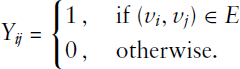Abstract
Mixture models are probabilistic models aimed at uncovering and representing latent subgroups within a population. In the realm of network data analysis, the latent subgroups of nodes are typically identified by their connectivity behaviour, with nodes behaving similarly belonging to the same community. In this context, mixture modelling is pursued through stochastic blockmodelling. We consider stochastic blockmodels and some of their variants and extensions from a mixture modelling perspective. We also explore some of the main classes of estimation methods available and propose an alternative approach based on the reformulation of the blockmodel as a graphon. In addition to the discussion of inferential properties and estimating procedures, we focus on the application of the models to several real-world network datasets, showcasing the advantages and pitfalls of different approaches.
Introduction
The underlying idea of a mixture model is rather simple. Instead of assuming that the target variable follows a plain distribution, one considers a mixture of multiple distributions. Specifically, for a random variable Y, one assumes
While most of the literature cited above deals with a univariate response variable Y, in this article we aim to look at multivariate data with
If we consider
The rest of the article is organized as follows: Section 2 presents some real-world network datasets together with the potential questions that we face in analysing them. Those datasets will be later used to demonstrate the capabilities of stochastic blockmodels. Section 3 describes the blockmodelling framework in more detail, and introduces some of its most prominent variants and extensions. Section 4 compares the different estimation routines that are available, and introduces a Monte Carlo-based EM estimation routine under graphon representation. The empirical analysis of the previously introduced datasets is then carried out in Section 5, making use of the previously described models to tackle the questions posed in Section 2. Finally, Section 6 ends the article with some comments and conclusive remarks.
In order to demonstrate the capabilities of stochastic blockmodels, we have chosen network datasets pertaining to three different domains, namely political science, biology and sociology. Despite the different domains, the networks share the presence of some form of underlying community structure, or at least the appearance thereof. They all therefore lend themselves to be modelled through the use of mixture components. General descriptive measures of the data examples, which consist of undirected graphs, are given in Table 1, which shows that all three networks are of medium size and range from very dense to relatively sparse.
Descriptive statistics for the studied networks

Descriptive statistics for the studied networks

The first network that we introduce is constructed using data from the Alliance Treaty Obligations and Provisions project (Leeds et al.(2002)). The dataset provides information on military alliance agreements pertaining to all countries of the world. For the analysis we consider alliances that were in force in the year 2016. The countries are taken as nodes, and an edge between two countries is present if the two countries take part in a ‘strong’ military alliance treaty. More specifically, the alliances that we consider strong are defensive and offensive ones. This means, respectively, ‘\textsl alliances in which the members promise to provide active military support in the event of attack on the sovereignty or territorial integrity of one or more alliance partners’ and ‘\textsl alliances in which the members promise to provide active military support under any conditions not precipitated by attack on the sovereignty or territorial integrity of an alliance partner, regardless of whether the goals of the action are to maintain the status quo’ (see Leeds et al.(2002)). Note that, in general, an alliance can involve more than two nodes: Representing the network using dyadic edges only thus leads to the loss of some information. For example, pairwise edges between countries
Butterfly similarity network
The second real-world instance is a butterfly similarity weighted network, constructed using the data presented by Wang et al.(2009) and available from Zitnik et al.(2018). Each node represents a butterfly, and valued edges depict visual similarities between them. More specifically, pairs of butterflies with some positive degree of similarity between them are connected by a weighted edge, while no edge is present if the similarity score between the two is zero. The absence of an edge is thus equivalent to the presence of an edge with weight zero. The similarity scores lie in the interval
Email exchange network
The last network considered consists of anonymized email data from a large European research institution, collected between October 2003 and May 2005 (Leskovec and Krevl (2014)). Each node in the network represents a person, and an edge between nodes
Stochastic blockmodels: formulations and variants
The standard stochastic blockmodel
As anticipated in the introduction, if we consider the network 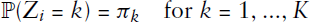
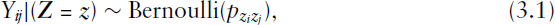
For estimation, a numerically simpler setting can result by approximating the binomial distribution through a Poisson distribution. This approximation is justified since the network density is usually low, implying that 
A well-known extension of the classical SBM is the degree-corrected stochastic blockmodel, introduced by Karrer and Newman (2011). In their work, the authors show how the standard stochastic blockmodel implicitly assumes the degree structure within communities to be relatively homogeneous. This, combined with the fact that many real-world networks exhibit extremely skewed degree distributions (Simon (1955); Barabási and Albert (1999)), leads the model to often only be able to find core-periphery type block structures, where node grouping is predominantly driven by degree similarity. To bypass this issue, Karrer and Newman (2011) introduced the idea of degree correction, making the probability of an edge depend not only on group membership, but also on node-specific heterogeneity parameters. More precisely, the original version of the degree-corrected SBM can be written in the same way as (3.2), but in this case

The stochastic blockmodel can also be formulated through the graphon model class, which recently received a lot of attention concerning the modelling of complex networks. Although the scope of the structures representable as a graphon is quite large, its formulation is rather simple. Let us therefore introduce 
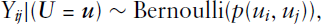
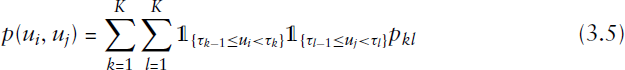
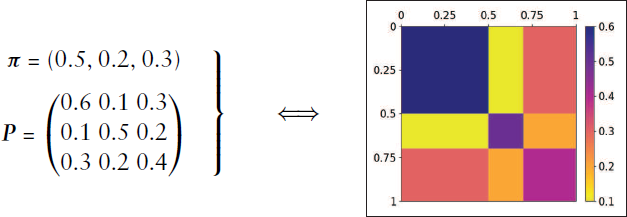
It is thus not difficult to see how this formulation of graphon models is equivalent to SBMs.
In this context, it should be noted that the graphon model suffers from major identifiability issues, which, with regard to the SBM representation, also include the label switching problem (we refer to the Appendix for more details and illustrations). This non-identifiability arises from the fact that any permutation of
Many other variants and extensions of SBMs exist. These include the mixed membership model (Airoldi et al.(2008)), in which nodes can belong to multiple communities simultaneously, and the hierarchical stochastic blockmodel (Peixoto (2017)), in which communities are comprised of meta-communities, leading to a hierarchical block structure. A matter of simplifying the model representation is what motivates the microcanonical variant of the SBM (see, e.g., Peixoto (2012)), where the structural pattern is strictly fixed in absolute values. This, in turn, allows for fitting more elaborate generative models, which usually require Markov Chain Monte Carlo (MCMC) techniques for evaluation, to larger networks and to an increased number of groups, as demonstrated by Peixoto (2017). The same author also proposed a nested hierarchical variant of the SBM (Peixoto(2014a)) in which the generative model inferred at an upper level serves as prior information to the one at a lower level, thus also providing an increased resolution when performing model selection. Despite its more elaborate formulation, this hierarchical model remains tractable, and it is feasible to apply it to very large networks. It is also possible to add covariates to the analysis, as initially proposed by Tallberg (2005) and further elaborated by Choi et al.(2012), Sweet (2015) and Huang and Feng (2018). A further extension is the mixture of experts SBM (see Gormley and Murphy (2010); White and Murphy (2016)), which allows covariates to enter the latent position cluster model in a number of ways, yielding different model interpretations. Extensions for more specific purposes have also been developed: Bouveyron et al.(2018) introduced the stochastic topic blockmodel, a probabilistic model for networks with textual edges. Their model addresses the problem of discovering meaningful clusters of vertices that are coherent with regards to both the network interactions and the text contents. Finally, another relevant approach that can be seen as a generalization of the SBM is the latent position cluster model proposed by Handcock et al.(2007) (originating from Hoff et al.(2002), see also Krivitsky et al.(2009)). It is worth noting that most of the mentioned specifications can be applied to binary data as well as to valued and count data (see, e.g., Nowicki and Snijders (2001)). In this article, we do not concentrate on these extensions, but focus on the more ‘classical’ SBMs.
Estimation techniques
Variational methods
The EM algorithm proved to be a powerful and numerically efficient way for estimating parameters in mixture models (see Aitkin (1980) or Friedl and Kauermann (2000)). Unfortunately, this does not extend to the estimation of stochastic blockmodels. The complete data log-likelihood resulting from (3.2) in the case of an undirected network equals
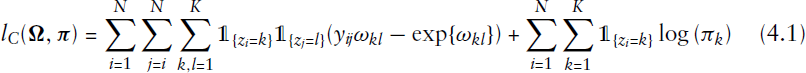
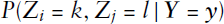
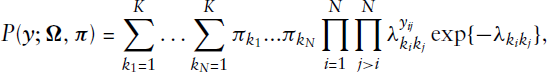
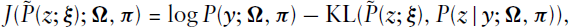
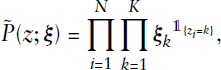
Another possibility for the estimation of stochastic blockmodels is to maximize the likelihood through vertex switching routines. The basic idea of this type of algorithms is the following: starting from an initial, possibly random group assignment, a starting value of the likelihood is computed. From there, one or more vertices are moved from one group into another, and the likelihood is computed again. The new allocation is then accepted or rejected based on a function of the previous and the subsequent likelihood, and such procedure runs iteratively until convergence is reached, that is until a maximum is found. Algorithms of this type include single-vertex Monte Carlo (see, e.g., Peixoto (2013), 2014b) and a local heuristic routine inspired by the Kernighan–Lin algorithm used in minimum-cut graph partitioning (Kernighan and Lin (1970); Karrer and Newman (2011)). In principle, computing the likelihood that many times may seem quite expensive. On the other hand, it is not always necessary to calculate the complete likelihood at each step. Depending on the model specification, it is often possible to write the change in the likelihood in a computationally efficient way, so that the algorithm becomes quite competitive in terms of speed. The chief issue with this type of algorithm is that, given the heuristic maximization routine, it is not possible to obtain a measure of uncertainty for group assignments. The procedure will only produce the graph partitioning that (locally) maximizes the likelihood, without any additional information. This is fine if the problem at hand is one of pure community detection, but can become problematic if the goal is proper mixture modelling, as the stochastic component of the mixture is lost. Another potential issue is the possibility to get stuck at local maxima, which usually is tackled by running the procedure several times with different (random) starting points.
Monte–Carlo-based EM estimation under graphon representation
A third and to some extent novel estimation routine is to estimate the block structure using its graphon representation. Although it is not quite clear how this approach competes with already existing methods, our ambition here is to demonstrate a further possible form of representing and estimating mixture models in networks. Such a model can be fitted appropriately by applying an EM-type algorithm including Gibbs sampling in the E-step. As mentioned above, EM-based algorithms are a common approach to estimate mixture models as well as other models involving latent variables, although in the case of networks the task becomes analytically intractable and numerically demanding. We therefore make use of MCMC techniques to approximate the complex posterior distribution of the latent quantities, which here reflect the group assignments. In this approach, we thus slightly reformulate the stochastic blockmodelling procedure, relating it to graphon estimation (see, e.g., Latouche and Robin (2016) or, for the reverse link, Olhede and Wolfe (2014) and Airoldi et al.(2013)). We here want to follow the estimation approach of Sischka and Kauermann (2019), applying it to SBMs. The idea is to make use of model (3.4) and estimate, in the M-step, the parameters of 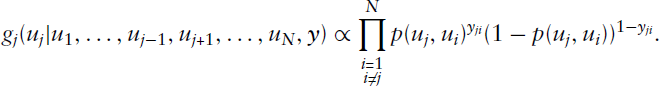
This allows for applying Gibbs sampling in a straightforward manner. Details on this sampling scheme, as well as remarks on the associated potential issues of label switching and non-identifiability, are given in the Appendix. In this context, we underline how the issue of label switching is prevented through the EM algorithm on the primary level of the estimation procedure (apart from the exceptional case of complete symmetry, as described in the Appendix). In comparison, label switching is a common problem when making use of an overall Bayesian estimation procedure (if the MCMC scheme is run for sufficiently long, see Stephens (2000)), where one randomly draws quantities from the corresponding posterior distributions in alternating fashion. This, in contrast, is circumvented in the EM framework, since in the E- and M-steps the results of the respective other step are kept fixed and, based on that, the ‘optimal’ solution is carried out to be used for the next iteration. Parameter estimates are thus not achieved by averaging over several iterations but are given for each iteration separately. Therefore, with regard to our estimation routine, no post-hoc relabelling is required, and assignments can be adopted as deduced from the subordinate Gibbs sampling scheme. Making use of the sampling sequence, we specify the posterior mode in the mth iteration using 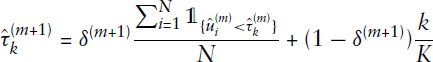
The M-step is then carried out by maximizing the likelihood conditionally on 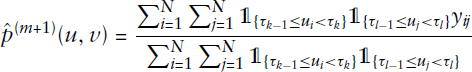
In contrast to non-stochastic estimation routines, such as the vertex switching algorithm discussed in Section 4.2, this modelling approach naturally yields information about the inherent uncertainty of the proposed group allocation. In order to achieve this, we run the E-step one more time after the algorithm has converged. The resulting Gibbs sampling sequence of this last iteration then reveals the distribution of the node allocation with respect to the model estimate
Choosing the number of blocks
A general big challenge in mixture models (and hence also in stochastic blockmodels) lies in the choice of the number of mixture components (blocks). In fact, most of the variants presented so far require that number to be known a priori. This is typically not true in real-world applications. In mixture models the question of choosing the number of mixture components is tackled, for instance, in Aitkin (2011). In the field of stochastic blockmodels, a comprehensive list of different approaches is provided by Lee and Wilkinson (2019). Approaches based on penalized likelihood criteria have emerged. In particular, Wang and Bickel (2017) consider an approach based on the log-likelihood ratio statistic, enabling the use of a likelihood-based model selection criterion that is asymptotically consistent. Other techniques are also available: Chen and Lei (2018) develop a network cross-validation approach which is based on a block-wise node-pair splitting technique, combined with an integrated step of community recovery using sub-blocks of the adjacency matrix. Mariadassou et al.(2010) base the choice on an Integrated Classification Likelihood criterion. The number of blocks can also be estimated using ‘collapsed’ approaches, where the model parameters are integrated out in a Bayesian formulation of the model. The model space and cluster allocations can then be estimated using a greedy search routine (Côme and Latouche (2015)) or using MCMC (McDaid et al.(2013)). Another possible approach is that of Peixoto (2013), who uses the Minimum Description Length principle, which seeks to minimize the total amount of information required to describe the network and avoid overfitting. This also allows to deduce general bounds on the detectability of any prescribed block structure, given the number of nodes and edges in the sampled network. Finally, Riolo et al.(2017) (see also Newman and Reinert (2016)) present a method for estimating the number of communities in a network using a combination of Bayesian inference and an efficient Monte Carlo sampling scheme. While other approaches have been proposed, we will not go into further detail here. For modelling the previously described networks, when possible we select K such that the resulting number of blocks coincides with the ground truth. If such ground truth is not available, we make use of the Akaike Information Criterion (AIC), which can be easily calculated when using the graphon representation-driven algorithm.
Application to real world networks
International alliances network
To model the network we use the standard version of the stochastic blockmodel, as in (3.1). Estimation was performed using the Monte Carlo-based EM routine under graphon representation. Applying the AIC yields seven communities as the optimal dimensionality of the blockmodel. The resulting fitted block decomposition is given in Figure 1. Network visualization, as for the rest of the examples in this section, is carried out through use of the open-source tool Gephi (Bastian et al.(2009)). The associated world map is shown in Figure 2, where countries are coloured by block. States coloured in grey on the map are isolates in the network, meaning that they were not involved in any strong military alliance in 2016. Moreover, China, Cuba and North Korea are only connected to each other, and are thus isolated from the rest of the network. Those countries have therefore been excluded from the model fitting.
Global network of political alliances in 2016. Two countries are connected if they have taken part in a strong alliance treaty. Labels indicate country codes, while nodes are coloured by block memberships found through the standard stochastic blockmodel
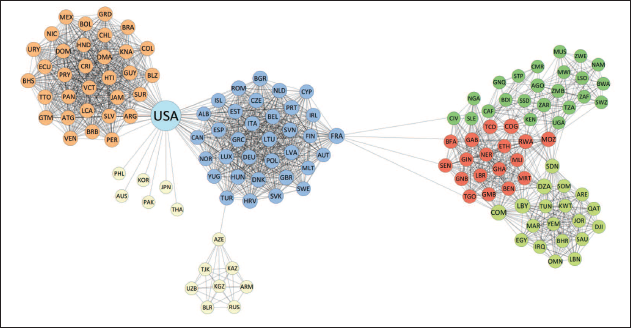
Global network of political alliances in 2016. Two countries are connected if they have taken part in a strong alliance treaty. Labels indicate country codes, while nodes are coloured by block memberships found through the standard stochastic blockmodel
The plots show how the blocks recovered by the stochastic blockmodel are very much related and in accordance with the geopolitical structure of the modern world, while also revealing some interesting patterns. The network representation can be visually split into two large components. In the first component, on the left side of the plot in Figure 1, the central block contains most European countries together with Canada. This block is very densely linked, as most of the countries inside it belong to NATO and other major alliances. The block on the very left pretty much coincides with Central and South America, and it is also quite dense. The European and the American block are linked by the USA, which, given its unique connectivity behaviour, constitutes a block on its own. The bottom block includes mostly Asiatic countries as well as some Pacific states, which share a very low edge density.
World map with countries coloured by block membership. Colours are kept consistent with Figure 1. Countries not included in Figure 1 are isolates, meaning that they were not part of any strong military alliance as of 2016, with the exception of China, Cuba and North Korea, which are only connected among themselves. Labels indicate the three countries with the highest uncertainty in block membership
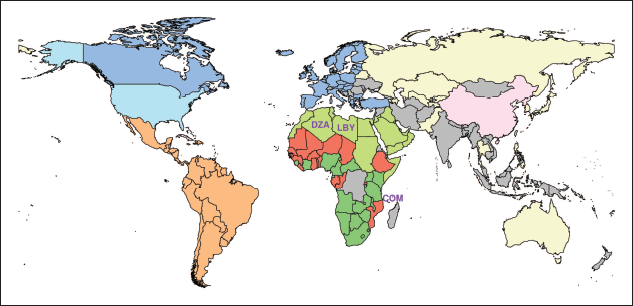
The other component of the network, on the right-hand side of Figure 1, is made out of three blocks. The block on the bottom contains all countries from the Middle East together with Northern African countries such as Libya, Tunisia, Egypt and Morocco. The middle block includes countries from Central and Western Africa. Finally, the upper block is composed of Southern African countries. The central block is well connected with both the northern and the southern blocks, mostly through countries that share borders, while the latter two blocks are instead only directly bridged by Sudan. As an additional note, we can observe that the two major components of the network are linked exclusively through France, that, while belonging to the European block, acts as a bridge between Africa and Europe itself. Finally, it is evident how transferring the group assignments to the world map in Figure 2 clearly reveals a general geographic proximity of countries belonging to the same community.
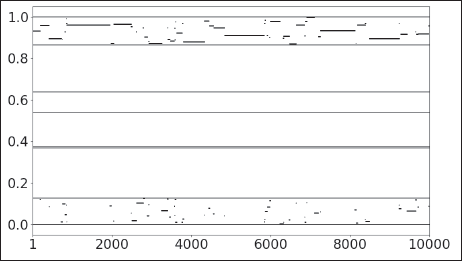
In addition to the detected block structure, we also investigate the uncertainty of the node allocation, using the Monte Carlo-based posterior samples. We therefore consider the last Gibbs sampling sequence after the algorithm has converged. More specifically, we take a look at the three countries with the lowest values of the normalized Gini coefficient calculated over the allocation frequencies, which in turn imply the highest uncertainty. These countries are Libya (LBY), Algeria (DZA) and Comoros (COM), which all belong to the Arabic block. The switching of communities exhibited by Libya throughout the posterior sampling is illustrated as an example in Figure 3. It shows how the sample for
Posterior sample of the latent quantity U for Libya plotted against the MCMC states. Horizontal lines represent community boundaries
Posterior frequencies for the three countries with the highest uncertainty in their community memberships. The corresponding normalized Gini coefficient is depicted in the rightmost column
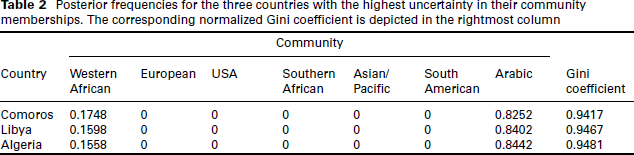
The standard SBM as showcased in the previous section is suitable for modelling binary networks. As described in Section 2, this dataset is, however, comprised of similarity scores which lie in the interval [0,1.55]. While it would be possible to binarize the data, for example defining a threshold within the domain as cut-off, this would lead to considerable information loss. We therefore use the Poisson version of the stochastic blockmodel as defined in (3.2), taking advantage of the fact that this variant is suitable to treat multi-edged networks as well as binary ones. To fit this model, we discretized underlying similarity measures into count data through binning. More specifically, each similarity measure was multiplied by 100 and rounded to the nearest integer, resulting in natural values between 0 and 155. Estimation on the resulting multi-edged network was performed using the Variational EM approach developed by (Mariadassou et al.(2010); see also Daudin et al.(2008)) and implemented in the
Comparison between ‘ground truth’ communities (species) and groups found by the Poisson stochastic blockmodel in a network of butterflies, with weighted edges representing the degree of visual similarity between them
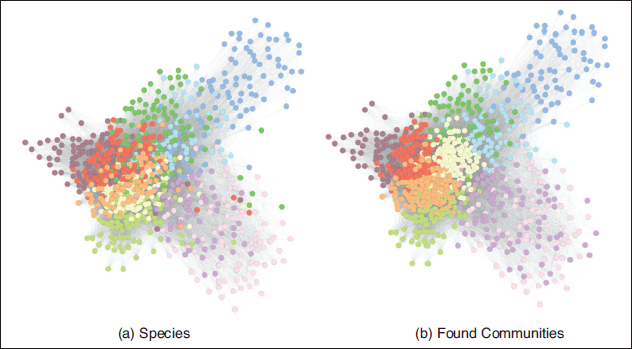

Comparison between ‘ground truth’ communities (species) and groups found by the Poisson stochastic blockmodel in a network of butterflies, with weighted edges representing the degree of visual similarity between them
At a first glance, we can see that the communities recovered mirror the real species relatively well. The most evident difference lies in the fact that two of the species (located towards the centre of the plot) are apparently really similar according to the utilized measure of visual similarity, and are therefore split up by the blockmodel. It is also interesting to note how communities found by the stochastic blockmodel seem to be visually clearer than ground truth ones. This is attributable to the fact that network visualization techniques and the clustering algorithm utilized are both based solely on the ties between the nodes, and thus tend to be more in accordance. The ground truth, on the other hand, is always given a priori, and can easily have outliers in terms of connectivity behaviour. In this specific case, visualization and blockmodelling are both based on the aforementioned measure of visual similarity between butterflies, while the ground truth communities are given by the classification of butterflies into species by biologists. This at least partially explains the discrepancy between the ground truth communities and the positioning of the nodes in the visualized graph. Despite this discrepancy, in general, the structure that was found does not appear to present major differences from the biological classification of the species. To quantify the goodness of the recovered block structure compared to the ‘ground truth’ communities, several measures are available (we refer to Jebabli et al.(2018) for a comprehensive survey). Here we opted for the Rand index, a measure of similarity between two data clusterings that can simply be described as the number of agreements in classifying pairs divided by the total number of pairs (Rand (1971)). The index takes values between
This network of emails within a research institution exhibits a skewed degree distribution, as shown in Figure 5. This type of degree distribution is typical of real-world social networks, and leads the classical SBM to often only be able to find core-periphery type block structures, with nodes grouped mostly on the basis of degree similarity.
Empirical degree distribution of the email exchange network
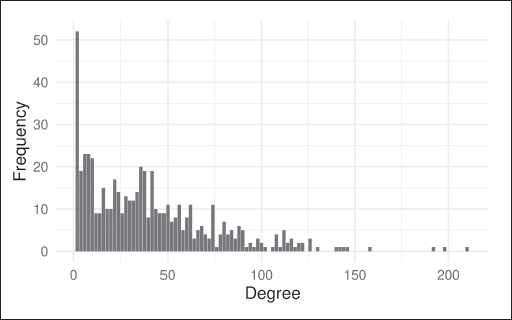
Empirical degree distribution of the email exchange network
As explained above, one way to circumvent this issue is to use degree correction. For this application, we therefore made use of the original version of the degree-corrected stochastic blockmodel as in (3.3) (Karrer and Newman (2011)). The results of the model fitting, together with the partitioning of the network into real departments, are visualized in Figure 6.
Comparison between ‘ground truth’ communities (departments) and groups found by the degree-corrected stochastic blockmodel in a network of email exchanges within a large European research institution
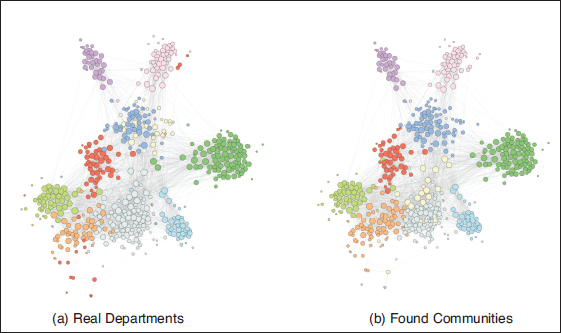
Looking at the plots, it is evident how the model with degree correction is able to recover the communities quite accurately. Comparing the partition discovered by the degree-corrected SBM with the actual departments, one small department (depicted towards the upper-centre of the figure) merges into another one close to it, and an additional block is therefore found at the bottom-centre of the plot, splitting the larger bottom department into two. Other than that, the structure found is remarkably similar to the partition induced by the departments, with some exceptions due to the existence of disconnected components within departments. In this case the Rand index is equal to 0.95, indicating a very high level of agreement among the partitions. For comparison purposes, we also fit a standard SBM to the same data, and computed the Rand index for the partition found with that as well. The resulting value of the index only amounts to 0.86, underlining the importance of applying degree correction in this case.
Mixture modelling can be extended to network data through stochastic blockmodels. Networks are rather complex structures, leading to computationally demanding estimation routines. Several algorithms specific for this class of problems have emerged over time, some of which are discussed in this article. We also provided an overview of different types of blockmodels by applying them to real-world network datasets. Among others, one of the models that we showcased is the degree-corrected stochastic blockmodel, which is particularly well suited for networks with a highly skewed degree structure.
Considering stochastic blockmodels (and community detection problems) as mixture models opens up a new avenue of extensions and novel models. Looking at the many model proposals in the field of mixture, ranging from mixing different distributions towards the mixture of experts, it is evident that these extensions can be brought forward in network modelling with mixtures as well. In fact, block-wise constant connectivity probabilities could be extended towards non-constant ones. Moreover, covariates could also be included. These extensions lie well beyond the scope of this article, but it is evident how the long history of mixture models, which started with Pearson (1894), has not come to an end, and extends promisingly in the realm of networks.
Appendix
Details on MCMC sampling scheme
Assuming 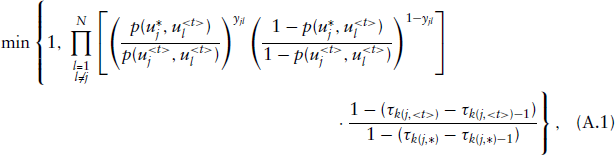
Label switching and non-identifiability
As has been extensively discussed in other works, approaching the conceptual formulation of mixture models by MCMC methods induces the label switching problem (see, for example, Stephens (2000)). This issue describes the invariance of the likelihood under relabelling of the mixture components. However, since in the proposed MCEM algorithm the model parameters are not part of the MCMC scheme but rather given as fixed based on the M-step, the label switching problem reduces to the exceptional case of symmetric parametrization. The two different situations can be exemplified by the configurations shown in Figure A.7.
Simple examples to illustrate the label switching problem in SBMs expressed through graphon representation. a) Two plausible blockmodels for the same seven-node network which are specified by
and
, respectively (illustrated by node colors), and a corresponding step function
(depicted in the right column). Both models yield the same value for the likelihood and can be transferred into one another through label switching. b) Two potential partitions of a six-node network, each forming a blockmodel. The partitions can again be transferred into one another through label switching, but in this case they both refer to the same function
. Only b) poses a label switching problem for the proposed MCEM algorithm
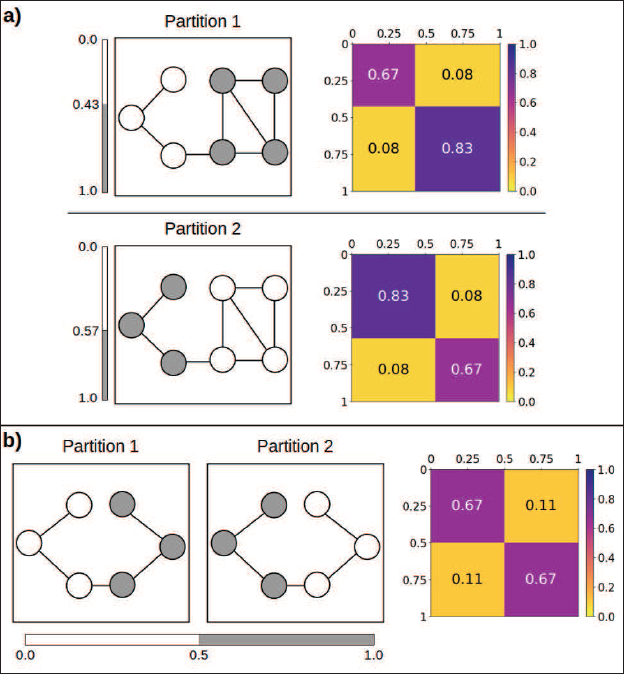
In both of the depicted cases (a) and (b), the two respective models describe and capture the exact same structure of the respective given network. That means none of them is preferable, and it is thus unclear beforehand to which label ordering the algorithm will tend. Nevertheless, regarding the non-symmetric configuration in (a), we point out that our MCEM algorithm will remain in either one of the partitions once that has been reached. At the stage of convergence, fixing the model parameters based on the M-step will leave the partition unchanged in the MCMC-based E-step (if the Gibbs sampling sequence is chosen to be sufficiently large). This is because the posterior distribution of the allocations is not invariant to label switching when
Another issue similar to the label switching problem which is inherent in graphon models is that of non-identifiability. This issue consists in the fact that different arrangements of the function
Simple example to illustrate the splitting of groups in SBMs expressed through graphon representation. The node colouring exhibits node assignments, with the corresponding graphon functions depicted on the right. Both of the models describe and capture the same block structure in the given network. Nevertheless, the upper model is preferable to the lower model with respect to the following three criteria: a monotonically non-decreasing marginal function, merging similarly behaving nodes, and parsimony in terms of the number of communities
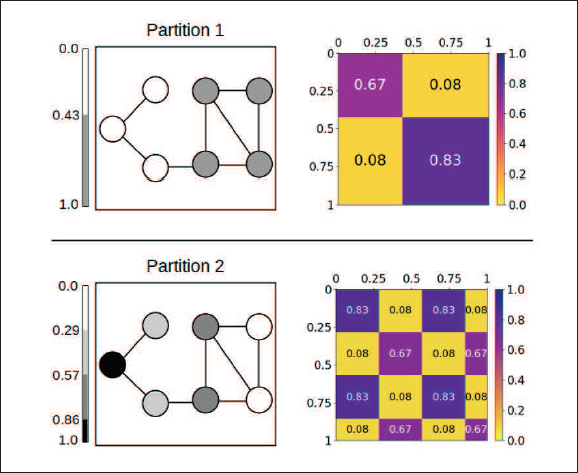
As has been mentioned in Section 3.3, the identifiability issue can be resolved by assuming a monotonically non-decreasing marginal function. This condition only applies to the upper model representation. However, considering our MCEM algorithm, the E-step aims to merge nodes with similar connectivity behaviour and therefore naturally prevents the splitting of groups. In addition, the lower representation is that of a blockmodel with four groups, a number which, compared to the upper representation, appears to be unnecessarily inflated. We hence argue that the identifiability issue in regards to the splitting of groups is a matter of the applied blockmodel dimensionality and can be also prevented through an appropriate choice of the number of mixture components. We therefore avoid the additional constraint of a monotonically non-decreasing marginal function.
Footnotes
Acknowledgments
We would like to thank the European Cooperation in Science and Technology [COST Action CA15109 (COSTNET)]. The authors of this work take full responsibility for its content. The first author would also like to thank Cornelius Fritz for invaluable comments and discussions. Finally, the last author would like to thank Murray Aitkin for his enthusiasm, ingenuity and open-mindedness with respect to statistics, and, last but not least, for his friendship.
Declaration of conflicting interests
Funding
The authors disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This work was also partly supported by the German Federal Ministry of Education and Research (BMBF) under Grant No. 01IS18036A as well as the Elite Network of Bavaria (ESG Data Science).