporating feature fusion modules, attention gating mechanisms, and triple-threshold fuzzy enhancement to improve performance; however, challenges remain in segmenting tumors attached to normal tissues, handling inter-modal discrepancies, and mitigating the high cost of acquiring paired images. Li et al 22 proposed a framework consisting of a cross-modal synthesis network (CICVAE, trained with semantic cycle-consistency loss using anatomical images only to synthesize auxiliary modalities) and a segmentation network (Res-Unet) for lung tumor segmentation. To address challenges such as intensity inhomogeneity and blurred tumor boundaries, Cho et al 23 designed a tumor region attention module that combines the high sensitivity of PET with Squeeze-and-Excitation Normalization (SE Norm) for multi-scale context fusion. Li et al 24 introduced a novel grayscale similarity term leveraging PET background and CT foreground information, combined with a fuzzy C-means clustering (FCM) edge stopping function to enhance the localization of ambiguous boundaries.
Swin-Transformers in Medical Image
In 2021, Liu et al 24 introduced the Swin Transformer—a Vision Transformer architecture designed to mitigate domain shift challenges in visual transfer learning. Its key innovations encompass two core mechanisms: a shifted window scheme enabling localized self-attention computation within sliding windows, and cross-window connectivity to maintain global coherence. That same year, Hatamizadeh et al 25 developed Swin-UNET, which incorporates a Swin Transformer encoder with shifted window self-attention to extract hierarchical features across five resolution levels. This architecture integrates skip connections and a fully convolutional decoder to enhance segmentation precision. Wei et al 26 constructed an HRNet-based framework by substituting convolutional layers with Swin Transformer blocks, facilitating continuous multi-resolution feature exchange to produce high-fidelity representations. While CNNs exhibit limitations in capturing long-range dependencies, Transformers often demand substantial computational resources. To address these constraints, He et al 27 upgraded Swin-UNETR through convolutional module integration, yielding the enhanced SwinUNETR-V2 model. Upadhyay et al 28 presented MaS-TransUNet, which amalgamates multiple attention mechanisms—including self-, edge-, channel-, and feedback attention—while employing Swin Transformer blocks to construct hierarchical features, thereby substantially improving performance metrics. However, the fixed window masking in Swin Transformer inherently constrains very long-range pixel interactions. To circumvent this limitation, Fu et al 29 engineered the U-shaped SSTrans-Net for multi-organ segmentation. This network preserves effective long-range dependencies via target distribution channels, while simultaneously eliminating irrelevant dependencies in local context channels to optimize feature interaction dynamics.
Multi-Scale Feature Extraction
Multi-scale feature methods are extensively utilized in image processing tasks—including segmentation, detection, and classification—to effectively address the limitations of single-scale approaches in simultaneously capturing fine details and global contextual information. 30 Sinha et al 31 developed a guided self-attention mechanism that integrates local features with global dependencies, enabling adaptive emphasis on channel interdependencies and discriminative regions; however, its restricted capacity for capturing long-range dependencies led to suboptimal feature map reconstruction. Poudel et al 32 combined an EfficientNet encoder with a UNet decoder and incorporated an attention mechanism to suppress noise and irrelevant features, thereby reconstructing fine details with improved fidelity. Peng et al 33 introduced the MShNet model, which employs a dual-decoder architecture—one decoder for preliminary segmentation information and another for fused feature representation—to enhance feature learning. They also designed a fused convolutional pyramid pooling module to mitigate information loss during downsampling. Zhang et al 34 constructed the MS-Net (multi-scale feature pyramid fusion network), which extracts multi-level details through skip connections featuring dynamically adjustable receptive fields. To adaptively enhance target feature weights, they implemented a deeply nested multi-scale strategy alongside a multi-layer feature perception module. Zhao et al proposed a basic subtraction unit (SU) to generate inter-layer difference features and developed a multi-scale multi-subtraction network framework. This extends the SU into same-layer multi-scale SUs to provide both pixel-level and structural-level difference information. By deploying SUs in a pyramid configuration with varying receptive fields, the model achieves cross-layer multi-scale feature aggregation, capturing rich multi-scale differential information for enhanced representation learning.
Methods
This section provides a comprehensive overview of the proposed EPIDSeg-Net framework, detailing its network architecture, the technical specifications of core components—including the Swin-Transformer, MPFE module, DAM module, and LKG-Gate module—and systematically examining the design principles of the loss functions utilized during training. As demonstrated in Figure 1, the EPIDSeg-Net framework is structured across three principal components: an encoder for hierarchical feature extraction, a decoder for refined reconstruction, and a fusion layer for multimodal information integration.

The architecture of proposed method.
Network Architecture
To address the challenging clinical task of segmenting targets in low-quality EPID images, this study proposes a multimodal-assisted segmentation model named EPIDSeg-Net. Constructed on a U-shaped Swin-Transformer architecture, 11 the model employs multimodal training data denoted as M = {DRR, EPID}. As illustrated in Figure 1, the framework comprises three sequential stages: encoder, multi-scale feature extraction, and decoder.
Encoder Stage: Taking multimodal data M as input, the Dual Attention Module (DAM) first performs spatio-temporal feature extraction from each modality. These features are subsequently fused through a convolutional module, generating a unified feature map that is then input to the Swin-Transformer block for modeling long-range pixel dependencies.
Multi-scale Feature Extraction Stage: Leveraging skip connections, two key modules—the LKG-Gate and MPFE module—are introduced. The MPFE module employs parallel multi-scale feature extraction to mitigate information loss during downsampling, thereby preserving object details across multiple scales and providing foundational feature support for the decoder. The LKG-Gate integrates large-kernel convolution for global context capture, grouped attention for computational efficiency, and a gating mechanism for dynamic feature refinement, facilitating deep integration and interaction of multi-scale features.
Decoder Stage: Utilizing upsampling modules, the decoder progressively aggregates multi-scale features received via skip connections. When combined with encoder information, the feature maps are eventually restored to the original input resolution, enabling pixel-level prediction of target regions.
The pursuit of architectural innovation must often be reconciled with practical computational constraints. To address this challenge, we strategically reduced the input resolution from the conventional 512 × 512 to 128 × 128. This resolution reduction yields over an order of magnitude decrease in computational and memory overhead, enabling efficient prototype development, hyperparameter tuning, and comprehensive ablation studies under limited hardware resources—a crucial advantage for exploratory research. Furthermore, the adopted network architecture inherently compensates for low-resolution inputs. Its core modules are specifically designed to extract robust semantic features: the Swin Transformer employs window-based multi-head self-attention to model global and local contextual dependencies while reducing computational complexity, thereby partially mitigating detail loss from downsampling. The dynamic attention mechanism adaptively enhances feature responses in critical regions, directing the network's focus toward task-relevant anatomical structures. Multimodal and multi-scale feature fusion mechanisms enable cross-hierarchical information integration, leveraging inherent multi-scale learning to improve robustness to input resolution variations. Finally, targeted designs ensure boundary segmentation accuracy. To counteract potential impacts of reduced resolution on boundary details, a dedicated joint loss function incorporating the L_FTLoss term explicitly enhances the model's sensitivity to boundary pixels, directly optimizing boundary segmentation quality during training. Consequently, the 128 × 128 resolution represents a thoroughly validated balance between computational cost and model performance, with subsequent experimental results confirming that this strategy achieves the intended objectives in both overall segmentation accuracy and boundary metrics.
Encoder Stage
During the encoder stage, the multimodal input is defined as 1. In the initial stage of multimodal fusion, the Dual Attention Module (DAM) achieves multi-level dynamic adjustment of feature representation by constructing a two-dimensional attention mechanism (as shown in Figure 2). Its core architecture consists of a collaborative channel attention branch and spatial attention branch. Max Pooling and Average Pooling are used for feature map downsampling—Max Pooling selects local area maximum values to retain salient features, while Average Pooling computes local area averages to capture global information. Their combination effectively enhances multi-scale information capture capability
36
;7 × 7 convolution kernel extracts local features from input feature maps and generates higher-order features. After processing with the Sigmoid activation function, it generates probability values to further regulate spatial and channel attention weights
37
;The Spatial Attention Mechanism (SAM) assigns pixel-level weights to focus on high-information regions, improving local detail capture and suitability for complex-structured image processing
38
; the Channel Attention Mechanism (CAM) assigns channel-level weights to identify key feature channels, enhancing accuracy with notable results in medical image segmentation
39
;The Shared Multilayer Perceptron (MLP) reduces model complexity through parameter sharing, enhancing nonlinear relationship modeling. Combining convolutional features with global MLP representations improves medical image segmentation performance.
40
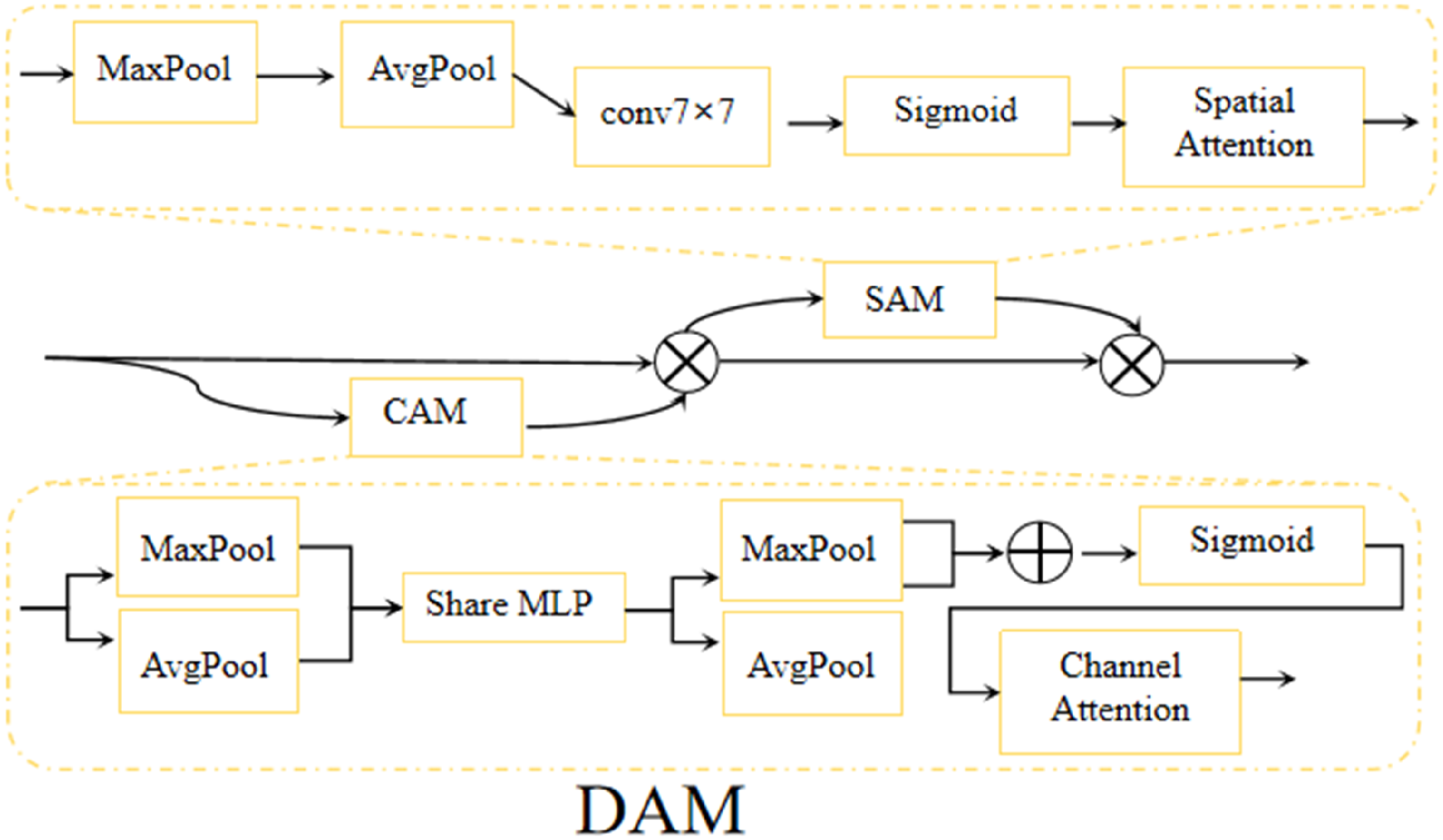
DAM block.
Furthermore, through end-to-end learning combined with appropriate loss functions and regularization techniques, dynamic learning strategies and hybrid attention mechanisms have been widely applied in medical image segmentation, significantly improving model segmentation accuracy.
41
This dual-modal attention fusion strategy and component combination overcome the limitations of traditional convolutional networks in non-selective extraction of spatial and channel features, enabling adaptive focus on discriminative local features and global semantic information.
2. The image patching operation partitions the high-resolution feature map into non-overlapping patches to enable subsequent linear embedding and Transformer-based processing. This module is implemented through a 4 × 4 convolutional layer integrated with transpose and flatten operations, adhering to a transpose → flatten → conv structural sequence. The resulting feature representation after this processing step is formally denoted as 3. In contrast to the conventional multi-head self-attention (MSA) mechanism, the Swin Transformer block introduces an innovative architecture grounded in shifted windows.
42
As depicted in Figure 3, the core processing unit comprises two consecutive Swin Transformer blocks. Each Transformer block adheres to a standardized architecture: it includes a LayerNorm (LN) layer, a multi-head self-attention module, residual connections, and a two-layer feed-forward network (MLP) with GELU non-linear activation. Notably, these two consecutive blocks implement window-based multi-head self-attention (W-MSA) and shifted-window-based multi-head self-attention (SW-MSA) respectively. Through the offset of windows between adjacent blocks, this shifting strategy facilitates dynamic cross-window information interaction.

Swin transformer block.
The input image is initially partitioned into four non-overlapping sub-regions, followed by spatial concatenation through a patch merging layer. This operation reduces the spatial resolution of the feature map to half its original dimensions (ie, 2×downsampling) while simultaneously expanding the feature dimensionality to four times the original input due to channel-wise concatenation. To harmonize the feature dimensions, a linear transformation layer is applied to the concatenated features, compressing the channel count to twice the original dimensionality. This process achieves a balanced adjustment between spatial resolution reduction and channel depth expansion.
Multi-Scale Extraction Stage
To address limitations of traditional networks—including semantic gaps, loss of fine details, and low computational efficiency—this study introduces a multi-scale feature fusion mechanism based on skip connections. This mechanism effectively captures multi-scale feature distributions, integrates information, and preserves the original characteristics of input data. The proposed approach is implemented through two key modules: MPFE and LKG-Gate. The former enhances feature fusion capability while expanding the receptive field; the latter dynamically adjusts feature representation. Together, they synergistically optimize multi-scale feature transmission to the decoder without introducing additional computational overhead in subsequent stages. The detailed structure is illustrated in Figure 4.
1. The MPFE module consists of four parallel branches. The first branch is composed of three cascaded 3 × 3 convolution layers; the second branch adopts a structure of 1 × 1 convolution layer followed by 3 × 3 convolution layer - this configuration is widely regarded as the standard way for local feature extraction. The convolution kernel effectively captures local and global information through stacking layers, while maintaining higher computational efficiency compared to larger convolution kernels with fewer parameters
43
; the third branch uses a single 1 × 1 convolution layer; the fourth branch retains the original input features through identity connection. After each convolution operation, a BN + ReLU activation unit is followed. The outputs of all branches are concatenated and processed through 1 × 1 convolution to generate the final feature representation. Residual connections are introduced by adding the output to the original input elements, thereby preserving global context information. The multi-scale convolution kernel enables multi-scale fusion and hierarchical feature extraction with deep abstraction of deep features, as 3 × 3 convolution kernels can effectively capture local-global information while reducing parameters
43
; while 1 × 1 convolution kernels help in channel-level information integration and dimension reduction, enabling the retention of key features while reducing computational costs.
44
Batch normalization (BN) alleviates internal covariate shift through input standardization, accelerates training convergence, and improves stability.
45
The ReLU activation function prevents gradient vanishing through negative value truncation, significantly improving the training efficiency of deep networks.
46
The multi-scale feature fusion module improves the effect of medical image segmentation through dense connections of multi-scale feature layers,
47
and channel-level weighted summation enhances the discriminative focus on significant features,
48
this comprehensive method improves model performance in complex tasks. The specific structure is shown in Figure 4(a), and its operation can be formally defined as follows: The (a) in the figure represents MPFE, and (b) represents the LKG-gate module.

In Equations (1)-(5), Z represents the input feature tensor,
In order to verify the impact of the parallel branch selection in MPFE on the model performance, this study also conducted branch ablation experiments on MPFE to validate the scientificity of the branch selection. As shown in Table 1.
MPFE Branch Ablation.

When all three branches (Top, Mid, and Under) are included, the model achieves the optimal comprehensive performance. Its DICE coefficient is 93.2%, IOU is 85.9%, and HD95 is 10.3. This result indicates that the parallel branch structure we designed can effectively capture multi-scale features ranging from deep semantics to shallow details, and its collaborative effect is the key to improving the segmentation accuracy.
2. The LKG-Gate module comprises two parallel 3 × 3 convolutional branches, each followed by a batch normalization (BN) operation. The outputs are merged and processed through a 1 × 1 convolution, then passed through BN and a Sigmoid activation function to generate adaptive weight coefficients. Leveraging the Sigmoid gating mechanism, this module dynamically adjusts feature weights—enhancing key features while suppressing redundant information. By integrating 3 × 3 convolution kernels, 1 × 1 convolution, batch normalization, and Sigmoid activation, the LKG-Gate module effectively extracts local and global image features while achieving robust feature fusion.
35
This design improves segmentation accuracy, particularly in complex tasks such as medical image analysis, enabling precise identification and segmentation of critical regions.43–45 The selective feature fusion capability emphasizes extraction of salient features, optimizing discriminative power for enhanced model performance. The formal definition of its operation is as follows:


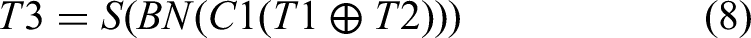

In Equations (6)-(9),
Decoding Stage
During the decoder stage, we first concatenate the outputs of two fusion blocks— U and 
Where
In constructing a decoder branch symmetric to the encoder architecture, this study utilizes a combination of dual Swin-Transformer blocks and a Patch Expanding (PE) module as the fundamental building block. This design progressively restores spatial resolution through a feature expansion mechanism. Specifically, the decoder pathway incorporates a feature reorganization module that integrates multimodal skip-connected features to reconstruct spatial dimensions, thereby transforming low-resolution feature maps into high-resolution representations. Concurrently, a channel compression strategy is applied to reduce the number of feature channels to half of the input dimensions. This mechanism functionally complements the encoder's feature aggregation module: while the encoder compresses spatial dimensions and expands channel capacity via downsampling, the decoder restores detailed information and optimizes feature dimensionality through upsampling. Together, they constitute an end-to-end feature transformation pipeline, ultimately achieving complete spatial resolution recovery and generating a pixel-wise segmentation probability map.
Loss Function
The performance and robustness of deep learning models are critical metrics for evaluating their effectiveness. In medical image segmentation tasks, the cross-entropy loss function is widely adopted as a baseline method. However, the standard cross-entropy loss alone often proves insufficient to address class imbalance issues. To mitigate this impact, this study incorporates the Dice loss function, which quantifies the overlap between predictions and ground truth annotations. Furthermore, to simultaneously enhance segmentation accuracy and preserve boundary details, the Focal Tversky Loss is introduced. Ultimately, a composite loss function is constructed to optimize model training performance. The multi-task loss function used in the end-to-end training process can be mathematically expressed as follows: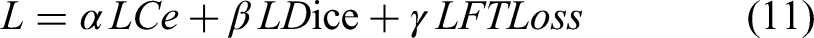



In this image segmentation task, the Focal Tversky Loss (FTLoss) is typically selected to address class imbalance issues. To demonstrate the advantages of optimal focal parameter selection within this loss function, ablation experiments were performed using grouping strategies proposed in 49 to verify the impact of different parameter combinations on segmentation performance, 50 as shown in Table 2. Following the approach of Abraham N et al, 49 when adopting the parameter set (q = 0.3, n = 0.7, m = 2), the model demonstrates enhanced focus on challenging regions in medical images—particularly in scenarios where foreground-background class imbalance is severe. This parameter configuration improves segmentation accuracy for target areas, 51 especially in tasks involving small lesion detection or complex regional segmentation, while simultaneously enhancing precision in EPID image segmentation.
FTLoss Focus Parameter Ablation Results.

To investigate the impact of weight variations, seven comparative configurations were designed, including:Component Dominance Experiment: Successively setting α, β, and γ as dominant weights to evaluate individual component influence on overall performance. Weight Variant Experiment: Testing cases with equal weights, moderate weights, and low-weight extremes to explore boundary effects of weight balancing. Gender Abandonment Experiment (as detailed in Table 3). Analysis of Table 3 reveals critical insights: excessively high weights (eg, “γ Dominance” or “Low-weight extreme”) significantly degrade model performance. Specifically, the Dice coefficient drops to approximately 89% while the boundary error HD95 deteriorates to over 26 mm. This demonstrates that overemphasizing boundary penalties disrupts gradient balance in segmentation optimization, causing the model to sacrifice overall accuracy and consistency in pursuit of perfect boundaries. Under “α Dominance” and “β Dominance” configurations, the model maintains high Dice coefficients (>92%), confirming the robustness of cross-entropy and Dice loss as segmentation foundations. However, compared to the baseline, these dominant configurations fail to improve performance and exhibit varying degrees of HD95 degradation. This indicates that relying solely on regional optimization losses is suboptimal.
Loss Weight Abandonment Experiment.

Comprehensive comparison across all configurations confirms that the baseline weight allocation (α = 0.5, β = 0.3, γ = 0.2) achieves optimal overall performance: it balances the Dice coefficient (93.2%) and boundary precision HD95 (9.42 mm). This success stems from its balanced strategy—higher weights for LCE and LDICE ensure precise capture of overall segmentation regions, while a moderate LFT boundary penalty refines edge details without disrupting global optimization. The configuration effectively integrates advantages of different loss terms, enabling the model to achieve optimal balance between segmentation accuracy and boundary quality.
In Equations (11) to (14)Our baseline model adopted the weight configuration of the hyperparameters are set as α = 0.5, β = 0.3, and γ = 0.2. Here, pi∈ [0,1] denotes the predicted probability of pixel i belonging to the foreground (class 1), and ti ∈{0,1} represents the ground truth label of pixel i (where 1 indicates foreground and 0 background). The hyperparameters q = 0.3 and n = 0.7 control the relative weighting of false positives (FP) and false negatives (FN), respectively. The focal parameter m adjusts the loss contribution of easy versus hard examples. A smoothing term ɛ = 1 is added to prevent division by zero. Finally, TP, FP, and FN denote the number of true positive, false positive, and false negative pixels, respectively.
Experiments and Results
Dataset and Evaluation Metrics
DRR-EPID-Seg Dataset
The dataset utilized in this study is an internal private dataset from the Radiotherapy Center of the Third Affiliated Hospital of Nanjing Medical University (Changzhou Second People's Hospital), with detailed characteristics presented in Table 4. The dataset comprises 350 paired samples, each containing EPID images, corresponding DRR images, and expert-annotated gold standard segmentation labels. These samples are categorized by tumor stage (I: 63 cases, II: 98, III: 87, IV: 105) and difficulty level (low: 152, medium: 113, high: 85). All images maintain a resolution of 1 × 512 × 512 pixels. EPID images were acquired using the Siemens Promise system under standardized positioning verification with beam parameters of 6 MV and 2 MU. DRR images were generated by the same treatment planning system from planning CT data using acquisition parameters of 120 kVp, 150 mAs, and L/H ratio 1199/2398. To protect patient privacy, all data underwent strict anonymization—retaining only essential imaging and geometric information while preserving texture and edge details.To ensure a clean and reliably labeled training cohort, rigorous inclusion/exclusion criteria were established:
Inclusion criteria: (1) EPID and DRR pairs with complete geometric consistency; (2) GTV contours validated by consensus of two senior radiotherapy physicians. Exclusion criteria: (1) Cases with severe motion/metal artifacts or reconstruction anomalies obscuring tumor boundaries; (2) Cases with complex postoperative cavities, multiple synchronous primaries, or irreproducible blurred boundaries; (3) Non-standard beam parameters (eg, non-6 MV beams, non-2 MU settings).
Parameters and Equipment Indicators of the DRR-EPID-Seg Dataset.

Challenging but interpretable cases (eg, varying tumor sizes/locations/complex shapes) were retained to reflect clinical practice, while only “unreadable” cases lacking expert consensus were excluded to prevent gold standard contamination.
Evaluation indicators
In our study, the Dice Similarity Coefficient (DICE) was employed as the primary evaluation metric for segmentation performance. The DICE quantifies the spatial overlap between the predicted segmentation and the ground truth, with higher values indicating superior segmentation accuracy. Mathematically, the DICE is defined as:
where P represents the predicted segmentation mask, T denotes the ground truth annotation, and ∣⋅∣ signifies the cardinality of the set. The DICE ranges from 0 (complete dissimilarity) to 1 (perfect overlap), providing a standardized measure for comparing segmentation results across different methodologies.
Data Pre-Processing
To ensure multimodal data reliability and consistency, this study implemented a comprehensive preprocessing pipeline encompassing physical calibration, intensity normalization, and spatial registration.
First, patient CT data was calibrated in Hounsfield units to ensure accurate physical intensity representation. Based on calibrated CT, orthogonal DRR images (0° anterior-posterior and 90° lateral views) were generated. Physical consistency between DRR electronic density and actual x-ray attenuation was verified to guarantee cross-modal physical property alignment.
To address contrast/noise disparities between DRR and EPID imaging devices, inter-modal histogram matching was applied. This remapped target modal grayscale distributions to the standard [0,1] interval, significantly reducing grayscale differences from imaging principles and providing standardized model input.
For precise anatomical alignment, a hybrid strategy combining non-rigid registration algorithms with SIFT-based anatomical landmark detection registered DRR and EPID images. This ensured high-precision cross-modal anatomical alignment, forming the foundation for subsequent feature comparison and fusion.
Pipeline effectiveness was verified through dual subjective-objective validation: clinical physicians conducted subjective quality assessments to ensure clinical compliance, while objective metrics like Hausdorff distance quantified registration errors and segmentation mask accuracy, strictly controlled within clinically acceptable limits.
Implementation Details
The EPIDSeg-Net model was implemented using the PyTorch framework and deployed on an NVIDIA A100 80GB GPU. Its backbone architecture utilized Swin-UNet. During training, the AdamW optimizer was employed for parameter optimization with an initial learning rate of 0.001. Input data was formatted as a single-channel tensor (1 × 512 × 512), while DRR images, EPID images, and corresponding label images maintained a resolution of 1 × 128 × 128. The batch size was set to 1, and the model underwent 200 training epochs.
The dataset was partitioned into training (270 samples), validation (40 samples), and test (40 samples) sets with a 270:40:40 ratio. To enhance model generalization, data augmentation techniques—including random rotation, cropping, and horizontal flipping 52 —were applied during training. Additionally, five-fold cross-validation was implemented to rigorously evaluate generalization performance and minimize randomness associated with single data partitions, with results presented in Table 5.
Table of Results from Five-Fold Cross-Test.
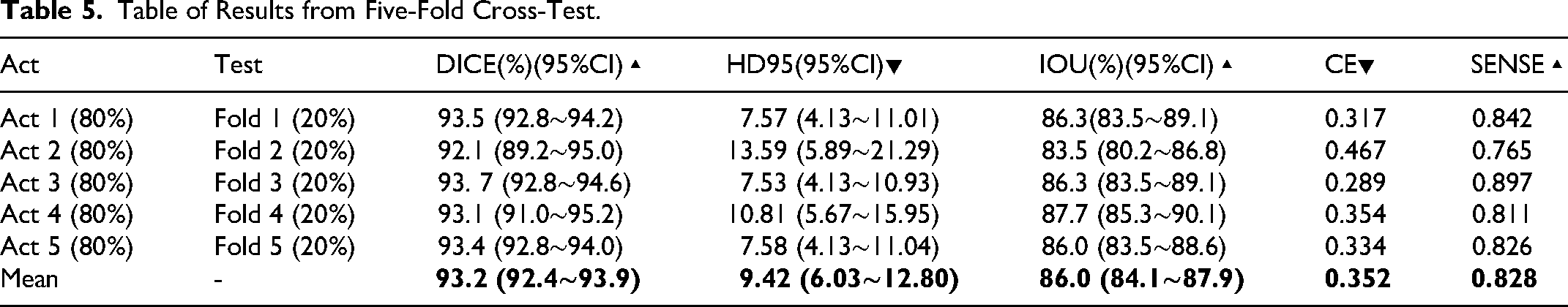

As shown in Table 5, through the five-fold cross-validation experiment, it can be effectively demonstrated that this model performs exceptionally well and stably in the core segmentation task, reaching a level that can be put into practical application or undergo further clinical verification. Furthermore, the consistency of the experimental results demonstrates that the model does not suffer from severe overfitting and can generalize well to unseen data.
To comprehensively evaluate the generalization capability and robustness of the proposed model across varying clinical stages and anatomical complexities, stratified experimental analyses were conducted based on tumor stages (I-IV) and segmentation difficulty levels (low, medium, high). Detailed quantitative results of these group-specific evaluations are presented in Table 6.
Group Experiment.
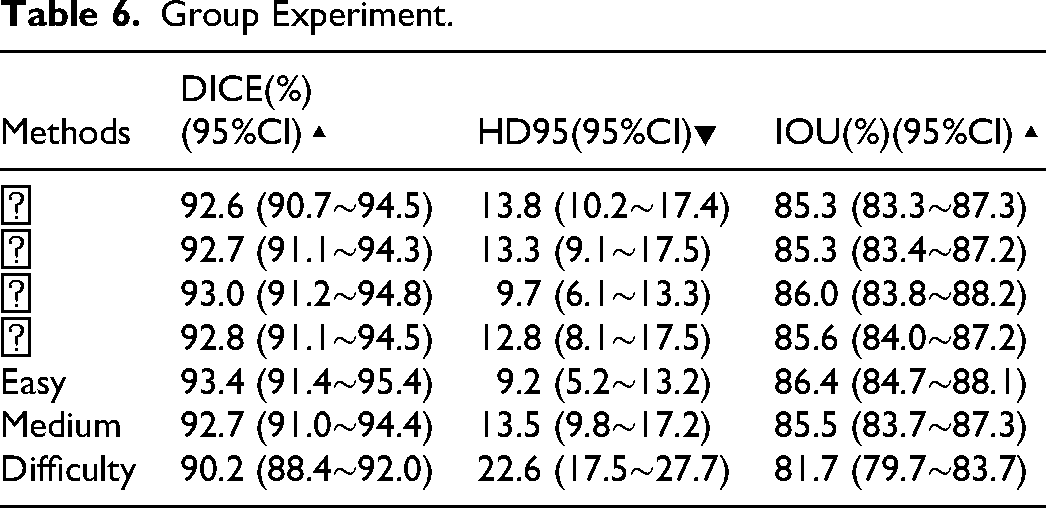
The analysis confirms that the proposed model maintains consistent superiority across all clinical stages and difficulty levels. For tumor stages I-IV, segmentation accuracy remained stable with DICE coefficients ranging narrowly from 92.6% to 93.0% and IOUs consistently above 85.3%. Notably, while DICE values showed minimal variation across stages, the model achieved optimal boundary precision for Stage III cases, evidenced by the lowest HD95 value (9.7) compared to other stages—demonstrating robust generalization across disease progression levels.
Performance boundaries were further delineated through difficulty-stratified analysis. In low-difficulty cases, the model attained peak performance (DICE: 93.4%, HD95: 9.2). Medium-difficulty cases exhibited only slight degradation (DICE: 92.7%), maintaining high accuracy. However, high-difficulty cases showed more pronounced metric declines—particularly in boundary error (HD95: 22.6)—consistent with clinical expectations. This trend validates the correlation between evaluation metrics and practical segmentation challenges, while identifying clear avenues for model enhancement in extremely complex scenarios.
Contrast Experiment
This section validates EPIDSeg-Net effectiveness through comparative experiments on the DRR-EPID-Seg dataset. Experimental setup: Mainstream segmentation models (Hiformer, 53 EANet, 54 TransUnet, 55 U-Net, 56 U-Net++, 57 U-Net3+ 58 ) served as benchmarks, trained under near-identical hyperparameter configurations with results reproduced via official codebases.
Comprehensive evaluations (Table 7) confirm our method achieves optimal performance across all core metrics, significantly outperforming all baselines. Specifically: DICE = 93.2%, IOU = 86.0%, HD95 = 9.42 (lowest), CE = 0.352 (lowest), SENSE = 0.828 (highest). These results robustly validate the superiority of our architectural design.Compared to state-of-the-art Transformer-based models (eg, TransUnet), our method shows clear advantages: DICE improved by 0.6% while HD95 reduced by nearly 4 units (13.4→9.42), demonstrating superior integration of global context and local details. Even against the closest competitor Hiformer, consistent improvements in DICE/HD95/CE validate synergistic enhancements from innovative modules (MPFE/LKG-Gate/DAM). Traditional U-Net variants (U-Net++/U-Net3+) improved baseline U-Net through dense connectivity but still lag behind Transformer-based architectures and our method. This underscores the necessity of advanced feature extraction/fusion mechanisms for complex segmentation tasks, aligning with our design philosophy.The visualized segmentation results of each comparison model are shown in Figure 5 and Figure 6.

A qualitative comparison of segmentation results across four representative samples is presented. The first column shows paired DRR-EPID input data, followed by seven columns displaying results from each comparative method. Visual analysis reveals that, unlike EPIDSeg-Net, baseline methods exhibit significant over-segmentation or under-segmentation in target regions. Color coding clarifies: navy blue indicates model predictions, crimson denotes ground truth targets, and purple highlights overlaps. Green contours outline segmentation results, while red contours mark gold standard boundaries.

Segmentation mask comparison.
Comparative Results of Different Methods on the DRR-EPID-Seg Dataset.

The training curve (Figure 7(c)) clearly illustrates the model's convergence behavior and performance evolution over 200 epochs. As shown, the loss value decreases rapidly with training epochs and gradually stabilizes, while the segmentation accuracy metrics (DICE and IOU) demonstrate a concurrent significant increase, eventually converging to high plateaus. These divergent yet complementary trajectories indicate a stable and effective optimization process. Specifically, after approximately 60 epochs, the improvement rates of all metrics noticeably decelerate, suggesting the model is approaching its performance ceiling. Ultimately, both DICE and IOU reach and maintain high levels, demonstrating the model's exceptional segmentation capability and robustness. The entire training process exhibits no significant oscillations, confirming appropriate learning rate configuration and optimization strategy without overfitting.

(a) shows the line graph of various indicators in the comparative test, (b) shows the line graph of HD95 in the comparative test, and (c) shows the trend graph of changes during the training process of EPIDSeg-net.
Ablation Experiment
To systematically evaluate the contribution of each proposed module, a rigorous stepwise ablation study was conducted (Table 8). The experimental trajectory clearly demonstrates a dual-dimensional improvement in segmentation accuracy and model efficiency from the baseline to the full model. For segmentation accuracy, all core evaluation metrics exhibit monotonic increases as modules are progressively integrated: DICE coefficient steadily rises from 91.2% to 93.2% (+2.0%), while HD95 decreases from 15.9 to 9.42 (−40.8%), validating enhanced overall segmentation consistency and boundary handling capability. The continuous decline in cross-entropy loss (CE) and steady increase in sensitivity (SENSE) further confirm the model's improved recognition capability.
Experiment of Sequential Ablation of key Components.

Detailed module contribution analysis reveals: FTLoss serves as the initial performance booster, improving DICE by 0.4% with minimal cost (0.01 M parameters/0.01G FLOPs increase), effectively addressing class imbalance. The LKG-Gate module achieves a significant +0.6% DICE improvement with efficient parameter utilization, demonstrating its effectiveness in feature integration and filtering through gating mechanisms. The MPFE module, despite adding 2.07 M parameters/1.45G FLOPs, contributes 0.3% DICE improvement and 0.5 HD95 reduction, highlighting its core value in multi-scale parallel feature extraction for rich contextual capture. As the final enhancement component, the DAM module plays a pivotal role in boundary refinement, achieving the largest single-step HD95 reduction (12.8→9.42). Although it introduces 3.13 M parameters/13.3G FLOPs, the accuracy and boundary quality improvements justify this cost.
Regarding efficiency, while the full model exhibits increased parameters, computational load (FLOPs), and inference time per image compared to the baseline, this trade-off is reasonable and efficient. FTLoss and LKG-Gate provide substantial initial performance gains with negligible cost, while MPFE and DAM, as the primary computational contributors, strategically allocate resources to address multi-scale perception and detail reconstruction challenges. Ultimately, the model achieves an excellent balance between accuracy and efficiency.
This ablation study robustly demonstrates the necessity and unique contributions of each proposed module. The complete architecture forms a logically coherent and progressively optimized system: FTLoss and LKG-Gate establish an efficient foundation for feature optimization, while MPFE and DAM build upon this to deliver powerful feature extraction and detail recovery capabilities. All modules work synergistically to achieve state-of-the-art segmentation performance.The visualization effect is shown in Figure 8.
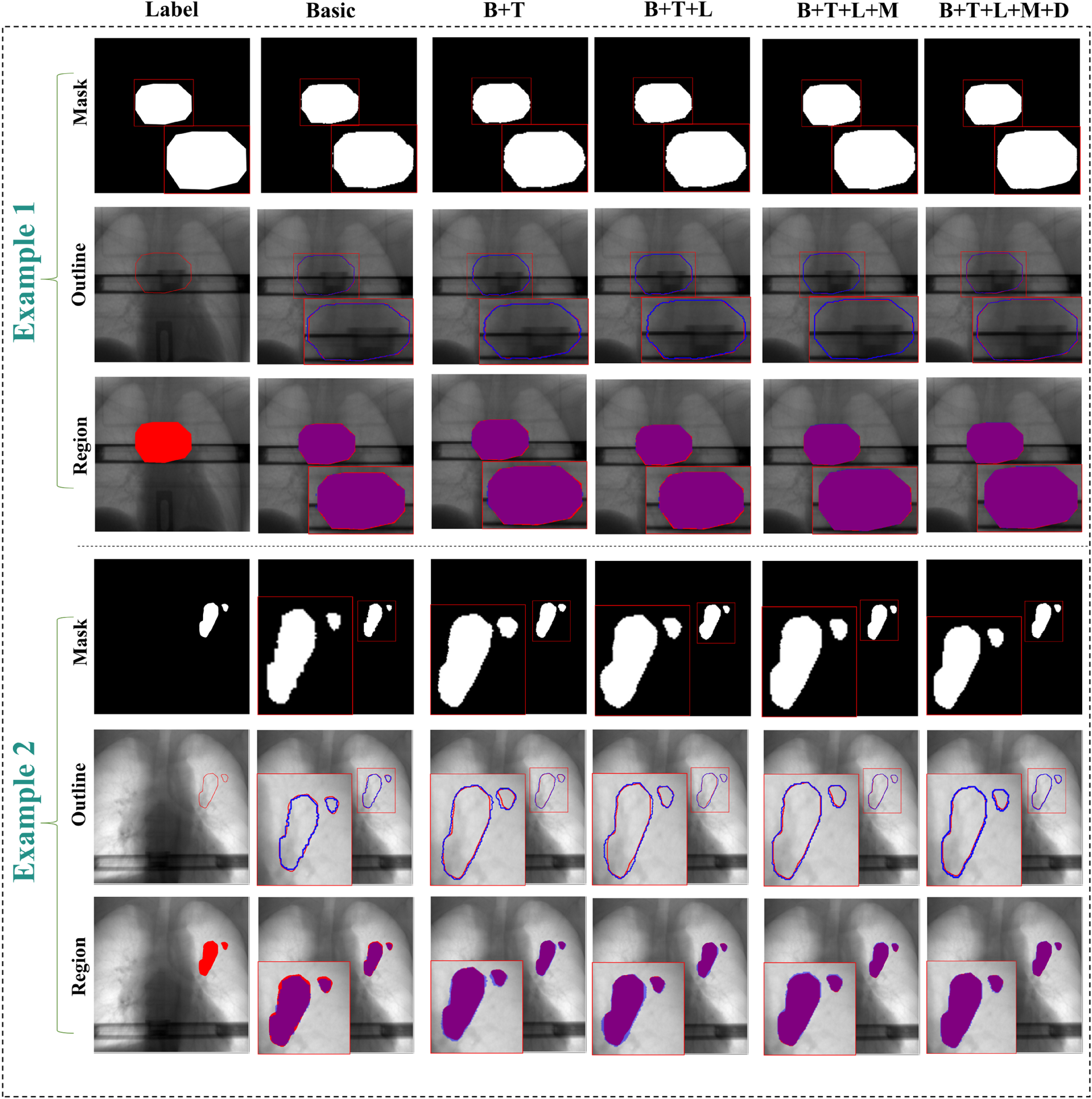
The results of the module-by-module ablation experiments. The figure visually demonstrates the ablation results through two representative cases. Each case pair presents comparative results in three distinct visualization formats (individual segmentation masks, overlaid edge contours, and overall region overlays). In these visualizations, the ground truth is indicated in red, the segmentation results in blue, while the overlapping regions in the overall region overlay are displayed in purple.
To rigorously evaluate the independent contributions of each component to model performance, ablation experiments were designed and conducted (Table 9). The results reveal distinct impacts of various modules on performance metrics. Regarding segmentation accuracy, the MPFE module exhibited the most critical improvement: when added to the baseline, it achieved a DICE coefficient of 92.0% and sensitivity (SENSE) of 0.816—the highest values among all single-module addition strategies. The LKG-Gate and DAM modules also contributed varying degrees of performance gains. However, the standalone integration of FTLoss failed to enhance the baseline model effectively, with both DICE and SENSE metrics showing deterioration. This suggests that the theoretical advantages of this loss function may require high-quality features from other modules to manifest fully.
Key Component Single Ablation Experiment.

In terms of model efficiency, each module introduction led to a controlled increase in computational overhead. Notably, the DAM module showed the most substantial growth in parameter count and GPU memory usage, while the LKG-Gate module achieved effective performance improvements while nearly maintaining computational load (FLOPs), demonstrating favorable efficiency characteristics.
These experiments confirm the effectiveness of MPFE, LKG-Gate, and DAM as standalone components while revealing potential dependencies and the necessity of synergistic operation for FTLoss in practical applications. These findings provide clear guidance for optimized design of the overall model architecture, emphasizing the importance of module interaction and balanced resource allocation.
Integration into the Clinical Workflow
Operational Integration and Validation of EPIDSeg-Net:Specifically, EPIDSeg-Net analyzes corresponding EPID images acquired during each radiotherapy session, automatically segments the target volume, and compares it with the phase-matched digitally reconstructed radiograph (DRR). This enables precise quantification of dynamic changes in target volume and shape. The process not only provides objective evidence of morphological changes but also supports treatment response assessment. The feature optimization capabilities and automated segmentation algorithm of EPIDSeg-Net effectively reduce latency and facilitate dynamic optimization of pre-treatment plans, thereby enhancing radiotherapy precision.
Latency and Acceptance Criteria:Integration of EPIDSeg-Net into the intra-treatment workflow significantly reduces latency associated with manual segmentation or processing. By automating target segmentation, it minimizes the time traditionally required for manual verification and image analysis, enabling faster adaptation and optimization of treatment plans. Acceptance criteria should encompass real-time performance, accuracy, and consistency metrics. The system must complete target segmentation within a defined time window and align with clinical requirements to ensure both immediate response and precision in radiotherapy delivery.
Human-in-the-Loop Quality Assurance and Prospective Validation:Human-in-the-loop quality assurance (QA) and prospective validation are critical during the integration of EPIDSeg-Net. Human QA ensures the alignment of automated segmentation with clinical standards, for instance, by verifying consistency between automatically segmented targets and those manually delineated by physicians. Prospective validation involves extensive testing across diverse patients and clinical scenarios to verify the stability and accuracy of EPIDSeg-Net under various treatment settings. Furthermore, human intervention can correct potential system errors and provide clinical validation, ensuring that EPIDSeg-Net effectively and reliably supports the development and implementation of individualized treatment strategies during dynamic adjustment of radiotherapy plans.
Discuss
EPIDSeg-Net establishes a precise radiotherapy support system that integrates automated efficiency with clinical reliability to ensure segmentation outcomes meet clinical real-time decision-making requirements. However, this study was conducted under controlled single-center, single-vendor, and fixed-energy conditions. Caution should therefore be exercised when extrapolating these findings to other devices, beam energies, or more complex motion scenarios. Evaluating cross-vendor and cross-energy generalization capabilities, along with performance validation in complex motion scenarios, will be critical directions for subsequent multi-center or prospective EPIDSeg-Net research. Furthermore, systematic exploration of integrating EPIDSeg-Net with advanced motion management frameworks represents a meaningful future research direction. This prudent and well-defined approach—compared to potentially insufficient analyses under current data constraints—more effectively ensures the reliability and reproducibility of our conclusions while providing a clear development path for future work. Subsequent efforts will focus on incorporating multi-institutional data and lightweight model designs to reduce dependency on auxiliary modalities while maintaining accuracy, extending the framework to achieve high-precision target localization, and investigating optimized multi-modal fusion strategies.
Conclusions
This study proposes a novel multi-modality aided segmentation network, named EPIDSeg-Net, for segmenting treatment targets from low-quality EPID images. The model integrates the strengths of CNN and Swin Transformer architectures to extract local image features effectively. Furthermore, it incorporates specifically designed modules—DAM, MPFE, and LKG-Gate—that enable sequential feature transformation, achieving multi-modal feature fusion, multi-scalef feature extraction, and large-field convolution optimization. These components collectively facilitate the modeling of long-range dependencies across modalities and the learning of multi-scale feature representations. Ablation studies confirm the effectiveness of each key component, and evaluations on the DRR-EPID-Seg dataset demonstrate superior performance over existing segmentation methods. Future work will focus on further optimizing the multi-modal segmentation framework to enhance its clinical applicability and reduce target segmentation response time.
Footnotes
Acknowledgements
Not applicable.
Ethics approval and consent to participate
All experiments involved in this study were conducted in strict accordance with relevant guidelines and regulations, and all study participants were informed and consented to the study. All the private data is studied retrospectively in this research approved by Changzhou NO.2 People's Hospital (MR-32-23-007615).
Consent for Publication
Not applicable.
Authors’ Contributions
Hqj conducts research projects, designs models for experiments, collects data, and drafts the initial version. Zh and Slt performs paper revision and provides innovative materials. Nxy and Jzq offers guidance on research directions, handles fund applications, and conducts the final review of papers.
Funding
This work is supported by the National NaturalScience Foundation of China (No. 62371243), National Key Research and Development Program of China (No. 2025YFC2427600), Jiangsu Provincial Medical Key Discipline Cultivation Unit of Oncology Therapeutics (Radiotherapy)(No. JSDW202237), Changzhou Social Development Program (No.CJ20244020).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of Data and Materials
Due to ethical and legal restrictions, the data supporting this study cannot be made publicly available. The dataset contains sensitive information related to Personal privacy, which is protected under ethical guidelines.