Abstract
Introduction
Rapid magnetic resonance imaging (MRI) plays an increasingly important role in radiotherapy. It improves the accuracy of delineation of target volumes and organs at risk (OARs), but generating accurate synthetic CT(sCT) from MRI remains challenging, and the lack of electron density information limits its further clinical application. Therefore, the purpose of this study was to develop and evaluate a CBAMPix2Pix model for MRI to CT synthesis.
Methods
We adapted the CBAMPix2Pix architecture, incorporating convolutional block attention module(CBAM), to synthesize CT images from 17260 MRI slices of 86 patients with metastatic brain cancer. The model analyzes local features to enhance image authenticity and is designed to map T1-weighted Contrast-enhanced (T1wc) to sCT. To address the data imbalance between normal tissue and bone, we introduce structural similarity loss(SSIM) to enhance local features of learning images, thereby better reducing differences in Hounsfield Unit(HU).
Results
We evaluate the performance of the model through quantitative and qualitative evaluations. Our proposed model achieves higher peak signal-to-noise ratio (PSNR) of 27.5 ± 3.3 dB, normalized mean absolute error (NMAE) of 0.019 ± 0.023, and structural similarity index (SSIM) of 0.857 ± 0.059 for sCT images in MR simulation sequences, and the average mean absolute error(MAE) was 74.48 ± 22.88 HU in body and 185.89 ± 21.59 HU in bone. The
Conclusion
We have developed a novel CBAMPix2Pix model that can effectively generate realistic sCT images comparable to real images, potentially improving the accuracy of MRI-based treatment planning.
Introduction
Magnetic Resonance imaging (MRI) is extensively employed in radiotherapy due to its superior soft-tissue contrast compared to Computed Tomography (CT) imaging. This enables more accurate delineation of tumor targets and organs at risk.1–4 The development of a novel radiotherapy modality based solely on MRI simulation, namely MR-only radiotherapy, is emerging as a prevalent research focus. 5 In head-and-neck cancer radiation treatment, some studies have shown that using MRI images can greatly reduce inter-observer variability in tumor delineation and improve treatment outcomes.6–11 This is attributed to the elimination of additional CT imaging requirements, thereby reducing patients’ radiation exposure. 12 But to implement MR-only radiotherapy, we need to derive synthetic CT(sCT) images from MR images to obtain electron densities for radiation dose calculation. Unlike CT, MR signals usually depend only on hydrogen atoms, not the entire material composition. So, MRI alone can't be directly linked to the key physical properties needed for dose calculations, like electron density in photon therapy. Moreover, MR images are often fused with simulated CT through inter-modality registration, and physicians delineate contours based on the fused MR view. However, this approach is likely to have systematic errors, which can cause geometrical uncertainties for the contours.13,14
The primary challenge in MR-only workflows lies in obtaining electron density data from MRI for radiation dose calculations. Unlike CT numbers, which can be directly changed into electron density, MRI pixel values only show the magnetic relaxation time of tissues and aren't directly related to electron density. However, tissue relaxation time can first be converted into CT numbers (Hounsfield Unit, HU) and then into electron density. 15 To solve this problem, many ways to get sCT images have been suggested. These are statistical modeling, 16 traditional machine learning,17,18 and multi-atlas based methods.19–23 Lately, there's been more interest in artificial intelligence-based methods, especially deep convolutional neural networks (CNN). With deep learning using CNN, the model can automatically extract features at different scales and combine them into an end-to-end network for prediction. This lessens the need for manual feature extraction.24–30 The performance of these methods can be affected by the accuracy of MR-CT registration, and even tiny voxel-wise misalignments may lead to blurring of synthesized images. To create a complex and nonlinear mapping mechanism between MR and CT image domains, self-learning and self-optimizing strategies can be used. Once the best Deep learning(DL) parameters are determined, sCT images can be easily obtained in a few seconds by putting new MR images into the trained model. This approach allows for efficient and accurate extraction of electron density information from MRI, thereby overcoming the limitations of single-modality imaging in clinical radiotherapy implementation and enabling more precise radiation dose calculations. Recently, increasing interest focused on generative adversarial network (GAN),31–33 and its variants,34–36 have shown promise in synthesizing high-quality sCT images with reduced blurriness compared to conventional CNN approaches.
In this study, we are dedicated to generating sCT images from MR images to optimize MR-based radiotherapy workflows. We propose a novel deep learning network called CBAMPix2Pix. This network introduces a convolutional block attention module(CBAM) based on the Pix2Pix framework to enhance the feature representation capabilities of the deep network and extract more effective features. Additionally, we employ a structural similarity(SSIM) loss function to ensure geometric consistency between MR and sCT images. This allows for a more comprehensive evaluation of image characteristics and provides guidance for optimizing radiotherapy plans and treatment effects.
Materials and Methods
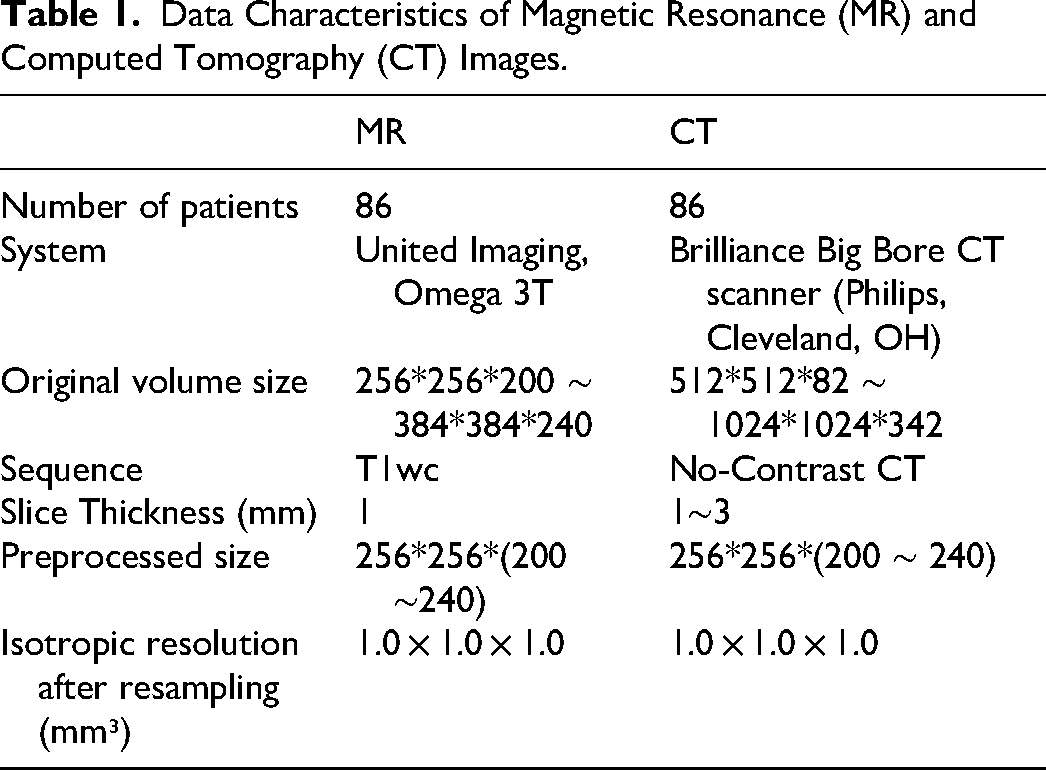
Table 1 summarizes the characteristics of the MR and CT datasets used in this study. A total of 86 brain cancer patients treated at the authors’ institutions between January 2023 and March 2025 were enrolled: 61 for model training, 10 for validation, and 15 for testing. The data split was performed at the patient level to ensure independence and prevent information leakage. Specifically, all MRI and CT slices from a given patient were assigned exclusively to one of the training, validation, or test sets. All MR and CT scans were acquired using identical patient positioning and thermoplastic masks on a dedicated radiotherapy simulation scanner. CT images were rigidly aligned to MRI and resampled to 1 mm³ isotropic resolution, MRI was deformably registered to CT using ANTsPy (v0.0.7) with the SyN algorithm. The field of view was cropped to 256 × 256 × (160–240) voxels. CT Hounsfield Units (HU) were clipped to [−1000, 3071], and both MR and deformed CT (dCT) underwent histogram matching and intensity scaling to [−1, 1]. MRI preprocessing included N4 bias field correction and histogram equalization. MRI preprocessing included: (1) N4 bias field correction 37 to address non-uniformity; (2) histogram equalization.
Data Characteristics of Magnetic Resonance (MR) and Computed Tomography (CT) Images.

All models (CycleGAN, Pix2Pix, CBAMPix2Pix) were implemented in PyTorch (v1.12.0) and trained on an RTX 2080 GPU (12 GB). A batch size of 1 was used with Adam optimizer (β₁=0.5, β₂=0.999) and initial learning rate of 1e−4 for both generator and discriminator. Training ran for up to 40 epochs, with early stopping based on validation loss, the best model was selected by validation performance.
Models
GAN have shown significant potential in image-to-image translation tasks, particularly in domain adaptation within medical image analysis. They have driven advancements in segmentation, cross-modal synthesis, anatomical registration, and radiation treatment planning, including dose calculations. However, their clinical application is hindered by challenges such as model instability from adversarial training, limited generalization across diverse imaging modalities, and reliance on high-quality paired datasets. Cycle-consistency models like CycleGAN, 38 can process misaligned images but may produce multiple solutions, making them less suitable for high-precision medical image translation. Pix2Pix, 39 a conditional GAN (cGAN) model, comprises a generator with residual blocks for capturing context and multi-scale features, and a discriminator for assessing image authenticity. While effective in many tasks, it struggles with complex scenes and fine details.
To enhance the classic Pix2Pix model, we introduced the CBAM to create the CBAMPix2Pix architecture, thereby boosting the model's feature capturing and representing abilities. The overall architecture is illustrated in Figure 1. CBAM, a lightweight attention module, combines channel and spatial attention mechanisms. It first employs channel attention to learn channel importance and then feeds the output into the spatial attention module to learn spatial position weights. This process adaptively recalibrates input feature maps, enabling the model to focus on key channels and spatial positions. This allows the network to better capture important features and improves its performance in image-to-image translation tasks. The generator in the original Pix2Pix model uses residual blocks to capture image context and multi-scale features, while the discriminator assesses the authenticity of generated images. However, Pix2Pix has limitations in handling complex scenes and fine-grained details. By incorporating CBAM into the Pix2Pix framework, our CBAMPix2Pix model addresses these limitations and achieves better results in generating synthetic CT images from MRI data.
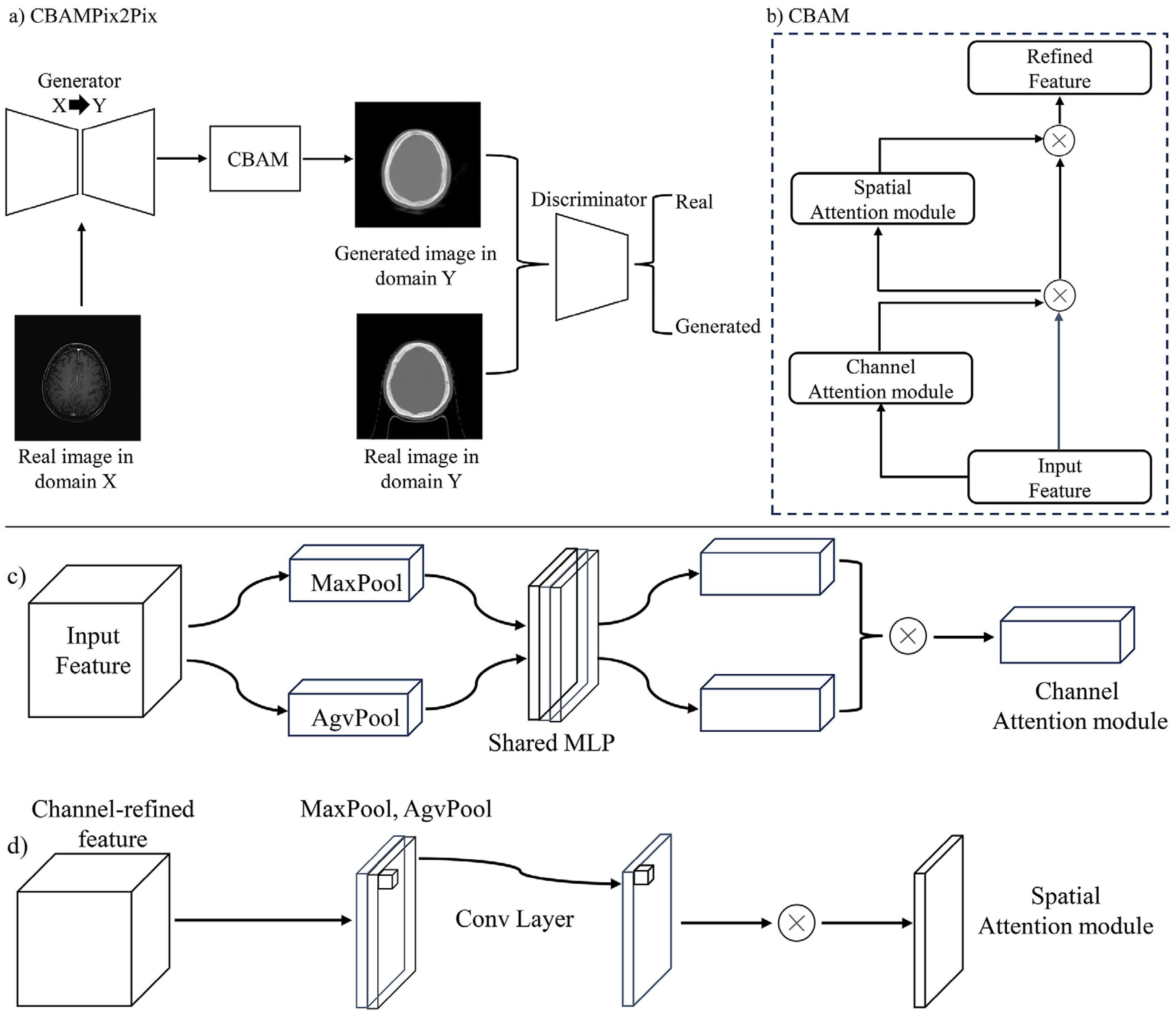
(a) Is the overall architecture of CBAMPix2Pix; (b), (c), and (d) represent the operation flowcharts of the channel and spatial attention mechanisms, respectively.
Figure 2 presents the architecture of the modified generator utilized in this study. In the CBAMPix2Pix generator, a CBAM was incorporated before the residual blocks to enhance feature extraction. The feature map first undergoes convolution to extract preliminary features, then enters the CBAM module. Within CBAM, the channel attention module employs global average and max pooling alongside a shared multilayer perceptron (MLP) to generate a channel attention map. This map weights the feature map's channels, amplifying important ones and suppressing less relevant ones. Subsequently, the weighted feature map enters the spatial attention module, which calculates the importance of each spatial position to produce a spatial attention map. This further emphasizes key spatial regions by weighting the spatial positions of the feature map. After undergoing CBAM processing, the feature map proceeds to the up-sampling stage. This design enables the generator to more accurately capture image details and semantics, resulting in higher-quality image generation. It enhances feature capture, focuses on key image regions, reduces noise and irrelevant features, and boosts the model's expressiveness. This allows it to better preserve image details and textures during generation, resulting in more realistic and accurate images.
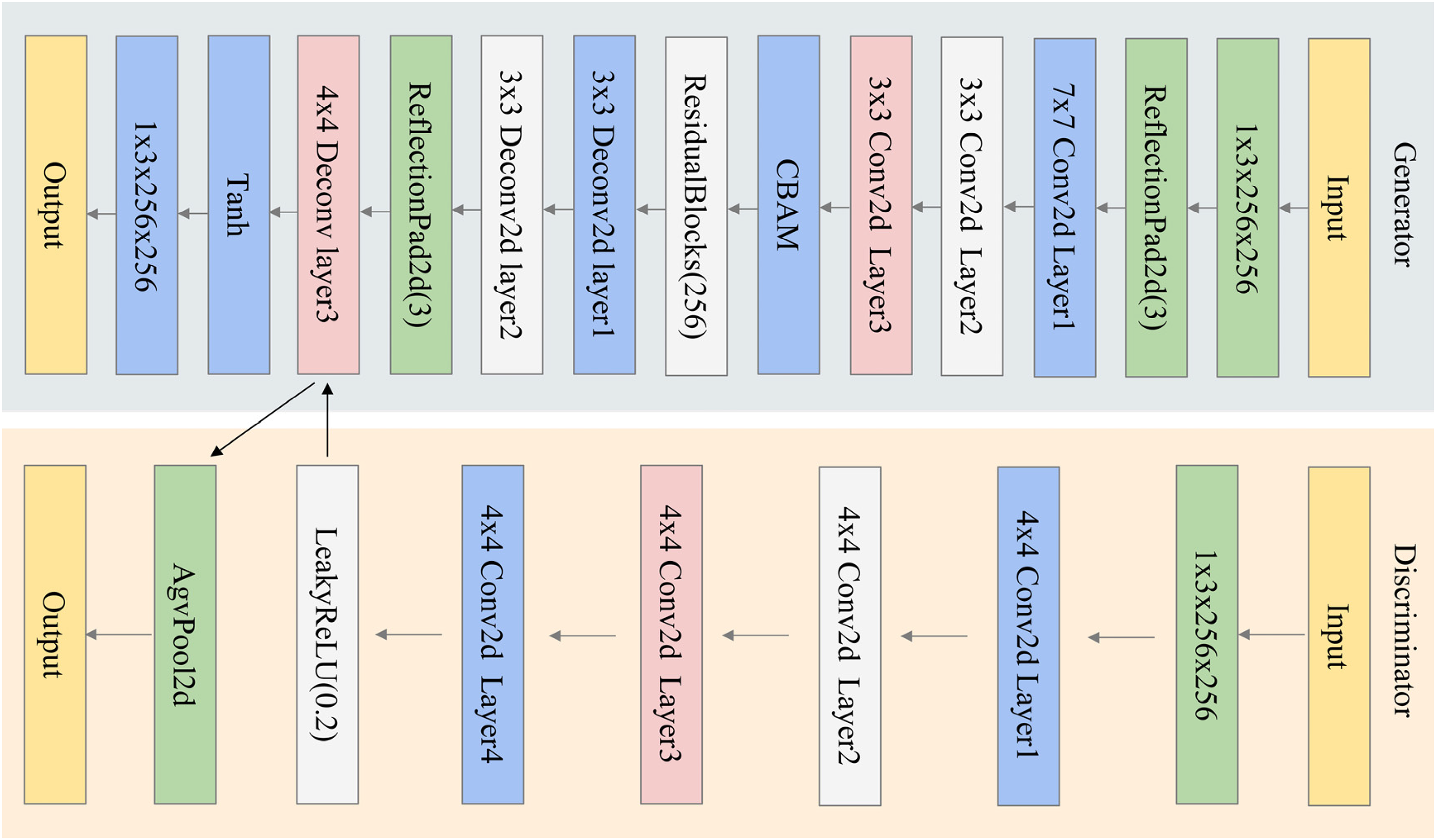
Illustration of CBAMPix2Pix. The model begins with a convolutional layer for input image processing, followed by CBAM.
Loss Functions
The proposed model employs a multi-loss(L1 loss, adversarial loss and structural similarity loss) framework to optimize CBAMPix2Pix architecture. The primary loss components include 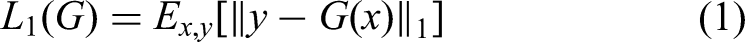
The adversarial loss(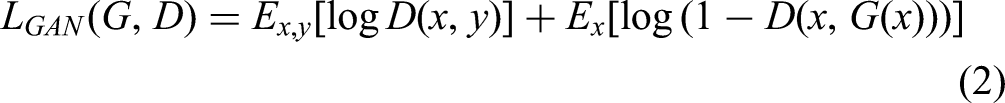
Equation (2) is the adversarial loss function for Pix2Pix, where
In order to effectively measure the similarity of the regional contours of MRI and CT images, a structural similarity(SSIM) loss calculation method was proposed. As show as Equation (3):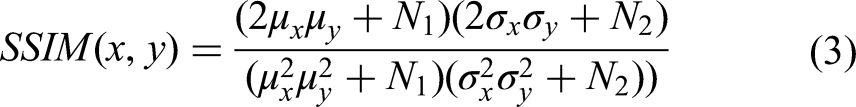
In Equation (3),
The Total loss is as follows Equation (4):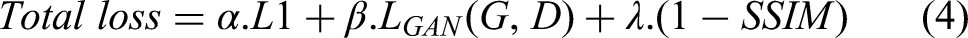
The hyperparameters
Model Evaluation
Model performance was quantitatively evaluated using four metrics: Peak Signal-to-Noise Ratio (PSNR), Structural Similarity Index (SSIM), Mean Absolute Error (MAE), and Normalized Mean Absolute Error (NMAE). The metrics (NMAE, PSNR, SSIM) were computed on a per-2D-slice basis across the entire test set, and then averaged across all slices.
PSNR is an important indicator for evaluating image or signal quality. It is mainly used to measure the similarity between the generated image and the real image, and is especially widely used in the field of image compression and reconstruction. Higher PSNR values indicate greater similarity to the original image, with the formula defined as: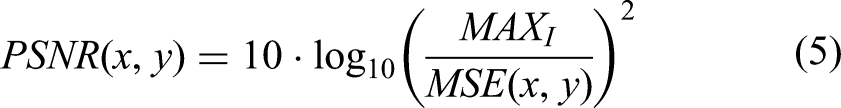
In Equation (5),
SSIM is an indicator used to measure the similarity between two images, mainly from three aspects: brightness, contrast and structure. Higher SSIM values indicate better perceived quality. The SSIM formula is as shown in Equation (3);
MAE is a widely-used metric for assessing image quality, quantifying the discrepancy between a generated image and its corresponding real image. It is calculated by taking the absolute differences between the true pixel values and the predicted pixel values, summing these differences, and then averaging them. A lower MAE value indicates a higher degree of similarity between the generated and real images, thereby reflecting the superior performance of the image generation model, The NMAE is computed after mapping model outputs and ground truth from the [−1, 1] preprocessing range to [0, 1]. Thus, NMAE ∈ [0, 1], where 0 represents perfect agreement and 1 indicates maximal deviation: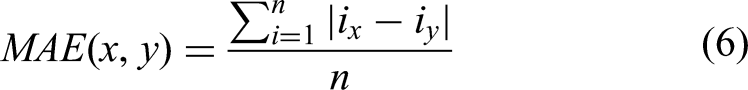
Where
Statistical Analysis
All statistical analyses were performed using Python (version 3.10.13; https:// www. python. org/) and SPSS Statistics 27. The differences between the sCT images and the dCT images, and between models were all assessed using the Wilcoxon signed-rank test, as the comparisons were performed on paired observations.
Results
Quantitative and Qualitative Evaluation
The three models were trained in 36 (CycleGAN), 16 (Pix2Pix), and 17(CBAMPix2Pix) hours, respectively. Once the models have been trained, it took a few seconds to generate sCT images from new MR volume image. Figure 3 shows the visual comparison at different anatomical locations, which contains the axial view of the original MR, the corresponding dCT, and sCT images generated from CycleGAN, the Pix2Pix, the proposed CBAMPix2Pix. In the visual inspection, the generated results of our method are more consistent with the dCT compared with other two methods in terms of global and local imaging regions. As can be seen in Figure 3, the sCT generated using CBAMPix2Pix showed sharper boundaries than the sCT generated using the original CycleGAN and Pix2Pix.
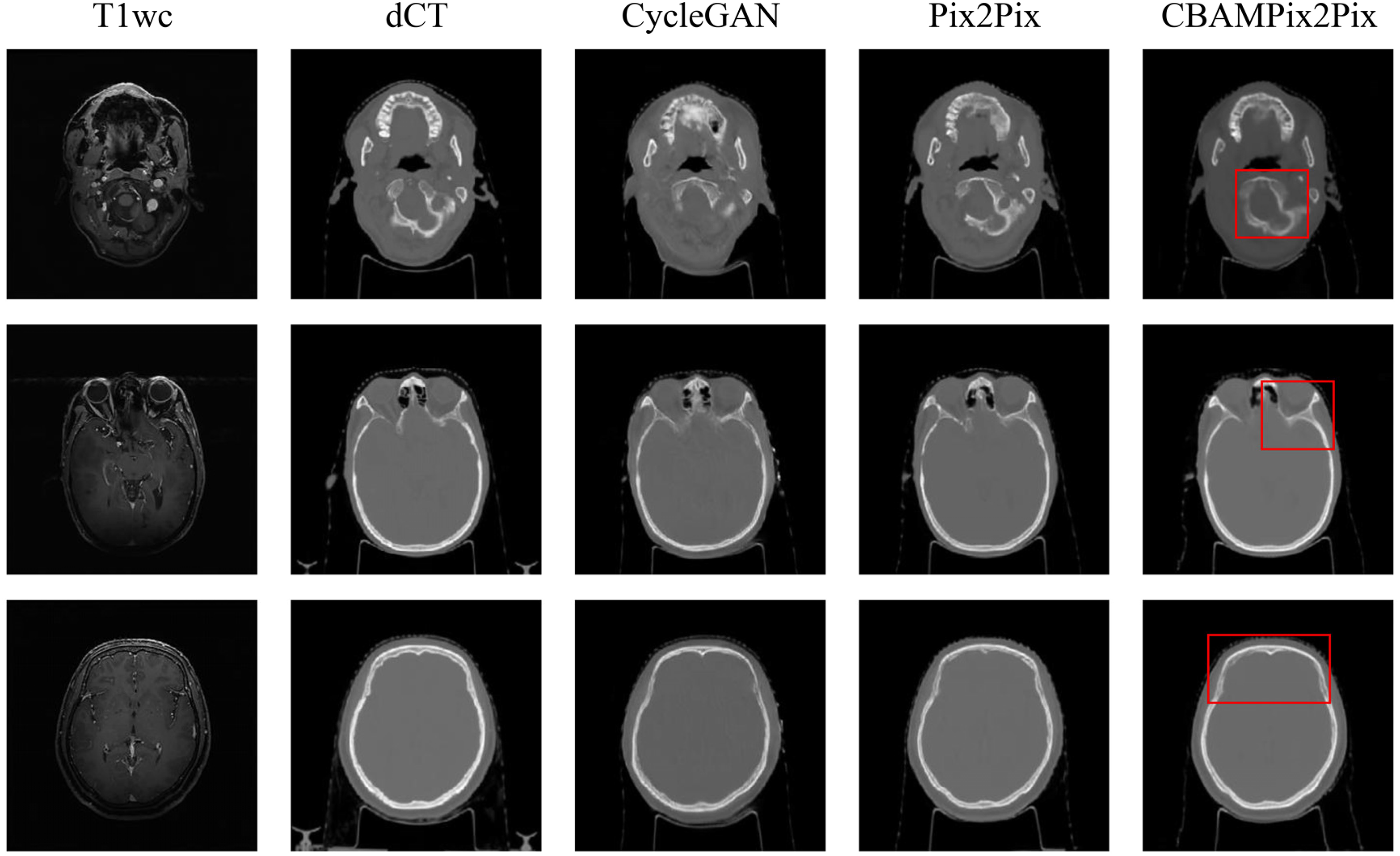
Comparison of deformed CT(dCT) and synthetic CT(sCT) images generated by different models. The first and second columns show MR images and dCT. And the third, fourth columns, and the fifth column respectively display sCT images generated by the CycleGAN model, the Pix2Pix, and CBAMPix2Pix. Red boxes highlight anatomical regions where CBAMPix2Pix shows the most significant improvement.
Table 2 summarizes the quantitative results of NMAE, SSIM, and PSNR for sCT images of all test patients relative to reference CT images. It shows that CBAMPix2Pix generated sCT images have smaller errors (NMAE) and higher similarity (PSNR and SSIM) to dCT than those from CycleGAN and Pix2Pix. For CycleGAN, Pix2Pix, and CBAMPix2Pix, the average ± SD of NMAE between in T1wc images are 0.023 ± 0.022, 0.020 ± 0.023, and 0.019 ± 0.023; the average ± SD of PSNR are 25.4 ± 2.4 dB, 27.1 ± 3.0 dB, and 27.5 ± 3.3 dB; the average ± SD of SSIM are 0.818 ± 0.061, 0.843 ± 0.067, and 0.857 ± 0.059, respectively. CBAMPix2Pix outperforms traditional CycleGAN in synthesis quality. In addition to slight improvements in NMAE (0.019), PSNR (27.5 dB), and SSIM (0.857) for T1wc images, our model demonstrated minimal but consistent enhancements across all structures.
Errors (NMAE) and Similarities (PSNR, SSIM) Relative Between sCT and dCT in the Test Group.
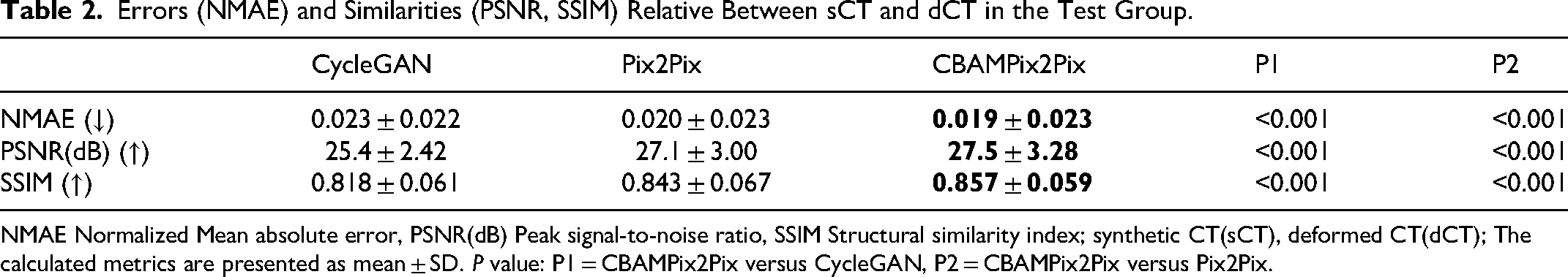

NMAE Normalized Mean absolute error, PSNR(dB) Peak signal-to-noise ratio, SSIM Structural similarity index; synthetic CT(sCT), deformed CT(dCT); The calculated metrics are presented as mean ± SD.
The sCT images generated by CBAMPix2Pix from T1wc MR images show smaller differences in Hounsfield Unit (HU) values compared to dCT. As indicated in Table 3, the MAE is 74.48 ± 22.88 HU for body and 185.89 ± 21.59 HU for bones, significantly lower than those of CycleGAN and Pix2Pix. These results highlight the enhanced accuracy of CBAMPix2Pix in synthesizing CT-like images from MRI data.
HU Discrepancies Between the Deformed CT(dCT) Image and the Synthetic CT Image Generated from the Models Trained with All Samples on 3D Volumes.

Abbreviations: HU = Hounsfield unit; MAE = mean absolute error; SD = standard deviation. P value: P3 = CBAMPix2Pix vs CycleGAN, P4 = CBAMPix2Pix vs Pix2Pix.
The
Discussion
Generating CT images from MRI data is inherently a style transfer challenge. Recent advancements in data-to-data translation within generative models have spurred research into improving sCT quality via GANs and their variants. Nevertheless, the exclusive utilization of the sigmoid cross-entropy loss function in the original GAN tends to result in unstable training procedures. Pix2Pix, functioning as a cGAN, has demonstrated efficacy in image-to-image translation assignments. It accomplishes this by acquiring a mapping from input to output images when trained on paired data. In contrast to CycleGAN, which operates using unpaired data, Pix2Pix necessitates paired datasets. Yet, owing to the direct supervision afforded by the paired MR and CT images, it can produce more precise and detailed sCT images. Notwithstanding this advantage, when employing Pix2Pix for generating sCT images from MR data, the L1 loss function is typically utilized to regulate the similarity between the generated CT images and the dCT images. Although the L1 loss imposes constraints at the pixel level and computes loss values on a pixel-by-pixel basis, it fails to precisely gauge structural similarity in terms of global structural cues such as contour or shape consistency. This limitation can give rise to deficiencies in the sCT images, particularly in the depiction of bone structures. For example, Peng et al 40 revealed that, in patients with nasopharyngeal carcinoma (NPC), the MAE of sCT images generated by cGAN was greater than that of CycleGAN. Specifically, the MAE values were 69.67 ± 9.27 HU (cGAN) and 100.62 ± 7.90 HU (CycleGAN) in soft tissues, and 203.71 ± 28.22 HU (cGAN) and 288.17 ± 17.22 HU (CycleGAN) in bone. Critically, despite these residual errors (particularly in bone), the same study reported 2%/2-mm γ passing rates were (98.68 ± 0.94)% and (98.52 ± 1.13)% for the cGAN and cycleGAN, respectively. Meanwhile, the absolute dose discrepancies within the regions of interest were (0.49 ± 0.24)% and (0.62 ± 0.36)%, respectively. These values substantially exceed the commonly accepted clinical threshold of 95% for γ passing rates. Furthermore, Kang et al 41 discovered that when utilizing CycleGAN, the mean SSIM and PSNR values were merely 0.90 ± 0.03 and 26.3 ± 0.7 dB in patients with pelvic, thoracic, and abdominal tumors.
In this study, we concentrated on producing sCT images from the MR images of metastatic brain cancer patients, utilizing paired 3 T MR data. We introduced a new deep-learning model named CBAMPix2Pix, designed to enhance the MR-based radiotherapy workflow. This model is an extension of the Pix2Pix framework, enhanced by the CBAM to strengthen feature representation. We also incorporated a structural similarity loss function to guarantee geometric consistency between MR and sCT images. Our experiments revealed that CBAMPix2Pix works effectively on brain cancer patients'MR images. The sCT images closely resemble real CT images in image quality and HU value accuracy. The HU values of the sCT images are nearly identical to those of real CT images, fulfilling the radiotherapy dose calculation requirements and reducing dose errors caused by inaccurate HU values. The generated sCT images bear a striking similarity to dCT images, especially in high-density bony tissues. This makes them ideal for direct use in quantitative applications like dose calculation and adaptive treatment planning, as shown in Figure 3. Unlike CycleGAN, which uses unpaired data and may fail to preserve local features, our method uses paired data for more accurate and detailed sCT generation. By integrating CBAM and a structural similarity loss function, CBAMPix2Pix overcomes the limitations of traditional Pix2Pix methods. It better captures global structural cues and ensures geometric consistency, producing higher-quality and more realistic sCT images. This is vital for clinical radiotherapy planning and treatment. Lately, many modified GAN-based methods have emerged to boost the accuracy and efficiency of sCT generation from MR. For example, Yang et al 42 created an extra structure-consistency loss based on the modality independent neighborhood descriptor for unsupervised MR-to-CT synthesis. Their method outperforms original CycleGAN in synthetic CT image accuracy and visual quality. A novel compensation-CycleGAN was proposed, modifying the cycle-consistency loss in traditional CycleGAN to simultaneously generate sCT images and compensate for missing anatomy in truncated MR images. Focusing on cervical cancer patients, Cusumano et al 43 applied a conditional GAN (cGAN) to 20 test patients. They used MAE and mean error (ME) to evaluate image metrics, achieving an MAE of 78.71 HU in the abdominal region and an ME of 10.83 HU. Fu et al 44 used cGAN and CycleGAN networks, validated via leave-one-out cross-validation on 12 abdominal tumor patients. They assessed image metrics using MAE and PSNR. The cGAN achieved an MAE of 89.80 HU and a PSNR of 27.4 dB, while the CycleGAN had an MAE of 94.10 HU and a PSNR of 27.2 dB.
The geometric and structural accuracy of sCT images is crucial for radiotherapy planning, as it ensures precise radiation dose delivery. Even minor inaccuracies in anatomy can lead to severe consequences in radiation therapy outcomes. To address this, we proposed the CBAMPix2Pix method, which effectively preserves structural characteristics and focuses on the human body region or the most critical part of the sCT. We evaluated our method on 15 independent test patients (see Table 2). The results indicated that while CycleGAN and Pix2Pix can produce realistic CT images, they struggle with regions of low MR signals, such as bone interfaces, leading to uncertainties. In contrast, CBAMPix2Pix outperformed the other two methods in terms of NMAE, PSNR, and SSIM, likely due to its integration of the advantages of Pix2Pix and the inclusion of channel and spatial attention mechanisms (see Figure 2). A common challenge in MR-to-CT synthesis is the less accurate HU values compared to dCT images. However, the three methods implemented in this study demonstrated a good ability to correct the CT values of MRI. Among them, CBAMPix2Pix achieved the best results, followed by Pix2Pix and then the original CycleGAN. Comparing our results with previous studies, Zhao et al6reported average MAE, PSNR, and SSIM values of 91.30 HU, 27.4 dB, and 0.94, respectively, for their Comp-CycleGAN models trained with body contour information. Our CBAMPix2Pix model showed better performance in quantitative metrics, achieving a PSNR of 27.5 dB and an MAE of 74.48 HU in body. The accurate sCT generation provided by our model aids oncologists in contouring precise targets, leading to more accurate radiotherapy plans for cancer patients and potentially improving survival rates.
CBAMPix2Pix, which integrates an SSIM loss function and channel-spatial attention mechanisms, has certain limitations. First, all data were acquired at a single institution using a fixed protocol (United Imaging 3 T MRI and Philips Big Bore CT), limiting generalizability across vendors, field strengths, or acquisition settings. Multi-center, multi-vendor validation is essential for clinical translation and will be pursued in future collaborations. Second, while our method reduces HU prediction errors, a full dosimetric evaluation was not performed. Task-based validation—such as dose recalculation in a commercial treatment planning system (TPS) using clinical plans—is critical to confirm clinical acceptability. Future work will quantify dosimetric accuracy and assess suitability for applications including dose calculation, online adaptive radiotherapy, and multi-center trials. Notably, as a fully MR-derived approach, our framework is inherently compatible with MR-guided radiotherapy(MRgRT) platforms, where it could enable CT-free daily replanning by eliminating manual bulk density assignment. Third, the current model uses only a single MR sequence; future studies will explore multimodal MR inputs to further improve sCT quality and robustness. Beyond these technical directions, our framework also aligns with emerging clinical paradigms that integrate artificial intelligence(AI), imaging, and real-world clinical data, As highlighted by Bilski et al 45 While our sCT model is developed for photon therapy, its use in proton beam therapy—where accurate HU-to- stopping power ratios (SPR) conversion is critical, especially for central nervous system(CNS) tumors—is not currently supported. Systematic discrepancies between sCT and true CT in bone and air-tissue interfaces remain a barrier to reliable SPR estimation.
Conclusion
This study proposes a CBAMPix2Pix network to generate sCT from MR images to optimize radiotherapy procedures. The network integrates CBAM mechanism, adopts SSIM loss, and combines T1wc to improve performance and generalization capabilities. Qualitative and quantitative experiments revealed that our approach outperformed both CycleGAN and Pix2Pix in terms of both structural preserving and HU accuracy. Our analysis shows that sCT images can provide a new perspective for radiotherapy plan optimization and efficacy prediction. In the future, the model will be optimized, the sample size will be expanded to verify stability, and the application potential of different tumor types will be explored.
Footnotes
Abbreviations
Acknowledgements
Not applicable.
Ethics Considerations and Consent to Participate
The studies involving humans were approved by Scientific Research Ethical Review of Ganzhou Cancer Hospital (Approval number 2025-71). The studies were conducted in accordance with the local legislation and institutional requirements. Written informed consent for participation was not required from the participants or the participants’ legal guardians/next of kin in accordance with the national legislation and institutional requirements.
Author Contributions
C.X., C.C., Q.H., F.X., L.L., W.L., and Y.X. contributed to the study conception and design, C.C., Q.H., F.X, L.L., and W.L. prepared materials and collected data. C.X. and Y.X. analyzed the experiments. The first draft of the manuscript was written by C.X. and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets generated and analyzed during the current study are not publicly available due to patient privacy and institutional policies but may be available from the corresponding author upon reasonable request and with permission from Ganzhou Cancer Hospital.
