Abstract
In many countries, soiled toilet paper is placed in trash bins rather than flushed down the toilet. We investigated the use of soiled toilet paper in Guatemalan markets to surveil for pathogenic sequence types (STs) of E. coli and third generation cephalosporin-resistant E. coli (3GCR-EC). We collected used toilet paper from trash bins in the restrooms of three Guatemalan public markets. We screened samples for E. coli and 3GCR-EC. Whole genome sequencing was used to characterize the isolates. E. coli was successfully cultured from all 30 toilet paper samples. Soiled toilet paper was an effective matrix for surveilling multidrug-resistant E. coli and could be an efficient method in certain contexts.
Introduction
Infections caused by antibiotic-resistant Enterobacterales have increased at alarming rates, resulting in greater investments in antibiotic resistance (ABR) surveillance to address the emergence and spread (Murray et al. 2022). Unlike other bacterial infections, like Mycobacterium tuberculosis, where incident events are monitored, describing ABR surveillance data from clinical isolates and subsequently monitoring risk factors is challenging. Bacteria within the Enterobacterales family have different antibiotic-resistant phenotypic profiles, causing varying degrees of pathologies among patients and resulting in surveillance data that may not be comparable within or across countries (Murray et al. 2022). One approach to overcome challenges in the diversity of sampling strategies is to monitor ABR Enterobacterales from commensal isolates colonizing the gut (i.e., typically non-pathogenic strains), which provides a more robust estimate of ABR prevalence across a broader population not limited to participants with infectious disease pathologies (Smith et al. 2021).
Studies using stool sample collection have estimated the prevalence of cephalosporin-resistant Enterobacterales colonization to be high in communities in low to middle-income countries, ranging from 29% to 78% in Botswana, Chile, Kenya, Guatemala, and Bangladesh. This is much higher than what has been identified in high-income countries (Chowdhury et al. 2023; Mannathoko et al. 2022; Odoyo et al. 2023; Ramay et al. 2023). In these studies, contact with healthcare facilities and hygiene-based factors facilitating the transmission of bacteria were associated with colonization (Caudell et al. 2023).
Findings underscore the value of studies using robust approaches to describe the prevalence and risk factors associated with community colonization not only as a reservoir for ABR bacteria but also as a mediator of community-acquired infections and subsequent risks for hospital-acquired infections. Although these studies provide powerful estimations, stool sample collection is time-consuming and costly. Alternative methods of stool sample collection may provide avenues for more efficient surveillance techniques while providing robust estimates of ABR colonization.
Toilet paper is widely used for anal cleansing across the globe, but the low biodegradability of toilet paper’s pulp fibers can cause sewer lines to clog (Elmas and Öztürk 2019). Because of this, in many areas worldwide such as East Asia and Central America, soiled toilet paper is placed in trash bins and disposed of through the solid waste cycle (Sun and Han 2021). In China, researchers have highlighted that there are approximately 196,000 public toilets across mainland China and that more than three-quarters have users place soiled toilet paper into the waste bins (Sun and Han 2021).
Used diapers and perianal or rectal swabs are commonly used to determine whether individuals are colonized with pathogens or antibiotic-resistant bacteria (Lau et al. 2015; Pathak et al. 2013; Rogers et al. 2013; Tapiainen et al. 2019). Given the increasing complexity in conducting human subject research, collecting environmental samples from public trash bins could carry many benefits similar to that of wastewater in the surveillance of fecal associated pathogens like SARS-CoV-2 (Hillary et al. 2020).
Soiled toilet paper presents an opportunity to conduct efficient surveillance of antibiotic resistance when the toilet paper is collected in a reasonable timeframe relative to its disposal. This type of sampling could also provide a more accurate prevalence estimate of antibiotic resistance in a community, given that the toilet paper samples would represent an individual versus an aggregate sample from wastewater. In this study, we set out to understand whether soiled toilet paper could be used to understand the current status of 3GCR-EC in individuals at three markets in Guatemala.
Materials and Methods
Study Design
We conducted a cross-sectional study at three large community markets relatively close to Guatemala City, Guatemala. The selected markets were the municipal market of San Martín Jilotepeque in the Department of Chimaltenango, a municipal market of Santa Catarina Pinula in the Department of Guatemala, and a commercial market in Guatemala City, Guatemala. We used convenience sampling to collect 30 total samples of toilet paper contaminated with feces from the public bathrooms at the markets from June to July 2022.
Sampling Methods
Two to six stalls in the bathrooms of the markets were visited before noon on each collection day. If visibly soiled toilet paper was present in the stalls’ trash bins, the toilet paper pieces were collected and then aseptically placed in a sterile Whirl-Pak® Bag (55 oz). Each stall had its own sample bag and was only sampled once per collection day to maintain differentiation between the samples. All samples were transported to the Universidad del Valle de Guatemala (UVG) in coolers using ice packs (4–8°C) and began processing within 6 hours of collection (Figure 1). Sampling schematic demonstrating basic sample collection and analysis steps. Steps include: (A) identification of toilet stalls where soiled toilet paper is placed in a trash receptacle; (B) removal of lid, if present, to observe contents and determine suitability for sampling; (C) cse of tongs or disposable gloves to collect soiled toilet paper sample; and (D) streaking fecal matter present on toilet paper sample onto a MacConkey agar plate for culture.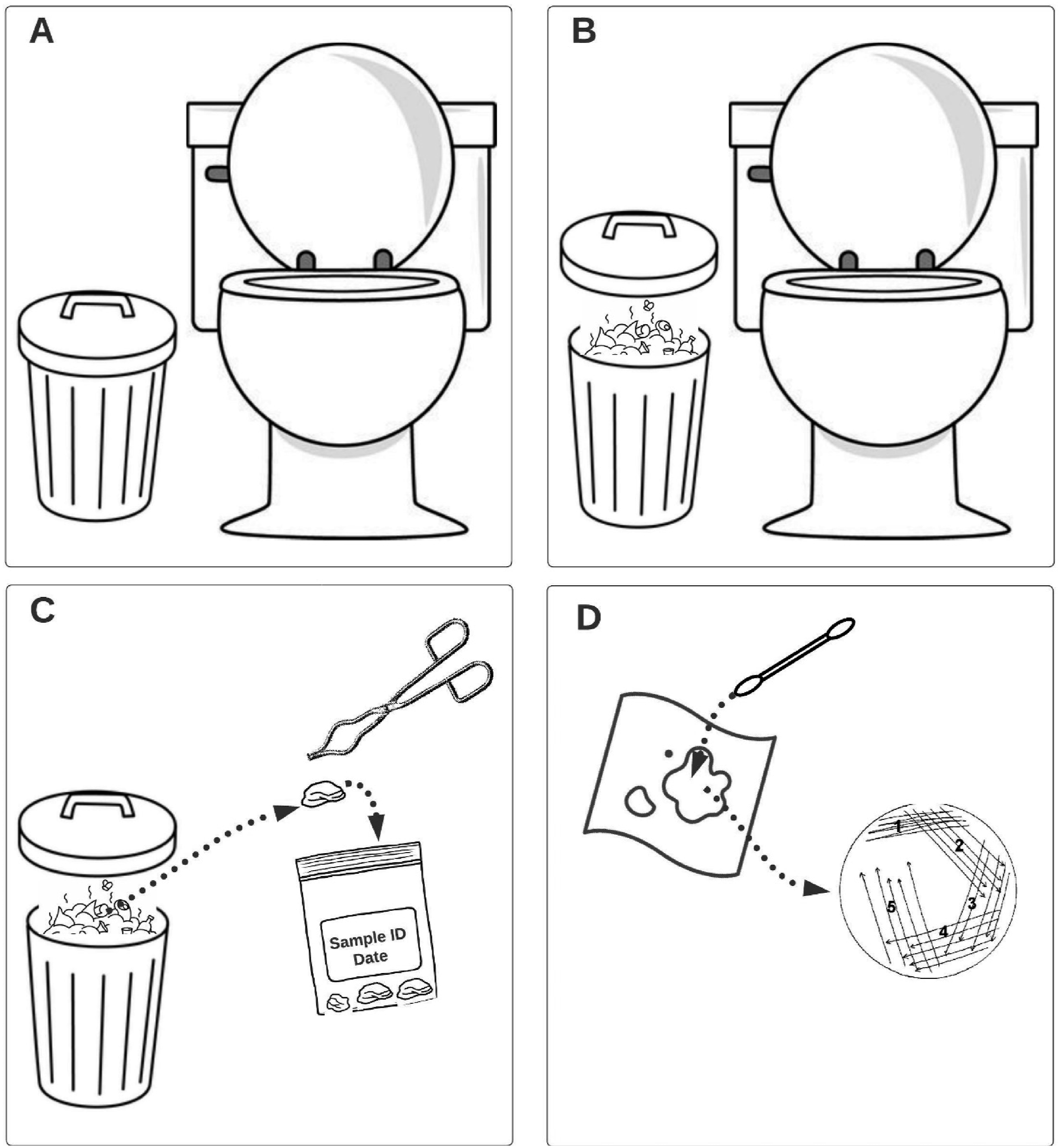
Isolation of 3GCR E. coli
A cotton swab saturated with Tryptic Soy Broth medium (Difco, USA) was rubbed on the areas of the toilet paper with the most observed fecal matter. The swab was then streaked on MacConkey agar (Difco, Maryland) absent of antibiotics and incubated at 37°C for 24 hours, after which ten lactose-positive colonies were selected from the MacConkey plate based on their pink or red appearance (Lautenbach et al. 2008). The putative E. coli colonies selected from their respective MacConkey plate were later plated on CHROMagar™ extended spectrum beta-lactam (ESBL) and incubated at 37°C for 24 hours (Hornsey et al. 2013). Up to two isolates from each toilet paper sample identified as putative 3GCR-E. coli based on their pink or dark red appearance and unique morphology, were selected and frozen at −80°C in Tryptic Soy Broth medium (Difco, USA) with 15% glycerol. We used the negative control, non-ESBL-producing E. coli ATCC 25922, and the positive control, ESBL-producing E. coli ATCC 13476, when culturing each batch of samples for quality assurance.
DNA Sequencing and Bioinformatic Analysis
Genomic DNA from the isolates was extracted using the DNeasy Blood and Tissue Kit (QIAGEN, Germany), according to the manufacturer’s instructions. One microgram of extracted DNA was stored in 1.5-milliliter microcentrifuge tubes at 4°C until they were ready to be sent for sequencing. The genomic DNA was shipped on dry ice to Novogene Corporation Incorporated for whole-genome sequencing via an Illumina NovaSeq 6000 platform (Illumina Inc., San Diego, USA) to create 150 base pair, paired length reads. Quality control of the raw sequence data was performed using FastQC version 0.11.9 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). De novo assembly was performed on the sequence data using Unicycler version 0.4.9 to generate draft genomes using paired short-read settings (Wick et al. 2017). The Unicycler assembly pipeline was run using a high-computing system, Savio, at UC Berkeley. The paired short-read sequences were evaluated using QUAST version 5.2 (Mikheenko et al. 2018). Antibiotic resistance genes were identified using the ABRicate tool version 1.0 (Zankari et al. 2012). The ResFinder database, one of the pre-downloaded databases in ABRicate, was cross-referenced to detect resistance genes in the sequences. The sequence data underwent multilocus sequence typing using the MLST package version 2.23.0 (https://github.com/tseemann/mlst) to identify unique clonal groups based on highly conserved bacterial genes (Jolley et al. 2018). Core genome multilocus sequence typing, which uses approximately 2,500 bacterial genes to assign sequence types to isolates, was completed with the assistance of the Center of Genomic Epidemiology (Clausen et al. 2018; Zhou et al. 2020).
Ethics Statement
This project did not involve any intervention or interaction with humans as research subjects and did not involve the collection of samples from live animals. All samples were collected as part of a study approved as nonhuman subjects research by the Ethics Committee of the Center for Health Studies of the Universidad del Valle de Guatemala (Protocol 267-05-2022).
Results
All 30 soiled toilet paper samples were positive for putative E. coli, and 10 of these samples were positive for putative 3GCR-EC, based on their growth on CHROMagar™ ESBL, which further confirmed their identification as E. coli. From the 10 positive samples, 13 isolates with distinct morphologies were sequenced and selected for whole genome sequencing.
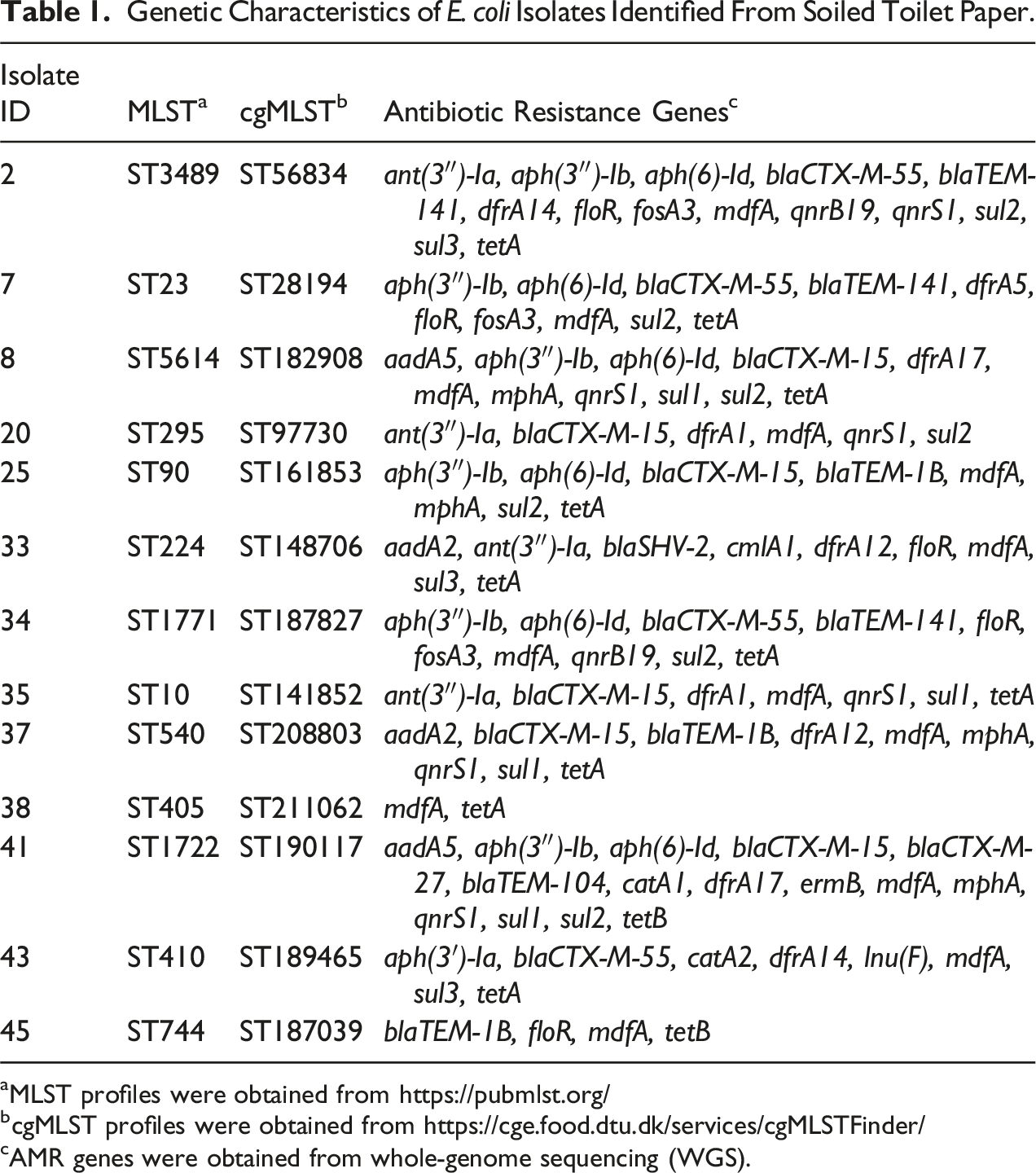
Thirty-four antibiotic resistance genes (ARGs) were detected among the 13 isolates sequenced (Figure 2). The most prevalent antibiotic resistance gene was mdfA, appearing in all 13 of the isolates (Figure 2). This gene confers resistance to a broad range of antibiotics, including aminoglycosides, beta-lactams, macrolides, and fluoroquinolones (Yardeni et al. 2018). Ten of the 13 isolates had tetA detected, which is part of the tet gene family involved in resistance to the tetracycline class of antibiotics (Figure 2) (Roberts 2005). There were 13 unique sequence types detected among the isolates, and many of them were “pandemic” lineages known to cause disease globally (Table 1) (Manges et al. 2019). Abundance of antibiotic resistance genes across E. coli sequence types (ST). Black stars indicate the ST is a well-recognized pandemic lineage, according to Manges et al. (2019). Genetic Characteristics of E. coli Isolates Identified From Soiled Toilet Paper.
aMLST profiles were obtained from https://pubmlst.org/
bcgMLST profiles were obtained from https://cge.food.dtu.dk/services/cgMLSTFinder/
cAMR genes were obtained from whole-genome sequencing (WGS).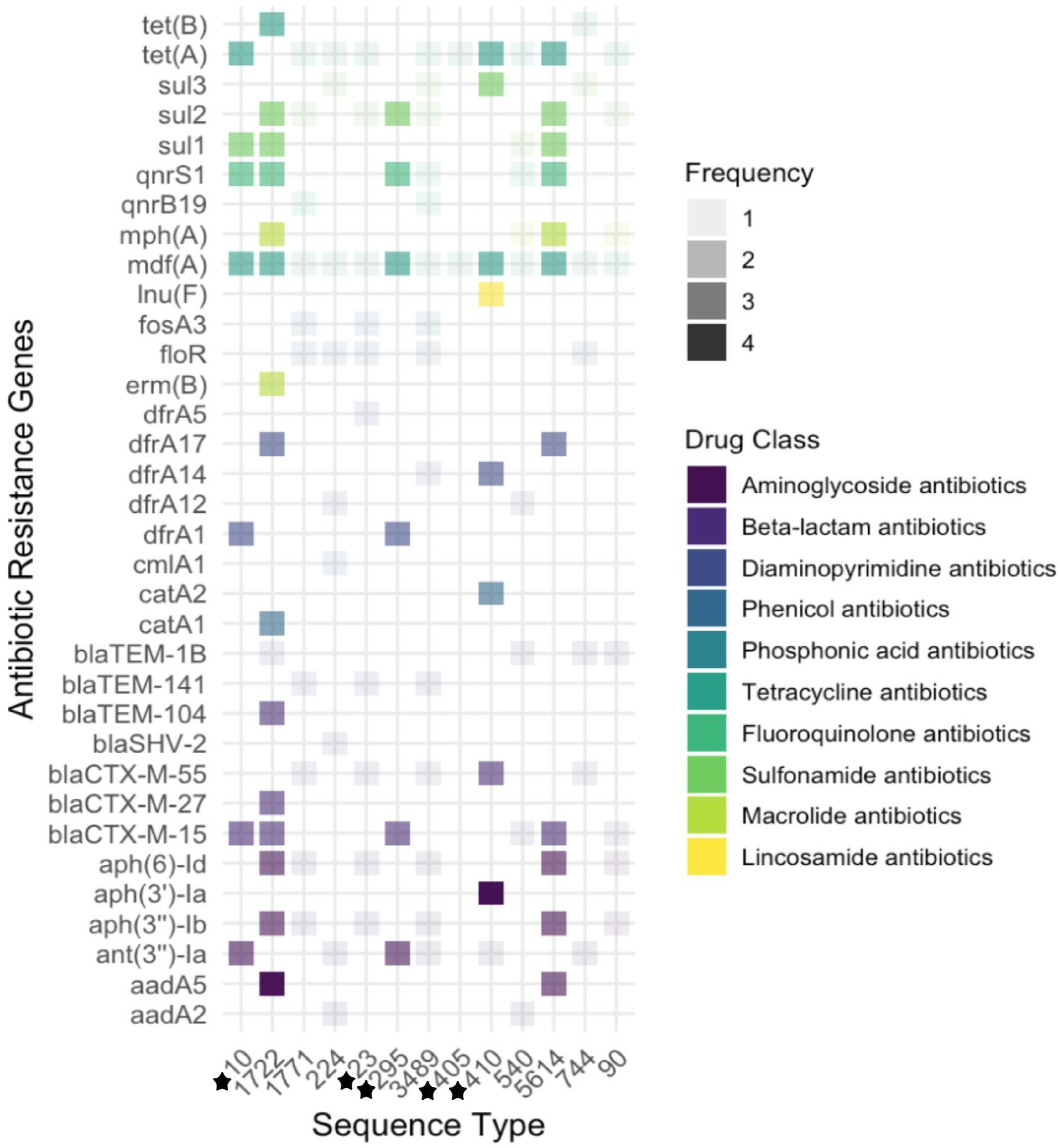
Discussion
In this study, we explored the use of soiled toilet paper as a method for antibiotic resistance (ABR) surveillance. Using limited resources, we were able to consistently culture E. coli from toilet paper, which then allowed for the subsequent detection of 3GCR-EC from the samples. Our methods provide insight on the utilization of low-cost culturing to screen soiled toilet paper for antibiotic resistance phenotypes before whole genome sequencing. All of the toilet paper samples that presented phenotypically for antibiotic resistance also had antibiotic resistance genes present based on genomic analyses.
Findings from this study demonstrate the effectiveness of collecting soiled toilet paper to identify ABR E.coli and may provide an alternative method for ABR surveillance. Wastewater surveillance, for example, is a promising, cost-effective method to obtain prevalence measures of antibiotic resistance among humans (Blaak et al. 2021; Chau et al. 2022) but is limited in its ability to report individual-level colonization of humans. Additionally, grab samples captured in wastewater surveillance systems are subject to selective pressures from contaminants in the wastewater (e.g., antibiotic residues) that may cause bias, resulting in identification of strain types that are more or less resistant to changes in pH, temperature, oxygen levels, and antibiotic residues. Toilet paper sampling has the capacity to capture a more precise estimate in the variation of antibiotic-resistant bacteria identifying dominant antibiotic-resistant strains among the sampled population. These methods expand on existing commercial clinical assays used for detection and diagnosis of gut microbiota, fecal occult blood and colorectal cancer (Al et al. 2018; Hsin-Yao et al. 2020).
Based on multi-locus sequence typing conducted on the isolates, we have identified several STs that are of concern; ST10, ST23, ST405, and ST410 have been indicated to be the most commonly reported pandemic lineages in studies of ExPEC (Manges et al. 2019). These sequence types have been found globally among hospital and community acquired infections, making them high-risk indicators for multidrug resistance.
ST10 is the most prevalent E. coli ST described in EnteroBase, a public database of variant strains of enteric bacteria, and the ST ranges from drug-susceptible to multidrug-resistant (Zong et al. 2018). ST10 has also been described as a uropathogenic E. coli associated with urinary tract infections found among college-aged women (Yamaji et al. 2018). During another clinical investigation of carbapenemase-producing E. coli in Denmark, ST10 was one of the most commonly detected sequence types among patients, highlighting its importance in hospital infections (Hammerum et al. 2020). ST10 and ST23 have been commonly found in clinical isolates in Spain during investigations of ESBL-producing E. coli (Oteo et al. 2010). A study of clinical isolates in French hospitals found ST23 to be a dominant clone expressing Extended-Spectrum AmpC β-Lactamases (Crémet et al. 2010). In a study of nursing home residents diagnosed with recurrent UTIs, ST23 was included among the identified sequence types (Hidad et al. 2022). ST405 has been described as an emerging uropathogenic E. coli, and in a study of ESBL-producing E. coli in Sweden from 2007 to 2011, 2%–6% of the isolates belong to this sequence type (Brolund et al. 2014). Along with ST10, ST405, and ST410 are included on a list of high-risk clones that are indicated to be highly transmittable, persistent in the environment, and known to cause infections in humans (Kocsis et al. 2022).
The repertoire of antibiotic resistance genes found among the selected isolates included genes known to confer resistance to broad-spectrum antibiotics, such as mdfA and tetA, but their detection in humans is limited. Studies have shown that tetA genes commonly exist in E. coli isolates collected from humans and animals, displaying their prevalence in these reservoirs (Bryan et al. 2004). mdfA genes are less commonly described in humans. In a study of giant pandas, mdfA was one of the widely found antibiotic resistance genes among E. coli isolates (Guo et al. 2015). In a clinical isolate from a urine sample collected in Turkey, mdfA was noted among the genetic determinants contributing to its resistance (Turumtay 2023). There have been limited analyses of antibiotic resistance genes circulating in Guatemala, and in Central America more broadly (O’Neal et al. 2020). One study by Villegas et al. (2016) identified carbapenemase genes in clinical isolates, which were absent in the isolates sequenced in this study (Villegas et al. 2016).
Many limitations remain to implementing this surveillance method. More research is needed to compare toilet paper samples to control samples (e.g., fresh human fecal samples) if public health researchers and practitioners move forward with this approach for surveillance. Characterizing data in comparison to a gold standard and defining variability in measurement are key steps for refining this surveillance approach. For example, it may be that the toilet paper environment allows certain E. coli strains to thrive and more dominant strains to die-off. This was observed in fecal matter studied over a 24-hour period, where major shifts in the population structure and ABR patterns were observed in dominant E. coli (Barrera et al. 2019). In this pilot study, there is a possibility of selection bias due to the use of convenience sampling. The toilet paper samples are likely representative of sub-populations within Guatemala City who access these urban markets and also use public toilets. More research would be needed to understand how samples of soiled toilet paper from trash bins vary from a more rigorous study enrollment methods (e.g., stratified random sample of individuals).
Despite the limitations described above, the collection and analysis of soiled toilet paper from wastebaskets have the potential to provide important data that can be used to prevent and control the spread of ABR at a relatively low cost, compared to studies that must enroll study participants, seek participant consent and collect human fecal samples. This method of surveillance could facilitate studies that aim to identify context-specific behaviors or risk factors for colonization with ABR or enteric pathogens. Identification of antibiotic resistance genes in bacterial isolates from toilet paper could also be used in tandem with samples from wild and domestic animals, the food supply and the broader environment to understand important reservoirs of ABR, which could inform targeted prevention and control efforts.
Footnotes
Acknowledgments
We are grateful to the Universidad del Valle de Guatemala staff for their time, resources, and participation in this research project. We want to thank the Center for Global Public Health for the additional funding used for research materials. Finally, we greatly appreciate Nazario Lopez for his assistance in the field and his hard work completing sample collection.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Research reported in this publication was supported in part by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health (NIH) under Award Number R01AI135118 and R01AI167989. The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
