Abstract
Introduction:
The human esophageal mucosa expresses various components of the renin-angiotensin system (RAS), e.g. the main effector peptide angiotensin II (AngII). The aim of this study was to investigate the esophageal presence of angiotensin III (AngIII) and angiotensin IV (AngIV) forming enzymes and the AngIV receptor (AT4R). The aim was also to study the actions of AngIV and to look for aberrations in patients with gastroesophageal reflux disease (GERD).
Materials and methods:
Esophageal biopsies were collected from healthy volunteers (n: 19) and individuals with erosive reflux disease (n: 14). Gene transcripts and protein expression of aminopeptidase A, -B and -M, and the AT4R were investigated by reverse transcriptase polymerase chain reaction (rt-PCR), western blot (WB) and immunohistochemistry (IHC). The functional impact of AngIV was examined in an Ussing chamber.
Results:
Aminopeptidase A, -B and -M and the AT4R were expressed in the esophageal epithelium. The AT4R was less prominent in certain areas in the mucosa of reflux patients. AngIV influenced the esophageal epithelial ion transport. The impact was lower in patients with GERD.
Conclusion:
The AT4R and formation enzymes of AngIII and AngIV are present in the human esophageal epithelium. Moreover, the present results suggest that AngIV exert regulatory impact on the epithelium and that RAS is involved in mucosal aberrations associated with GERD.
Keywords
Introduction
The renin-angiotensin system (RAS) is important for body fluid balance and blood circulation, but the system can also be involved in regulation of cellular growth, proliferation, inflammation and carcinogenesis.1,2 In addition to the classical endocrine mode of action, RAS is locally present in various tissues including the gastrointestinal tract.3,4 Previous studies in our laboratory have shown that several components of RAS are expressed by the human esophageal mucosa and that this system influences the epithelial resistance and ion transport.5,6 In addition, it was shown that components related to the main effector peptide, angiotensin II (AngII), were significantly altered in relation to gastroesophageal reflux disease (GERD), indicating involvement of the system in the pathogenesis of the disease. However, degradation-fragments of AngII also exist. These fragments can exert biological activity, for example angiotensin III (AngIII) and angiotensin IV (AngIV). 7 The possible presence or functions of AngIII and AngIV in the human esophagus have never been investigated. AngIII is formed by aminopeptidase A (AP-A), cleaving the aspartic acid from the N-terminus of AngII.2,8 Alternatively, AngIII can be formed from angiotensin I by the combined actions of AP-A and angiotensin-converting enzyme (ACE). 8 AngIV is formed when arginine is cleaved from the N-terminus of AngIII by the actions of aminopeptidase B (AP-B) or -M (AP-M; also termed aminopeptidase N and CD13), as well as by actions of dipeptidyl aminopeptidases (DAPs).7–9 AngIII binds to the angiotensin II type 1 receptor (AT1R) and type 2 receptor (AT2R). The former is known as the receptor mediating the classical AngII actions of RAS (blood pressure regulation, water homeostasis, cell growth and apoptosis etc.), whereas the latter generally counteracts these effects. 1 AngIV stimulates the angiotensin IV receptor (AT4R: equivalent to oxytocinase, OTase, and insulin-regulated aminopeptidase, IRAP)2,7 with known physiological effects in memory retrieval,9,10 renal sodium reabsorption, 11 cerebral and renal vasodilatation, 12 insulin-regulated glucose uptake 10 and intracellular signaling.7,13,14
We have previously reported that the AT1R and AT2R influence esophageal transepithelial transport.5,6 However, nothing is known regarding potential AngIV and AT4R effects in the human esophagus. The first aim of the present study was to investigate if prerequisites exist for formation of the AngII fragments AngIII and AngIV as represented by the presence of AP-A, AP-B and AP-M. A second aim was to examine the presence of AT4R and whether exposure to the ligand AngIV influenced the functional state of the human esophageal epithelium. In view of the previous findings that some RAS components were altered in relation to GERD, a third aim was to elucidate whether the above-mentioned formation enzymes of AngIII and AngIV, as well as the AT4R expression and AngIV functionality, were influenced by this condition.
Materials and methods
Human subjects
Esophageal biopsies were collected from healthy volunteers (n: 19, mean age 26 years, range 21–37 years, male 13) and from individuals with erosive reflux disease (ERD; n: 14, mean age 42, range 21–65, male 8; of Caucasian origin) during an upper endoscopy with a high-resolution magnification video endoscope using biopsy forceps (Endotherapy disposable biopsy forceps FB-234U Olympus, Hamburg, Germany). The enrolled ERD patients abstained from medication with proton pump inhibitors for two weeks before screening. All individuals were scored to be Grade A according to the Los Angeles classification for reflux esophagitis. 15 The biopsies were collected within 1 cm from the gastroesophageal junction (GEJ) at the three o’clock position. 16 One biopsy was collected from the ERD patients in the mucosal break area (ERD m.b.) and one from the endoscopically normal epithelium (ERD n.e.), 1 cm lateral to the mucosal break at the same distance from GEJ.
The study was performed in accordance with the Declaration of Helsinki. All study individuals signed a consent form and were informed in writing as well as verbally. The Regional Ethical Review Board in Gothenburg and the Ethics Committee at the University of Gothenburg approved the study.
RNA extraction and complementary DNA (cDNA) synthesis
Reverse transcriptase polymerase chain reaction (rt-PCR) was used to investigate AP-A, AP-B and AP-M gene transcription, in healthy, endoscopically normal as well as mucosal break area epithelial biopsies, to support the concept of local production of AngIII and AngIV. The primer sequences and PCR products sizes are listed in Table 1.17,18
Reverse-transcriptase polymerase chain reaction (PCR) primer sequences and product sizes.
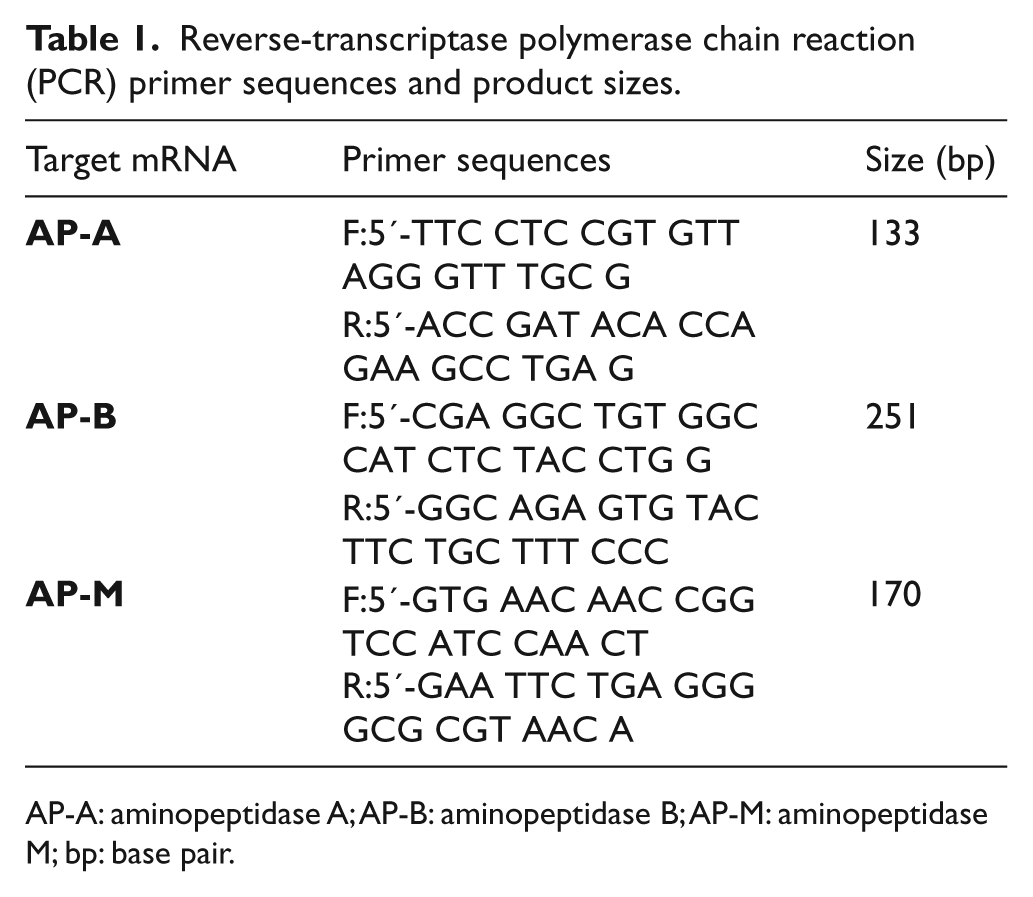
AP-A: aminopeptidase A; AP-B: aminopeptidase B; AP-M: aminopeptidase M; bp: base pair.
The biopsies were put in RNA STAT-60 (Nordic Bio Site AB, Stockholm, Sweden) and snap frozen in liquid nitrogen immediately after collection. Total RNA was extracted according to the manufacturer’s instructions using phenol-chloroform extraction and ethanol precipitation. Reverse transcription from 2.5 µg of total RNA was carried out using the SUPERSCRIPT™ First-Strand Synthesis System (Invitrogen, Lidingö, Sweden) with Oligo (dT) Primers (Life Technologies, Täby, Sweden). Resulting cDNA was stored at −20°C until use. Real-time PCR was performed in a Lightcycler Q-PCR (Roche Diagnostics AB, Stockholm, Sweden) using the hot-start technique and FastStart DNA Master SYBR Green I (Roche Diagnostics). The MgCl2 concentration was optimized to 4 mM and the cycling condition were as follows: 95°C, 10 s; 60°C, 10 s; 72°C, 10 s ×45. The length of the PCR products was confirmed by gel electrophoresis on a 2% agarose gel with 0.5 µg/ml ethidium bromide.
Western blot analysis
Western blot (WB) was used to investigate the protein expression of AP-A, AP-B, AP-M and the AT4R in the epithelium of healthy individuals and ERD patients, in and outside their mucosal break area. The biopsies were snap frozen in liquid nitrogen immediately after collection. Upon analysis, the biopsies were defrosted, sonicated and homogenized in a buffer (10 mM potassium phosphate buffer, pH 6.8, 1 mM EDTA; Complete-protease inhibitor tablet, Riche Diagnostics, Stockholm, Sweden; and CHAPS detergent 10 mM 3-((3-cholamidopropyl)dimethylammonio)-2-hydroxy-1-propanesulfonate, Boehringer Mannheim, Mannheim, Germany). The homogenates of each sample were centrifuged (10,000 g, 10 min at 4°C) and the supernatants analyzed regarding the protein content by using the Bradford method. 19 The diluted samples (sodium dodecyl sulfate buffer), positive controls (human kidney extract, rat adrenal cell lysate and human liver neoplasm cells) and a prestained molecular weight standard (SeeBlue, Novex, San Diego, California, USA) were heated to 70°C before being loaded onto a gel and the proteins in each sample separated with gel electrophoresis (NuPage 10% Bis-Tris gel and MOPS-buffer; Invitrogen, Lidingö, Sweden). The separated proteins were transferred to a membrane (polyvinylidene difluoride membrane; Amersham, Buckinghamshire, UK), which was incubated with the respective primary antibody (AP-A sc-18065, AP-M sc-166270 and AT4R sc-365051 from Santa Cruz Biotechnology, Heidelberg, Germany; AP-B ab119761 from Abcam, Cambridge, UK; loading control GAPDH IMG-5143A from Nordic BioSite AB, Täby, Sweden) overnight at 4°C. To enhance detection signal, a secondary antibody (goat anti-rabbit or mouse/donkey anti-goat) conjugated with alkaline phosphatase was linked to the primary. The secondary antibody was detected by a chemoluminescent light (reagent CDP-Star, Tropix, Bedford, Massachusetts, USA), which was captured (Chemidox XRS cooled charge-coupled device camera) and analyzed using the software program Quantity One (camera and software from Bio-Rad Laboratories, Hercules, California, USA). The membrane was incubated with a new primary antibody after that previous antibody had been removed with a re-blot mild antibody stripping solution (Millipore, Solna, Sweden).
IHC
The intraepithelial locations of AP-A, AP-B, AP-M and the AT4R were demonstrated by IHC. On collection, the tissue specimens were fixed (neutral buffered 4% formaldehyde), dehydrated (ethanol followed by xylene) and embedded in paraffin. Three µm thick paraffin sections were cut and placed on glass slides, after which they were deparaffinized (xylene followed by ethanol and deionized H2O) and boiled in citrate buffer (10 mM, pH 6.0 for 10 min followed by cooling for 20 min) for antigen retrieval. The slides were incubated in peroxidase block and serum block, before the respective primary antibody (see WB for antibody details; n: 4–7 per antibody; negative control sections were incubated with normal mouse immunoglobulin G (IgG)) was applied overnight, accordingly to the manufacturers’ instructions. Thereafter, the slides were incubated with a secondary antibody (anti-mouse immunoglobulin G conjugated with horseradish peroxidase (IgG-HRP) or anti-goat IgG-biotin), an amplification system (fluorescyl-tyramide hydrogen peroxidase or HRP-streptavidin) and the chromogen 3,3’-diaminobenzidine (CSA II kit K1497, Dako, Stockholm, Sweden; or sc-2053, Santa Cruz Biotechnology, Heidelberg, Germany).
Ussing chamber experiments
Ussing chamber experiments were performed to elucidate the in vitro effects of AngIV. Biopsies from healthy individuals and from the endoscopically normal epithelium of ERD patients were placed in oxygenated Krebs solution (mM: 118 NaCl, 4.7 KCl, 2.5 CaCl2, 1.2 MgSO4, 1.0 NaH2PO4, 25 NaHCO3 and 11 glucose), after which they were mounted in vertical mini Ussing chambers (insert with square area 0.018 cm2; Warner instruments, Hamden, Connecticut, USA). The luminal and serosal Ussing chamber reservoirs were filled with 5 ml Krebs. The solution was oxygenated and stirred with a gas flow (95% O2 and 5% CO2) and kept at 37°C. After an equilibration period of 30 min, baseline measurements were performed (exclusion of biopsies not reaching a potential difference (PD) of >0.3 mV lumen negativity) before AngIV (A0230, Sigma-Aldrich, Stockholm, Sweden) was added to the serosal reservoir.
The PD was measured using voltage electrodes (Ag/AgCl electrodes; Warner instruments) and the epithelial electrical resistance (Rep) using the Ussing pulse method (UPM). The UPM utilizes the capacitor properties of polarized cells and the method estimates the epithelial resistance specifically. The method has been described in detail elsewhere. 20 The PD and Rep were collected using an amplifier and software developed in LabView (National Instruments, Austin, Texas, USA). Ohm’s law (I=U/R) was used to calculate the epithelial electrical ion current (Iep), by Iep=PD/Rep.
Data analysis and statistics
Kruskal-Wallis and Mann-Whitney U-test were used for analyzing protein expression and Student’s t-test for analyzing Ussing data (SPSS 18.0, Chicago, Illinois, USA), where a p-value of ≤0.05 was considered significant. Individuals were denoted n and preparations/observations N.
Results
Gene transcription
The rt-PCR demonstrated gene transcripts of the angiotensin forming enzymes AP-A, AP-B and AP-M in all biopsies (n: 10). Representative examples of the PCR products on the agarose gel are shown in Figure 1.

Polymerase chain reaction (PCR) products of aminopeptidase A (AP-A), aminopeptidase B (AP-B) and aminopeptidase M (AP-M).
Protein expression and localization
All proteins investigated by WB – AP-A, AP-B, AP-M and the AT4R – were detected in both healthy individuals and in ERD patients (Figures 2 and 3). The AT4R expression was significantly lower (p-value 0.014) in the mucosal break area than in the normal epithelium of ERD patients and attained border significance (p-value 0.055) compared with the epithelium from healthy individuals (Figure 2(d)).
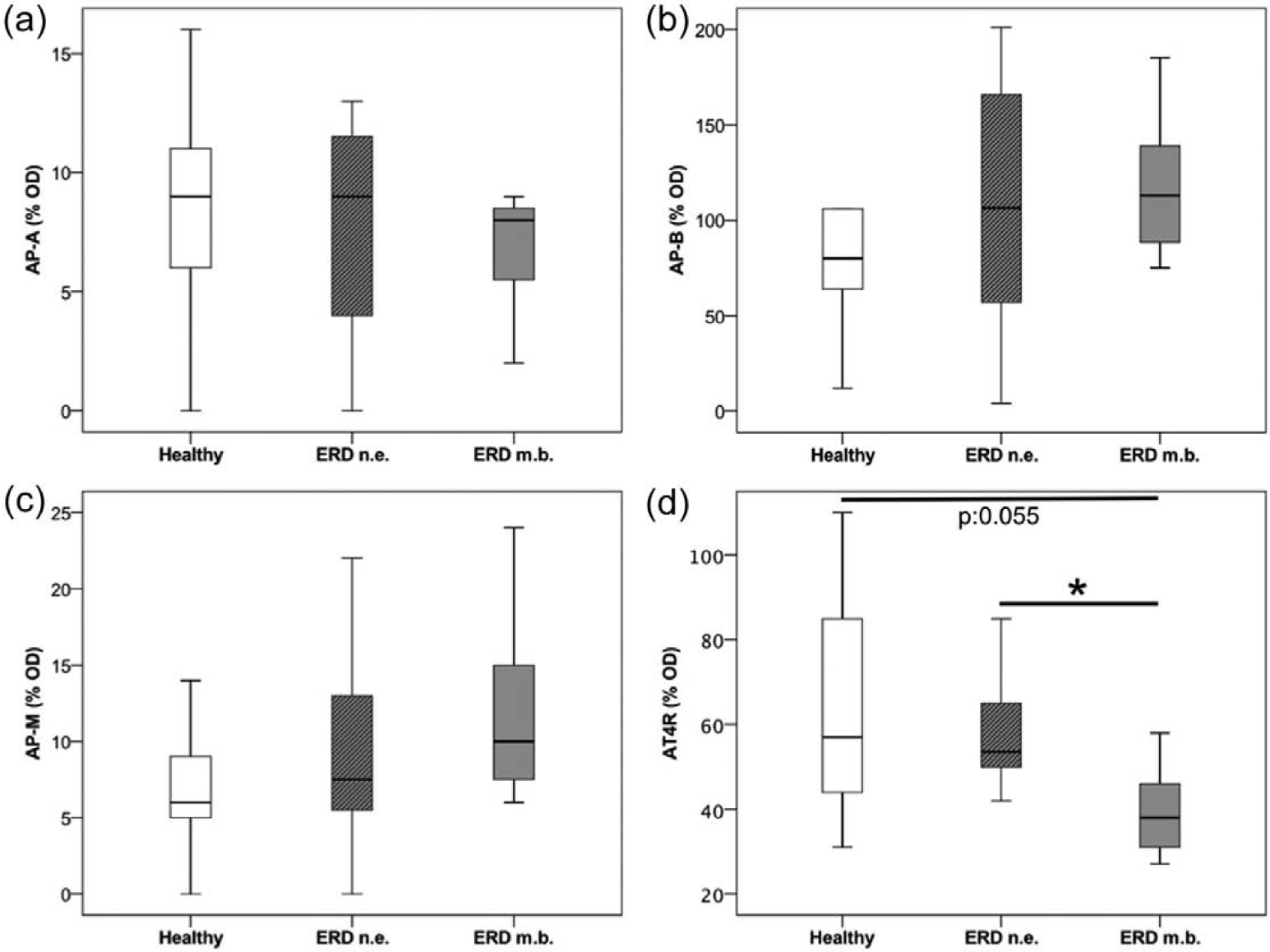
Protein expression of the angiotensin IV receptor (AT4R) and formation enzymes of angiotensin III (AngIII) and angiotensin IV (AngIV).

Protein bands on the Western blot (WB)-membrane.
The immunohistochemical analyses were performed in specimens from four to six subjects for each antibody and representative pictures are shown in Figure 4. The investigated components were detected throughout the epithelium. Particularly dense staining was observed around (AP-B: Figure 4(a) and (b), the AT4R: Figure 4(g)) and in the papillae (AP-M: Figure 4(d) and (e)). The intraepithelial localization seemed to be more cytosolic in the surface epithelium and more cell and nuclear membranously orientated in the basal and intermediate epithelium. In addition, AP-M was detected in small vessels (Figure 4(d) and (e)) and the AT4R in the nucleus (Figure 4(g) and (h)). No immunoreactivity was detected for AP-A or in the negative control sections (Figure 4(c), (f) and (i)).

Localization of angiotensin III (AngIII) and angiotensin IV (AngIV) enzymes and the angiotensin IV receptor (AT4R) in the esophageal epithelium.
Basal electrical parameters and AngIV effects in vitro
The basal epithelial electrical parameters are displayed in Table 2. The endoscopically normal ERD epithelium (n: 8, N: 27) had significantly lower Rep (p-value 0.002) compared with the epithelium from healthy subjects (n: 13, N: 43). The basal Iep was more than doubled in ERD patients (p-value ≤0.001) compared with healthy subjects. The epithelia from ERD patients also demonstrated a trend towards a more lumen negative PD, but the difference did not attain statistical significance (p-value 0.055).
Baseline characteristics of the esophageal electrical parameters.

Iep: epithelial electrical current; n: numbers of individuals; N: number of preparations; PD: potential difference; Rep: epithelial electrical resistance. Values are given as means±standard error of the mean (SEM).
Denotes differences between groups by p≤0.05.
The effects of AngIV addition after 10 min are shown in Figure 5. AngIV increased the PD (i.e. became more lumen negative, Figure 5(a)) and net Iep (Figure 5(b)) in a dose dependent manner in both the epithelium from healthy individuals and ERD patients. However, the PD and Iep increased more from baseline in the healthy individuals than in the ERD patients upon the highest AngIV dose (1×10–5 M). No changes in Rep were observed upon AngIV stimulation (data not shown).
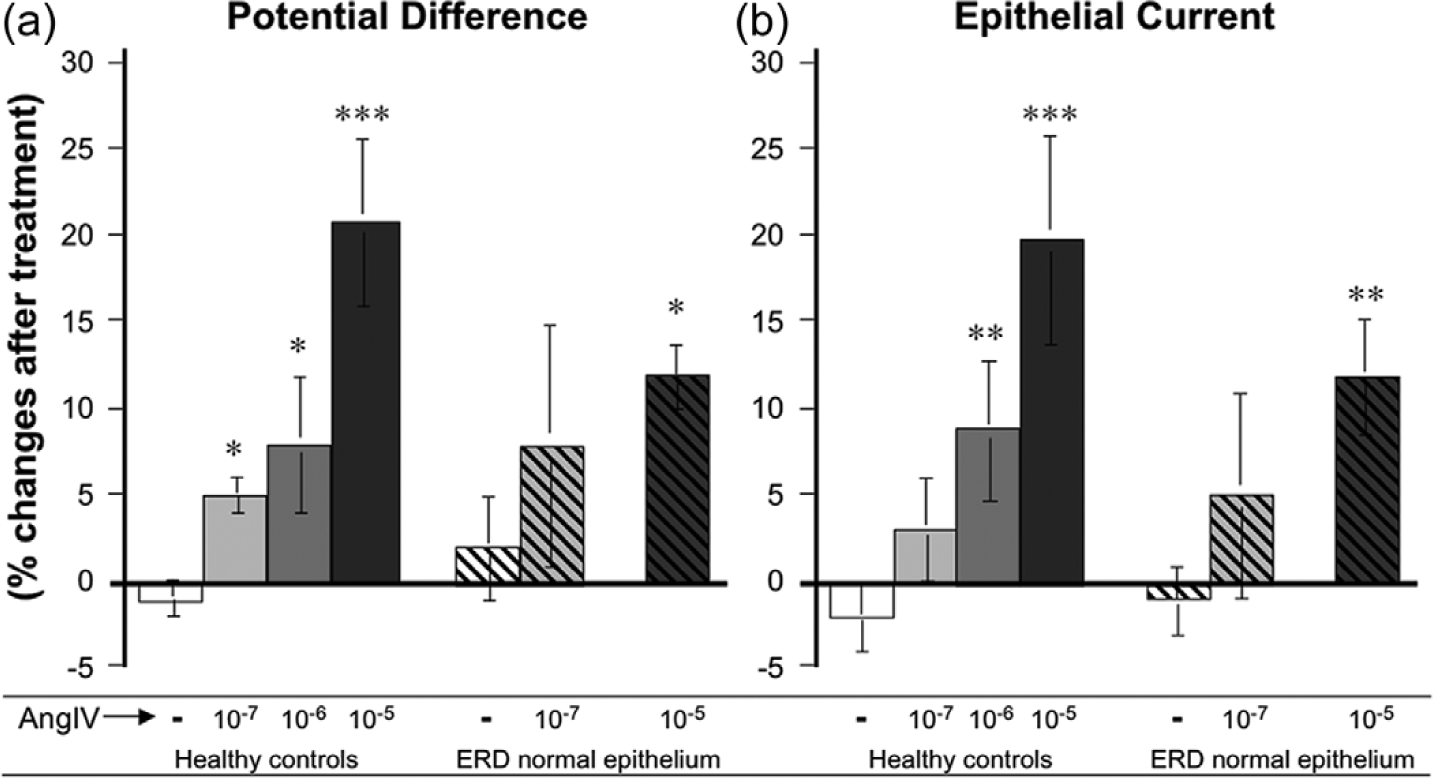
In vitro effects of angiotensin IV (AngIV).
Discussion
Immunoreactivity of the enzymes AP-A, AP-B and AP-M was observed in the human esophageal mucosa. The presence of these enzymes together with the substrate peptide, AngII,5,6 indicates local production of the peptide AngIV. Furthermore, the detection of gene transcripts corresponding to these angiotensin-forming enzymes strongly supports the concept of local AngIV synthesis in the esophageal epithelium. In addition, the receptor of AngIV, the AT4R, was expressed in the epithelium and addition of the peptide ligand AngIV was shown to affect epithelial ion transport in vitro. Moreover, the expression and activity were altered in the mucosa of GERD patients. Taken together, the results strengthen the view that the local RAS exerts regulatory impact on the human esophageal epithelium.
Recent studies from our laboratory indicate that the human esophageal mucosa harbors a local RAS, involving the effector peptides AngII and Angiotensin 1–7 (Ang1–7).5,6,21 The present finding of the three aminopeptidases AP-A, AP-B and AP-M suggest formation of AngIII and AngIV by local degradation of AngII. Both the present and previous5,6,21 data show that the angiotensin-forming enzymes are in peripapillary locations and in the blood vessel walls, and thus in close proximity to potential substrate delivered by the blood circulation. The present IHC stained for AP-B and AP-M in all epithelial layers, indicating that AngIV can be formed throughout the epithelium, from the papillae to the superficial layers near the lumen. AP-A was detected by WB, but not by IHC, which could be due to various technical reasons (e.g. antibody concentration or deficiency and antigen retrieval). Interestingly the localization pattern was similar for the AT4R that, in addition, was present in the nuclei. This topography thus suggests that locally produced AngIV easily can reach its epithelial receptor.
In an attempt to approach the functionality of AngIV, in vitro investigations were performed on mucosal samples from the distal human esophagus. These Ussing chamber experiments showed that AngIV influenced both the transepithelial PD and Iep in a dose-dependent manner, while the Rep remained unchanged, suggesting action mainly on the transcellular ion transport. 20 In other tissues, the AT4R has been reported to induce cellular influx of Ca2+ and Na+,13,14 and reduce the level and activity of the membranous basolateral Na+/K+ ATPase pump. 11 In the esophageal epithelium the primary ion transportation is Na+ absorption, creating a gradient that in turn regulates the intracellular acidity by extruding H+ or moving HCO3– into the cell. 22 It is possible that AngIV increased the Na+ influx in the present study, since the PD became more lumen negative. If this is the case, AngIV might influence the regulation of the epithelial intracellular acidity. This hypothesis is an attractive starting point for future studies of GERD, being a condition characterized by frequent epithelial exposures to acid refluxate that in turn demands optimal intracellular pH regulation to prevent tissue damage.
When adding AngIV, the PD and Iep responses from baseline were lower in ERD epithelia than in control. This is interesting since no difference in AT4R levels were shown by WB. Hence, these Ussing chamber results suggest that the receptor activity is reduced in the epithelia of ERD patients. This could be due, for example, to faster endogenous cleavage of the AngIV peptide or to malfunction, modifications and internalization of the AT4R. However, the low AngIV response could also be due to the ERD epithelium already having a higher net ion transport at baseline, and speculatively this could be due to acid or other refluxed compounds that had stimulated the mucosa.
Thus, the roles of RAS in general, and AngIV in particular, in relation to GERD remain to be clarified. The complexity is illustrated by the highly dynamic behavior of RAS, with various enzymatic pathways, receptors and bioactivities. For example, AngIV is suggested to have opposite effects depending on concentration 14 and to also bind the AT1R. 23 The AT4R is, in turn, suggested to have various subtypes 14 and to possess enzymatic capacities. 7 Moreover, AngIV can inhibit the enzymatic activity of the AT4R, meaning that some effects of the peptide might be due to reduced activity of the AT4R processing various peptides, increasing their half-life. 7 In addition to influencing cellular ion transport, the RAS has also capacity to modulate inflammation,1,2,4,24,25 this being another factor of relevance to the pathophysiology of GERD. We have previously shown that epithelia from ERD patients had increased gene transcript activities of ACE and AT1R and reduced protein levels of mas oncogene receptor (MasR), 5 thus a picture that is compatible with pro-inflammatory signaling. On the other hand, anti-inflammatory AT2R was shown to be elevated in ERD mucosa. 5 The present investigation demonstrated low protein expression of AT4R in the mucosal break area, suggesting involvement of this receptor in response to gastroesophageal reflux. However, the present exploratory study is limited by its small size and potential functional significance cannot be firmly concluded. How, and to what degree, the various RAS components influence the inflammatory picture in ERD remains to be analyzed in detail.
In summary, the present results indicate that all enzymes necessary for AngIII and AngIV formation are present in the human esophageal epithelium. Addition of AngIV stimulated the transepithelial PD and net ion currents in vitro. These responses were different in the endoscopically normal epithelium of ERD patients, and the AT4R protein levels were down-regulated in the mucosal break area. The present results support the hypothesis that the RAS exerts regulatory impact on the human esophageal epithelium and is involved in mucosal aberrations associated with GERD. More research is needed to elucidate functional details and the clinical relevance of AngIV in the human esophagus.
Footnotes
Acknowledgements
The authors acknowledge the assistance of the nurses Niclas Björnfot, Eva-Lotta Eén, My Engström, Diana Lustgarten and Maria Wållberg in assembling the study individuals and collecting the esophageal biopsies. The authors also thank Christina Ek for her brilliant WB and IHC laboratory work and Sören Lundberg for his technical support regarding the Ussing pulse method and various software programs.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Swedish Research Council (grant number K2010-55X-21432-01-2); and the University of Gothenburg and the Sahlgrenska University Hospital (ALF grants).
