Abstract
Keywords
Introduction
Acute myeloid leukemia (AML) is a highly heterogeneous malignant disease of myeloid precursor cells. 1 There are no substantial changes in standard strategies for initial therapy in the last several decades. Cytarabine (Ara-C) in combination with anthracyclines are still the main induction regimens. 2 Although nearly 70% AML patients achieved complete remission after Ara-C based chemotherapy, 60% to 65% of elderly patients relapsed within 3 years of diagnosis. Ara-C resistance was one of the crucial reasons in the process of relapse in those AML patients.
The mechanisms of resistance to Ara-C have not yet been fully clarified in leukemia cells. It may be the result of multi-factor interactions. Overactivation of drug resistance-related signal pathways and genetic changes are the main factors. 3 Previous research found that a large proportion of AML had activating mutations in NRAS or KRAS genes,4,5 but the prognostic value of RAS mutations in AML remained inconclusive. Several studies indicated that RAS mutations were not significantly associated with the prognosis of AML patients.6,7 Another large sample study found that there was a tendency of longer survivals in patients who carried RAS mutations and specific molecular markers such as FLT3-LM, MLL-PTD, and NPM1. 8 A research in Japan showed that RAS mutations were frequent in AML, especially in French-American-British (FAB) types M4 and M5. And patients with RAS mutations often relapse after the treatment course. 9 In addition, the RAS family is also correlated with the activation of mitogen-activated protein kinase (MAPK) pathway, thus participating in the regulation of cell cycle, apoptosis, and differentiation. 10 DUSP1, a phosphatase, plays an important role in the cellular response to DNA damaging agents. DUSP1 regulates the dephosphorylation of MAPK pathways, which are the key RAS effectors. 11 A recent research suggested that DUSP1-p38 pathway activation was associated with chemotherapy sensitivity in myeloid leukemia. 12
All these findings suggested that RAS as well as DUSP1 signaling pathway may play a role in the growth and survival of leukemic cells and could be identified as a target for drug resistance. In this study, we detected the associations of DUSP1 and RAS mutations, as well as clinical outcomes in AML patients. We also explored the function and possible mechanisms of DUSP1 pathway in regulating Ara-C sensitivity in AML.
Methods
Patients
Between March 2017 and March 2019, we selected a cohort of 15 AML patients based on World Health Organization (WHO) criteria, alongside 15 controls carefully matched for baseline characteristics, all from Taizhou Central Hospital (Taizhou University Hospital). The control group consisted of individuals without malignant diseases. No controlled interventions were administered to either the AML patients or the controls following the study design. Bone marrow (BM) specimens were obtained via biopsy at the time of diagnosis, and consent was obtained from all participants. To safeguard privacy, all personal information was de-identified in this study. The collected BM samples were utilized for immunohistochemistry (IHC) staining and the analysis of DUSP1 expressions. This study was approved by the Institutional Research Ethics Committee (TZYXY2019-525). The reporting of this retrospective study conforms to STROBE guidelines. 13
Reagents and Cell Culture
Ara-C (Solarbio, China) was dissolved in phosphate buffer saline and stored at 4 °C. The erythroleukemia (AML-M6) KG-1 cell line, the acute monocytic leukemia (AML-M5a) MOLM-13 cell line, and acute myelomonocytic leukemia (AML-M4) OCI-AML2 cell line in this study were derived from cell lines of the Chinese Academy of Sciences (Shanghai, China). All cell lines were maintained in RPIM 1640 medium (Thermo Fisher Scientific, USA) supplemented with 10% heat-inactivated fetal bovine serum (FBS; Gibco, Thermo Fisher Scientific, USA), 1% penicillin-streptomycin (Thermo Fisher Scientific, USA). HEK293T cells were maintained in Dulbecco's Modified Eagle's Medium with 10% FBS, 1% penicillin, and 1% streptomycin. All cells were cultured at 37 °C in 5% CO2.
IHC Staining
BM samples were incubated with DUSP1 antibody (Abcam, UK) overnight at 4 °C. Polymer-HRP secondary antibody (Dako, Denmark) was added at room temperature for 1 h. The sections were then incubated with the DAB kit (ZSGB-BIO, China). The images were acquired by Leica Imaging Software and then evaluated by two experienced pathologists. The IHC score composed of the color and positive cell frequency. Scores of positive cell frequency: 0, < 25%; 1, 25%-50%; 2, 50%-75%; 3, > 75%. Scores of stained color: 0, no staining; 1, yellow; 2, tan; 3, brown.
Western Blotting
Protein samples were transferred to PVDF membranes (Millipore, USA). The membranes were blocked with BSA (Millipore, USA) for 1 h at room temperature and then treated with the primary antibodies of DUSP1 (dilution 1:1000; EMD Millipore, Catalog No. 07-535), p38 (Cell Signaling Technology, Catalog No. 9212), p-p38 (CST, Catalog No. 9211), JNK (CST, Catalog No. 9259), p-JNK (CST, Catalog No. 9258), ERK (CST, Catalog No. 9102), p-ERK (CST, Catalog No. 9101), or β-actin (dilution 1:3000; Abcam, UK). After washing, membranes were incubated with the goat anti-rabbit IgG H&L (HRP) (dilution 1:5000; Abcam, UK) for 1 h at room temperature.
Real-Time PCR
Total RNA was purified according to the instructions of GenElute™ Total RNA Purification Kit (Abcam, UK). RNA products were converted to cDNA by reverse transcription system (Takara, Japan). Real-time PCR cycles were implemented with SYBR green fluorescent dye kit (Takara, Japan). The primers used in RT-PCR were listed as follows: DUSP1 forward, 5’-TGCAGTACCCCACTCTACGA-3’; DUSP1 reverse, 5’-CCTCCACAGGGATGCTCTT-3’. NRAS forward, 5’-TGAGAGACCA ATACATGAGGACA-3’; NRAS reverse, 5’-CCCTGTAGAGGTTAATATCCGC A-3’. GAPDH forward, 5’-CAACGAATT TGGCTACAGCA-3’; GAPDH reverse, 5’- AGGGGTCTACATGGCAACTG-3’.
siRNA Interference
The siRNA transfection experiment was conducted using Invitrogen's Lipofectamine TM RNAi Max transfection reagent (Thermo Fisher Scientific, US). DUSP1 protein and mRNA levels were confirmed by western blot and Q-PCR. The DUSP1 siRNA interference sequences were as follows: DUSP1 siRNA sense, 5’-GCUUACCUUAUGAGGACUATT-3’; DUSP1 siRNA antisense, 5’-UAGUCCUCAUAAGGUAAGCTT-3’.
Generation of Stable Transformed Cell Lines
Full-length NRASG13D and NRAS wild type coding sequences were amplified by PCR according to sequences in the NCBI consensus coding sequence database and then cloned into pCDH + 3XFlag and copEGFP vectors. pCDH-CMV-NRAS-G13D and pCDH-CMV-NRAS recombinant lentiviral vectors were successfully constructed (Figure S1A). The sequence was verified by full-length sequencing identification (Figure S1B). HEK293T cells were transfected with lentivirus vectors. At 48 h post-transfection, medium was harvested and used to infect cell lines in presence of polybrene. Stable transfected cell lines were then selected using 1 µg/ml puromycin for 7 days. The infection efficiency was confirmed by GFP expression with fluorescence microscope (Figure S1C).
Cell Viability Assays
Cells were plated in 96-well plates with the concentration of 2 × 104 cells/well (100 μL). After treatment with Ara-C for 72 h, MTT was added to each well according to the instructions of CellTiter 96 Aqueous One Solution Cell Proliferation Assay (Promega, USA). The mixture was cultured in an incubator for 4 h. Cell viability was tested by reading absorbance (OD 450 nm).
Apoptosis Assay
The flow cytometry was used to evaluate cell apoptosis. Cells were stained with Annexin V-FITC/PI detection kit (Beyotime, Shanghai, China) following the manufacturer's instructions. 10,000 cells were collected for analysis. FlowJo software was used to calculate and analyze the percentage of the cells.
Gene Expression Profiling Analysis
Gene Set Enrichment Analysis (GSEA) software was used to analyze the RNA sequencing and ChIP sequence data obtained from GEO Datasets and the experiment included in this study. MAPK target gene sets database was downloaded from GSEA website (https://www.gsea-msigdb.org/gsea/msigdb/index.jsp). Database for Annotation, Visualization and Integrated Discovery (DAVID) was used to perform the GO pathway enrichment. Survival analysis was performed by R2: Genomics Analysis and Visualization Platform (https://hgserver1.amc.nl/cgi-bin/r2/main.cgi) and Gene Expression Profiling Interactive Analysis (GEPIA, http://gepia.cancer-pku.cn/index.html).
Immune Interaction Network and Immune Infiltration Analysis
The immune genes were downloaded from the website (https://www.immport.org/shared/home). The protein-protein interaction (PPI) was analyzed in the platform of Pathway Commons (http://www.pathwaycommons.org/). CIBERSORT is used to provide an estimation of the immune cell in 2 kinds of cell population divided by gene expression levels of DUSP1 (GSE43176). Median value of DUSP1 was chosen as the cut-off value of the 2 groups.
Statistical Analysis
SPSS, GraphPad Prism, and R language were used in this study. Statistical analyses were performed using 1-/2-way ANOVA or Student T-test. Statistical significance was set as P-value <.05. Statistically significant differences are marked by asterisks (*P < .05; **P < .01, ***P ≤ .001).
Results
DUSP1 was Highly Expressed and Correlated With Poor Prognosis in AML
IHC staining results showed that the protein levels of DUSP1 were over-expressed in the AML patients compared with controls (mean IHC scores, 2.3 vs 1.2 with AML vs control; P = .03; Figure 1A-B). The analysis of 172 AML patients from The Cancer Genome Atlas (TCGA) database (Project: TCGA-LAML) suggested that Transcripts Per Million (TPM) of DUSP1 was higher in AML (Figure 1C). Further survival analysis of TCGA indicated that AML patients with high DUSP1 expression level had poor prognosis (Figure 1D; HR = 2.7; P = .015). We also verified the finding in Gene Expression Omnibus (GEO) databases (Series GSE37642). The results in 422 AML patients showed that AML patients with high DUSP1 expression levels (ID: 201041_s_at) had a poor clinical outcome (P = .02; Figure 1E), which was consistent with the results from TCGA.

DUSP1 expression levels in AML. (A) DUSP1 expression detected by IHC in AML and control (×400). (B) DUSP1 IHC scores in AML and control. (C) DUSP1 expression in AML and controls from TCGA. (D) Survival analysis of AML from TCGA. (E) Survival analysis of AML from GEO datasets.
DUSP1 was Activated in AML With NRAS Mutations
Analysis of gene mutations in 531 AML patients from the public database 14 indicated that the frequency of NRAS mutations in AML was 13% (71 cases). Mutations in G13D were one of the most common mutation types (18 cases; Figure 2A). The frequency of KRAS mutations was 5% (24 cases) in AML. The most common mutation types among AML patients with KRAS mutations were mutations in G12D (6 cases; Figure 2B). The results from GEO database (GSE43176) showed that DUSP1 expressions were significantly increased in NRAS mutant AML patients compared with controls (P < .001; Figure 2C). Further analysis from GEO database (GSE6891) showed that DUSP1 expressions were significantly increased in NRAS mutated (NRASMUT) AML patients compared with NRAS WT (NRASWT) AML patients (log transferred DUSP1 expression, 9.3 vs 12.2 with NRASWT vs NRASMUT; P < .001; Figure 2D). As indicated in Figure 2A-B, NRASG13D mutation is one of the most frequent subtypes among RAS mutations in AML. Thus, we set to establish NRASG13D mutated AML cells for further study. Ectopic overexpression of NRAS was confirmed by qPCR assays (1.0 vs 7.1 vs 14.0 with control vs NRASWT vs NRASG13D mutation; Figure 2E). Western blot result showed that flag proteins were positive in AML with NRASWT and NRASG13D when compared with negative control (Figure 2F).

The relations between DUSP1 and RAS mutations in AML. (A) The frequency and types of NRAS mutations in AML. (B) The frequency and types of KRAS mutations in AML. (C-D) DUSP1 expressions in NRASMUT AML, NRASWT AML, and HC. (E) qPCR results of NRAS expression in KG-1 cells with NC, NRASG13D, and NRASWT. (F) Western blot results of FLAG-TAG in KG-1 cells with NC, NRASG13D and NRASWT.
Knocking Down DUSP1 Increased the Ara-C Sensitivity in AML
DUSP1 silencing was conducted in AML cell lines with NRASWT and NRASG13D mutation. Compared with the control, mRNA and protein expression levels of DUSP1 were significantly downregulated in DUSP1 knockdown KG-1 cells, MOLM-13, and OCI-AML2 cells (Figure 3A-B, Figure S2A-C). No significant difference of cell proliferation was found in NC, NRASWT, and NRASG13D mutated AML cell lines with Ara-C treatment (Figure S2D). MTT assays showed DUSP1 knockdown could sensitize KG-1 cells to Ara-C treatment (72 h Ara-C IC50, control 0.67 µM vs DUSP1 siRNA 0.53 µM in RASWT; control 0.69 µM vs DUSP1 siRNA 0.39 µM in RASG13D AML; P < .05; Figure 3C). Ara-C treatment in DUSP1 siRNA KG-1 cells induced significant apoptosis activation evidenced by an increased Annexin V-positive staining. DUSP1 siRNA in NRASG13D KG-1 cells significantly increased apoptosis rate compared in DUSP1 siRNA NRASWT KG-1 cells (apoptosis rate, WT + NC Ara-C 0 nM 6.6% vs 300 nM 19.5%; NRASG13D + NC Ara-C 0 nM 7.0% vs 300 nM 11.4%; WT + siDUSP1 Ara-C 0 nM 4.6% vs 300 nM 39.3%; WT + siDUSP1 100 nM 2.0% vs 300 nM 51.6%; P < .05; Figure 3D). The similar effect could also be found in MOLM-13 cells. DUSP1 knockdown could sensitize MOLM-13 cells to Ara-C treatment (72 h Ara-C IC50, control 0.61 µM vs DUSP1 siRNA 0.59 µM in RASWT; control 0.63 µM vs DUSP1 siRNA 0.43 µM in RAS G13D mutated AML; P < .05; Figure S2E). DUSP1 siRNA in NRASG13D MOLM-13 cells had significantly increased apoptosis rate compared to DUSP1 siRNA NRASWT MOLM-13 cells (apoptosis rate, WT + NC Ara-C 0 nM 11.0% vs 300 nM 31.2%; NRASG13D + NC Ara-C 0 nM 9.0% vs 300 nM 35.4%; WT + siDUSP1 Ara-C 0 nM 8.6% vs 300 nM 49.2%; WT + siDUSP1 100 nM 11.9% vs 300 nM 60.4%; P < .05; Figure S2F). The similar effects could also be found in OCI-AML2 cells (Figure S3A-B).

Ara-C treatments in DUSP1 silencing AML cells. (A) DUSP1 mRNA expression in DUSP1 silencing and control NRASG13D KG-1 cells. (B) DUSP1 protein expression in DUSP1 silencing and control NRASG13D KG-1 cells. (C) MTT results of DUSP1 knockdown KG-1 cells with Ara-C treatment. (D) Apoptosis of DUSP1 knockdown KG-1 cells with Ara-C treatment.
DUSP1 Knockdown Induced Phosphorylation of MAPK Pathway in NRASG13D AML
Gene enrichment analysis indicated that MAPK signaling pathway was significantly enriched in AMLRAS patients (NRASMUT vs NRASWT, P = .072, Figure 4A; NRAS mutations vs health control (HC), P = .001, Figure 4B; KRASMUT vs KRASWT, P = .041, Figure S3C). Leading edge genes, which represent the most enriched subset of genes, were shown in Supplemental List 1-3. We further tested total and phosphorylation levels of MAPK subfamilies that include extracellular signal-regulated kinases (ERK), p38, and c-Jun N-terminal kinases (JNK) in DUSP1 downregulated NRASG13D AML cell lines by western blot. The western blot results showed that the phosphorylation levels of MAPK signaling pathway were significantly upregulated in DUSP1 knockdown NRASG13D KG-1, MOLM-13, and OCI-AML2 cells (Figure 4C, S3D). RNA sequencing was performed in DUSP1 siRNA and control siRNA KG-1 cells, and the genes of all the comparison groups were collected and unitized as the different gene sets (Figure S4). After overlapping the differently expressed genes in DUSP1 knockdown KG-1 cells and in RASMUT AML patients samples (GEO Datasets GSE43176), 71 genes were found to be differentially expressed both in those 2 groups (Figure 4D). Among them, LDHB, HNRNPD, S100P, LYZ, HNRNPAB, TSPO, IQGAP1, NEAT1, S100A9, PNISR, CEBPE, PAK2, INSIG1, S100A8, RBPJ, COPS5, SON, RANBP2, MEF, PTK2B, and PIK3CA, marked in red, have been reported to be related to chemotherapy resistance (Figure 4E). Further Gene Ontology (GO) analysis showed that apoptotic process, RNA splicing, mRNA processing, cell-cell adhesion, protein transport, innate immune response signaling pathways were both enriched in DUSP1 knockdown KG-1 cells and in RASMUT AML patients samples. Among them, innate immune response signaling pathways were further analyzed (Figure 4F).
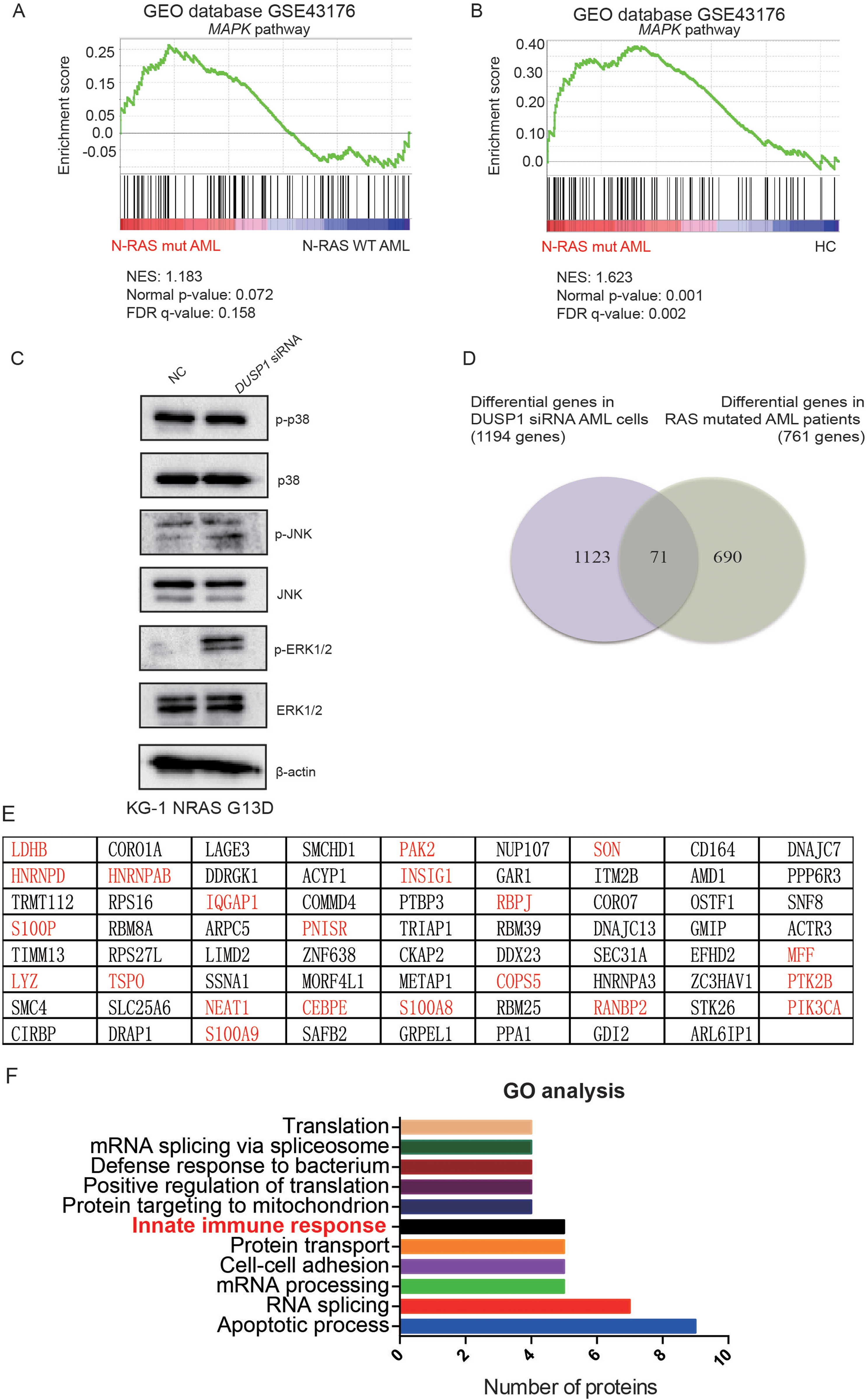
Downstream pathways in DUSP1 silencing AML cell lines and NRASMUT AML patients. (A) Gene set enrichment analysis of MAPK pathway in NRASMUT AML compared to NRASWT AML. (B) Gene set enrichment analysis of MAPK pathway in NRASMUT AML compared to HC. (C) Phosphorylation levels of MAPK signaling pathway in DUSP1 knockdown NRASG13D KG-1 cells. (D) Overlap of different genes in DUSP1 siRNA KG-1 cells and RASMUT AML patients. (E) The gene list of overlapped common genes. The genes in red indicated that the genes were reported to be chemotherapy-resistant. (F) GO analysis of overlapped differential genes in DUSP1 siRNA KG-1 cells and RASMUT AML.
Immune Interaction and Tumor-Infiltrating Immune Cells Associated With DUSP1
GO analysis showed that innate immune response signaling pathways were enriched both in DUSP1 knockdown KG-1 cells and RASMUT AML patients. To investigate innate immune response signaling pathways in RASMUT AML, a DUSP1 immune PPI network was performed to uncover the role of DUSP1 in TIME (tumor immune microenvironment). The PPI network was composed of 148 immune genes. The platform of Pathway Commons was used to analyze genetic interactions between DUSP1 and immune genes as previously reported. 15 Among those genes, DUSP1 acts directly on immune gene CREB1 and CXCL8 in NRASMUT AML. As shown, some immune related genes were further interacted with CREB1 and CXCL8 (Figure 5A). CIBERSORT is a tool to characterize cell compositions by gene expression profiles. 16 In this study, CIBERSORT was used to assess the links between DUSP1 and tumor-infiltrating immunocytes in RASMUT AML. Among 19 types of immunocytes, the immune infiltrating score of memory B cells, regulatory T cell, follicular helper T cells, and activated NK cells were differently expressed according to the levels of DUSP1 (Figure 5B). Memory B cells and regulatory T cells were highly expressed in AML patient with the lower levels of DUSP1. Follicular helper T cells were highly expressed in AML patient with the higher levels of DUSP1. The matrix, as shown in Figure 5C, indicated the links between different immunocytes in RASMUT AML.

Immune interaction of DUSP1 in RASMUT AML patients. (A) DUSP1-immune PPI networks. (B) Immune infiltrating differences in RASMUT AML patients with high and low DUSP1 levels. (C) Correlation matrix of immunocytes.
Discussion
Our previous research found that nuclear factor erythroid 2-related factor 2 (NRF2) underlies chemo-resistance in malignant hematological diseases. DUSP1 is a target gene of NRF2, which plays an important role against oxidation and DNA damage. 17 Some studies had showed that DUSP1 is over-expressed in many tumors and associated with cancer progression. 18 DUSP1 may serve as a predictive biomarker. With the development of gene sequencing, some genetic mutations with important prognostic significance have been identified and played important roles in guiding clinical treatments of AML. RAS gene mutation had been identified as an activated oncogene in tumors for nearly 40 years. RAS gene mutation, with a frequency of 15% to 40%, was one of the most common genetic alterations in AML patients.19–21 The frequency of NRAS mutations is higher than KRAS mutations in AML. Although the prognostic significance of RAS mutations is inconclusive in AML patients, researches have found that RAS mutation was associated with late relapse and Ara-C resistance. Thus, RAS mutation may serve as a novel therapeutic target for AML.
Our research showed that NRASMUT AML patients had significantly higher DUSP1 expression. It was previously shown that inhibition of Notch signaling could activate DUSP1 in myeloid progenitors of KrasG12D mice. 22 The relation between DUSP1 and all subtypes of RAS mutations in AML, as well as the involved mechanisms warrant further investigation in the future. And further external validation cohort is imperative to be included to enhance the reliability of the TCGA and GEO datasets analysis. Our study indeed showed that AML cells were sensitive to Ara-C by knocking down the expression of DUSP1. DUSP1 downregulation induced phosphorylation of MAPK pathway in NRASG13D AML cells. It has been reported that in BCR/ABL-positive leukemia patients, knocking down C-FOS and DUSP1 at the same time can eliminate TKI-resistant residual leukemia cells by activating p38 activity. 21 In addition, one research in AML patients found that inducible cyclic adenosine monophosphate early repressor could repress DUSP1 and activate the p38 pathway and then enhance caspase-dependent apoptosis, thus enhancing sensitivity of drug in myeloid leukemia. 23 In breast cancer, DNA damage induced the activation of the MAPK signaling pathway, which further induces cell cycle arrest at the G2 phase by affecting the p53 activity. 24 The effect of phosphorylated MAPK signaling pathway on cell cycle in NRASMUT AML with Ara-C treatments needs to be further explored. Some limitations in our study should be mentioned. The effect of Ara-C treatment on DUSP1 downregulated AML cell lines in this study cannot rule out the effect of siRNA induced genetic modifying process. To determine the roles of DUSP1 in chemotherapy, in vivo study of murine AML models with genetic deletion of DUSP1 need to be further investigated. Other analogs of deoxycytidine that resembles Ara-C (such as fludarabine and gemcitabine) should also be further tested in future study.
RNA-sequencing results showed that 71 genes were upregulated both in RASMUT AML patients and DUSP1 downregulated KG-1 cells. GO analysis suggested that the upregulated genes were significantly enriched in 11 pathways pertaining to innate immune responses. Similar as solid tumor, AML also has deregulated innate and adaptive immune responses, induced regulatory T cells and reduced T and natural killer (NK) cell function and cytotoxicity.25–27 All these factors are involved in the induction of immune tolerance and subsequent treatment failure as well as relapse. 28 Our analysis suggested that there were significant differences in memory B cells, regulatory T cells, and NK cell activation between AML patients with low and high expression of DUSP1, indicating that DUSP1 may be involved in the regulation of immune tolerance in AML patients.
Conclusion
Our research found that DUSP1 was upregulated and predicted poor prognosis in AML patients. DUSP1 knockdown could sensitize AML cells to Ara-C treatment. DUSP1 regulated Ara-C sensibility and decreased the phosphorylation levels of MAPK in NRASG13D AML. DUSP1 correlated with immune gene CREB1 and CXCL8. These results indicated that DUSP1 signaling pathway may regulate Ara-C sensitivity in AML. Our study shed lights on the drug resistance mechanisms in AML.
Supplemental Material
sj-pdf-1-tct-10.1177_15330338231207765 - Supplemental material for DUSP1 Signaling Pathway Regulates Cytarabine Sensitivity in Acute Myeloid Leukemia
Supplemental material, sj-pdf-1-tct-10.1177_15330338231207765 for DUSP1 Signaling Pathway Regulates Cytarabine Sensitivity in Acute Myeloid Leukemia by Huali Sun, Yanling Ren, Xinping Zhou, Qi Chen, Yanmei Liu, Chumeng Zhu, Yanyun Ruan, Hongli Ruan, Hongyan Tong, Shenpeng Ying and Peipei Lin in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-2-tct-10.1177_15330338231207765 - Supplemental material for DUSP1 Signaling Pathway Regulates Cytarabine Sensitivity in Acute Myeloid Leukemia
Supplemental material, sj-docx-2-tct-10.1177_15330338231207765 for DUSP1 Signaling Pathway Regulates Cytarabine Sensitivity in Acute Myeloid Leukemia by Huali Sun, Yanling Ren, Xinping Zhou, Qi Chen, Yanmei Liu, Chumeng Zhu, Yanyun Ruan, Hongli Ruan, Hongyan Tong, Shenpeng Ying and Peipei Lin in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Author Contributions
PPL, SPY, and HYT helped to design the study. PPL, HLS, and YLR executed the experiments and wrote the manuscript. XPZ, QC, YML, CMZ, YYR, and HLR provided technical help.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Statement
15 AML patients and 15 controls were enrolled in this study. Patients were selected based on availability of BM specimens at Taizhou Central Hospital. The study was conducted in accordance with the Declaration of Helsinki. Consents were obtained from all patients and controls. This study was approved by the Institutional Research Ethics Committee of Taizhou Central Hospital (TZYXY2019-525).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by grants from National Natural Science Foundation of China (81800121), Zhejiang Provincial Natural Science Foundation of China (LQ20H080003), Medical Health Science and Technology Project of Zhejiang Provincial Health Commission (2020KY1053), Fund of the Science and Technology Department of Taizhou City (1901ky34), Foundation of Taizhou University (2019PY057).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
