Abstract
MicroRNAs (miR) are a class of non-coding endogenous RNA molecules that suppress the translation of protein-coding genes by destabilizing target mRNAs. The MiR-574-5p has been reported to be involved in the several types of cancer. However, the expression of miR-574-5p and its mechanism in nasopharyngeal carcinoma (NPC) remain unclear. We found that the expression level of miR-574-5p was significantly increased in the NPC cell lines. We further demonstrated that Forkhead box N3 (FOXN3) was a target gene of miR-574-5p. FOXN3 overexpression and inhibition reversed the promoting or suppressing effect, respectively, of NPC cell proliferation, migration and invasion caused by miR-574-5p. Furthermore, miR-574-5p enhanced the β-catenin and TCF4 protein expression by repressing FOXN3 expression, resulting in the activation of the Wnt/β-catenin signaling pathway, but the activity of the Wnt/β-catenin signaling pathway was inhibited by a miR-574-5p inhibitor or FOXN3 overexpression, which reversed the effect of miR-574-5p. Wound-healing and Transwell assays also showed that miR-574-5p promotes the cell migration and invasion of NPC cells, whereas the promoting effect of miR-574-5p was also reversed by a miR-574-5p inhibitor or FOXN3 overexpression. Collectively, these data suggested that miR-574-5p promotes NPC cell proliferation, migration, and invasion at least partly by targeting the FOXN3/Wnt/β-Catenin signaling pathway.
Introduction
Nasopharyngeal carcinoma (NPC) is characterized by a distinct geographical distribution, and it is a high-frequency disease in East and Southeast Asia, but a low-frequency disease in western countries. 1, 2 An estimated 42,100 new cases and 21,320 deaths were attributed to NPC in China in 2013, accounting for 1.14% of all new cancer cases and 0.96% of all cancer-related deaths in 2013 in China. 3 NPC is different from other tumors as 70% of newly diagnosed NPC patients present with locally advanced disease. 4 The first-line treatment for nasopharyngeal cancer is radiation therapy due to the anatomical location of nasopharyngeal carcinoma and its sensitivity to irradiation. In the early stages of NPC, radiation therapy increases the survival rate to 84.7-87.4%. 5 In advanced stage NPC patients, the survival rate can be increased to 50%-70% by combining chemotherapy and radiotherapy. 6 However, the lack of biomarkers for early diagnosis results in many patients being diagnosed at advanced stages. Thus, it is important to understand the mechanisms and pathway in the underlying pathogenesis of NPC as well as to identify biomarkers for diagnosis of NPC patients.
In recent years, it has been demonstrated that microRNAs (miRs) play an important regulatory role in the occurrence and development of diseases, especially cancer. MicroRNAs are a class of small RNA molecules that regulate the translational level of genes, and they can inhibit the translation of protein-coding genes to repress protein translation by binding to target mRNAs. 7 -9 A previous study has demonstrated that miRNAs are associated with the development, progression and treatment of cancer. 10 MiR-574-5p is an important small RNA molecule that plays important roles in regulating the migration of tumor cells, including breast cancer, colorectal cancer, thyroid cancer and non-small cell cancer. 11 -13 A previous study has indicated that miR-574-5p is a serum biomarker for early-stage non-small cell lung cancer. 12,14 Additionally, miR-574-5p promotes the invasion of small-cell lung cancer by regulating β-catenin signaling. 15 These results reveal a novel and important mechanism for regulating miR-574-5p and tumor cell metastasis, including colorectal cancer, thyroid cancer and non-small cell cancer. 11 -13 However, the expression of miR-574-5p and its mechanism in nasopharyngeal carcinoma (NPC) remain unclear.
Forkhead box N3 (FOXN3) belongs to the forkhead box (FOX) protein family, which has transcriptional inhibitory activity and is involved in tumorigenesis. 16,17 Previous studies have indicated that the FOXN3 gene is markedly downregulated in 11 different malignancies, including liver cancer, lung cancer, colon cancer, prostate cancer, laryngeal cancer and glioblastoma multiform. 18 -22 The FOXN3 gene as a biomarker has been demonstrated to inhibit the growth, migration and invasion of cancer cells by overexpressing and knocking down the FOXN3 gene in a colon cancer cell line. 23 FOXN3 silencing resultes in decreased expressions levels of Snail, N-cadherin and c-Myc (3 downstream genes of the β-catenin/TCF signaling pathway), and it promotes the β-catenin/TCF pathway by increasing the amount of TCF4 that binds to β-catenin to initiate transcription in the nucleus, thereby promoting the migration and invasion of colon cancer cells. 24 β-catenin as a classical regulator of cancer, which can simultaneously stimulate the Wnt and E-cadherin pathways, and the interaction between miRNAs and the β-catenin pathway has been widely investigated in cancer. 25 In addition, a previous study has demonstrated that the capability of TCF4 to form complexes with β-catenin is weaker when miR-378 targetes FOXN3, which blocks the catenin/TCF signaling pathway, resulting in reduced epithelial mesenchymal transformation and metastasis in melanoma. 26 However, FOXN3-specific expression changes in NPC have not been reported and the regulatory relationship between miR-574-5p and the Wnt/β-Catenin signaling pathway remain unclear. Thus, the aims of the present were to investigate whether miR-574-5p regulates NPC cell invasion by targeting FOXN3 and to validate the regulatory relationship between miR-574-5p and the Wnt/β-catenin signaling pathway.
Materials and Methods
Cell Line and Cell Viability Assay
The nasopharyngeal epithelial (NPE) cell line, NP69, and the nasopharyngeal carcinoma (NPC) cell lines, TW03 and C666-1, were purchased from Shanghai Institute of Biochemistry and Cell biology of Chinese Academy of Science (Shanghai, China) and cultured in IMEM medium (Gibco®, Grand Island, NY) containing 10% heat-inactivated fetal bovine serum (FBS, PAN Biotech). Cells were maintained at 37 oC in a humidified 5% CO2 incubator (SANYO, Osaka, Japan). Fresh growth medium was changed every day until cells were confluent.
Cell viability was determined by the CCK-8 assay using the Cell Counting Kit-8 kit (CK04, Dojindo). Cells were plated at 4 × 104 cells/mL in 96-well plates and allowed to attach for 0, 24, 48 and 96 h at 37ºC and 5% CO2. CCK-8 (10 μL) was then added to each well. After incubating for 1h at 37oC, absorbance values were detected at 450 nm using a Biotek Synergy 2 plate reader (Winooski, VT).
Reverse Transcription-Polymerase Chain Reaction (RT-PCR)
NP69, TW03 and C666-1 cells were plated into flat-bottom cell culture plates and incubated overnight at 37 oC with 5% CO2. Cell supernatants were collected from each group after centrifugation at 1000 r/min and 4 oC for 10 min. Total RNA was extracted from NP69, TW03 and C666-1 cells using TRIzol reagent (Invitrogen, USA), and a miRNeasy Mini kit (Qiagen, Germany) was used to isolate total RNA. Reverse transcription and qPCR mixtures were prepared on ice according to the manufacturer’s instructions as previously described. 27 For real-time PCR, 2.0 µL of reverse transcription product was mixed with 10.0 µL of qPCR SYBR® Green Master Mix (A6001, Promega, USA), 0.5 µL of forward and reverse primers and 7.0 µL of nuclease-free water to a final volume of 20 μL. Subsequently, the ABI 7500 PCR system (ABI, USA) was used to detect mRNA expression and the 2-△△Ct method was used to calculate the relative mRNA expression levels according to a previous study. 28 The qPCR conditions were as follows: 95 °C for 5 min; and 40 cycles of at 95 °C for 10 s, 60 °C for 30 s and 72 °C for 30 s. The primers were designed using Primer 5.0 software and synthesized by Sangon Biotech (Shanghai, China). The following primer sequences were used: miR-574-5p, CGCGTGAGTGTGTGTGTGTGA (Forward) and AGTGCAGGGTCCGAGGTATT (Reverse); RT, GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACACACAC. U6, CTCGCTTCGGCAGCACA (Forward) and AACGCTTCACGAATTTGCGT (Reverse); and RT, AAAATATGGAACGCTTCACGAATTTG.
Plasmid Construction and Mimics/Inhibitors/siRNA
The full-length open reading frame of FOXN3 was cloned into pcDNA3.1 to generate expression vectors. Wild-type (WT) FOXN3 3′-UTR was cloned into the psiCHECKTM-2 basic vector (Promega Corporation, USA). Site-directed mutagenesis of the miR-574-5p seed sequence in the 3′-UTR (Mut) was performed using the QuikChange™ Site-Directed Mutagenesis kit (Stratagene, USA). The following
mimics, inhibitors and siRNA were designed and constructed by RiboBio (Guangdong, China).: miR-574-5p mimic (sequence: 5′-UGAGUGUGUGUGUGUGAGUGUGU -3′); mimic NC (sequence: 5′- UCUACUCUUUCUAGGAGGUUGUGA-3′); miR-574-5p inhibitor (sequence: 5′- ACACACUCACACACACACACUCA-3′); inhibitor NC (sequence: 5′-UCUACUCUUUCUAGGAGGUUGUGA-3′); si-NC (sense:5-CGGUCCGUGAAGGAUAUAAUG-3′ and antisense:5’-UUAUAUCCUUCACGGACCGUG-3′); and si-FOXN3(sense:5’- GUACCUUCUUCAAGAGAAAUG-3′and antisense:5’-UUUCUCUUGAAGAAGGUACUG-3′). Cells were transfected with mimics, inhibitors or siRNA using Lipofectamine 2000 (Invitrogen, L3000015) following the manufacturer’s protocol with final concentrations of 20 nM, 40 nM, respectively. And 50 nM. Cells were subjected to further experimentation at least 48 h after transfection. The transfection efficiencies of all mimics and inhibitors were verified.
Luciferase Reporter Assays
The binding site of miR-574-5p was predicted using TargetScan (http://www.targetscan.org). The FOXN3 3’-UTR fragment with the putative (FOXN3-WT) or mutated binding site (FOXN3-Mut) of miR-574-5p was inserted into the psiCHECK-2 luciferase reporter vector (Promega, Madison, USA). C666-1 cells were grown in a 96-well plate and cotransfected with miR-574-5p mimic or negative control and the FOXN3-WT or FOXN3-Mut reporter plasmid. After 48 h of transfection, the relative luciferase activity was assessed with a dual-luciferase reporter assay and a fluorescence reader (Promega, USA). All experiments were performed independently 3 times.
Wound-Healing Assays and Transwell Invasion Assays
For the wound-healing assays, C666-1 cells (3 × 105 cells/mL) were suspended in 2 mL of IMEM medium containing 10% FBS, plated in 6-well plates and cultured at 37 °C for approximately 24 h. Subsequenly, the 70 µL cell suspension was added into the Ibili-cell plug-in. Once cell monolayers formed, the Ibili-cell plug-in was removed using tweezers, and the culture was added to serum-free medium at 37 °C for 0, 12 or 24 h in an incubator containing 5% CO2. Cells that had migrated into the wound area were imaged using an inverted microscope (Olympus, CKX41). The distance was measured using Image J software.
For the Transwell invasion assays, C666-1 cells (1 × 105 cells/mL) were plated in the Transwell chamber with extracellular matrix gel (Sigma, E1270-coated membranes; 24-well insert, pore size of 8 μm, BD Biosciences) and cultured in medium containing 10% FBS. After 24-48h, the Transwell chamber was removed. The cells that did not migrate through or invade the pores were removed with a cotton swab. After washing 3 times with the PBS the cells on the lower surface of the membrane were fixed with methanol, stained with 0.1% crystal violet (Sigma) and counted.
SDS-PAGE and Western Blot Analysis
Protein was extracted from C666-1 cells using radio-immunoprecipitation assay (RIPA) buffer containing PMSF. Samples were placed on ice for 20 min at 4°C and then centrifuged at 12 000 rpm for 20 min. The supernatant (200 μL) was collected and placed in an EP tube. Total protein concentration was determined using the BCA Protein Assay Kit PC0020-500 (Solaibao Technology Co. LTD, Beijing, China). Protein were separated by a SDS-polyacrylamide gel according to previously reported methods. 28, 29 After blocking, blots were washed with TBS containing 0.5% Tween-20 (Beyotime Institute of Biotechnology, Jiangsu, China) and incubated overnight at 4°C with the anti-FOXN3 mAb (1:1500, Abcam, Beijing, China), anti-β-Catenin mAb (1:1500, Abcam, Beijing, China) or anti-TCF4 mAb (1:1000, Abcam, Beijing, China). After several washes with Tris-buffered saline containing 0.1% Tween-20, the membranes were incubated in a 1:5000 dilution of an anti-mouse HRP-conjugated secondary antibody for 2 h. Subsequently, the signal was detected by chemiluminescence using the Thermo ECL Substrate (Bio-Rad). The same membrane was incubated with a β-actin antibody (1:1000, Abcam, Beijing, China), which served as a loading control. The results were collected via the Versa DocTM imaging system (Peiqing Technology Co. LTD, Shanghai, China) and analyzed with Image J software.
Statistical Analysis
All statistical analyses were conducted using SPSS 22.0 statistical software. The data are expressed as mean ± SD. Student’s t-test was performed to compare 2 groups, and 1-way ANOVA was performed for experiments with the more than 2 groups. P < 0.05 was considered statistically significant.
Results
MiR-574-5p and FOXN3 Expression in NP69, TW03 and C666-1 NPC cells
For miR-574-5p and FOXN3 expression, both NPE and NPC cells were evaluated. As shown in Figure 1, miR-574-5p was significantly overexpressed in TW03 and C666-1 NPC cells compared to the NP69 NPE cells (P < 0.05 or P < 0.01). The FOXN3 protein levels in TW03 and C666-1 cells were lower than those in NP69 cells ((P < 0.05 or P < 0.01). In C666-1 cells, the miR-574-5p gene expression level was the highest and the FOXN3 protein level was the lowest. Therefore, C666-1 cells were used to evaluate the functional effect of miR-574-5p in the subsequent analyses.

MiR-574-5p Transfection Promotes NPC Migration and Invasion In Vitro by Targeting FOXN3
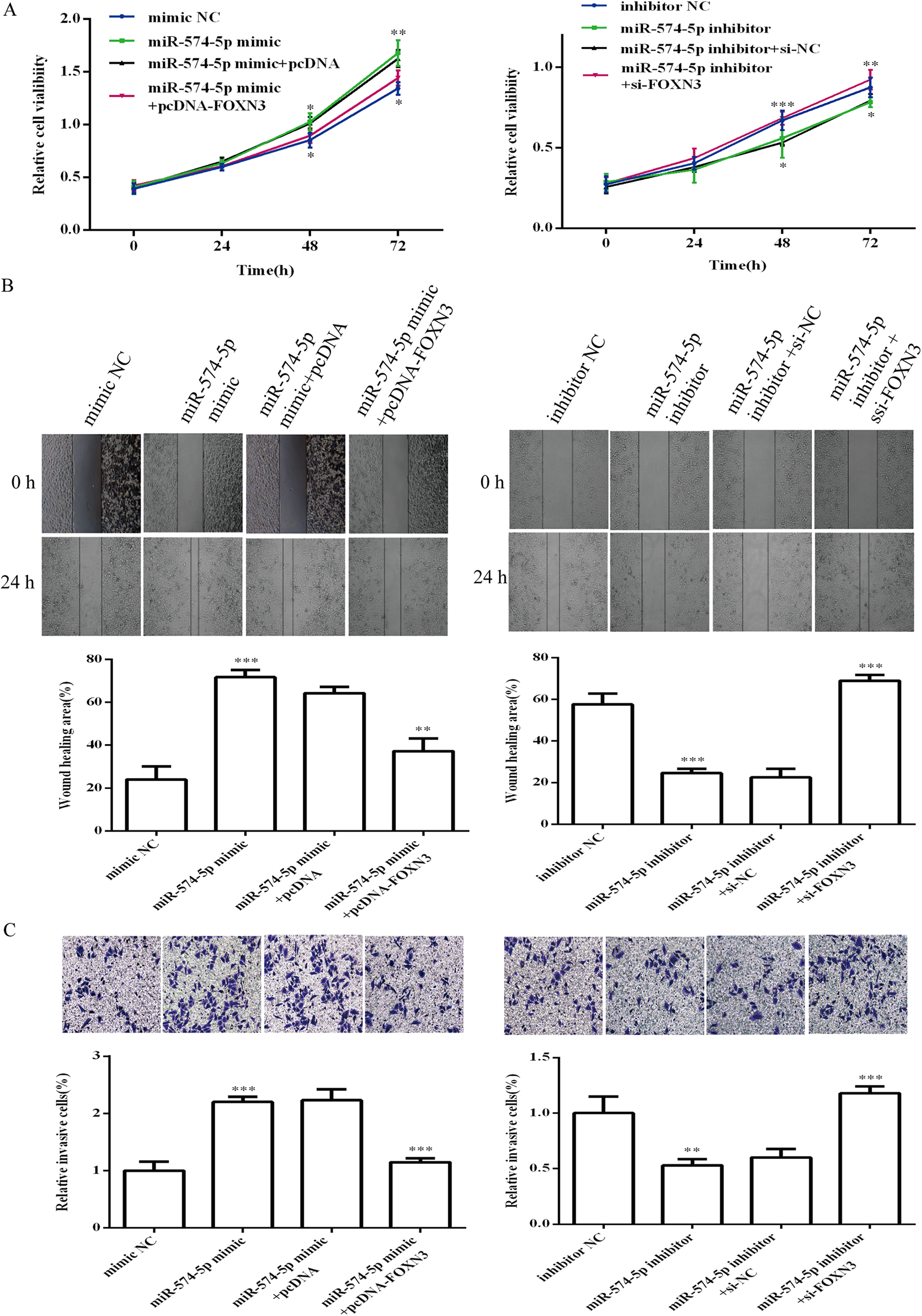
To determine whether miR-574-5p regulates cell invasion, cells were infected with the miR-574-5p mimic to establish stably miR-574-5p-overexpressing C666-1 cells or transfected with the miR-574-5p inhibitor to inhibit miR-574-5p expression. The expression level of miR-574-5p was verified by qRT-PCR. MiR-574-5p was upregulated in the miR-574-5p mimic group but downregulated in the miR-574-5p inhibitor group (Figure 2A). Cell viability of C666-1 cells was detected by CCK-8 (Figure 2B). Cell viability was significantly increased in the miR-574-5p mimic group, but it was reduced in the miR-574-5p inhibitor group (Figure 2B). To determine the target genes of miR-574-5p, candidates were searched using Target-Scan (http://www.targetscan.org/). To analyze the relationship between miR-574-5p and FOXN3, a luciferase reporter assay containing the wild-type (wt) or mutant (mut) miR-574-5p target sites in the FOXN3 3′-UTR was performed using C666-1 cells (Left panel of Figure 2C). Overexpression of miR-574-5p significantly decreased the luciferase activity of the FOXN3 3′-UTR compared to mimic control transfected-cells (Right panel of Figure 2C), indicating that miR-574-5p reduces FOXN3 expression by binding to target sites. In addition, overexpression of miR-574-5p decreased FOXN3 protein levels in C666-1 cells, while transfection of the miR-574-5p inhibitor increased FOXN3 protein levels in C666-1 cells (Figure 2D). These results indicated that miR-574-5p specifically regulated FOXN3 expression at the posttranscriptional level, indicating that FOXN3 may be a direct target of miR-574-5p. In addition to changes in FOXN3, transfection with the miR-574-5p mimic enhanced the β-catenin and TCF4 protein levels according to western blot analyses (Figure 2E). As shown in Figure 2E, transfection with the miR-574-5p inhibitor reduced β-catenin and TCF4 protein levels. Because the β-catenin signaling pathway plays an important role in cell adhesion and invasion, we performed wound-healing and invasion assays. The results showed that transfection of the miR-574-5p mimic significantly enhanced the migration of C666-1 cells. However, C666-1 cells transfected with the miR-574-5p inhibitor showed reduced cell migration (Figure 2F). Transwell invasion assays were used to verify these findings, and similar results were obtained (Figure 2G). Overall, these results showed that miR-574-5p promotes C666-1 cell migration and invasion in vitro by targeting FOXN3.

FOXN3 Overexpression Inhibits Cell Migration and Invasion Through the Wnt/β-Catenin Pathway
To ascertain the role of FOXN3 in the NPC cell invasion process, pcDNA-FOXN3 and si-FOXN3 were utilized to overexpress and knockdown FOXN3, respectively (Figure 3A). The cell viability of C666-1 cells was significantly enhanced after FOXN3 was overexpressed, but the cell viability was decreased after transfection with si-FOXN3 (Figure 3B). The wound-healing and Transwell invasion assays demonstrated that FOXN3 overexpression significantly inhibited the migration and invasion of C666-1 cells (Figure 3C and D). In contrast, knockdown of FOXN3 enhanced the cell migration and invasion of C666-1 cells (Figure 3C and D). We next investigated the mechanism of the inhibition of NPC cell invasion by FOXN3-induced inactivation of the Wnt/β-catenin signaling via repressing β-catenin expression. Western blot analysis revealed that knockdown of FOXN3 significantly promoted β-catenin and TCF4 protein expression (Figure 3E). In addition, FOXN3 overexpression markedly decreased β-catenin and TCF4 protein expression (Figure 3E). These results indicated that FOXN3 is a biomarker of activated Wnt/β-catenin signaling in C666-1 cells.

Co-transfection of miR-574-5p and FOXN3 Regulates Cell Migration and Invasion Through the Wnt/β-Catenin Signaling Pathway
To explore whether FOXN3 is involved in the biological roles of miR-574-5p in C666-1 cells, miR-574-5p and FOXN3 were co-transfected into C666-1 cells. The cell viability (Figure 4A), migration (Figure 4B) and invasion (Figure 4C) of C666-1 were significantly enhanced in the miR-574-5p mimic group, but co-transfection of the miR-574-5p mimic and pcDNA-FOXN3 inhibited the promoting effect of the miR-574-5p mimic. . In contrast, C666-1 cell viability, migration and invasion were also enhanced in the miR-574-5p inhibitor and siRNA-FOXN3 co-transfection group compared to the miR-574-5p inhibitor transfection group. Furthermore, transfection of the miR-574-5p mimic activated the Wnt/β-catenin signaling pathway, whereas co-transfection of the miR-574-5p mimic and pcDNA-FOXN3 inhibited the promoting effect of the miR-574-5p mimic (Figure 5). Transfection of the miR-574-5p inhibitor repressed the Wnt/β-catenin signaling pathway, whereas co-transfection of the miR-574-5p inhibitor and si-FOXN3 attenuating the effect of the miR-574-5p inhibitor on the Wnt/β-catenin signaling pathway in C666-1 cells. These results suggested that miR-574-5p promots cell migration and invasion by targeting FOXN3 in C666-1 cells via the Wnt/β-catenin signaling pathway.
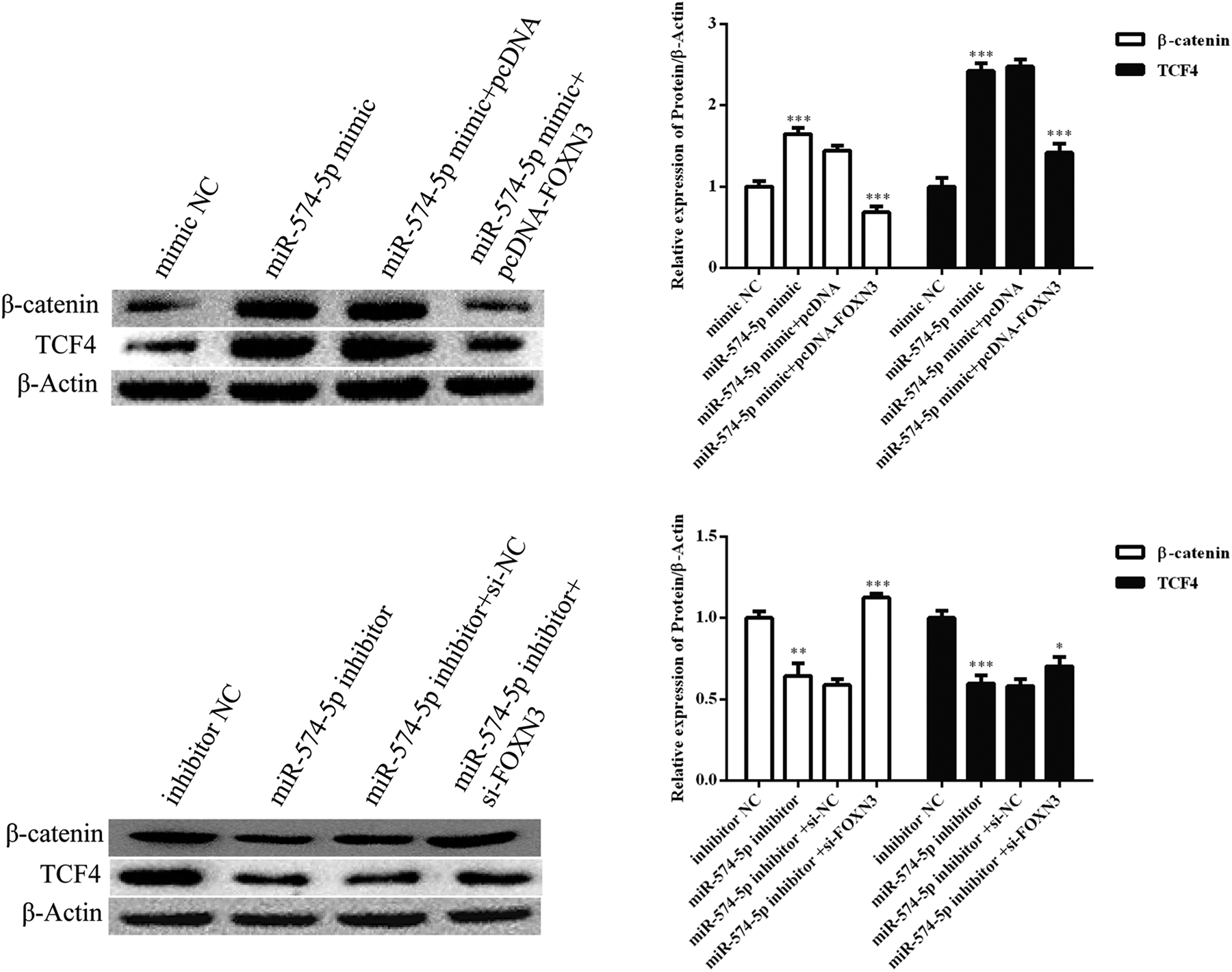
MiR-574-5p transfection and FOXN3 overexpression regulate Wnt/β-catenin signaling pathway. *P < 0.05, **P < 0.01 and ***P < 0.001.
Discussion
NPC is the most common squamous cell carcinoma, and the pathogenesis of NPC involves multiple processes, including genetic factors, Epstein-Barr virus infection and environmental impacts. 30 At present, the most effective treatment for NPC is radiotherapy and chemotherapy. However, these therapies do notcompletely inhibit NPC development. 31 Monotherapy can control the development of resistance, but the efficacy and quality of life for NPC patients is not guaranteed. Therefore, novel molecular therapeutic targets that can control the development of NPC are urgently needed. In recent years, the inhibition of cell proliferation and invasion as well as the induction of apoptosis have been suggested for anticancer activities. Previous studies have showed that almost all patients with NSCLC eventually relapse due to the activation of cancer cell invasion, resulting in metastatic disease and death. 32, 33 Previous studies have also shown that microRNA expression profiling is associated with tumor development, progression and response to therapy.
The present study showed that miR-574-5p expression was frequently increased in human NPC cell lines compared to the NP69 nasopharyngeal epithelial cell line, which agreed with previous studies. 11,13 -15 Thus, miR-574-5p may be a crucial factor for further prognosis of NPC. We next investigated the biological role of miR-574-5p in the cell invasion ability of C666-1 cells. Transfection of the miR-574-5p inhibitor transfection significantly reduced cell invasion, suggesting that low miR-574-5p expression has an inhibitory effect on cell invasion. MiR-574-5p, which functions as a tumor-promoting factor, has been shown to be downregulated, which suppresses cancer cell proliferation, migration and invasion. 34 The present study demonstrated that miR-574-5p was upregulated and served as a positive regulator for cell invasion in NPC cells. One possible explanation is that miR-574-5p enhances target gene expression. 35
Based the target gene candidate analysis, the FOXN3 was further evaluated. Previous results have indicated that FOXN3 is an important tumor suppressor gene. 18 -21 Previous experiments using overexpression and knockdown of FOXN3 have shown that FOXN3 is a biomarker that inhibits the growth, migration and invasion of cancer cells. 14,15 Interestingly, Li et al. reported that the FOXN3 complex was essential for cancer. 36 ChIP-Seq experiments have found that the FOXN3 complex suppresses a group of genes to regulate cell differentiation, leading to breast cancer development and metastasis. 36 Sun et al. reported that FOXN3 inhibits the proliferation of liver cancer cells both in vivo and in vitro. 23 Our results suggested that FOXN3 is a target gene of miR-574-5p. The expression of FOXN3 was downregulated in C666-1 cells, and enforced overexpression of FOXN3 inhibited C666-1 cell invasion. Our results also showed that co-transfection of the miR-574-5p mimic and pcDNA-FOXN3 inhibited the promoting effect of the miR-574-5P mimic, demonstrating that FOXN3 is inhibited during miR-574-5p-induced NPC cell invasion. In addition, the present study demonstrated that inhibition of FOXN3 expression enhanced β-catenin and TCF4 protein expression, thereby activating the β-catenin signaling pathway.
The present study also demonstrated that the activity of the β-catenin pathway plays an essential role in the process of regulating NPC cell invasion by miR-574-5p. β-catenin is a classical regulator of cancer, and it stimulates the Wnt pathway in the pathogenesis of cancer. 25 In detail, activation of the Wnt/β-catenin pathway causes an accumulation of β-catenin in the cytoplasm and its eventual translocation into the nucleus to regulate various developmental processes, including oncogenesis. 37 -39 In addition, previous studies have shown that TCF4 combined with β-catenin forms complexes, which are weakened when miR-378 targets FOXN3, resulting in reduced cell metastasis in melanoma. 26 Thus, miR-574-5p promotes NPC cell invasion, at least in part, through the targeting FOXN3, which activates the Wnt/β-catenin axis.
Collectively, these data demonstrated that miR-574-5p promotes NPC cell proliferation, migration, and invasion by targeting FOXN3, resulting in activation of the Wnt/β-catenin pathway. Our results suggested that miR-574-5p has potential as a novel therapeutic target for NPC.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Our study did not require an ethical board approval because it did not contain human or animal trials.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from Natural Science Foundation of FuJian Province (grant number 2017J01393) and Fujian medical innovation project (grant number 2018-CXB-22).
