Abstract
Introduction
Over the past decade, with the development of imaging technologies and efficient learning strategies, radiomics has emerged and is widely accepted in precision medicine by extracting mineable high-dimensional features from multimodal medical images, for example, computed tomography (CT), positron emission tomography (PET), MRI, or ultrasound (US).1,2 Studies demonstrated that radiomics features could characterize the internal structure of tissue—such as tumor heterogeneity to enhance the diagnostic, prognostic, and therapeutic outcomes for a broad range of diseases.3,4 Studies also indicated that with advanced analysis of images using artificial intelligence, radiomics could correlate imaging phenotypes with genomic and proteomic signatures,5,6 and subsequently complement the information of tissue sampling and circulating biomarkers to reform clinical decision-making.7,8
However, one major shortcoming of radiomics is that features could be influenced by many factors, such as the type of scanner, imaging settings, reconstruction parameters, delineation of the tumor, etc,9–11 which results in a low reproducibility of radiomics features and makes it difficult to compare and interpret radiomics studies for clinical application.9,10 Standardization in imaging acquisition and analysis had been proposed to improve feature stability, but these standardization approaches are not always feasible as most radiomics studies are retrospective in nature. 3 Awareness of the shortcomings of the radiomics features and studies is of critical significance for the feasibility and applicability of these radiomics studies.
Thanks to its advantages of nonionizing radiation, portability, accessibility, and cost-effectiveness, US has been one of the most used imaging modalities for screening and diagnosis in obstetrics and gynecology. 12 Radiomics studies indicated that US radiomics features were highly associated with breast biologic characteristics, 13 gestational age, 14 neonatal respiratory morbidity, 15 etc. Studies also demonstrated that US-based radiomics was able to predict lymph node metastasis (LNM) for patients with papillary thyroid carcinoma and cervical cancer (CC).16,17 However, US images also suffer from some limitations, such as low imaging quality caused by noise and artifacts, high inter- and intra-observe variability across different scanners and institutes, and highly operator or diagnostician experience dependent, which will hinder the clinical application of US-based radiomics models. 18 The purpose of this study is to investigate the influence of different US scanners on the reproducibility of radiomics features and the accuracy of LNM prediction for patients with CC.
Materials and Methods
Patients
By searching electronic medical records, a total of 1723 CC patients underwent radical hysterectomy and pelvic lymphadenectomy in our hospital between January 2014 and November 2018 were retrospectively reviewed. Inclusion criteria: (a) patients should have standard ultrasonography within 2 weeks before hysterectomy and (b) patients had confirmed histological characteristics and lymph node status after the operation. Exclusion criteria: (a) patients with incomplete or incorrect clinical data, (b) patients who had other malignant tumors, (c) patients who were treated by chemotherapy or radiotherapy before the operation, and (d) patients with unclear or missing images. A total of 536 patients were enrolled, including 148 with HDI5000, 75 cases of Voluson E8, 100 cases of MyLab classC, 110 cases of ACUSON S2000, and 103 cases of HI VISION Preirus. This retrospective study was conducted following the Declaration of Helsinki and approved by the Ethics Committee in Clinical Research (ECCR) of Wenzhou Medical University First Affiliated Hospital (ECCR No. 2019059). The written informed consent was waived by the ECCR due to the retrospective nature of this study with confirmation of following the Declaration of Helsinki with patient data confidentiality. Besides, all of the patient details in this study have been anonymized. The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guideline. 19
US Imaging and Machines
Transvaginal US images were acquired when patients lie in a lithotomy position with an empty bladder. In this study, the following 5-color Doppler ultrasonic machines were used to acquire the US images: ATL HDI 5000(Philips)using the transducer C8-4v at 4-8 MHz; Voluson-E8 (GE Healthcare) using the transducer RIC5-9-D at 5-9 MHz; Mylab classC (Esaote) using the transducer EC1123 at 3-9 MHz; ACUSON S2000 (Siemens) using the transducer MC9-4 at 1.5-6.0 MHz and HI VISION Preirus (Hitachi Ltd) using the transducer EUP-U531 at 4-8 MHz. All images were stored in PNG format and archived in the hospital's DICOM system.
Segmentation and Feature Extraction
Usually, each patient will have 10 to 20 standard US images collected. Regions of interest were delineated manually by a radiologist with 10 years of experience in gynecological US diagnosis using the LIFEx package (http://www.lifexsoft.org), 20 and confirmed by another senior radiologist with 15 years of experience in gynecological US diagnosis. A typical transvaginal US image and contoured target volume are shown in Figure 1. Image preprocessing was performed before feature extraction. A total of 449 radiomics features were extracted from the contoured volumes after intensity normalization using Pyradiomics package (PyRadiomics, https://www.python.org), which includes shape features, first-order histogram statistics, gray-level co-occurrence matrix (GLCM), neighborhood gray-level different matrix, gray-level run length matrix (GLRLM), gray level size zone matrix (GLSZM), and wavelet filtered first-order, GLCM, gray-level different matrix (GLDM), GLRLM, and GLSZM, respectively.21,22

A typical transvaginal ultrasound image with contoured target volume.
Feature Selection and Model Building
Optimal features correlated with LNM were further selected with a 2-step method in the training cohort. First, the Mann-Whitney U test was used to select the radiomics features which had a potential correlation with LNM with a significant p-value <.05. Second, the least absolute shrinkage and selection operator was applied to select the optimal features with 5-fold cross-validation to adjust the elastic network parameters to select the best combination of feature parameters to avoid overfitting. The dependent variables in the study are dichotomous and we use the parameters family="binomial”, α=1, nlambda=100, type.measure=“auc.”</div><div class="p" xmltag="p">To avoid overfitting the data and achieve stable and superior performance, radiomics models for different machines were built with 2 machine learning classifiers: support vector machine (SVM) and logical regression, which are currently used most commonly in radiomics research studies. The DeLong test was adopted to compare the discrimination performances of differential radiomics models. A total of 10 models were built with radiomics features from 5 different ultrasonic machines. The receiver operating characteristic (ROC) curves with area under the curves (AUCs) were applied to evaluate the maximum diagnostic effect of these 10 models for the prediction of LNM for patients with CC in both the training and validation cohorts.
Statistical Analysis
Statistical analyses were performed with the R analysis platform (version 4.0.4, MathSoft, http://www. Rproject.org) and SPSS 19.0. The selection of key features and logistic regression model building was performed with the “glmnet” software package. SVM model building with the confusion matrix was performed using the “caret” package in the “e1071” software package. For all tests, a p<.05 was considered statistically significant.
Result
A total of 536 CC patients with confirmed histological characteristics and lymph node status after radical hysterectomy and pelvic lymphadenectomy were enrolled in this study. Patients were randomly divided into training and validation cohorts for each US imaging machine at a ratio close to 7:3. There were 148 patients (102:46) scanned in machine HDI5000, 75 patients (53:22) in machine Voluson E8, 100 patients (69:31) in machine MyLab classC, 110 patients (76:34) in machine ACUSON S2000, and 103 patients (73:30) in machine HI VISION Preirus, respectively. The flowchart is shown in Figure 2. Detailed characteristics of enrolled total patients and patients scanned with different machines are presented in Table 1.

The flow chart of patient selection
The Characteristics of Enrolled Patients in the Training and Validation Data Sets.

Abbreviations: LNM, lymph node metastasis; −, negative; +, positive; SD, standard deviation.
Notes: (1) p-value is calculated from the univariate association test between subgroups; (2) Fisher's exact test and chi-square test were used for categorized variables; (3) 2-sample t-test was used for continuous variables.
Figure 3 shows the selection of optimal radiomics for LNM prediction for different machines. Based on the elastic-net method through tuning the parameter (λ) with a 5-fold cross-validation via maximum AUCs in the training cohorts, radiomics features with nonzero coefficients were selected. Eventually, 4 radiomics features with potential correlation with LNM were selected from machine HDI 5000, 3 features from machine Voluson E8, 6 features from machine MyLab classC, 5 features from machine ACUSON S2000, and 5 features from machine HI VISION Preirus, respectively. Details of the selected radiomics features are presented in Table 2.

The selection of optimal radiomics for lymph node metastasis prediction based on the elastic-net method through tuning the parameter (λ) with a 5-fold cross-validation via maximum area under the curves: (a, b) for machine HDI 5000; (c, d) for machine Voluson E8; (e, f) for machine MyLab classC; (g, h) for machine ACUSON S2000; (i, j) for machine HI VISION Preirus; and (k, l) for all the machines.
List of Selected Radiomics Features with Potential Correlation with Lymph Node Metastasis for Patients with Cervical Cancer from Different Ultrasound Machines.

The performance of LNM prediction models with radiomics features from different machines is presented in Figure 4. The AUCs ranged from 0.75 to 0.86 and 0.73 to 0.86 in the training cohorts and from 0.71 to 0.82 and 0.70 to 0.80 in the validation cohorts for SVM and logistic regression models, respectively. Detailed performance of these models for different machines and total patients are shown in Table 3. DeLong test found that the performances of differential radiomics models were not significantly different (both, p>.05), as shown in Tables 4 and 5.

The performance of radiomics models with ultrasound images from different ultrasonic machines (a, b) using support vector machine and (c, d) using logistic regression with training and validation cohorts, respectively.
The Performance of SVM and Logistic Regression Models with Radiomics Features from Different Machines for the Training and Validation Cohorts.
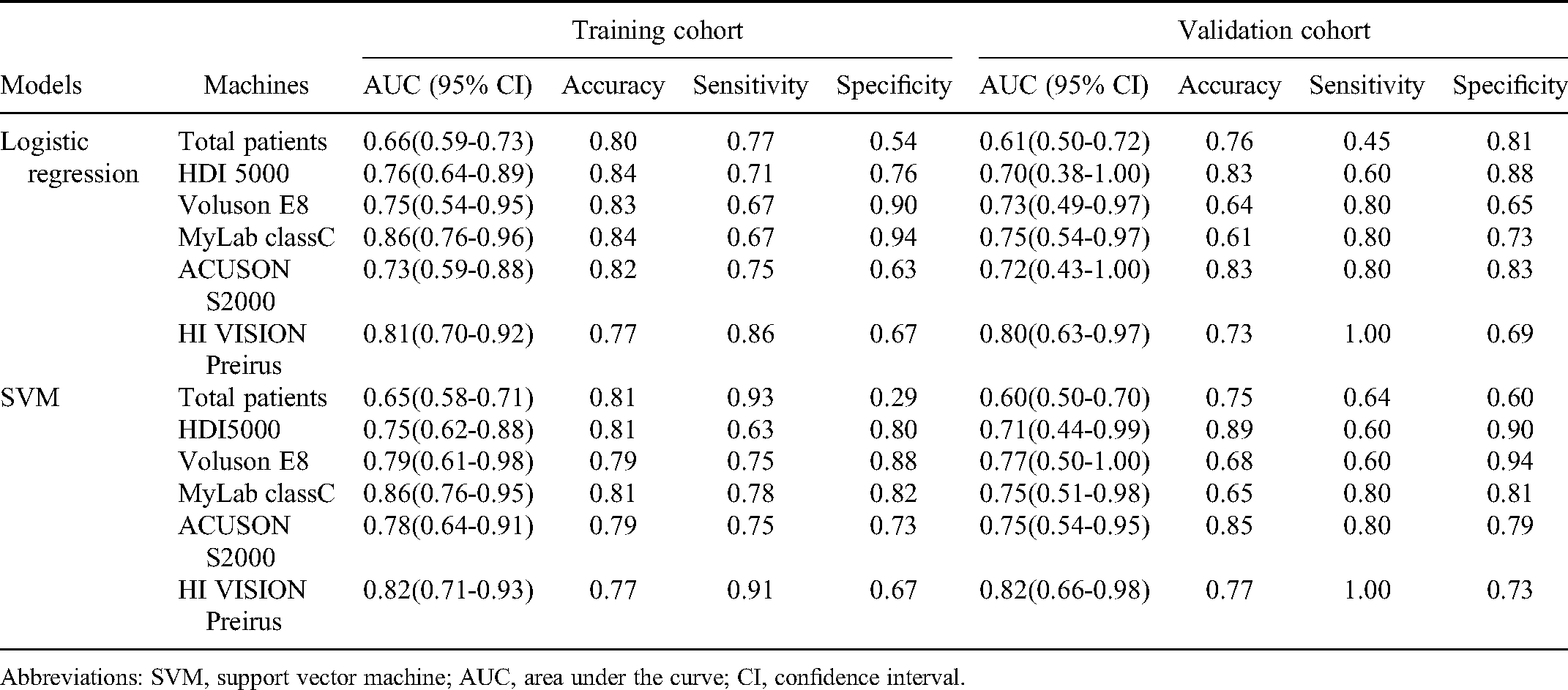
Abbreviations: SVM, support vector machine; AUC, area under the curve; CI, confidence interval.
The DeLong Test of Support Vector Machine Models from Different Machines for Validation Cohorts.

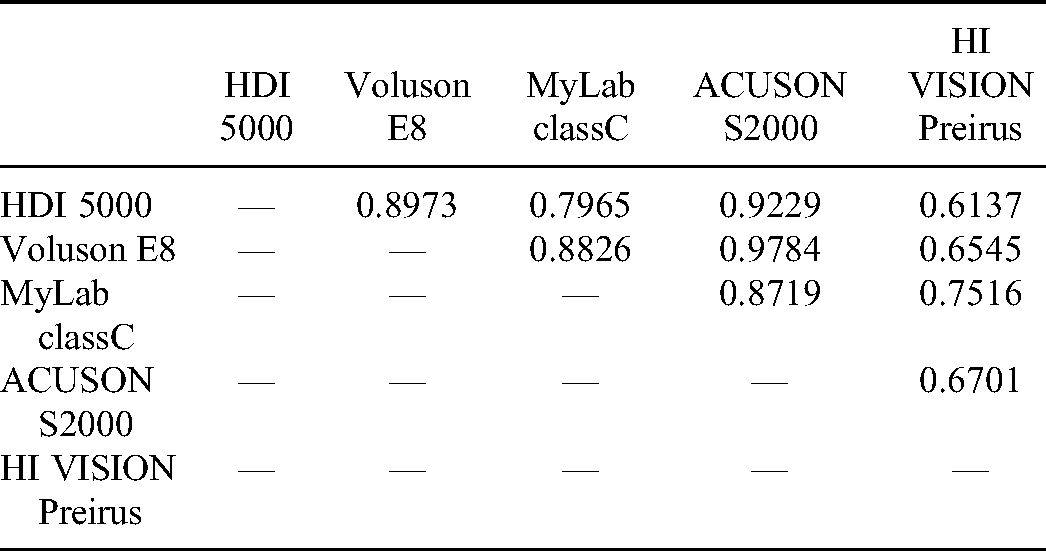
The DeLong Test of Logistic Regression Models from Different Machines for Validation Cohorts.
Discussion
LNM is one of the most important risk factors for recurrence and survival for patients with CC.23,24 Undiagnosed or inaccurately assessed LNM is a major cause of suboptimal treatment. For instance, 90% of routinely resected lymph modes are not metastatic, and unnecessary pelvic lymphadenectomy leads to complications, such as prolonged surgery, blood loss, infection, nerve or vascular injury, lymphocyst formation, etc.25,26 Therefore, an accurate noninvasive technique for lymph node assessment is vital in the clinical management of patients with cervical. A noninvasive LNM prediction method with radiomics has been intensively investigated for CC preoperatively with MRI,27,28 18F-fluorodeoxyglucose PET/CT,29,30 as well as US images. 17 In this study, the effects of US machines on the reproducibility of radiomics features and radiomics models for the prediction of LNM for patients with CC were investigated.
A noninvasive LNM prediction method using radiomics is of critical clinical benefit in the management of CC. Jin et al 17 reported an AUC of 0.79 and 0.77 in the training and validation cohorts with a US images-based radiomics model from one Philips machine (IU22) in predicting LNM for early-stage CC preoperatively. Our study demonstrated similar AUCs around 0.70 to 0.82 for radiomics models with US images from different machines. However, the highest difference in AUCs for different machines reaches 17.8% and 15.5% in the training and validation cohorts, respectively, as shown in Figure 4.
As shown in Table 2, the selected optimal features from different US machines were variable. Few radiomics features were reproducible among different machines. Only features of wavelet.HL_glszm_LargeAreaLowGrayLevelEmphasis repeated in 3 machines (HDI 5000, ACUSON S2000, and HI VISION Preirus) and wavelet.LH_glszm_SmallAreaHighGrayLevelEmphasis repeated in 3 machines (Voluson E8, MyLab classC, and HI VISION Preirus). This indicated that radiomic features from US images are highly scanner-dependent. Previously, Yasaka et al 31 demonstrated that radiomics features from CT images were also scanner-dependent based on a phantom study due to differences in imaging acquisition parameters and the scanner design. Mackin et al 32 indicated that the variation of radiomic features among different CT scanners was similar to their variation in CT images for 20 nonsmall cell lung cancer patients. Li et al 33 assessed the reproducibility of radiomics features from different US scanners, acquisition parameters, segmentation location, and extraction platforms, and also found that the wavelet features showed the best reproducibility among different scanners. The problem of scanner dependency on radiomics features should be considered, and their effects should be minimized in future studies for US images.
Table 3 also demonstrated that radiomics models with combined features from different machines performed inferior to models with features from 1 machine only. This is also a dilemma for current radiomics studies. A large cohort of patients from multiple centers is needed in order to generate convincing results to transfer radiomics as a prognostic tool for potential clinical application.34,35 However, data from multiple centers is unavoidable with variability in the scanner, models, acquisition protocols, reconstruction settings, etc, which further affects the reproducibility of radiomics features and models.9,36 As is similarly shown in this study, combined radiomics features from multiple US machines are inferior to the performance of radiomics models. By using the same image acquisition parameters, reconstruction technique, image segmentation, and method of feature extraction, the radiomics features repeatability problem caused by different scanners would be solved. Although harmonizing images and features had been aggressively investigated, there is much more work needed to realize the potential clinical value of radiomics in the future.37,38 Application of deep learning to harmonize US images from different scanners would be a solution for future radiomics studies.
There were several limitations in our study. First, lack of power analysis for estimation of sample size. Second, radiomics feature reliability assessed by the intraclass correlation coefficient was not considered in this study. Finally, the amount of dataset in several scanners is inadequate. Commonly, a larger amount of dataset will improve the confidence and performance of our model.
Conclusion
The effects of different US machines on the performance of radiomics models based on US images in the prediction of LNM status for CC preoperatively were investigated. The optimal features for prediction models were scanner dependent. The maximum AUC differences for models with images from the different scanners were 17.8% and 15.5% in the training and validation cohorts, respectively.
Footnotes
Acknowledgments
My sincere thanks to everyone who worked on this paper.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by funding from Wenzhou Municipal Science and Technology Bureau (Y20190183), Zhejiang Engineering Research Center of Intelligent Medicine (2016E10011), and Science and Technology Bureau (Y2020917).
Ethics Statement
The ECCR of the authors’ hospital approved this retrospective study (ECCR no. 2019059). The written informed consent was waived by the ECCR due to the retrospective nature of this study with confirmation of following the Declaration of Helsinki with patient data confidentiality.
