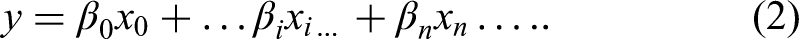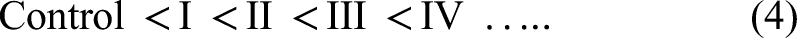Abstract
Early detection of cancers and their precise subtyping are essential to patient stratification and effective cancer management. Data-driven identification of expression biomarkers coupled with microfluidics-based detection shows promise to revolutionize cancer diagnosis and prognosis. MicroRNAs play key roles in cancers and afford detection in tissue and liquid biopsies. In this review, we focus on the microfluidics-based detection of miRNA biomarkers in AI-based models for early-stage cancer subtyping and prognosis. We describe various subclasses of miRNA biomarkers that could be useful in machine-based predictive modeling of cancer staging and progression. Strategies for optimizing the feature space of miRNA biomarkers are necessary to obtain a robust signature panel. This is followed by a discussion of the issues in model construction and validation towards producing Software-as-Medical-Devices (SaMDs). Microfluidic devices could facilitate the multiplexed detection of miRNA biomarker panels, and an overview of the different strategies for designing such microfluidic systems is presented here, with an outline of the detection principles used and the corresponding performance measures. Microfluidics-based profiling of miRNAs coupled with SaMD represent high-performance point-of-care solutions that would aid clinical decision-making and pave the way for accessible precision personalized medicine.
Keywords
Introduction
MicroRNAs (miRNAs) are small non-coding regulatory RNAs that bind to specific target gene transcripts perturbing their correct translation. 1 Given this epigenetic function, the dysregulation of miRNAs is a significant mechanism promoting the development and progression of various cancers. Several factors contribute to miRNA dysregulation, including errors in miRNA biogenesis machinery,2,3 abnormal epigenetic changes,4,5 copy number changes in genes coding miRNAs,6,7 and aberrant transcription factor control of miRNA genes. 8
The dysregulated miRNA drives the progression of cancer,9,10 and based on the association with the specific hallmarks of cancer,
11
it could be classified; namely:
Significant altered expression of miRNAs is frequently reported in the literature for various cancers; for eg, miR-221, miR-301, miR-376a, and miR-21 in pancreatic cancers,23,24 and miR-155 and let-7a-2 in lung cancer.
25
MiRNAs are present in body fluids such as plasma, urine, cerebrospinal fluid, pleural effusions, and colostrum, circulating tumor cells (CTCs), exosomes, and stool26,27; for eg, miR-21 in the serum samples of B cell lymphoma patients.
28
Furthermore miRNAs are stable,29,30 and circulating miRNAs with signal aberrant expression constitute an important class of biomarker candidates for screening, diagnosis, subtyping, and prognosis of cancer. In a recent study, liquid biopsy has been used to derive early-stage miRNA biomarkers for diagnosing esophageal squamous cell carcinoma.
31
Liquid biopsy is a radical non-invasive method for tumor sampling in clinical oncology, and facilitates the simultaneous profiling of multiple biomarkers, paving the way for precise targeted therapy and optimal patient outcomes.
32
Figure 1 shows the different dysregulation classes of miRNAs that could be detected using a liquid biopsy. In the first part of this review, we summarize data-driven approaches for establishing an optimal panel of miRNA biomarkers for the desired objectives.

MiRNAs and cancer hallmarks. Aberrations in miRNA functions are crucial to tumorigenesis and the development of cancer hallmarks. Such miRNAs could serve as early-stage biomarkers detectable in a liquid biopsy.
Though circulating miRNAs make attractive early-stage biomarker candidates, they pose substantial detection challenges due to their very low prevalence (∼0.01% of total RNA in human body), small size (∼20-30 nucleotides), low copy number (∼10 000 copies per cell), and poor sequence differentiation (miRNAs in a given family differing in just one base). 33 In addition, miRNA concentrations between plasma and serum are different, and thus the collection source of circulating miRNAs exerts a critical role. 34 These constraints require the development of detection strategies with single-nucleotide specificity to a minute target RNA population against a background of abundant non-target RNA molecules. Quantitative Reverse Transcriptase Polymerase Chain Reaction (qRT-PCR), next-generation sequencing methods like miRNA-Seq, and Northern blotting constitute standard methods for profiling miRNA transcript abundance.35,36 Emerging techniques for detecting miRNAs include using synthetic DNA and RNA nanostructures with better performance profile.33,37,38 However, a viable alternative technology to miRNA detection and quantification in body fluids would be microfluidics-based strategies 39 that afford real-time point-of-care solutions with high specificity, high sensitivity, low detection limits, quicker response, and less sample volume. Other features of microfluidics devices include advantages in cost, time, and user-friendliness, which help in fostering the adoption of such devices in the development of biomarker-based analytical diagnostics.40,41 In the latter section of this review, we discuss different options for incorporating microfluidic strategies towards profiling an identified miRNA biomarker panel.
AI Assisted miRNA Biomarker Panel Discovery
miRNA Biomarker Discovery
Genome-wide –omics big data have marked a paradigm shift in the process of biomarker discovery from the traditional hypothesis-driven knowledge-based approach to a hypothesis-agnostic data-driven modern approach. The data-driven approach is systematic, ideally unbiased (up to assumptions in experimental design), and cost-efficient with respect to the identification of biomarkers optimal for a phenotype of interest. It must be noted that bias in experimental design will induce an implicit hypothesis in the collected data. Well-known public repositories of big omics data include the Cancer Genome Atlas (TCGA), 42 the Gene Expression Omnibus (GEO), 43 and the International Cancer Genome Consortium (ICGC). 44 TCGA houses multiple omics big data (including miRNA-Seq) with > 20 000 cancer and matched normal samples for over 33 cancer types. GEO is a global repository supported by the National Center of Biotechnology Information (NCBI) that archives high-throughput gene expression and genomics data. ICGC is an international cancer research collaboration for sharing –omics data pertaining to nearly 86 cancer types. Dedicated databases of miRNA expression profiles include miRmine, 45 DIANA-miTED, 46 TissueAtlas, 47 and Chinese Glioma Genome Atlas. 48 Other databases supporting the investigation of miRNA roles in cancers include dbDEMC, 49 miRCancer, 50 miR2Trait, 51 and CancerMIRNome. 52
MiRNA-Seq –omics data could be obtained using a variation of RNA-Seq, where larger RNA fragments (mRNAs and rRNAs) are initially excluded by size selection, read counts obtained, and transformed to an expression matrix. Differentially expressed miRNAs could be found from the miRNA-Seq –omics expression matrix using a log fold change between the cancer samples and the controls, along with a significance test. More complex workflows could yield specific DE miRNAs pertaining to some clinical metadata of interest like clinical or pathological stage, survival status, response to treatment, etc. The clinical annotation of the samples may be used to design regression models that could yield specific biomarkers containing predictive utility with reference to the classes in the clinical annotation.
Classes of miRNA Biomarkers
A prime interest in cancer research is the identification of early-stage biomarkers. Below, we illustrate the use of clinical annotation in the form of AJCC staging to identify early-stage biomarkers. Our two-tiered protocol has been described earlier53,54 in summary:
(i) Tier-I: The miRNA features specific to a clinical variable can be identified by using a two tier contrast that was developed in our lab. A contrast between the controls on the one hand and each class in the clinical variable of interest on the other could identify class-specific miRNA biomarkers. In the case of AJCC staging data, this contrast is performed using the following design matrix: (ii) Tier-II: Contrast between the various classes of the clinical variable of interest could identify class-salient biomarkers. This between-class analysis is performed for all pairs of classes, using the results from the following design matrix:
The design matrix of contrasts is then applied to find the log fold change and associated P-value for each contrast in both tier-I and tier-II. This could be done by supplying the contrast matrix as an argument to the contrastsFit function in limma. The contrast against the control demands a stronger significance (adj. P < .001) than the pairwise contrasts (P < .05). Using an |lfc| threshold ∼ 2 for each contrast, tier-I constraints yield significant upregulated or downregulated DE miRNAs. Application of tier-II constraints to the stage-specific miRNAs (from tier-I) yields stage-salient miRNAs.
In addition, an ordinal model could be used to model miRNA expression, treating the cancer stage as a numeric variable. This is given by the following equation:
Classes of miRNA Biomarkers That Could Be Useful in Building AI-Based Cancer Diagnosis and Prognosis Models.

Construction of ML Model
The identified biomarkers could be used in the construction of an ML model for screening, early-stage diagnosis, and subtyping purposes. Machine learning models are generally characterized by the {T,P,E} formalism, where they learn from experience E of –omics expression data with respect to the class of tasks T of interest, improving on some performance measure P.55,56 The generalized workflow for building such models for oncology applications is presented in Figure 2. The response variable of interest is extracted from clinical metadata, and appended to the miRNA expression data matrix. In a pre-processing step, miRNAs with minimal variance in expression (expression σ < 1) and samples with missing values in the clinical variable of interest are removed. The dataset is log2-transformed through voom, which stabilizes the variance in expression in the remaining genes. 57 The dataset is split into train dataset and test dataset, and biomarker identification is performed with the train dataset to prevent leakage into the test dataset. Potential biomarkers could belong to one of the classes enumerated in Table 1, especially stage-salient and ME miRNAs. Feature selection is then performed to optimize the feature space up to 16 biomarkers (usually three or four) for detection with multiplexed microfluidics. This may be done using techniques like Recursive Feature Elimination (RFE), 58 which uses backward variable selection, and Boruta wrapper algorithm, 59 which uses top-down elimination of features irrelevant to the learning problem. The consensus between the feature selection techniques could be used as the final feature space for ML model development. A wide range of algorithms that work with tabular data could be used for model construction, including SVM, 60 neural networks (shallow and deep), 61 and ensemble methods such as Random Forest 62 and XGBoost. 63 Model hyperparameters may be optimized with ten-fold cross-validation on the train dataset using libraries such as caret 64 and KerasTuner. 65 The optimized models are evaluated on the unseen test set and the best model then determined. The performance of the best model may be assessed on an external out-of-domain dataset. Comprehensive information on developing machine intelligence in oncology could be found in contemporary reviews of the state of the art.66–71 Successful models represent a Software-as-Medical-Device (SaMD) endpoint 72 that could then be refined with real-time clinical data for continuous deployment and integration. Examples of such models built using miRNA biomarkers have been recently reported.73–75

A general workflow for the development of an ML model with miRNA biomarker features for varied diagnostic applications. The model is created with miRNA –omics data and could lead to the production of SaMD.
Construction of Prognostic Model
The development of prognostic models offers another application for miRNA biomarkers to improve the practice of medicine. Prognostic models are combinations of predictor variables that score the risk of encountering a well-defined endpoint, like death or relapse of disease, in a specified period for individuals with a given state of health.76–78 Parametric techniques to model time-oriented data include logistic regression, Kaplan–Meier curves, and Cox proportional hazards models. Neural networks represent an emerging class of nonlinear nonparametric prognostic models.
79
Prognostic models could be calibrated and evaluated for discrimination between patients in two groups in a given period of time using the concordance index.80,81 A generalized statistical survival analysis for developing a prognostic model based on miRNA biomarkers would involve the following steps (Figure 3):
constructing univariate Cox regression models for each biomarker of interest, including stage-salient and ME miRNAs, and identifying the prognosis-significant miRNAs (P < .05) applying robust feature selection methods such as LASSO cox regression or SVM-RFE to reduce the dimensionality constructing a multivariate Cox regression with backward variable selection to arrive at an optimal prognostic signature of three to four miRNAs. This step yields an expression for the risk score in terms of the optimal biomarker panel:

Microfluidic Systems for Profiling miRNA Signatures
The miRNA biomarkers comprising the final diagnostic or prognostic final models could be detected and quantified using microfluidic-based multiplexed technologies; Figure 4 summarizes a scheme for translating biomarker discovery into a microfluidics-based SaMD. Microfluidics platforms are combinations of fluidic operating units including channels (1-1000 mm in size), chambers, and nanoscale structures like pillars, wires, and tubes. They serve to integrate, automate, multiplex, and thereby streamline biochemical processes, offering significant advantages in performance and cost. Microfluidics have been used to study the unique “fluidic signatures” of cancer, including growth, metastases,85,86 induction of angiogenesis,87,88 and the tumor microenvironment.87,88 In particular, they have been used to study CTCs, which carry essential cancer biomarkers but possess short half-life, posing a detection challenge that is overcome with the use of probes such as aptamers, peptides, immune-affinity proteins, and cytokines.35,37

A schematic for the development of a prognostic model with miRNA biomarker features. The model is created with miRNA –omics data and could lead to the production of SaMD.

Microfluidic detection of miRNA biomarkers. A scheme for developing and deploying the microfluidics-based SaMD is shown.
Microfluidics-based devices are perfectly suited for converting miRNA biomarker signatures of cancers into clinical outcomes such as screening, diagnosis, and prognosis. These devices could be characterized in terms of the design of the microfluidic platform as well as the detection principle used. Platform designs could be based on nanomaterials, pico-droplets, lab-on-a-chip, and paper-based analytical devices (μPADs). The detection methods could be varied and encompass optical, electrochemical, and conventional PCR readouts. Table 2 summarizes various microfluidics-based devices that have been used for detecting miRNA cancer biomarkers, along with their respective detection principles.
A Classification of Microfluidics Strategies by Their Detection Principle. The Specific Biomarkers Identified and the Performance (if Available) of the Respective Strategies are Noted, Along with the Primary Reference Source. Performance is Given in Terms of Limit of Detection (LOD), Analysis or Detection Time, Sample Volume, Sensitivity, Specificity, and/or Linear Detection Range

The optical detection methods include Surface Enhanced Raman Spectroscopy (SERS), 109 Surface Plasmon Resonance (SPR), 110 upconversion nanoparticles, 111 fluorimetry, Förster resonance energy transfer (FRET), 112 and colorimetry. 113 All these methods share very high sensitivity relative to other detection principles. SERS achieves enhancement of the regular Raman spectroscopy signal of analytes (including single-molecule miRNAs) by a factor of ∼ 1010-1011, either directly via adsorption onto metal nanostructured surfaces or indirectly via a reporter mechanism, and with a spectral profile tailored for multiplexing and stable quantitative readouts. 114 SPR refers to the collective coherent oscillations of excited delocalized electrons on a thin metal surface due to light incident at a particular angle to the surface. The specific angle of incidence is a function of the characteristic target such as miRNAs, leading to differential detectable signals. 115 SPR and SERS display the lowest detection limits (< 1 aM miRNAs); however, these techniques pose challenges in scaling and reproducibility. Upconversion nanoparticles absorb low-energy photons and emit high-energy fluorescence post target capture, with an ability to detect 0.1 nM, but remain limited by conversion efficiency. Fluorescence-based detection remains the most sought-after readout method, due to the ease of detection and sensitivity. Laser-induced fluorescence (LIF) is a special type of fluorescence detection method with an ultra-sensitive detector, and works with samples of different dimensions and states (solid, liquid, gas). Further, the LIF instrument is small, cheap, and portable making it the ideal detector for microfluidic paper-based analytical devices (μPADs). 109 μPADs are microfluidics-based systems that predominantly make use of fluorescence and colorimetric detection principles. μPADs are advantageous for numerous reasons: one, in terms of microfluidics, fluid flow could be easily controlled by capillary forces on them, ruling out the requirement of any external driving forces; two, the cellulose paper substrate remains compatible to carry out various chemical and biochemical reactions and furnishes passive liquid transfer owing to its hydrophilicity, porosity, and homogeneous structure; three, PADs are highly disposable due to their low sample consumption, cheap cost, and low weight; four, paper allows for preserving various biomolecular reagents like antibodies, enzymes, nucleic acids, and cells by simple freeze drying of the storing areas.36,112 In the work of 96, LIF was used for the detection of miR-21 and miR-31. 97 Nanomaterial-based designs utilize colorimetric properties of gold nanoparticles (AuNPs). Two-dimensional nanomaterials like graphene oxide, MoS2, 110 and WS2 111 have been employed for miRNA capture, leading to a readout based on FRET. FRET refers to the non-radiative energy transfer within a light-sensitive donor–acceptor molecular couple, and is extremely sensitive to extremely small changes in donor–acceptor distance, their relative orientation, and the lifetime of the donor excited state. Complementary techniques for miRNA capture could be used: (i) the target miRNA binds to a complementary RNA sequence, which is coupled to a FRET pair and immobilized in a micro-channel, thus giving rise to a fluorescence signal 89 ; (ii) the target miRNA acts to separate the donor and acceptor fluorophores in a FRET pair, resulting in fluorescence quenching; and (iii) two different oligonucleotide probes, with donor and acceptor fluorophore dyes, hybridize to adjacent regions of the target miRNA, yielding the FRET signal of proximity of fluorophore dyes. 116 Despite the gains in sensitivity and reliability, the clinical utility of colorimetric and FRET-based microfluidic devices could be circumscribed by the requirement that each target miRNA have its own detection element.
Electrochemical detection provides highly sensitive detection with quick response times. 113 The use of microelectrodes relaxes the necessary analyte volume, and the electrochemical system directly transduces the biological process into an output signal. Thus, by bringing the best of both the worlds of microfluidic systems and electrochemical readout systems, diverse strategies for biomarker detection have been invented (Table 2). For instance, Sun et al coupled μPADs with electrochemical readout, designing a microfluidic device with (i) gold (Au) nanorods for improving the conductivity of the cellulose paper substrate; and (ii) cerium dioxide—Au conjugated with glucose oxidase enzyme for serving as the electrochemical probe. 104 The synergistic composite of cerium dioxide—Au conjugated with glucose oxidase enzyme helps in improving the device's sensitivity and specificity.
In addition to the above, microfluidics-based strategies offer refinements to the conventional PCR-based detection method, overcoming inefficiencies in heat transfer and reaction times. Microfluidics afford controllable thermal cycling, which could be used in droplet-based PCR systems for detecting miRNA biomarkers. 108 Droplet-based microfluidic systems have been used for nucleic acid amplification-free single-molecule RNA detection, via Cas13a assays with fluorescence readout. 117 Such devices could be used to count cell-free miRNA biomarkers from serum samples, which could be used for cancer screening applications (for eg,[ 34 ]). New directions for microfluidics-based strategies include the demonstration of loop mediated isothermal amplification (LAMP) followed by fluorescence measurement for multiplexed detection of multiple strains of pathogen DNA. 118 DNA markers have been shown effective in multicancer early detection tests based on liquid biopsies, 119 a result that invites further research into mining the omics biomarkers driving cancer hallmarks.
In summary, the key drivers for adopting microfluidic systems for miRNA detection include: (i) modular nature, with the freedom to mix and match the detection platform and readout physics; (ii) requirement of low sample volume; (iii) affordance of spatiotemporal control; and (iv) multiplexed sub-pM sensitivity, all of which help in the design of efficient portable high-performance systems for the automated detection of miRNA signatures of cancer. On the other hand, some major limitations include: (i) charge-based or affinity-based non-specific binding of detection probes (eg, to device surfaces), and fouling of microchannels, especially in the case of nanoparticle/polymer-based microfluidic detection systems; (ii) lack of unified protocols and standardized benchmarks, raising quality control issues that frustrate clinical validation; (iii) high costs of development and clinical trials, especially in the case of diagnostics in oncology; and (iv) regulatory compliance and market adoption issues that tend to plague novel maturing technologies, especially pronounced in the case of diagnostic medical devices. 120 Most of these challenges are common to emerging high-throughput technologies, and microfluidics-based SaMD-coupled miRNA profiling could be no exception. Potential drawbacks notwithstanding, continued research of microfluidics-based devices would overcome the obstacles, and deliver increased optimization and adoption of such quantitative diagnostics.
In conclusion, miRNAs are an effective and compelling class of biomarkers for the early detection of cancers towards clinical management of these deadly killers. AI-based methods enable the discovery of early-stage miRNA signatures for cancer screening, diagnosis, and prognosis, yielding SaMDs that could be deployed in conjunction with microfluidic sensing platforms for delivering real-time diagnostic and prognostic information. The power and precision of these systems is likely to drive their increasing currency in cancer diagnosis especially as a point-of-care solution in resource-constrained settings.
Footnotes
Abbreviations
Acknowledgments
We would like to thank Profs. K.S. Rajan and R. John Bosco Balaguru for helpful discussions. We are grateful to the management of SASTRA Deemed University for infrastructure and support. We would like to thank reviewers for the careful reading of our manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.