Abstract
Introduction
Soft tissue sarcoma (STS) is a group of highly heterogeneous malignant tumors originating from non-epithelial extraskeletal tissues. 1 With an incidence of 2.38/100 000, STS accounts for approximately 0.8% of all cancers in China. 2 STS arises most commonly in the extremities, followed by the retroperitoneum, trunk, head, and neck.3,4 Depending on its morphology and biological behavior, STS can be classified into more than 50 subtypes. 1 The most common isoforms include undifferentiated polymorphic sarcoma (UPS), liposarcoma, leiomyosarcoma, and synovial sarcoma.4–6 At present, the main treatment for early STS is surgery, while anthracycline chemotherapeutic agents such as adriamycin and isocyclophosphamide are the standard care for advanced STS patients.7–9 There is no standard treatment after the failure of first-line chemotherapy.
The emergence of targeted therapy and immunotherapy has revolutionized cancer therapy. In recent years, the investigation of novel sensitive biomarkers such as MDM2 and tumor mutational burden (TMB) for predicting the efficacy of targeted therapy or immunotherapy has been of great interest in the field of oncology.10,11 Adequate research into the molecular profile of STSs is critical to the development of more effective diagnostic tests and therapeutic approaches. Significant advances in next generation sequencing (NGS) technology have reduced the cost of testing and facilitated comprehensive screening of different genetic alterations to provide reliable and effective treatment options for individualized therapies. In recent years, there have been a number of genomic profiling studies of STS based on NGS. Studies have found that the most commonly mutated genes in 2539 cases of STS were TP53 (26.3%) and BRCA2 (16.7%). 12 Similarly, in studies such as Xu et al, TP53 (27.94%), PIK3C2G (22.06%), NCOR1(20.59%), and KRAS (20.59%) were the most frequent mutations in Chinese patients with STS who underwent surgical resection and received conventional adjuvant therapy. 13 Lucchesi et al performed high-throughput sequencing to study the molecular characteristics of 584 STS patients and found that the 10 genes most often altered were TP53, MDM2, CDK4, RB1, ATRX, CDKN2A, PTEN, NF1, CDKN2B, and KMT2D. Further analysis revealed that 41% of patients carried at least one clinically targeted genetic alteration. 14
The vast majority of the previous studies have been carried out in Western populations with differences in the distribution of STS subtypes in China. Besides, the studies in Chinese groups have largely used small panel molecular assays, so the extensive genetic analysis in the Chinese population is still lacking. In addition, there is little clinical evidence on whether patients with STS can benefit from NGS testing. To determine whether genomic profiling could improve diagnostic precision, inform prognosis, or aid in therapeutic selection, we used the NGS platform to characterize the comprehensive genomic profile of 65 Chinese patients with STS and assess whether patients with potentially actionable alterations achieved clinical benefit.
Materials and Methods
Patients
A total of 65 Chinese STS patients who received surgery or biopsy at Zhejiang Cancer Hospital from August 2017 to March 2020 were enrolled in this study. All patients were diagnosed via pathological, imaging, and clinical findings. Both formalin-fixed paraffin-embedded (FFPE) tumor tissues and matched blood samples from these 65 Chinese patients were collected. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Institutional Review Board of Zhejiang Cancer Hospital (registration number: IRB-2021-402), and individual consent for this retrospective analysis was waived, patients below 18 years old were provided with signed informed consent by their legal parents/legal guardians for the research use of clinical data and publication.
DNA Isolation and Targeted Sequencing
Genomic DNA of tumor samples and paired white blood cells was extracted from the FFPE and blood samples using QIAamp DNA FFPE Tissue Kit (Qiagen, 180134) and QIAamp DNA Blood Mini Kit (Qiagen, 51306) according to the manufacturer's instructions, and fragmented for constructing a library using KAPA Hyper Prep kits (KAPA, KK8504). Fragments 200 to 400 bp from the sheared tissue DNA were purified (Agencourt AMPure XP Kit, Beckman Coulter). Prepared libraries (600 ng) were hybridized with 2 different hybridization reagents and blocking agents in the SureSelectXT Target Enrichment System (Agilent Technologies). The enriched libraries were amplified with the P5/P7 primer. After qualification by a 2200 Bioanalyzer and quantification with a Qbit3 and a QPCR NGS library quantification kit (Agilent Technologies), the libraries were sequenced on NovaSeq6000 platform (Illumina) with paired-end reads and a mean coverage depth of greater than 700 × for 825 cancer-related genes (Genetron Health).
Bioinformatics Analysis
Sequence data processing: Primary processing of NGS data for tissue samples was performed using Trimmomatic methods (0.36), including demultiplexing and masking of dual-index adaptor sequences. Sequence reads were aligned against the human reference genome (version GRCh37/hg19) using BWA (version 0.7.10). Variation identification and annotation: Candidate somatic mutations, consisting of point mutations, small insertions, and deletions, were identified by SAMtools (version 1.3.1) and Pindel (version 0.2.5b8, 20151210) across the targeted regions of interest. Somatic copy number variants (CNVs) were called using Control-FREEC (version 10.5) with default parameters. DNA translocation analysis was performed by using Tophat2 (http://ccb.jhu.edu/software/tophat/index.shtml). Effects of variants were annotated using Variant Effect Predictor (version 83, parameters “–everything, –fork 8, –buffer_size 1000”). Each somatic variant was manually confirmed using Integrative Genomics Viewer (version 2.3.34). TMB was calculated in non-synonymous somatic mutations including single-nucleotide variants (SNVs) and INDELs in coding regions per megabase.
Pathway Enrichment Analysis
Kyoto Encyclopedia of Genes and Genomes pathway enrichment analysis was processed by using KOBAS 3.0 (http://kobas.cbi.pku.edu.cn/kobas3), where P value <.05 was set as the threshold.
Statistical Analysis
All types of genomic mutations including SNVs, frameshift variants, insertions/deletions, CNVs, and gene fusions were precisely analyzed. Statistical analyses were conducted using the Graphpad (Prizm 8). A P value of <.05 was considered as statistically significant.
Results
Demographic and Clinicopathological Data of the STS Patients
The demographics and clinicopathological data of patients in the cohort are summarized in Table 1. Genetic landscape of 37 male and 28 female Chinese patients with STS was analyzed. The median age of enrolled patients at the time of sequencing was 55 years (range, 16-81 years), and a higher proportion of STS patients aged over 60 was in male than in female (26.2% vs 12.3%, P = .201). AJCC staging evaluation demonstrated stage I disease in 5 patients, stage II disease in 16 patients, stage III disease in 25 patients, and stage IV disease in 19 patients. At the time of genetic testing, 58% of patients had recurrent or metastatic lesions. In terms of histological subtype distribution, the cohort contained multiple STS subtypes including fibrosarcoma (n = 10), UPS (n = 9), myofibroblastic sarcoma (n = 8), liposarcoma (n = 5), clear cell sarcoma (n = 4), epithelioid sarcoma (n = 4), rhabdomyosarcoma (RMS; n = 4), malignant peripheral nerve sheath tumors (n = 3), alveolar soft part sarcoma (n = 2), Ewing sarcoma (n = 2), leiomyosarcoma (n = 2), synovial sarcoma (n = 2), malignant mixed Müllerian tumor (n = 1), and unknown types (n = 9).
Clinical Characteristics of the Chinese STS Patients (n = 65).

Abbreviations: STS, soft tissue sarcoma; UPS, undifferentiated pleomorphic sarcoma.
Patients with unknown histology indicated that a clear pathological subtype could not be defined based on pathological testing.
Profiling of Genomic Alterations in Chinese STS Patients
A total of 218 genetic variants were identified in the 65 STS patients, with an average of 3.35 variants per patient (range, 0-28). In total, 95.4% (62/65) of patients harbored at least one detectable oncogene alteration. Of the 218 genetic variants, 137 (62.8%) were SNVs, 42 (19.3%) were small insertions or deletions (Indels), 7 (0.3%) were fusions, and 32 (14.7%) were copy number alterations, all of which were amplification. The most frequently mutated genes in STS patients were TP53 (21.5%, 14/65), MDM2 (15.4%, 10/65), CDK4 (6.2%, 4/65), EWSR1 (6.2%, 4/65), KDR (6.2%, 4/65), and NF1 (6.2%, 4/65). Other mutated genes that occurred in more than 3 patients included ATRX, IGF1R, IGF2R, KIT, and RB1 (Figure 1).
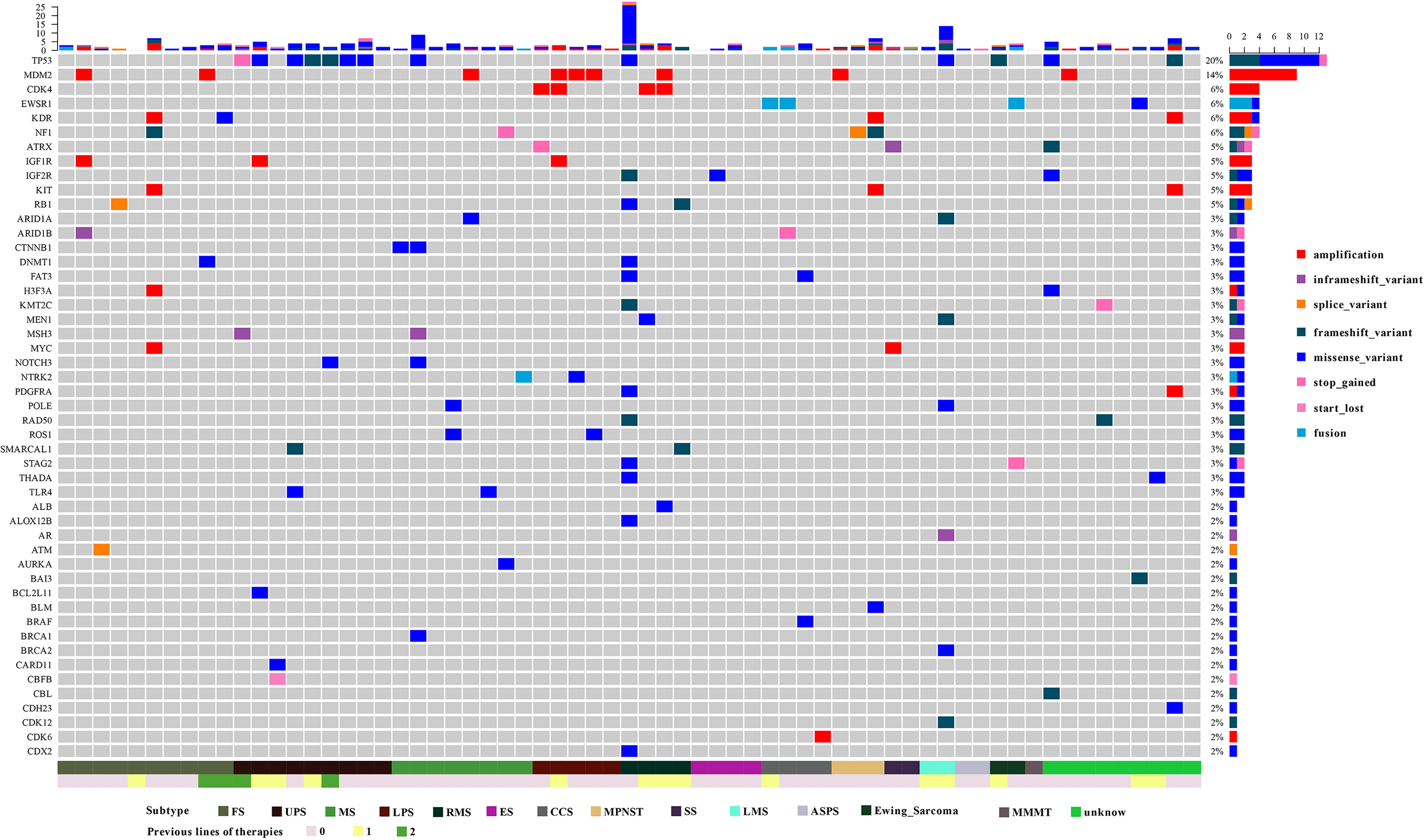
Landscape of genomic alterations in the 65 Chinese soft tissue sarcoma (STS) patients. Each row represents a gene and each column represents a patient. The mutational spectrum is grouped according to the histological subtype and previous lines of therapies of each patient. On top of the mutation heatmap, the histogram represents the number of mutations per patient. The mutation frequency of each gene is shown on the right side of the mutation heatmap. ASPS, alveolar soft part sarcoma; CCS, clear cell sarcoma; ES, epithelioids sarcoma; FS, fibrosarcoma; LMS, leiomyosarcoma; LPS, liposarcoma; MMMT, malignant mixed Müllerian tumor MPNST, malignant peripheral nerve sheath tumors; MS, myofibroblastic sarcoma; RMS, rhabdomyosarcoma; SS, synovial sarcoma; UPS, undifferentiated pleomorphic sarcoma.
Signaling pathway studies revealed that gene mutations were most common in the PI3K-Akt (n = 29), RAS (n = 19), RAP1 (n = 18), and ErbB (n = 16) signaling pathways for Chinese STS patients (Figure 2). Across all sarcoma subtypes, the most common alterations in the PI3K-Akt signaling pathway included TP53 (n = 13), MDM2 (n = 9), KDR (n = 4), and CDK4 (n = 4).

Mutation-enriched signaling pathways in Chinese soft tissue sarcoma (STS) patients. The vertical axis shows the most affected signaling pathways, and the horizontal axis shows the number of mutated genes in each signaling pathway.
Tumor mutational burden, defined as the number of somatic mutations per Mb in the tumor genome, has recently been found to be useful for predicting the efficacy of immunotherapy. Pan-solid tumor patients with a high TMB had a higher response rate to immunosuppressive drugs such as pembrolizumab. Several previous genomic studies identified STS as a relatively low TMB solid tumor type in the Caucasian population.15,16 The median TMB was 3.17 mutations/Mb (range 0-17.5) in the Chinese subgroup when being calculated and analyzed. A low TMB (<6 mutations/Mb) was presented in the majority of patients (86.2%, 56/65), and a high TMB (>10 mutations/Mb) was found in only 3 patients (4.6%, 3/65) (Figure 3). Microsatellite instability data were available for the enrolled patients, and all patients were microsatellite stable (data not shown).

Distribution of TMB stratified by histological subtypes in the Chinese soft tissue sarcoma (STS) cohort. TMB values were shown by median and quartile. Horizontal lines in each box denote the median. The bottom and top of each box denote the lower and upper quartile, respectively. ASPS, alveolar soft part sarcoma; CCS, clear cell sarcoma; ES, epithelioids sarcoma; FS, fibrosarcoma; LMS, leiomyosarcoma; LPS, liposarcoma; MMMT, malignant mixed Müllerian tumor; MPNST, malignant peripheral nerve sheath tumors; MS, myofibroblastic sarcoma; RMS, rhabdomyosarcoma; SS, synovial sarcoma; TMB, tumor mutational burden; UPS, undifferentiated pleomorphic sarcoma.
Targetable Genomic Alterations in STS and Clinical Impact of Alterations Targeting
In the cohort of 65 STS patients, 26 potentially actionable genetic variants were detected in 24 patients (36.9%). These clinically beneficial alterations included MDM2 amplification (n = 8), CDK4/6 amplification (n = 5), and NF1 deletions (n = 2). Other potentially targetable mutant genes included COL1A1/PDGFB fusion, BRAF oncogenic mutation, MAP2K1 oncogenic mutation, etc (Table 2). Among them, 6 patients underwent drug-matched clinical trials, and the efficacy of 4 patients could be evaluated. One patient with inflammatory myofibroblastoma carrying MAP2K1 activating mutation achieved a 2-month relapse-free survival (RFS) with oral Trametinib. One patient with CDK4 amplified liposarcoma received RFS for 4 months after receiving Palbociclib treatment. One patient with DFSP who was diagnosed with COL1A1/PDGFB fusion received first-line Imatinib targeted treatment and obtained RFS for 10 months. After NGS, the patient diagnosed with CCS was treated with Vemurafenib, which targeted the BRAF V600E mutation, and brain metastasis occurred 21 months after targeted therapy (Table 3).
Summary of Actionable Mutations and Potentially Beneficial Drugs.

Outcome of Targeted Therapy Inspired by NGS Test Results in 4 Patients.

Abbreviation: NGS, next generation sequencing.
Discussion
STS is a rare group of heterogeneous tumors with different histological manifestations and overlaps. Accurate molecular classification of patients with STS is necessary because the prognosis of STS groups varies greatly, and individual patients may benefit from different targeted therapeutic strategies.17,18 Targeted NGS is an accurate, cost-effective, and clinically available method for measuring targeted gene changes; it allows for deeper sequencing with fewer samples and less time than whole exome sequencing. The advent of NGS has enabled a much higher genomic resolution for the study of unreported and operable mutations that may affect patients' clinical treatment decisions. Over the past few years, some studies have described the genomic landscape of patients with STS in China through NGS, but the conclusions of these studies were limited by sample size or coverage subtypes, and the actual clinical value of NGS testing for STS patients has not been evaluated.13,19 The main objective of our study is to uncover the genomic profile of Chinese STS patients and to evaluate the potential clinical impact of NGS testing on the clinical management of these patients. In the Chinese STS cohort, 95.4% (62/65) of patients possessed at least one genetic mutation. The most frequently mutated genes in patients included TP53, MDM2, CDK4, KDR, etc. The vast majority (86.2%, 56/65) of patients had a lower TMB of less than 6 mutations/Mb. Our findings are consistent with previously reported results in patients with Caucasian STS. Cote et al performed comprehensive genomic profiling in 133 STS patients to determine the incidence of operability changes in STS. 14 At least one mutation was detected in 88% of the patients tested, and the most frequently altered genes were TP53, CDKN2A/B, and RB1. However, this study did not report data on the results of NGS findings used in clinical treatment. Interestingly, consistent with previous research, our study also shows that STS is a relatively low TMB solid tumor type (<6 mutations/Mb, 86.2%; >10 mutations/Mb, 4.6%). As a result, most STS patients may be less likely to benefit from immunotherapy. In contrast, high TMB had been shown to be associated with inferior OS in RMS, Ewing sarcomas, and some other subtypes.13,20,21 Therefore, TMB may be used as a prognostic factor for routine clinical treatment in STS patients rather than as a predictor of immunotherapy efficacy.
Conventional treatments for STS include surgery, radiation therapy, and chemotherapy. As STS is a heterogeneous tumor, different histological types, sites, pathological grades, and tumor sizes may have different effects on traditional treatment.3,22–25 In the era of precision medicine, targeted therapies have brought new options for STS treatment, which can further improve the efficacy and life quality of patients.16,19,26,27 NGS testing has shown promising applications in the precision treatment of STS patients. Groisberg et al performed a retrospective analysis of 102 sarcoma patients who underwent NGS testing. 28 About 61% (62/102) of patients had potentially actionable mutations, including 14% (14/102) with the approved drug-targeted treatment of sarcoma, and 45% (46/102) with drug targeting therapy approved for other diseases. In their study, the molecular profile assays guided the treatment of 16 patients, with 50% of patients achieving significant clinical benefit. In another study of 114 patients with STS, Boddu et al conducted NGS-based molecular profiling on tumors from those patients with sarcoma. 29 They reported that 49.1% of patients harbored actionable alterations after review by the molecular tumor board. Fifteen patients underwent a change in drug therapy based on NGS assays, of which 4 patients won partial response or stable disease more than 6 months. Similar to previous findings, variations potentially amenable to clinical trials were present in 36.9% (24/65) of patients in our STS cohort. Among them, 4 patients who received molecularly guided therapy achieved at least 4 months of RFS.
There are some limitations in our study. First of all, although patients with multiple types of STS were included, we did not cover all types of STS because of the large number of classifications. Secondly, some of these subtypes included only 1 or 2 individuals; thus the further study needs to enroll more patients. Despite those limitations, we have shown positive and promising results. To the best of our knowledge, this is a relatively large extensive study on the genomics of Chinese STS patients to date. We demonstrate that targeted capture NGS provides a framework for a comprehensive review of actionable and near-actionable mutations in STS.
Footnotes
Acknowledgments
The authors thanks for all patients who supported this study.
Authors' Note
Gu Jin designed this study; Quanyu Yang performed the procedure for detection; Gu Jin and Chunming Yin collected the clinical data; Chunyang Wang and Danhua Wang performed data analysis; Aiwen Zheng and Tao Li gave critical comments and suggestions; Gu Jin and Chunyang Wang drafted the manuscript. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the Institutional Review Board of Zhejiang Cancer Hospital (registration number: IRB-2021-402), and individual consent for this retrospective analysis was waived, patients below 18 years old were provided with signed informed consent by their legal parents/legal guardians for the research use of clinical data and publication.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
