Abstract
Background
Transforming growth factor-β1 (TGF-β1) is an immunosuppressive cytokine that is highly expressed in the tumor microenvironment (TME) of lung adenocarcinoma (LUAD). TGF-β1 plays important roles in regulating tumor metastasis and chemotherapy resistance. However, the specific molecular mechanisms by which TGF-β1 regulates cisplatin resistance in the TAM of LUAD remain unclear.
Materials and methods
THP-1 induced macrophages were co-cultured with A549 and H1975 cells, and subsequently transfected with silencing TGF-β1 (siTGF-β1), GLI2 (siGLI2), a GLI2 overexpression plasmid, and their negative controls. Cellular activity was measured by CCK-8 and colony formation assays. Cell apoptosis was evaluated by flow cytometry and TUNEL staining. Transwell assays were performed to assess cell migration and invasion capabilities. The levels of Smad2/3, GLI2, cyclin D, and cyclin E expression were evaluated by qPCR, western blotting, and immunofluorescence methods. TGF-β1 levels were determined by ELISA.
Results
Macrophages suppressed the apoptosis and promoted the migration and invasion of LUAD cells. TAM siTGF-β1 downregulated the Smad2/3 signaling pathways and GLI2 expression, deceased cell proliferation, and promoted apoptosis. SiGLI2 increased apoptosis and decreased the proliferation of LUAD cell lines. GLI2 decreased cisplatin resistance in LUAD cells.
Conclusion
High expression of TGF-β1 in the TAM positively activates GLI2 expression via the Smad2/3 pathway, which subsequently regulates cyclin D and cyclin E expression, and promotes the cisplatin resistance of LUAD.
Introduction
Lung cancer is the second leading cause of cancer-related death, and approximately 350 people die from lung cancer each day. 1 Smoking directly accounted for 81% of the lung cancer deaths in 2023 and 3560 deaths were caused by second-hand smoke. 2 As the most prevalent subtype of non-small-cell lung cancer (NSCLC), the incidence of lung adenocarcinoma (LUAD) continues to increase among ex-, current-, and even non-smokers. 3 LUAD is often diagnosed during in an advanced stage, and most patients die in < 5 years after diagnosis. 4 The chemotherapy drugs cisplatin and carboplatin remain the first-line treatments for lung cancer. 5 Compared with other types of lung cancer, LUAD shows a stronger inflammatory response and is more resistant to chemotherapy. The cisplatin resistance of LUAD poses great challenges for the treatment of this disease. 6 Therefore, understanding the molecular mechanism for cisplatin resistance has become a hot spot of research on LUAD.
The tumor microenvironment (TME) is considered to be a harsh environment for tissue growth, but still can promote tumorigenesis and tumor development. 7 The TME is closely related to tumor cells and regulates cellular signaling molecules, stimulates tumor angiogenesis, and induces immune tolerance.7,8 One type of tumor-associated immune cell, the macrophage, has been shown to promote cancer cell proliferation, metastasis, drug resistance, and immune escape in a variety of tumors.9,10 However, the specific pathways through which tumor-associated macrophages (TAMs) affect LUAD cells are not fully understood.
Transforming growth factor-β1 (TGF-β1) is an immunosuppressive cytokine that promotes tumor immune escape and distant metastasis. It has been found to induce the formation of dense connective tissue around solid tumors. The connective tissue than acts as a physical barrier that limits cytotoxic T-lymphocyte infiltration into tumors. 11 During the past three decades, TGF-β has been further studied, and especially for its dual role in the TME, where it was shown to promote apoptosis and inhibit cell proliferation in early-stage tumors. Moreover, TME TGF-β has also been shown to promote tumor formation by regulating genomic instability, epithelial-mesenchymal transition (EMT), neoangiogenesis, immune evasion, cell motility, and metastasis in advanced stage tumors. 12 Additionally, the downstream pathways activated by TGF-β1 may also be associated with drug resistance in cancer cells. Therefore, the specific molecular mechanisms by which TGF-β1 regulates chemotherapy drug resistance in the TAM of LUAD are of great importance.
As reported, GLI family zinc finger 2 (GLI2), but not GLI1, is responsible for activating SHH, which is involved in TGF-β1/Smad2/3 signaling and regulates the migration and invasion of pancreatic carcinoma cells. 13 Genes coding for the cell cycle proteins cyclin D and cyclin E have been reported as key target genes of GLI2.14,15 They also have been shown to play a regulatory role in lung cancer. 16 However, it remains unknown whether those cell cycle proteins exert their essential roles through GLI2 and TAM-derived TGF-β1. Therefore, our present study investigated the molecular mechanism by which TAM-derived TGF-β1 might regulate LUAD cell progression and cisplatin resistance.
Materials and Methods
TAM Cells and co-Culture Experiments
THP-1 cells are widely used for studying macrophage-related mechanisms, and were obtained from (ECACC, Salisbury, UK). The THP-1 cells were cultured in antibiotic-free RPMI media containing 10% fetal bovine serum (FBS). A549 and H1975 cells were obtained from the Institute of Cell Biology (Shanghai, China) and cultured in DMEM supplemented with 10% FBS and penicillin/streptomycin. THP-1-induced macrophages were co-cultured with A549 and H1975 cells; after which, all the cells were maintained at 37 °C in a humidified tissue culture incubator containing 5% CO2.
To identify the effect of macrophages on LUAD cells, the following 4 co-culture systems were constructed: A549 (upper layer) + A549 (bottom layer) (A549 control), macrophage (upper layer) + A549 (bottom layer) (mφ+A549), H1975 (upper layer) + H1975 (bottom layer) (H1975 control), and macrophage (upper layer) + H1975 (bottom layer) (mφ+H1975). The supernatants were collected after 2 days of culture.
Transfections
The macrophage cells were transfected at 37°C with silencing RNA TGF-β1 (siTGF-β1), GLI2 (siGLI2) and their negative controls (NCs), as well as the GLI2 overexpression plasmid and vector by using Lipofectamine® 2000 (Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA, USA). The plasmid and silencing RNA were acquired from GenePharma Co., Ltd (Shanghai, China). Four co-culture systems were constructed, including macrophages treated with siNC (upper layer) + A549 (bottom layer) (mφ siNC + A549), macrophages treated with siTGF-β1 (upper layer) + A549 (bottom layer) (mφ siTGF-β1 + A549), macrophages treated with siNC (upper layer) + H1975 (bottom layer) (mφ siNC + H1975), and macrophages treated with siTGF-β1 (upper layer) + H1975 (bottom layer) (mφ siTGF-β1 + H1975). Six other co-culture systems were also constructed, including A549 cells transfected with siNC (A549 siNC), A549 cells transfected with siNC and stimulated with TGF-β1 (A549 siNC + TGF-β1), A549 cells transfected with siGLI2 and stimulated with TGF-β1 (A549 siGLI2 + TGF-β1), H1975 cells transfected with siNC (H1975 siNC), H1975 cell transfected with siNC and stimulated with TGF-β1 (H1975 siNC + TGF-β1), and H1975 cells transfected with siGLI2, and stimulated with TGF-β1 (H1975 siGLI2 + TGF-β1). Four overexpression transfections were constructed, including GLI2 overexpression in both A549 (A549 GLI2) and H1975 cells (H1975 GLI2), and vector in A549 (A549 vector) and H1975 cells (H1975 vector).
Transwell Assays
Transwell assays were conducted for cell migration and invasion experiments. The two assays were conducted in similar manners, except for the supplementation of culture medium and use of DMEM in the invasion assays. First, approximately 1.0 × 105 cells in serum-free DMEM were added to the top Transwell chamber and medium containing 10% FBS was added to the lower chamber. After 6 h of incubation at 37°C, cells in the lower chamber were washed, fixed with glutaraldehyde, and stained with 0.1% crystal violet (Sigma-Aldrich; Merck KGaA). Finally, the numbers of cells in 5 visual fields were counted under a x400 microscope.
Colony Formation Assay
The cells were trypsinized, seeded into 96-well plates, and then cultured in RPMI-1640 supplemented with 10% FBS. The cells were then fixed with 4% paraformaldehyde, stained with 0.1% crystal violet, and observed under a light microscope (magnification, × 40).
Flow Cytometry Analysis
An Annexin V FITC/PI staining Apoptosis Detection Kit (Sigma-Aldrich, Germany) was used to measure cellular apoptosis. Briefly, approximately 3 × 104 cells were suspended in a binding buffer and treated with annexin V-FITC and PI staining solutions in the dark at room temperature. Next, the cells were treated with binding buffer and analyzed by flow cytometry (Agilent, Santa Clara, CA, USA), and the resultant data were analyzed using winMDI 2.9 software. In addition, the cell division cycle was also analyzed by flow cytometry with a Cell Cycle Analysis Kit (C1052, Beyotime, Shanghai, China), The analysis process was performed as previously described. 17 Briefly, approximately 1 × 105 cells were collected and fixed in 70% ethanol at 4 °C for 2 h. After washing and re-suspension, each aliquot of cells was treated with 0.5 mL of propidium iodide staining solution in a dark bath at 37 °C for 30 min. Finally, the fluorescence of each sample was detected by flow cytometry (Agilent) at an excitation wavelength of 488 nm.
Enzyme Linked Immunosorbent Assay (ELISA)
Cells were collected, lysed with RIPA, and then centrifuged at 12,000 rpm for 15 min at 4 °C. The levels of TGF-β1 were measured with commercial kits (Shanghai Boyun Biotech Co., Ltd) according to the manufacturer's instructions.
Cell Viability Assay
Approximately 3 × 103 cells were seeded into a 96-well plate, and then treated with cisplatin (0, 1, 2, 5, 10, 20, and 30 μM) for 48 h. Next, the cells were cultured with Cell Counting Kit-8 (CCK-8, Solarbio, Beijing, China) solution, and the cell numbers were determined by spectrometry at 450 nm. The IC50 values were then calculated.
Quantitative Real-Time PCR (QPCR)
The total cellular RNA was extracted using TRIzol reagent (Invitrogen, USA). Primer sequences for GLI2 and GAPDH were designed and synthesized by Sangon Biotech (Shanghai, China) (Table S1). QPCR was performed by using a HiScript II One Step qRT-PCR SYBR Green Kit (#Q221-01) on an ABI 7900 qPCR system (Applied Biosystems, Foster City, CA, USA). Relative levels of gene expression were calculated using the 2–ΔΔCt method.
Western Blot Analysis
The total cellular proteins were extracted with RIPA lysis buffer (#R0278, Sigma) and subsequently separated by a standard sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) method that used an upper stacking gel at 120 volts and a lower separation gel at 120 V. Next, the separated protein bands were transferred onto membranes that were subsequently washed and blocked overnight. The membranes were then incubated with primary antibodies against Smad2 (#cat: ab280888, Abcam, Cambridge, UK), Smad3 (#cat: ab74062, Abcam,), GLI2 (#cat: ab187386, Abcam), cyclin D (#cat: ab134175, Abcam), cyclin E (#cat: ab33911, Abcam), and β-actin (#cat: ab8226, Abcam). Next, the membranes were incubated with a secondary antibody (HRP-labeled goat anti-rabbit IgG; #cat: ab205718, Abcam) at room temperature and placed inelectrogenerated chemiluminescence (ECL) solution (ECL808-25, Biomiga, San Diego, CA, USA). The immunostained protein bands were observed under an x-ray machine (36209ES01, Shanghai, China), and the net density value of each band was calculated using Image-Pro Plus 6.0 software. All protein staining levels were normalized to those of β-actin, and all experiments were performed in triplicate.
Immunofluorescence (IF) Analysis
Briefly, cells were cultured to 60%-70%, confluence and then incubated with anti-GLI2, followed by incubation with a secondary anti-FITC antibody (Sigma, Darmstadt, DE). For nuclear/cytosolic localization, the cells were fixed again in paraformaldehyde and stained with 4’, 6-diamidino-2-phenylindole (DAPI). Finally, GLI2 fluorescence was observed under a confocal laser microscope (Olympus Optical, Tokyo, Japan).
TUNEL Staining
An In-Situ Cell Death Detection Kit (Roche) was used for TUNEL assays. Briefly, cells were treated with red fluorescent-tagged enzyme solution and examined under a fluorescence microscope (DM2500; Leica Microsystems, Germany). Image-Pro Plus version 6.0 software (Media Cybernetics, Rockville, MD, USA) was used to measure the red fluorescence of TUNEL-positive cells. Five randomly selected high-power lens visual fields were evaluated in each section. Images of cells stained with DAPI and TUNEL solution were recorded and merged.
Statistical Analysis
All statistical analyses were conducted using GraphPad Prism 9 software (San Diego, CA, USA). An unpaired student's t-test or one-way ANOVA analysis was performed depending on the groups. Data are presented as a mean value ± standard deviation (SD). A P-value < 0.05 was considered to be statistically significant.
Results
Co-Culture with Macrophages Promoted GLI2 Expression and Suppressed the Inhibitory Effect of Cisplatin on LUAD Cells
In both A549 and H1975 cells, the levels of TGF-β1 were significantly increased in the macrophage co-cultured systems when compared with control cells (P < 0.001,

Macrophages suppressed cell apoptosis and promoted the migration and invasion of LUAD cells. (A) The levels of TGF-β1 in 4 co-culture systems were measured by ELISA. (B, C) The levels of Smad2, Smad3, and GLI2 protein expression in 4 co-culture systems were assessed by western blotting. (D) The levels GLI2 in 4 co-culture systems were assessed by qPCR. (E) IF staining was performed to evaluate GLI2 expression in 4 co-culture systems. (F) TUNEL staining was performed to assess apoptosis in LUAD cells treated with cisplatin (5 μM). (G) Flow cytometry was used to detect apoptosis in cells treated with cisplatin (5 μM). (H, I) Transwell assays were performed to assess the migration capability of cells treated with cisplatin (5 μM). (J, K) Transwell assays were performed to assess the invasion capability of cells treated with cisplatin (5 μM). *** P < 0.001.
To evaluate the effect of macrophages on cisplatin resistance, 5 μM cisplatin was added to the co-culture systems. Two days later, the cells in the bottom layer were collected and analyzed by TUNEL staining and flow cytometry. When compared with control groups the numbers of TUNEL-positive cells were significantly decreased in the mφ+A549 and mφ+H1975 groups (Figure 1F), indicating a weakened role for macrophages in regulating LUAD cell apoptosis. Results of a flow cytometry analysis were consistent with that of TUNEL staining, which showed a significantly decreased apoptosis rate in the mφ+A549 and mφ+H1975 groups when compared with the control group (P < 0.001, Figure 1G). Finally, the migration and invasion of LUAD cells were assessed. The numbers of migrated and invasive cells were significantly increased in the mφ+A549 and mφ+H1975 groups when compared with the control group (P < 0.001, Figures 1H-K), suggesting a promotive role for macrophages in regulating LUAD cell metastasis. When taken together, these results indicated there were high levels of TGF-β1, Smad2, Smad3, and GLI2 in the TAM, in which cell apoptosis was suppressed and cell migration and invasion were promoted.
Macrophage-Derived TGF-β1 Promoted Cisplatin Resistance in LUAD Cells by Activating the Smad2/3-GLI2 Pathway
Because TGF-β1 was present at high levels in the TAM, we investigated how it influences LUAD. The levels of TGF-β1 in the four co-culture systems were measured, and those results showed lower levels in the mφ siTGF-β1 + A549 and mφ siTGF-β1 + H1975 groups than in the mφ siNC + A549 and mφ siNC + H1975 groups (P < 0.001, Figure 2A), demonstrating the successful transfection of siTGF-β1. Regarding the siTGF-β1 transfections, the levels of Smad2, Smad3, and GLI2 proteins were significantly decreased when compared with the si-NC group (P < 0.001, Figures 2B). The levels of GLI2 as tested by qPCR (P < 0.001, Figure 2C) and IF staining (Figure 2D) were consistent with those shown by western blotting. Thus, our experiments revealed that siTGF-β1 downregulated the Smad2/3 signaling pathways and GLI2.

TAM siTGF-β1 downregulated the Smad2/3 signaling pathways and GLI2 expression, and thus promoted cell apoptosis and decreased cell proliferation. (A) The levels of TGF-β1 expression in 4 co-culture systems were measured by ELISA. (B) The levels of Smad2, Smad3, and GLI2 protein expression in 4 co-culture systems were assessed by western blotting. (C) The levels GLI2 in 4 co-culture systems were assessed by qPCR. (D) IF staining was conducted to evaluate GLI2 expression in 4 co-culture systems. (E) TUNEL staining was performed to assess the apoptosis of LUAD cells treated with cisplatin (5 μM). (F) Flow cytometry was used to detect the apoptosis rates of cells treated with cisplatin (5 μM). (G) Colony formation assays were performed to evaluate the proliferation of cells treated with cisplatin (5 μM). *** P < 0.001.
LUAD cells were cultured with cisplatin, and their levels of apoptosis, migration, invasion, and proliferation were assessed. As shown in Figure 2E, the numbers of TUNEL-positive A549 and H1975 cells were significantly increased in the mφ siTGF-β1 group when compared with mφ siNC group. Flow cytometry was also used to evaluate cell apoptosis (P < 0.001, Figure 2F) and the results were consistent with those of TUNEL staining, as both assays revealed higher levels of cell apoptosis among macrophages treated with siTGF-β1 than among macrophages treated with the si-NC. Colony formation assays showed lower cell numbers in the mφ siTGF-β1 group than in the mφ siNC group (Figure 2G). These findings indicated that TAM siTGF-β1 downregulated the Smad2/3 signaling pathways and GLI2 expression, and promoted cell apoptosis, thereby decreasing cell proliferation.
Knockdown of GLI2 Affected the Cisplatin Resistance of LUAD Cells Stimulated with TGF-β1
While our data showed that siTGF-β1 downregulated GLI2 expression, promoted LUAD cell apoptosis, and decreased cell proliferation, it remained unknown whether GLI2 plays an essential role in LUAD cell progression. First, we assessed the levels of Smad2, Smad3, GLI2, cyclin D, and cyclin E expression in the different groups (Figures 3A-D). We found that the levels of Smad2 and Smad3 expression in both A549 and H1975 cells were significantly increased in the siNC + TGF-β1 and siGLI2 + TGF-β1 groups when compared the siNC group (P < 0.001; Figure 3B). No significant differences were observed between the siNC + TGF-β1 and siGLI2 + TGF-β1 groups, indicating that siGLI2 did not affect the Smad2/3 signaling pathway (P > 0.05). When the expression levels of GLI2 were assessed by western blotting (P < 0.001, Figure 3C), qPCR (P < 0.001, Figure 3E), and IF (Figure 3F), the data showed that GLI2 was significantly increased after TGF-β1 treatment, and decreased by si-GLI2 transfection, demonstrated a successful transfection of GLI2. The levels of cyclin D and cyclin E expression were increased along with those of GLI2, which were significantly increased after TGF-β1 treatment, and decreased by si-GLI2 transfection (P < 0.001, Figure 3D). SiGLI2 showed no effect on the Smad2/3 signaling pathway, but significantly decreased cyclin D and cyclin E expression.

SiGLI2 increased cell apoptosis and decreased cell proliferation in LUAD cell lines. (A-D) The levels of Smad2, Smad3, GLI2, cyclin D, and cyclin E protein expression in 6 groups were assessed by western blotting.(E) The levels of GLI2 in 6 groups were assessed by qPCR. (F) IF staining was performed to evaluate the levels of GLI2 expression in 6 groups. (G) Flow cytometry was used to detect the apoptosis rates of cells treated with cisplatin (5 μM). (H) The cell cycle was assessed using a flow cytometry assay. (I) TUNEL staining was performed to assess the apoptosis of LUAD cells treated with cisplatin (5 μM). (J) Colony formation assays were performed to evaluate the proliferation of cells treated with cisplatin (5 μM). *** P < 0.001.
After co-culture with cisplatin, LUAD cell apoptosis was significantly decreased by TGF-β1 and subsequently increased by si-GLI2 (P < 0.001, Figures 3G and 1). In addition, TGF-β1 decreased the numbers of cells in the G0/G1 phase of division and increased the numbers of cells in the S and G2/M phases, while knockdown of GLI2 weakened that effect (P < 0.001, Figure 3H). LUAD cell proliferation was significantly increased by treatment with siNC + TGF-β1 and reduced by treatment with siGLI2 + TGF-β1 when compared to cells treated with the siNC (Figure 3J). Therefore, we concluded that siGLI2 increased the apoptosis and decreased the proliferation of LUAD cells, which was consistent with the effects of siTGF-β1.
Overexpression of GLI2 Promoted Cisplatin Resistance in LUAD Cells by Activating Cell Cycle Related Gene Expression
Additionally, we overexpressed GLI2 and evaluated its role in LUAD cell progression under conditions of cisplatin treatment. Four transfections were constructed, including overexpressed GLI2 in both A549 (A549 GLI2) and H1975 (H1975 GLI2) cells, and vector in A549 (A549 vector) and H1975 cells (H1975 vector). The levels of GLI2, cyclin D, and cyclin E proteins were significantly increased in cells transfected with the GLI2 overexpression vector when compared to cells transfected with the empty vector (P < 0.001, Figures 4A and B). We found that the expression levels of cell cycle proteins (cyclin D and cyclin E) showed a trend consistent with that of GLI2. Therefore, the role of GLI2 in regulating cell progression might be dependent on cyclin D and cyclin E. Finally, there were higher levels of GLI2 expression in the GLI2 overexpression groups than in the vector groups (P < 0.001) (Figures 4C and D).

GLI2 relieved cisplatin resistance in LUAD cells. (A, B) The levels of GLI2, cyclin D, and cyclin E protein expression in transfected cells were assessed by western blotting. (C) GLI2 expression in transfected cells was assessed by qPCR. (D) IF staining was performed to evaluate the levels of GLI2 in transfected cells. (E) TUNEL staining was performed to assess the apoptosis of LUAD cells treated with cisplatin (5 μM). (F) Flow cytometry was used to detect the apoptosis of cells treated with cisplatin (5 μM). (G) The cell cycle was assessed using a flow cytometry assay. (H) Colony formation assays were performed to evaluate the proliferation of cells treated with cisplatin (5 μM). (I, J) CCK-8 assays were performed to evaluate the viability and IC50 values of transfected LUAD cells treated with 0, 1, 2, 5, 10, 20, and 30 μM cisplatin. *** P < 0.001.
As for the effect of GLI2 on cell progression, our experiments on cell apoptosis demonstrated lower apoptosis rates (P < 0.001, Figures 4E and F) and higher cell proliferation rates (Figure 4H) in the GLI2 groups than in the vector groups. Overexpression of GLI2 decreased the numbers of cells in G0/G1 phase, and increased the number of cells in S and G2/M phases (P < 0.001, Figures 4G). Finally, we stimulated the LUAD cells with gradual cisplatin and examined their viability and IC50 values. When compared with the vector groups, cell viability gradually decreased with increasing concentrations of cisplatin. The cell viability and IC50 values for the GLI2 overexpression groups were higher than those for the vector groups (P < 0.001, Figures 4I and J). These results indicate that GLI2 helps to alleviate cisplatin resistance.
Discussion
In our present study, we elucidated the molecular mechanism by which TGF-β1 regulates LUAD cell progression under conditions of cisplatin treatment. Our data showed the following results: (1) High levels of TGF-β1, Smad2, Smad3, and GLI2 expression were observed in the TAM, in which cell apoptosis was suppressed and cell migration and invasion were promoted. (2) The Smad2/3 signaling pathways and GLI2 expression were downregulated by interference with TGF-β1 expression in the TAM, inducing cell apoptosis and decreasing cell proliferation. (3) GLI2 influenced cell progression by regulating the cell cycle-related genes of cyclin D and cyclin E via the Smad2/3 signaling pathway. (4) GLI2 promoted cisplatin resistance in the TAM cells of LUAD. When taken together, our data showed that high levels of TGF-β1 expression in TAMs activates GLI2 expression via the Smad2/3 pathway, which subsequently regulates cyclin D and cyclin E expression, and promotes the cisplatin resistance of LUAD (Figure 5).
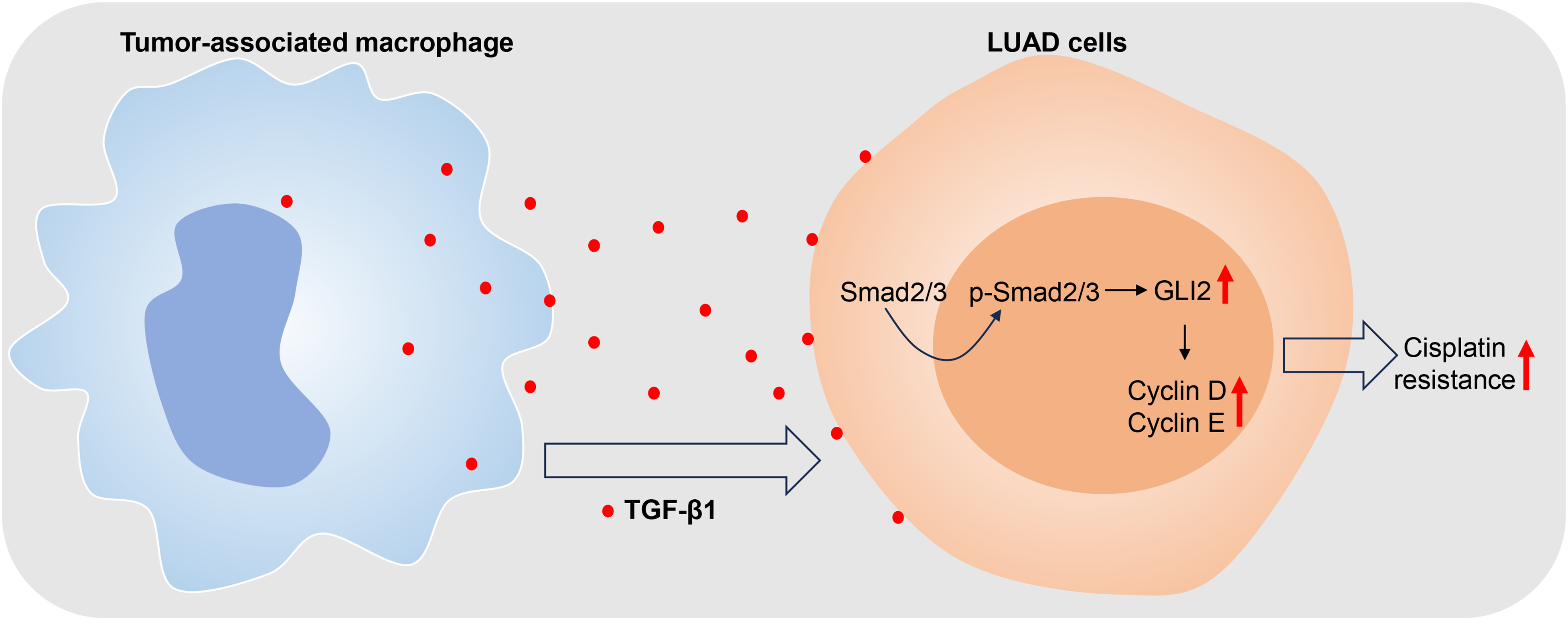
TAM-derived TGF-β1 regulated GLI2 via the Smad2/3 pathway, and thereby induced tumor cell progression and cisplatin resistance. A high level of TGF-β1 in TAMs induced GLI2 expression via the Smad2/3 pathway, which in turn, regulated cyclin D and cyclin E expression, and thus promoted the cisplatin resistance of LUAD.
TGF-β1 functions in the pathogenesis of lung cancers.18–20 We found that TGF-β1 was highly expressed in the TAMs of LUAD cells and promoted cell progression. Results from our present study were consistent with those of other studies of lung cancer.21–24 For example, myelin protein zero like 1 (MPZL1)/TGF-β1 was found to promote LUAD progression and suppress immune function, 21 and TGF-β1 was reported to be upregulated by lncRNA miR4435-2HG and promote the migration and proliferation of NSCLC cells. 22 Although those previous studies revealed the regulatory role of TGF-β1, none of them were specific for TAMs.
In our present study, we found that TGF-β1 in TAMs positively regulates cell progression via GLI2 in LUAD cells. These regulatory effects have also been demonstrated in other cancers.25,26 For example, TGF-β was found to induce GLI2 expression in a Smad3-dependent manner in primary leukemia cells. 26 However, in our present study, we found that the GLI2 regulates LUAD via TGF-β1 in a non-Smad3-dependent manner. Chen et al 27 described the GLI2/TGF-β1 feedback loop that exists between cancer cells and TAMs, and its role in maintaining EMT and endoplasmic reticulum homeostasis in cholangiocarcinoma. Those investigators demonstrated that TGF-β1 secreted by TAMs further activates SHH/GLI2 signaling in cholangiocarcinoma cells, which agreed with our present study on LUAD. TAMs promote the migration and invasion of human hepatoma Huh-7 cells via GLI2 by secreting TGF-β1. 28 These studies emphasize the molecular mechanism of TAM-secreted TGF-β1 in regulating cancer cell progression.
We further investigated how GLI2 regulates cell progression, and found that cell cycle proteins were involved. As reported, GLI2 is accountable for the transcription of cyclin D and cyclin E, by serving as an activator of transcription. 29 GLI2 and its downstream genes (cyclin D and cyclin E) are associated with activation of the hedgehog pathway in glioblastoma multiforme cells. 30 So far, the roles of GLI2/cyclin D and cyclin E in regulating cell progression have rarely been reported. Our present revealed for the first time that TGF-β1 in TAMs positively regulates LUAD cell progression via GLI2/cyclin D and cyclin E. However, it remains to be determined how cyclin D and cyclin E function in that regulation.
Cisplatin drug resistance is a major clinical impediment in treating LUAD patients, and numerous studies have indicated the importance of the TME for cisplatin resistance. 31 Previous studies revealed that suppression of TGF-β1 enhances the chemosensitivity of cisplatin-resistant lung cancer cells by inhibiting the synthesis of drug-resistant proteins, 32 which is consistent with our present study. It was reported that high levels of TGF-β1 expression significantly enhanced the chemo-resistance of oral squamous cell carcinoma (OSCC) cells by promoting their proliferation. 33 Cisplatin-induced activation of TGF-β signaling contributes to drug resistance. 34 Similarly, our present study also concluded that TGF-β1 in TAMs positively regulates cell progression and promotes cisplatin resistance.
Conclusion
In summary, we elucidated the molecular mechanism by which TGF-β1/GLI2/cyclin D and cyclin E in TAMs function to overcome cisplatin resistance. Our data showed that high levels of TGF-β1 in TAMs induce GLI2 expression via the Smad2/3 pathway, which regulates cyclin D and cyclin E expression, and thereby promotes the cisplatin resistance of LUAD.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Statement
N/A. No human or animal experiments were involved.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by 2023 Guangxi University Young and Middle aged Teachers Research Foundation Ability Enhancement Project (Guijiao Scientific Research [2023] No. 2)
