Abstract
Introduction:
Among the genetic factors for coronary artery diseases, PAI-1 4G/5G and ACE I/D polymorphisms can be noted. This study was carried out to investigate the association of these two polymorphisms and their synergism in coronary artery disease (CAD) from a sample of the Iranian population.
Materials and methods:
Sixty-one patients with a history of CAD and 92 healthy controls participated in our study. After DNA extraction from leukocytes, PCR was performed to characterize PAI-1 4G/5G and ACE I/D polymorphisms, using an amplification refractory mutation system technique.
Results:
In the studied patients, PAI-1 polymorphisms were 24.6%, 45.9%, and 29.5% for 4G/4G, 4G/5G and 5G/5G, respectively; the values for controls were 20.7%, 42.2% and 37.0%. The distribution rates of genotypes I/I, I/D and D/D in patients accounted for 29.5%, 45.9% and 24.6%; in the control group these figures were estimated to be 40.2%, 40.2% and 19.6%.
Conclusion:
Single and multivariate analyses showed a significant difference for the conventional risk factors, including hypertension, diabetes, hyperlipidemia, smoking and family history, for CAD between patients and controls (p value ≤ 0.001). However, no significant correlation was demonstrated considering ACE and PAI-1 polymorphisms either in association with 4G/4G or D/D genotypes or a combination of them in the Iranian population in the current study.
Keywords
Introduction
Coronary artery disease (CAD) is one of the main causes of morbidity around the world. Although diagnosis and treatments have improved, its prevalence shows a tendency to increase. 1 A set of environmental and acquired risk factors such as hypertension, diabetes, obesity, hyperlipidemia, smoking and family history have been accepted as the leading causes of CAD.2,3 However, some studies on family history 4 and twin investigations 5 have led researchers to propose a role for some genetic contributions to CAD. Therefore, some genetic factors have been proposed to trigger CAD, especially polymorphisms in genes regulating blood coagulation, blood pressure and lipid metabolism. 6 Since an occlusive or nonocclusive thrombus formation is the key point of CAD from an unstable angina to an ST-elevated myocardial infarction (MI), fibrinolysis has an important role in CAD. 7 Crucially, 50%–70% of cases of sudden death due to ischemic heart disease are the result of primary thrombotic events according to coronary Arteries. 8
Plasminogen activator inhibitor-1 (PAI-1) is one of suggested genetic factors predisposing patients to CAD. 9 PAI-1 has an important prothrombotic role and its expression is increased in sclerotic plaques. Also, high levels of this protein are detectable in the serum of post-MI patients. 10
One of the known polymorphisms of the PAI-1 gene is the 4G/5G single base pair (bp) polymorphism at 675 bp upstream from the transcriptional start site in the promoter region. This polymorphism and its association to CAD were first described by Dawson et al. in 1993. 11
Homozygosis for 4G has a 25% increase of PAI-1 in the plasma compared to 5G homozygous, while the 4G/5G genotype presents an intermediate level. 12 There is evidence supporting both effective relationships between PAI-1 and CAD,13–15 and a lack of this association16–18 based on different epidemiologic studies.
Angiotensin-converting enzyme (ACE) is the major component of the renin angiotensin system (RAS), which some studies indicate is important for CAD development. ACE converts angiotensin I to angiotensin II. In 1990, Rigat et al. described the insertion/deletion (I/D) polymorphism in ACE whose homozygosity in its deletion produces more ACE in serum and, therefore, more angiotensin II, which is a vasopressor and aldosterone-stimulating peptide. 19 Additionally, some studies show that angiotensin II increases PAI-1 expression based on transcriptional implication on its messenger RNA (mRNA). 20 For the first time, in 1992, Cambien et al. showed the association of the D allele with CAD. 21 Since then, numerous22–27 studies attempted to demonstrate this relationship, but it remains controversial. In this study we examine the association of PAI-1 and ACE polymorphisms as well as their combination effect on CAD, considering the conventional risk factors in case and control groups.
Materials and methods
Study population
A total of 153 individuals living in Tehran participated in our case-control study comprising 61 patients (45 men, mean age 55.3±8.6 years and 16 women, mean age 53.5±9.7 years) having experienced coronary heart disease from an unstable angina to ST elevated MI. These patients underwent angiography in Shahid Modarres Hospital. Ninety-two healthy controls who were also recruited for this study were referred to the Tehran blood transfusion center for voluntary blood donation (74 men, mean age 53.5±9.2 years and 18 women, mean age 54.4±8.9 years old).
CAD was confirmed in all patients by having coronary artery stenosis more than 50%, angiographically. This study was approved by the blood transfusion organization and the Shahid Modarres Hospital Ethics Committee. All participants were informed and signed the corresponding consent form. Demographic data and medical history such as conventional risk factors for CAD were collected in a questionnaire following a standard procedure of our research group.
Genotyping
Three ml of whole blood were collected from each participant in k2 ethylenediaminetetraacetic acid (EDTA). The DNA was isolated from leukocytes based on the selective detergent-mediated DNA precipitation from crude lysate according to the manufacturer’s instructions (Thermo Scientific). DNA concentration was measured at 260 nm and quality controls were performed.
PAI-1 and ACE genotypes were determined by polymerase chain reaction (PCR) using the amplification refractory mutation system (ARMS) technique. For ACE I/D polymorphism, as described, 28 a sense 5’GACCTGCTGCCTATAGACT3’ and anti-sense 5’GGGTAAAACTGGAGGATGGGT3’ primers that formed a 233 bp band for D and 521 bp band for I were used, respectively. If a sample was heterozygote both for 233 bp and 521 bp, bands should be detected. To avoiding mistyping of the DD genotype, another specific primer pair for I was applied: sense 5’GATTACAGGCGTGATACAGT3’ and antisense 5’GGGTAAAACTGGAGGATGGGT3’.
For the PAI-1 4G/5G polymorphism, as described, 29 two reactions were performed for each sample: one reaction for 4G containing its own specific primer and another one for 5G, respectively. The primers were the following: a sense control primer 5’AAGCTTTTACCATGGTAACCCCTGGT3’, a 4G allele-specific primer 5’ AGAGTCTGGACACGTGGGGA3’, a 5G allele-specific primer 5’AGAGTCTGGACACGTGGGGG3’ and a common anti-sense primer 5’ TGCAGCCAGCCACGTGATTGTCTAG3’. In both reactions, a 257 bp band was amplified as a control that formed by control sense and common anti-sense primers. In addition, a 136 bp band was observed for each 4G and 5G reactions if their alleles were present, yielded with allele-specific sense and common anti-sense primers. In the 4G/5G heterozygote the 136 bp band was amplified in both reactions and in the homozygote it was presented in only one reaction tube. All primers were synthesized by Bioneer (Korea). For both ACE and PAI-1, all reactions were performed in 25 μl volume containing 100 ng DNA, 10 mmol/l Tris-HCl (pH: 8.3), 50 mmol/l KCl, 1 mmol/l MgCl2, 0.2 mmol/l dNTP, 10 pmol of each primer (for PAI-1, 5 pmol of each sense allele-specific primers and control, and 10 pmol of common anti-sense primer.), 1 U Taq polymerase (Genet Bio, Korea). PCR reactions were carried out in the following thermal conditions: initial denaturation at 95°C for five minutes, afterwards for ACE it was followed by 35 cycles including denaturation at 94°C for 35 seconds, annealing at 52°C for one minute and extension at 72°C for one minute. For PAI-1 32 cycles were performed, including denaturation at 94°C for 35 seconds, annealing at 64.5°C for 30 seconds and extension at 72°C for 30 seconds followed by final extension at 72°C for five minutes for all reactions. PCR products were electrophoresed on 2% agarose and visualized under ultraviolet (UV) light after staining with ethidium bromide as shown in Figures 1 and 2.

PAI-1 4G/5G polymorphism detection in agarose gel. Gel (a) is for 5G and gel (b) is for 4G. Number 1 is a 5G/5G, number 2 is 4G/4G and number 3 is a 4G/5G sample. The 257 bp band is for control and the 136 bp band is yielded by 4G and 5G allele-specific primers. PAI-1: plasminogen activator inhibitor-1; bp: base pair.

Gel (a) shows the specimens that have insertion yielded a 521 bp band and those that have deletion yielded a 234 bp band. For heterozygotes both 521 and 234 bp bands are presented. Gel (b) shows a 233 bp band is formed from confirmatory primers for insertion for the same specimens. bp: base pair.
Statistical analysis
Genotype distribution and allele frequencies were represented as absolute numbers and percentages. Comparisons between patients and controls were performed by the Chi-square test. The associations between conventional risk factors and genotypes with CAD were reported using odds ratio (OR) with 95% confidence interval (CI). To determine crude OR and adjusted OR for other risk factors, simple and multiple logistic regressions were applied, respectively. To evaluate the interaction between studied genotypes, an interaction factor was introduced in the logistic regression model. The p value <0.05 was considered as significant correlation. Genotype distributions were compatible with Hardy-Weinberg equilibrium assessed by Chi-square test. All analyses were carried out using SPSS software for Windows version 20.
Results
A total of 153 individuals participated in our study; a greater percentage of patients (73.8%) and controls (80.4%) were male, with a mean age of 58.5±10.8 and 51.5± 6.4 years, respectively. The body mass index (BMI) of 29.5% of patients versus 19.6% of controls was ≥30.
As expected, differences between patients and controls were significant (p value ≤ 0.001) for conventional CAD risk factors such as hypertension (OR = 17.98; 95% CI: 6.41–50.45), diabetes (OR = 7.52; 95% CI: 2.82–20.09), hyperlipidemia (OR = 4.84; 95% CI: 2.03–11.52), smoking (OR = 4.15; 95% CI: 1.83–9.42) and family history (OR = 3.48; 95% CI: 1.72–7.05). Sex and BMI did not significantly differ between patients and controls. ACE and PAI-1 genotypes in patients did not significantly differ from controls (p value = not significant) (Table 1). To adjust OR for confounding factors, a multiple logistic regression was performed. Also other classical risk factors apart from sex and BMI contributed to CAD development. However, ACE and PAI-1 genotypes were not apparently involved in CAD susceptibility (Table 1).
Patient and control group demographic and genotypic characteristics.

Total cholesterol≥200 mg/dl, LDL ≥ 130 mg/dl, HDL≤ 40, Trigyserid ≥ 150 mg/dl or patent uses lipid-lowering drugs. b≥ 140/90 mmHg. cFasting blood sugar ≥120 or patient given hypoglycemic or insulin treatment. dEx- or persistent smoker. BMI: body mass index; LDL: low-density lipoprotein; HDL: high-density lipoprotein. NS: not significant; CI: confidence interval; OR: odds ratio; PAI: plasminogen activator inhibitor; ACE: angiotensin-converting enzyme; I: insertion; D: deletion.
As shown in Table 2, the distribution of different types of PAI-1 and ACE genotypes with their allele frequencies in case and control groups are shown with respect to the PAI-1 genotype; 24.6%, 45.9% and 29.5% of patients show the 4G/4G 4G/5G, and 5G/5G genotypes, respectively. The corresponding percentages for healthy blood donors were 20.7%, 42.2% and 37.0%. The allele frequency of 4G and 5G was 57% and 62.6% among patients, and 43% and 37.4% for controls.
PAI-1 and ACE genotype distribution and allele frequencies in patients and controls.

PAI: plasminogen activator inhibitor; ACE: angiotensin-converting enzyme; I: insertion; D: deletion.
Regarding the ACE genotype; I/I, I/D and D/D in patients amounted to 29.5%, 45.9% and 24.6%, respectively; in the control group they were 40.2%, 40.2% and 19.6%. Allele frequencies for the ACE gene were 55.7% and 64.55% for I and D in patients, and 44.3% and 35.5% for controls.
We also performed a study on the co-expression of various conditions of ACE and PAI-1 genotype in CAD development. The results are shown in Table 3; CAD risk was not increased in any condition significantly.
All condition of ACE and PAI combination for CAD development.
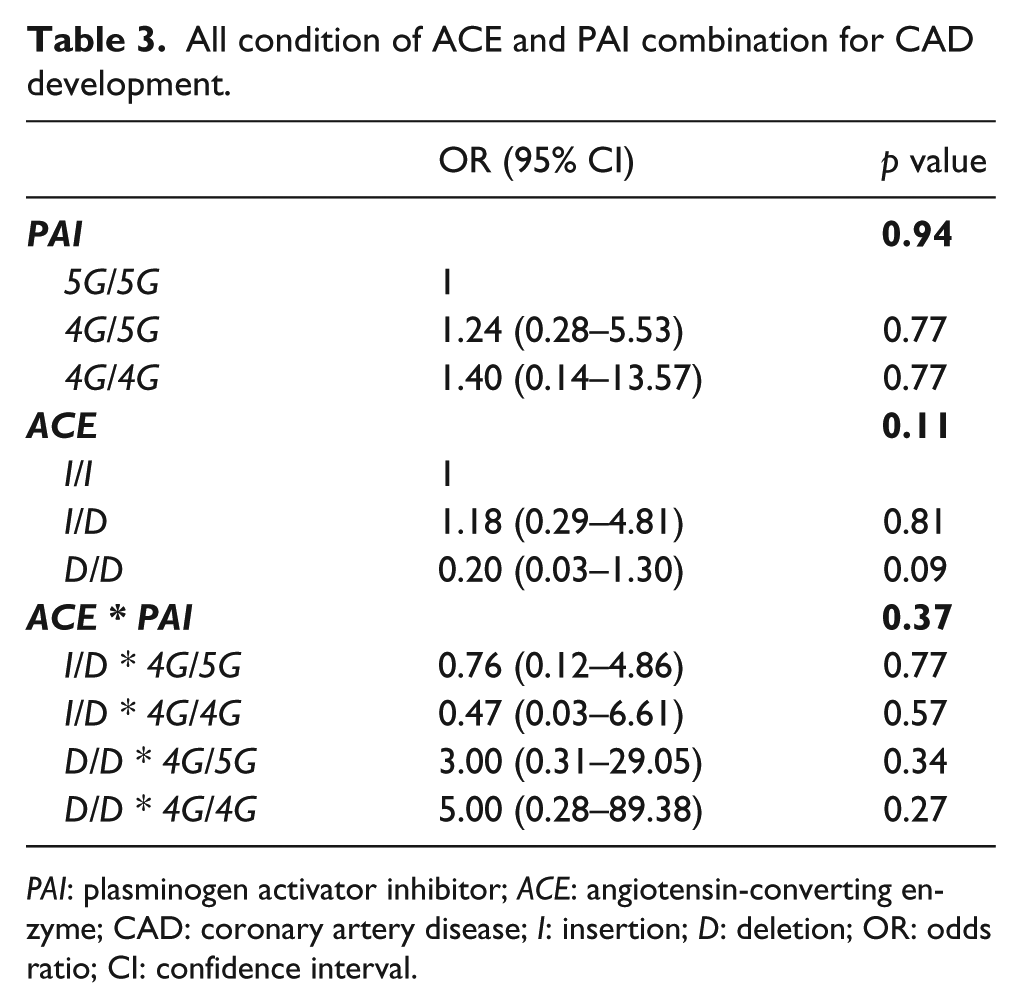
PAI: plasminogen activator inhibitor; ACE: angiotensin-converting enzyme; CAD: coronary artery disease; I: insertion; D: deletion; OR: odds ratio; CI: confidence interval.
Discussion
In this study, the ACE and PAI genotypes and their combinations did not show a significant correlation with CAD. Moreover, there was no significant genotype difference between patients and control groups. Therefore, our investigation supports that conventional and environmental risk factors are the principal causes of CAD2,3 by univariate and multivariate regression studies. Among conventional risk factors, hypertension, hyperlipidemia, diabetes, smoking and family history enhanced the hazard rate for CAD through crude and adjusted OR. On the other hand, sex and BMI did not have an indicative effect on hazard rate in the current study, although the selection of controls was biased toward males with standard BMIs. Therefore, our study cannot conclude on sex or BMI. Additionally, the number of paricipants involved in this study also limits the conclusions in this regard.
ACE I/D polymorphism is one of the genetic factors linked to CAD because of its critical role in atherosclerosis and also hypertension. 30 Since 1992, when the association between the DD genotype in ACE and CAD 21 was published, several researchers reported the prevalence of the ACE I/D polymorphism in different populations and ethnicities, including the positive effect of the DD polymorphism on CAD development.22–24 However, some studies did not find an association between the ACE polymorphism and CAD susceptibility.25–27 A meta-analysis was performed by Agerholm-Larsen and colleagues 31 to clarify the association of the ACE I/D polymorphism with ACE activity in plasma, blood pressure, risk of MI, ischemic heart disease and ischemic cerebrovascular disease through large and small studies. They selected 46 studies, including those investigating 32,715 white individuals. They concluded that the I/D polymorphism increases ACE activity in plasma, but has no effect on blood pressure. In addition, it was not an independent risk factor for both ischemic heart disease and ischemic cerebrovascular disease in large studies, in contrast to results from small studies.
Recently, another meta-analysis was carried out 32 to investigate the involvement of ACE I/D in the development of coronary heart disease in the Chinese population. The researchers surveyed 46 studies consisting of 5215 cases and 4782 controls. They showed a significant association between the DD genotype and vulnerability to coronary heart diseases in the Chinese population.
Something similar is observed for the PAI-1 gene. The PAI-1 4G/5G polymorphism in its promoter regulates the amount of this protein in cells. 4G/4G carriers produce more PAI-1 and possess less capacity for fibrinolysis. 11 Thus, it is thought that the PAI-1 4G/5G polymorphism is another genetic factor leading to thrombotic events in deep veins and coronary arteries; the association of the PAI-1 4G/5G polymorphism and MI 11 has been previously described. However, this association is still controversial owing to contradictory experimental results published by different groups. The studies carried out by Eriksson, 12 Song, 13 Abboud 14 and Margaglione 15 support the correlation of the PAI-1 4G/4G genotype with CAD development in contrast to the studies by Ye S, 16 Anderson 17 and Crainich. 18 Overall, in 2012 Gong 33 and colleagues performed a meta-analysis to investigate this association and showed that PAI-1 4G/5G is an independent genetic risk factor for CAD.
To our knowledge, these studies have never been carried out in the Iranian population, more specifically the association of the ACE I/D and PAI-1 4G/5G polymorphisms and their combination with CAD.
Additionally, there are a few studies that suggest the interaction between these two polymorphisms on CAD. In 2006, Loew and colleagues conducted a cohort study 34 on 907 patients with coronary heart disease to examine the association of ACE and PAI-1, and their combination with early onset coronary heart disease. This study divided patients into two groups: ≤55 and over 55 years, and concluded that no increased risk was observed for each isolated allele. On the contrary, the 4G/4G homozygosity combined with the D/D genotype and the adjusted OR from confounding factors trebled the risk for early onset of coronary heart disease. However, in our current study we did not find any elevated risk for CAD in association with either PAI-1 or ACE polymorphisms singly or in combination according to crude and adjusted OR. The control group, however, did not undergo any invasive or non-invasive method to recognize their arterial abnormalities, which could be a limitation of this study.
Considering the divergent results from different studies worldwide, even some meta-analysis on determining the impact of the PAI-1 4G/5G and ACE I/D polymorphisms on coronary heart disease, it would seem that the dependence of these two polymorphisms on race and location is significant. Therefore, it is difficult to obtain a certain conclusion and generalize it to a universal population. It would be more appropriate to use local databases to plan preventive or screening measures on cardiovascular disease.
According to our current study and most investigations on CAD risk factors to date, conventional and environmental risk factors such as hypertension, diabetes, hyperlipidemia and smoking, embodied in lifestyle, are the most prominent in CAD development. This is in contrast to genetic risk factors, for which their contribution remain controversial.
Moreover, further studies should be carried out to assess interactions between genes and environment, with a precise selection of homogenous cases and controls to determine this association definitely.
Footnotes
Acknowledgements
We would like to thank the Tehran Blood Transfusion Center, Ethics Committee of Iranian Blood Transfusion Organization and PayvandLab, clinical and specialty laboratory, for help with this project. We express our sincere gratitude to the Modarres Hospital staff and the ethics committee for the recruitment and phlebotomy of CAD patients.
Conflict of interest
None declared.
Funding
This study was supported by a grant from the High Council Institute of Iranian Blood Transfusion Organization (IBTO).
