Abstract
We investigated the association of polymorphisms in two renin-angiotensin system-related genes, expressed as angiotensinogen (AGT) and angiotensin II type 1 receptor (AGTR1), with blood lead levels and lead-related blood pressure in lead-exposed male workers in Korea. A cross-sectional study involving 808 lead-exposed male workers in Korea was conducted using a restriction fragment length polymorphism-based strategy to differentiate the various genotypes of polymorphisms in the AGT and AGTR1 genes. The association of clinical characteristics with genotypes as modifiers was estimated after adjustment for age, smoking status, drinking status, body mass index and job duration of each subject. Genotype and allele frequencies of the M235T polymorphism in AGT were associated with lead-related high blood pressure status. Moreover, blood lead levels were associated with allele frequencies of the AGT M235T polymorphism. These results suggested that the M/M genotype and M allele of AGT are risk factors for lead-related high blood pressure.
Introduction
Hypertension is associated with stroke and cardiovascular disease, and is one of the most common medical issues leading to significant morbidity and mortality.1–3 Identifying hypertensive patients with potentially treatable causes is of great importance: however, most patients have idiopathic or essential hypertension. Although several epidemiological studies have reported that environmental factors could contribute to the risk of essential hypertension, 4 many studies have suggested that essential hypertension is a complex disease resulting from the interactions of genes and environmental factors. 5 The renin–angiotensin system (RAS) may play a crucial role in the regulation of blood pressure and pathogenesis of essential hypertension. Therefore, genetic polymorphisms in the RAS genes, such as those of angiotensinogen (AGT), angiotensinogen II type-1 receptor (AGTR1) and angiotensin-converting enzyme (ACE) have been investigated extensively as potential genetic factors involved in essential hypertension. 6
Lead is a heavy metal pollutant in some areas and is distributed widely in the human environment. 7 Blood lead levels in humans are affected by exposure to environmental sources such as the manufacture of batteries, solder, ceramic glazes, plastics and sheet lead. 8 Lead influences mainly the central nervous system and the renal system. Additionally, several studies have reported a relationship between lead exposure and kidney disease.8–11 The toxic effects of lead on the cardiovascular system and its association with hypertension have been reported in animals12–14 and humans.15–17 Several mechanisms have been reported as causes of lead-induced hypertension, such as changes in calcium exchangeability, 18 increases in central sympathetic activity,13,19 enhancements in plasma catecholamines, 20 blockage of the Na+/K+-ATPase, 21 activation of smooth muscle protein kinase C 22 and enhanced activity of the RAS. 23 Although several mechanisms have been proposed to explain lead-induced hypertension, its etiology remains unclear.
Various studies have suggested that RAS gene polymorphisms, such as the M235T single nucleotide polymorphism (SNP) in AGT, the A1166C SNP in AGTR1, and the insertion/deletion (I/D) polymorphism in ACE, are genetic risk factors for hypertension and hypertension-related diseases.
A previous study reported that the TT genotype in AGT correlates with a higher risk for lacunar infarction 24 and history of hypertension. 25 Jin et al. 26 showed that the CC genotype in AGTR1 is associated with significantly increased left ventricular mass index, and He et al. 27 reported that A1166C in AGTR1 plays an important role in determining the blood pressure response to potassium intake. Subjects with the DD genotype in ACE carry a significantly increased risk of myocardial infarction. 28 In addition, we showed previously that the II genotype I allele in ACE is associated with a higher risk of lead-related hypertensive status. 29
No evidence of an association of lead-related hypertension with AGTR1 A1166C and AGT M235T polymorphisms has been presented. Thus, we investigated associations between genetic polymorphisms in the AGTR1 A1166C and AGT M235T and blood lead levels and lead-related blood pressure in lead-exposed workers in Korea.
Subjects and methods
Study population
This study included 808 workers employed in eight industrial facilities (five storage battery and three secondary smelting/litharge-making industries) during 2004. Workers were designated as lead workers based on their potential for exposure to lead during the manufacturing process. No medical exclusionary criteria were applied, and this study included subjects already identified to have hypertension. The study protocol was approved by the Institutional Review Board at the Soonchunhyang University College of Medicine. The DNA samples were provided by the Soonchunhyang University Biobank.
Data collection
Data collection was completed either at the Institute of Industrial Medicine of the Soonchunhyang University in Cheonan or at the study plants using methods reported previously.30,31 A standardized, interviewer-administered questionnaire was used to elicit information from each subject regarding the demographics, job duration, smoking and alcohol use. Blood pressures were measured with the IntelliSense blood pressure monitor (Model HEM-907; Omron, Vernon Hills, Illinois, USA). Measurements were taken in seated individuals after 5 min of resting in the left arm at 30 s intervals. The procedure was repeated three times, with systolic blood pressure (SBP) and diastolic blood pressure (DBP) defined as the mean of three independent measurements. Data included height and weight measurements to assess body mass index (BMI, kg/m2). We applied the criteria from the 2004 Korean Hypertension Treatment Guideline published by the Korean Society of Hypertension; the prehypertension group (SBP, 120–139 mm Hg; DBP, 80–89 mm Hg) was included in the high blood pressure (HBP) group 32 .
Blood lead levels
Blood lead levels were measured with a Hitachi 8100 Zeeman background-corrected atomic absorption spectrophotometer (Hitachi Instruments, Tokyo, Japan) 33 at the Institute of Environmental and Occupational Medicine, a certified reference laboratory for lead in South Korea. As an external quality assurance and control program, the Institute passed the German External Quality Assessment Scheme operated by Friedrick Alexander University, which is a standard for institutions measuring chemicals at low concentrations. Commercial reference materials were obtained from Bio-Rad (Lyphochek Whole Blood Metals Control) for the internal quality assurance and control program (CA 94547, USA).
Genotyping
DNA was extracted from fresh whole-blood samples anticoagulated with Ethylenediaminetetraacetic acid (EDTA), using a GenEx B DNA purification kit (GeneAll Biotechnology, Seoul, Korea). The sense and the antisense oligonucleotide primers of AGTR1 were 5’-TTC CCC CAA AAG CCA AAT CCC AC-3’ and 5’-CAG GCT AGG GAG ATT GCA TTT CTG TCA G-3’ as published previously by Hilgers et al. in 1999. 34 DNA samples (100 ng) were subjected to 35 cycles of polymerase chain reaction (PCR) amplification in an Exicycler™ 96 (Bioneer, Seoul, Korea) under the following conditions: 94°C for 2 min; 94°C for 20 s, 63°C for 30 s, 70°C for 2 min, repeated for 35 cycles; then 68°C for 20 min.
The PCR product was subjected to restriction digestion with 1 U of the enzyme DdeI (New England Biolabs, Inc. Ipswich, MA 01938-2723 978-927-5054) at 37°C for 16 h and subsequent gel electrophoresis. In addition to the genotype-specific DdeI site created by the 1166 C polymorphism, the 428-bp PCR fragment contained a second DdeI site present in all genotypes that was used as an internal control for the completeness of the restriction digestion. For the A allele, DdeI cleaved the 428-bp PCR fragment into 65- and 363-bp fragments, whereas three fragments were generated for the 1166 C allele; these were of 65, 220, and 143 bp.
The M235T variant of the AGT gene was detected by PCR restriction enzyme digestion using the restriction endonuclease Tth111I (Enzynomics, Inc., Daejeon, Korea). Segments of AGT were amplified from each DNA sample as described above for the AGTR1 variant, but using the 5′-CCG TTT GTG CAG GGC CTG GCT CTC T-3′ sense and 5′-CAG GTG TGT CAC CTG ACC C-3′ antisense primers as published previously by Yuan et al. in 2009. 35 PCR conditions were: 94°C for 10 min; 94°C for 15 s, 60°C for 45 s, 72°C for 45 s, repeated for 35 cycles; then 72°C for 10 min.
The specific mismatches incorporated into the antisense primer create a Tth111I site in variants having the T235. Subsequent digestion with this enzyme at 65°C resulted in 141- and 24-bp diagnostic fragments.
Statistical analysis
The Hardy-Weinberg equilibrium (HWE) was assessed by SNPStats (bioinfo.iconcologia.net/index.php) and SPSS 18.0 (SPSS Inc., Chicago, Illinois, USA). Associations between the AGTR1 A1166C and AGT M235T and the HBP group (DBP>80 mm Hg or SBP>120 mm Hg) were estimated by computing the odds ratios (ORs) and 95% confidence intervals (CIs) with logistic regression analyses, controlling for age and sex as covariables. In the logistic regression analysis for each SNP, models assuming codominant inheritance, dominant inheritance or recessive inheritance were used. The χ2 test was used to compare allele frequencies between groups. The descriptive statistics of participants were obtained by frequency distribution of categorical data and expressed as mean and standard deviation (SD) of continuous variables (age, blood lead levels, DBP, SBP, BMI and job duration). Significant differences in categorical variables and continuous variables were analyzed with the chi-square test and Student’s t-test between both genotype and allele groups. To obtain the statistical sample power, the required case size for each SNP was estimated using the genetic power calculator (pngu.mgh.harvard.edu/~purcell/). To avoid chance-findings due to multiple testing, the Bonferroni correction was applied by decreasing the significance levels to p=0.025 (p=0.05/2) for each of the two SNPs.
Results
Descriptive statistics of the study variables
Table 1 illustrates the blood lead concentrations and blood pressures in the study population. The mean and SD of blood lead concentrations in the normal blood pressure (NB) and HBP groups were 21.04±8.87 and 23.04±11.12 µg/dl, respectively (Table 1). The mean (±SD) and frequencies for the selected demographic, exposure and outcome variables are presented by genotype and allele (Tables 2 and 3). As shown in Table 2, a total of 190 (25.4%) participants had the MM genotype, 350 (46.9%) had MT, and 207 (27.7%) had TT for the AGT M235T. Regarding the allelic distribution of the AGT M235T polymorphism, 730 (48.9%) had M and 764 (51.1%) had T. Additionally, the genotype and allele frequencies of AGTR1 A1166C were analyzed (Table 3); a total of 656 (89.5%) participants had genotype AA, 75 (10.2%) had AC and 2 (0.3%) had CC for the AGTR1 A1166C, respectively. Regarding the allelic distribution in the AGTR1 A1166C polymorphism, 361 (94.6%) had A and 29 (5.4%) had C. For AGT T235M, significant differences in the blood lead level, SBP, DBP and HBP groups were observed according to genotype frequencies (p<0.05, Table 2). Comparisons within the groups showed that the TT genotype or T allele groups of AGT T235M had higher blood lead concentrations than the MM genotype or M allele groups. No other variables were significantly different between the groups (Table 2). For AGTR1 A1166C, no significant difference was observed between the groups (Table 3). To obtain the required sample size, we calculated the sample power using a genetic power calculator. The sample power of each SNP was as follows: 0.9221 for AGT T235M (α=0.05, genotype relative risk=twofold, for 80% power, n=369) and 0.9013 for AGTR1 A1166C (α=0.05, genotype relative risk=twofold, for 80% power, n=400), respectively. Therefore, the numbers of cases were sufficient to obtain a positive correlation for each SNP.
Blood lead concentration and blood pressures in the normal and high blood pressure group.
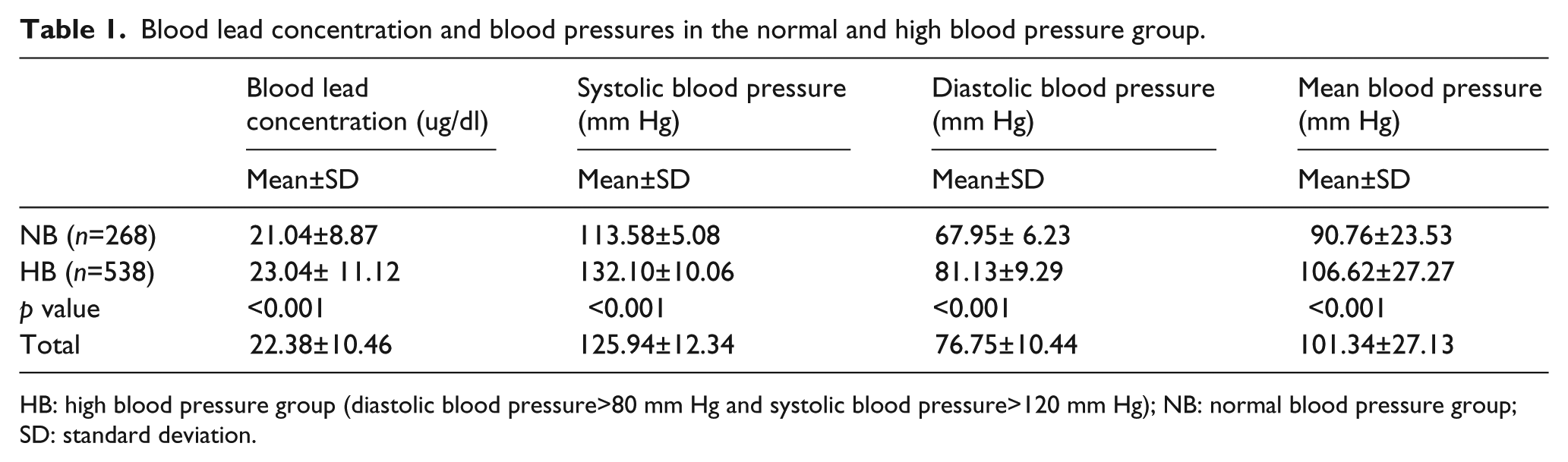
HB: high blood pressure group (diastolic blood pressure>80 mm Hg and systolic blood pressure>120 mm Hg); NB: normal blood pressure group; SD: standard deviation.
Descriptive statistics of the study variables.
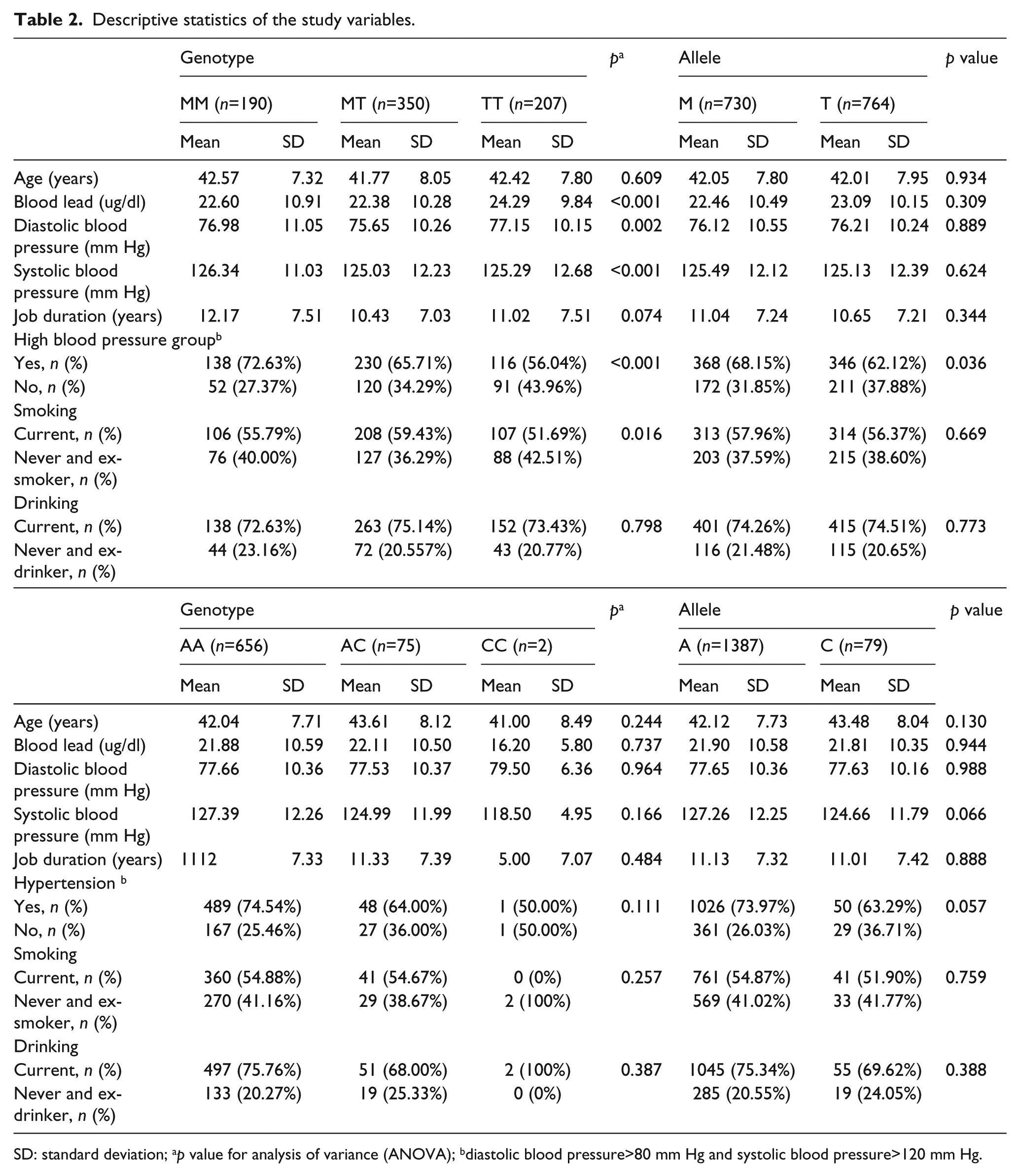
SD: standard deviation; ap value for analysis of variance (ANOVA); bdiastolic blood pressure>80 mm Hg and systolic blood pressure>120 mm Hg.
Association between AGTR1 and AGT polymorphisms and lead-related high blood pressure.
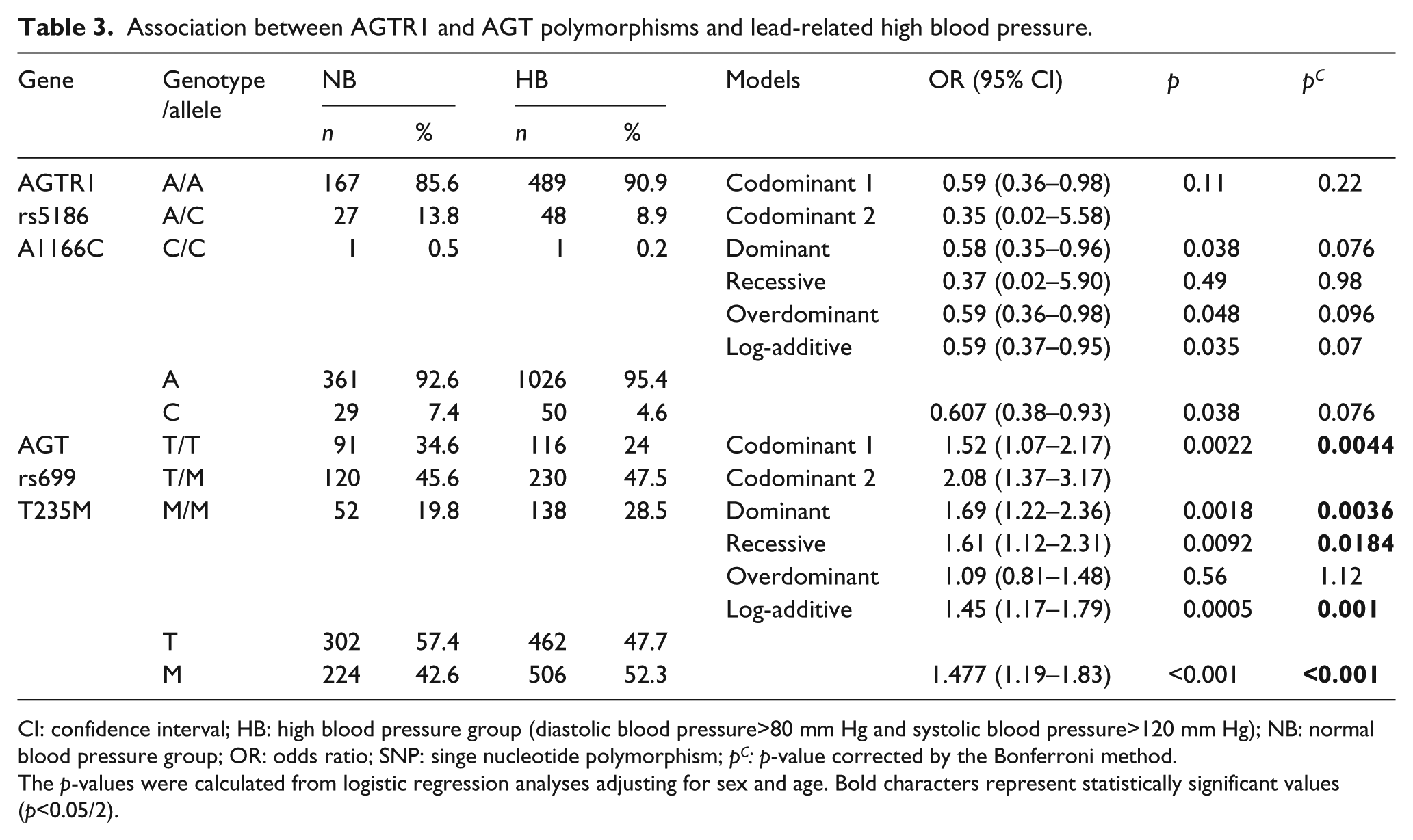
CI: confidence interval; HB: high blood pressure group (diastolic blood pressure>80 mm Hg and systolic blood pressure>120 mm Hg); NB: normal blood pressure group; OR: odds ratio; SNP: singe nucleotide polymorphism; pC: p-value corrected by the Bonferroni method.
The p-values were calculated from logistic regression analyses adjusting for sex and age. Bold characters represent statistically significant values (p<0.05/2).
Association of AGT and AGTR1 polymorphisms with lead-related high blood pressure in lead-exposed male workers
The genotype distributions of two SNPs in AGT and AGTR1 (AGT T235M and AGTR1 A1166C) were in HWE (p>0.05). As shown in Table 3, the genotype frequency of AGT T235M was significantly associated with HBP in codominant, dominant, recessive and log-additive models (p<0.05) after the Bonferroni correction.
In the codominant model for AGT T235M, the TT and MM genotype frequencies were 34.6 and 19.8% in the normotensive group and 24.0 and 28.5% in the HBP group, respectively (OR=1.52 (2.08), 95% CI=1.07–2.17 (1.37–3.17), p=0.0022, pc=0.0044). The MM genotype was associated with an increased risk of HBP. In the dominant model, the genotype frequencies containing the T allele (TT/TM) and not containing the T allele (MM) were 80.2 and 19.8% in the control group and 71.5 and 28.5% in the HBP group, respectively (OR=1.69, 95% CI=1.22–2.36, p=0.0018, pc=0.0036). In the recessive model, the genotype frequencies containing the M allele (TM/MM) and not containing the M allele (TT) were 65.4 and 34.6% in the control group and 76.0 and 24.0% in the HBP group, respectively (OR=1.61, 95% CI=1.12–2.31, p=0.0092, pc=0.0184). The allele frequency of AGT T235M was associated with hypersensitive status (OR=1.477, 95% CI=1.19–1.83, pc<0.025). The M allele frequency was higher in the HBP group (52.3%) than in the normotensive group (42.6%). The AGTR1 A1166C SNP (rs5186) was not associated with HBP status in lead-exposed workers (Table 3).
Discussion
In this study, we investigated associations between blood lead levels and lead-related blood pressure and polymorphisms in the renin-angiotensin system genes, AGT and AGTR1. Blood lead levels were positively associated with SBP and DBP: however, the correlations were not modified by the AGTR1 polymorphism. In contrast, blood lead levels were associated with AGT T235M allele frequencies. The T allele group showed slightly higher blood lead levels without statistical significance, whereas the TT genotype group showed significantly higher blood levels (Table 2). In addition, the AGT M allele and M allele carrier (MM or TM+MM) were risk factors for lead-related HBP status in lead-exposed male workers in Korea. Several recent reports have shown that lead affects the cardiovascular system in animals12–14 and humans;15–17 however, its etiology or/and pathological mechanisms remain unclear. To identify additional evidence of the etiology, we investigated the associations of lead-related hypertension with the AGT and AGTR1 RAS genes.
Hypertension is a multifactorial disease influenced by several genes and environmental factors. The genetic contribution is estimated to be 30–40% of all blood pressure variations. 36 The RAS is an important regulatory mechanism for maintaining blood pressure within the normal range, and AGT and AGTR1 encode two of the key enzymes in this system. Many studies have shown that these genes may play crucial roles in the pathogenesis of hypertension. Bonnardeaux et al. reported an increased prevalence of the C allele of AGTR1 A1166C in hypertensive subjects. 37 These results were replicated in some studies,38–40 but not in others.41–44 In the present study, we did not find an association between AGTR1 A1166C and lead-related hypertension. However, Spiering et al. 45 demonstrated an association of angiotensin II with the AGTR1 A1166C in essential hypertensive subjects. They reported no differences in active plasma renin concentration, aldosterone and atrial natriuretic peptide in response to angiotensin II among the three genotype groups. They concluded that the C allele of the AGTR1 A1166C polymorphism is associated with increased sensitivity but not reactivity to angiotensin II. An augmented response to angiotensin II may well be responsible for the increased incidence of cardiovascular abnormalities found in patients with one or two C alleles of AGTR1 A1166C. 45 The authors suggested a possible mechanism for the association of the C allele with increased sensitivity to angiotensin II and increased incidence of cardiovascular disease, as follows. The AGTR1 A1166C is located in the noncoding region of the AGTR1 gene and is likely to be in linkage disequilibrium with a nearby mutation that may affect AGTR1 mRNA stability, protein structure and/or function of the AGTR1, number of AGTR1s, and/or the process of receptor internalization. 44 Jeunemaitre et al. 46 reported a genetic association with essential hypertension in affected siblings, with a greater prevalence of the T variant in AGT T235M among hypertensive subjects than among controls. Additionally, plasma angiotensinogen concentrations were higher in subjects carrying the T allele in AGT T235M. 46 Hata et al. reported that the T allele of AGT T235M occurs at a higher frequency in essential hypertensive subjects than in normotensive controls in Japan. 47 Kamitani et al. also reported that the TT genotype was significantly more frequent in 108 hypertensive than in 104 normotensive subjects compared with the two other genotypes (MT and MM). 48 AGT is the substrate of the potent vasoactive angiotensin II, and its circulating plasma level has been positively related to blood pressure. Plasma AGT is elevated in hypertensive individuals and in the offspring of hypertensive parents compared with normotensive parents. 46 In the light of the previous studies, we investigated whether the hypertension-related SNP is associated with lead-related hypertension. Our results showed that the M allele frequency was higher in the HBP group (52.3%) than in the normotensive group (42.6%). Therefore, we conclude that the genetic association of lead-related HBP differs from that with essential hypertension. While some studies have confirmed the high prevalence of this variant and its association with hypertension,47–48 several studies in other populations have not shown such an association.35,49–51 This discrepancy suggests that ethnic and environmental differences may play a role in blood pressure control mechanisms.
Previously, we investigated the association between the ACE gene, an important gene in the renin-angiotensin system and lead-related hypertension. 29 In that report, we executed a cross sectional study involving 786 male lead-exposed workers from Korea, in which a PCR-based strategy was used to differentiate the genotypes of ACE I/D polymorphism. The distribution of genotypes among 786 male lead-exposed workers was in HWE and was similar to that in a previous study in Korean subjects (genotype distribution: II 36.5%, ID 48.7%, DD 14.9%). The results showed that the I/I+I/D genotypes and the I allele of the ACE gene were risk factors for lead-related hypertensive status; however, they were not directly associated with blood lead levels.
This study had several limitations. First, the expression levels of the genes (AGT and AGTR1) were not analyzed; therefore, the effect of variation on their expression and the susceptibility could not be determined. Second, this study presented the statistical differences in blood lead levels between the AGT T235M genotypes and alleles, which showed that the SNP is associated with HBP; however, we did not reveal why/how the variations of the associated SNP (AGT T235M) affect the blood lead levels and did not find any clues regarding the physiological relationship. Furthermore, no study has reported a direct biochemical relationship between lead levels and the AGT gene or AGT SNPs. We propose that the SNPs may be valuable as a possible marker for the lead-related HBP group, even if the blood lead levels do not act directly on a biochemical pathway.
In this study, we investigated a narrowly defined group consisting of male lead-exposed workers with HBP; therefore, it is difficult to compare the AGT and AGTR1 polymorphism results of our study population with those of other populations with hypertension. This report may increase our understanding of the genetic effects of candidate genes in lead-induced disease or/and clinical symptoms, particularly hypertension.
In conclusion, our data clearly indicate a strong association of the M allele and M allele carrier (M or MM+TM) in the AGT gene with HBP status in lead-exposed male workers. However, lead concentrations in blood were not associated with the genotype and allele frequencies of the AGT A1166C polymorphism. Further studies are needed to clarify the genetic association between candidate genes and lead exposure.
Footnotes
Funding
The present study was supported by the Soonchunhyang University Research Fund.
Conflicts of interest
All authors declare that they have no competing financial interests.
