Abstract
Keywords
Introduction
With the rapid development of digital healthcare technology, Electronic Health Records (EHR) and Electronic Medical Records (EMR) are important to modern medical information management.1,2 These records contain critical clinical information such as medical measurement data, diagnostic reports, and medication history records, which are very important for doctors in diagnostic decision-making, treatment selection, and patient health monitoring. However, as the amount and complexity of medical data continue to increase, doctors often need to spend a lot of time retrieving and organizing EHR and EMR when making clinical decisions,1,2 thus affecting the efficiency and effectiveness of diagnosis and treatment.
Current large language models (LLM) technology has shown potential in medical information extraction and summarization, 1 but there are still many challenges, such as insufficient accuracy, heterogeneity in data formats in different healthcare institutions, difficulties in standardization, and the issue of hallucination,1,3,4 which may cause the model to generate erroneous or inaccurate information, thereby affecting the credibility of clinical decision-making.
To address these issues, this study aims to develop a practical medical record summarization system and propose innovative X-RAG technology to assist doctors in making clinical decisions more efficiently and effectively. The specific objectives are as follows:
1. Development of a Medical Record Summarization System • Design a practical medical record summarization system that can process medical records in multiple formats (e.g., Word, PDF, and images) and standardize them into FHIR5–7 format to facilitate cross-institutional integration and application. • Generate structured and highly readable patient reports to help doctors quickly obtain patient information.
2. Development of X-RAG for medical data extraction • Enhance the existing RAG framework by adding page-based chunking, chunk filtering, and guided extraction prompting to improve the accuracy of extracting medical measurement data, diagnostic reports, and medication history records from EHR and EMR.
Related works
Extraction and summarization of medical information from EHR and EMR
Early EHR and EMR extraction techniques rely on rule-based matching and dictionary methods. However, these methods have limited generalization capabilities and have difficulty handling EHR and EMR in different formats.1,4 Machine learning methods (e.g., SpaCy 8 ) train classifiers through feature engineering and annotated data sets. Although these methods can improve accuracy, they are often limited by data heterogeneity and high annotation costs when dealing with large-scale EHR and EMR.2,9
In recent years, BERT variants designed for extracting specific medical information have been widely applied. For instance, CancerBERT 10 is designed to capture cancer phenotype information. In addition, some studies also use BERT models to extract tumor treatment outcomes from Japanese EHR. 11
With the development of LLM, they are increasingly used in EHR and EMR information extraction and summary generation. For example, RIEEL 12 has been applied to extract normalized clinical information from Chinese radiology reports. Few-shot learning1,13,14 and zero-shot learning1,13 allow models to effectively extract key medical information even in the lack of large-scale labeled datasets, thereby reducing data labeling costs and improving model generalization ability. Recently, retrieval-augmented generation (RAG) has shown potential in alleviating the hallucination problem of LLM, further enhancing the application value of LLM in the medical field.3,15,16
Retrieval-augmented generation (RAG)
The core concept of RAG 17 is to combine external retrieval mechanisms with generative models to provide more accurate content. Current RAG research primarily focuses on improving retrieval precision and the correctness of generated answers. Sentence Window Retrieval18,19 and Document Summary Index18,20 enhance retrieval precision by reducing noise in the retrieval process. Hypothetical Document Embedding (HyDE)18,21 and Multi-query18,22 try to solve the limitations of single-query retrieval by expanding the retrieval scope. Maximal Marginal Relevance (MMR)18,23 prevents retrieval results from being overly concentrated on a single topic by balancing relevance and diversity. Last, Cohere Rerank18,24 and LLM Rerank18,25 further enhance the relevance and quality of retrieved content by re-evaluating, filtering, and re-ordering.
Methods
System overview
The overall architecture of the system is shown in Figure 1. Each module is responsible for a specific task. First, the file format processing module converts various formats of EHR and EMR files (including .docx, .pdf, and images) into plain text format. Next, the X-RAG module extracts medical measurement data, diagnostic reports, and medication history records from the converted texts. Finally, the data management module uploads the extracted data to a local FHIR server for storage and generates a summary report to assist doctors in clinical decision-making. The architecture of the system.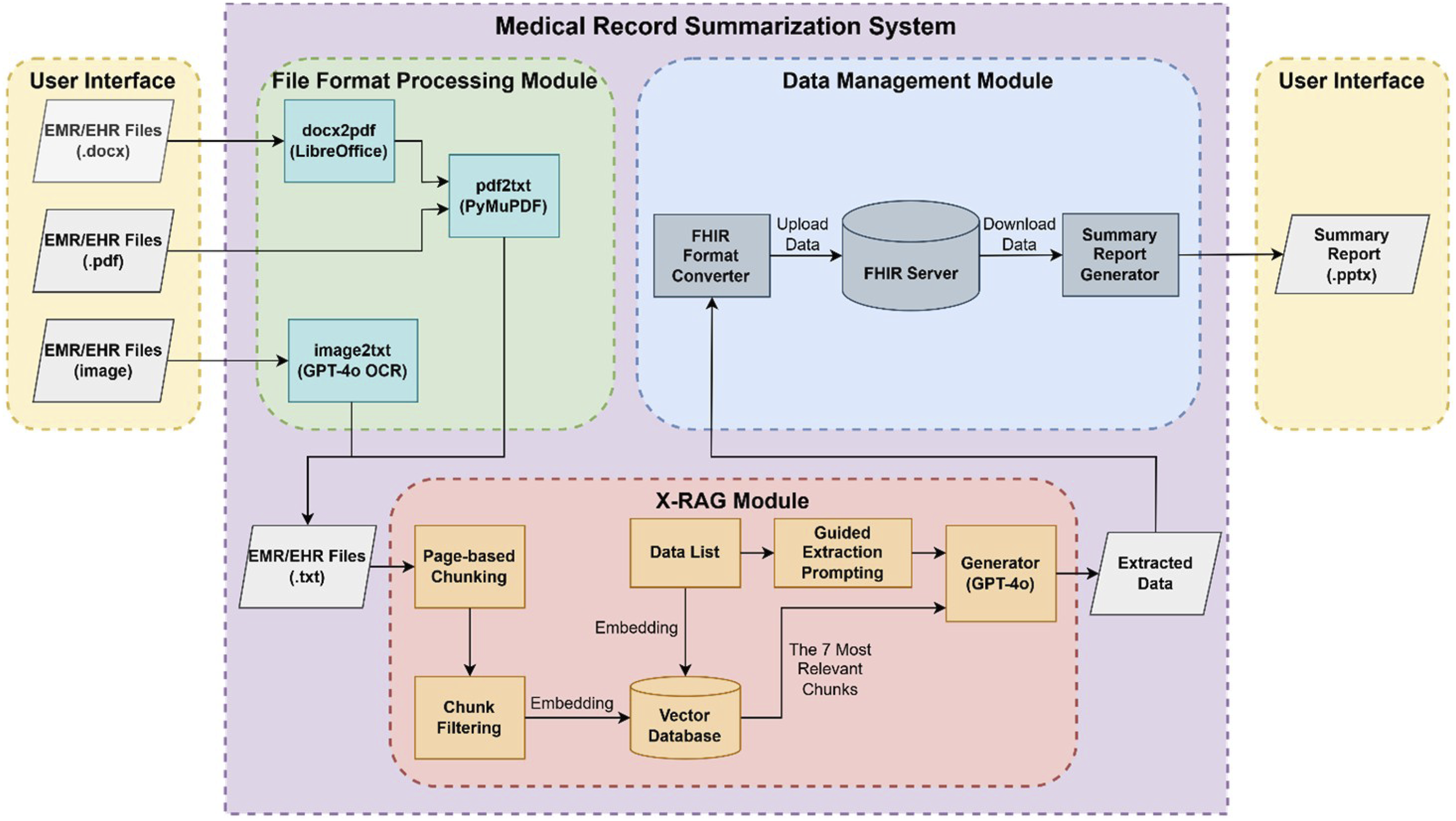
File format processing module
To facilitate subsequent processing, it is necessary to convert various medical record formats into a unified plain text format. This module first calls LibreOffice to convert .docx files into .pdf and then extracts text from PDF by using the PyMuPDF package. To ensure that the chunking process can accurately recognize page boundaries, we mark the beginning and end of each page with “Page Start” and “Page End”. Furthermore, we use GPT-4o’s built-in OCR function to convert image medical records into text and mark “page start” and “page end” to ensure the consistency of the data structure.
X-RAG module
Page-based chunking
We segment texts into multiple smaller chunks based on the “Page Start” and “Page End” tags to maintain the integrity of the text structure and content. When mapping back to the original format, each chunk corresponds to a single page in a Word or PDF file. If the medical record consists of multiple images, each chunk corresponds to one image. This page-based chunking approach ensures that retrieval accuracy is not affected by overly long or short text chunks.
Chunk filtering
Medical records often contain a lot of irrelevant content, such as administrative information or other redundant descriptive text, which may affect the retrieval accuracy. To address this issue, we use GPT-4o as a filter to remove chunks that are irrelevant to the target information by analyzing semantics. The filtering process is conducted separately for medical measurement data, diagnostic reports, and medication history records, ensuring that each dataset contains only the relevant and valid information. After chunk filtering, we obtain three refined datasets corresponding to medical measurement data, diagnostic reports, and medication history records (Figure 2). Chunk filtering illustration. (Red: Chunks of medical measurement data, Green: Chunks of diagnostic reports, Blue: Chunks of medication history records, Black: Irrelevant chunks).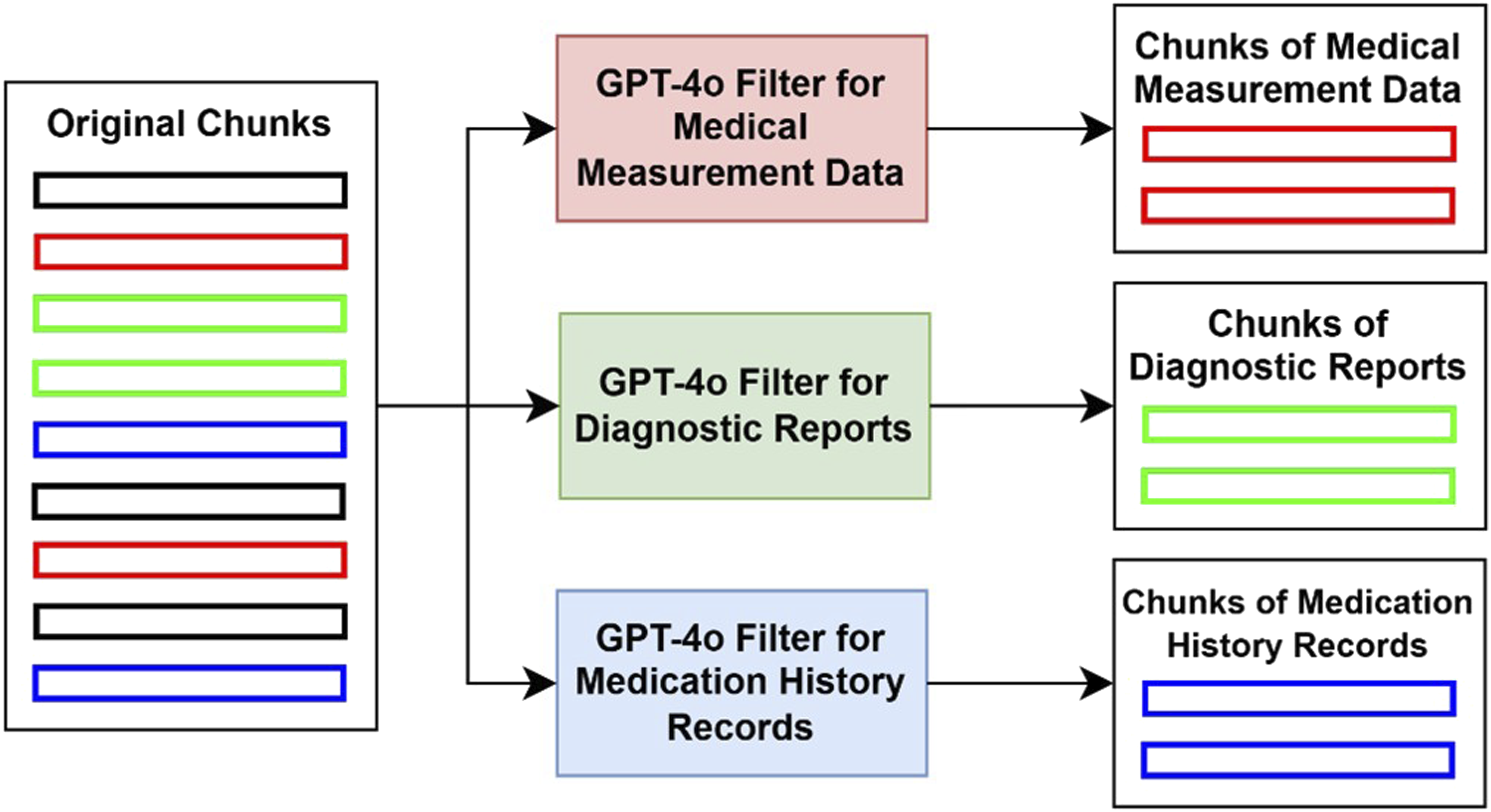
Embedding
We use multilingual-e5-large, 26 the best-performing open-source embedding model in the Traditional Chinese retrieval capability evaluation, 27 to convert text into vector representations. These vectors capture the semantics of the text, matching similar content in a high-dimensional space. We establish separate vector databases for the three data types: medical measurement data, diagnostic reports, and medication history records. This ensures that different types of medical information can be retrieved accurately.
Retrieval
The purpose of this study is to use RAG to extract information from medical records rather than general question-answering. Therefore, we compare item names with chunks stored in the vector database. We subdivide medical measurement data, diagnostic reports, and medication history records into several different subcategories. Specifically, medical measurement data is classified into 24 subcategories, diagnostic reports into nine subcategories, and medication history records into a single category. In each retrieval process, all item names within a subcategory are embedded into the vector space and compared with the chunks using cosine similarity. The system then retrieves the seven most relevant chunks from the corresponding vector database for a specific category.
Guided extraction prompting and text generation
We use GPT-4o as a text generator and design a Guided Extraction Prompt as its input. Guided Extraction Prompting consists of two parts: rules and examples. The rules define the expected content and format that GPT-4o should generate, and we have designed specific rules for three major categories: medical measurement data, diagnostic reports, and medication history records (Figure 3). The examples follow the principle of guided one-shot prompting,
13
and tailored examples are designed for each subcategory. By combining rules and examples, GPT-4o can better understand the requirements for information extraction, ensuring content generation accuracy and format consistency. Rules for medical measurement data, diagnostic reports, and medication history records.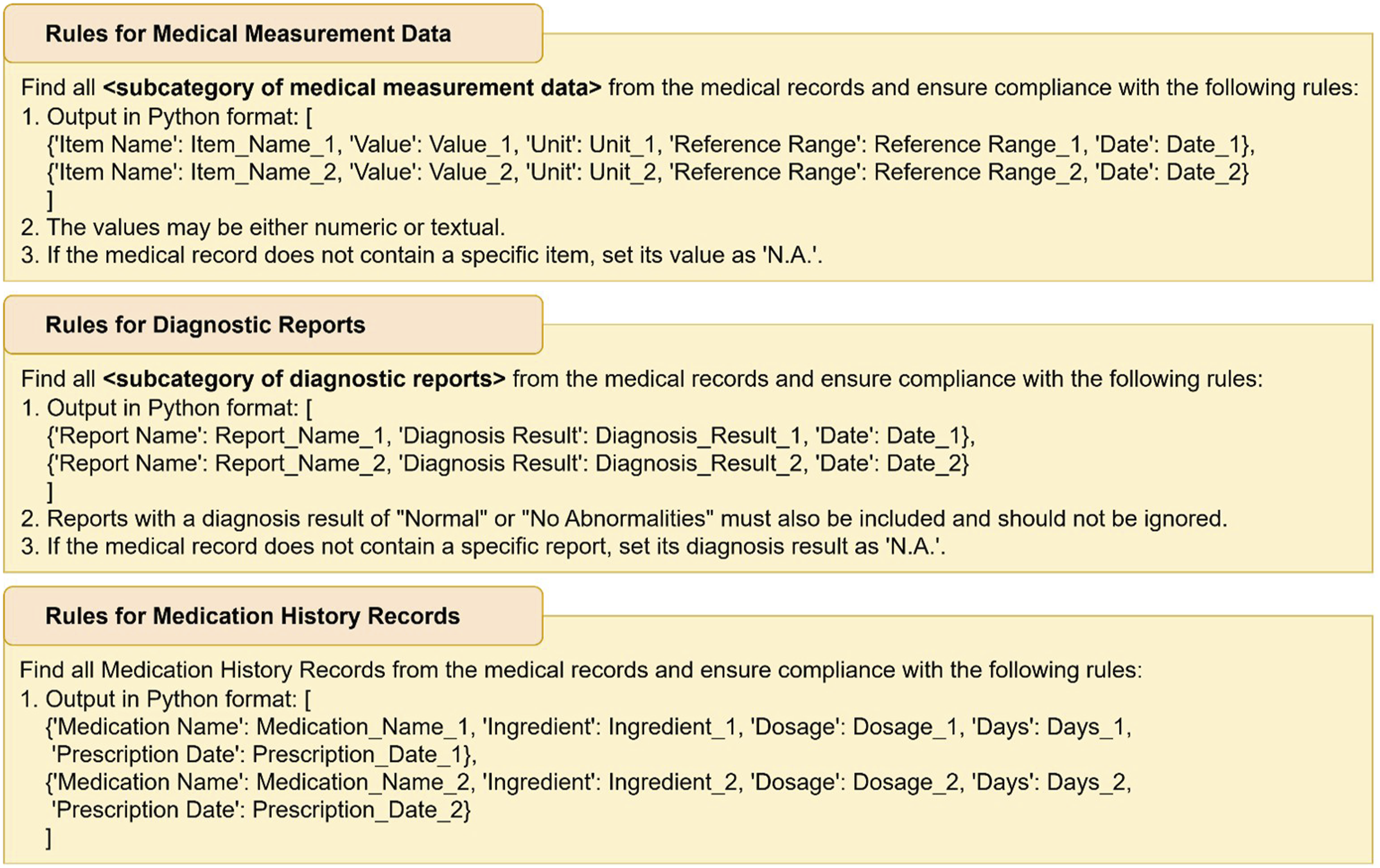
Mathematical formalization of X-RAG
To formalize the X-RAG process, we define the overall input document as a sequence of pages: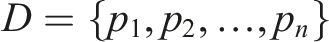
Each page 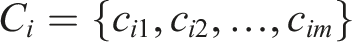
The complete set of chunks from the document is defined as: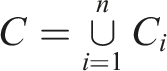
For a target information type 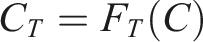
Each filtered chunk 
Let 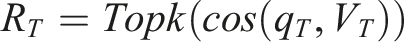
Finally, a guided extraction prompt 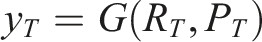
Data management module
The extracted medical data is converted into a structured FHIR R4 format by the FHIR format converter and then uploaded to the local FHIR server. After that, doctors can read and download medical measurement data, diagnostic reports, and medication history records stored in the local FHIR server. Finally, we use Python to automatically generate easy-to-read summary reports, which are formatted as PowerPoint presentations, so doctors can quickly understand the patient’s health status during diagnosis and treatment.
Results
Comparison of RAG methods for medical information extraction
To evaluate the performance of the proposed X-RAG technology in extracting medical data from EHR and EMR, we compared it with seven RAG methods. These methods encompass commonly used and representative design strategies in current RAG systems, focusing on different aspects of optimization, including enhancing retrieval precision with Sentence Window Retrieval 19 and Document Summary Index 20 ; expanding the retrieval scope with HyDE 21 and Multi-query 22 ; and improving result diversity and ranking quality with MMR, 23 Cohere Rerank, 24 and LLM Rerank. 25
Experimental dataset
The experimental dataset used in this study comprises 23 de-identified medical records collected from 11 healthcare institutions of various levels, including two medical centers, three regional hospitals, two district hospitals, and four primary care clinics. All medical records were obtained with informed consent from the patients and were de-identified to remove any personally identifiable information to protect privacy.
These medical records were written in a mix of Traditional Chinese and English, and the formats include 15 Word documents, 6 PDF files, and 2 JPEG scanned images. To facilitate model processing, all medical records were first converted into plain text through the File Format Processing Module. Specifically, Word documents were converted to PDF via LibreOffice and then processed with the PyMuPDF package to extract text, whereas scanned images were converted into text using the built-in OCR function of GPT-4o. The resulting text data retained the original page structure, with each page demarcated using “Page Start” and “Page End” markers.
In addition, the medical records used in this study were sourced from multiple different platforms and systems, reflecting practical differences in medical record management across institutions. To achieve data standardization and cross-platform integration, all extracted data were converted into the FHIR format and uniformly stored in a local FHIR server. This design not only enhances data interoperability but also lays the groundwork for future integration with different institutions and systems.
The data processing pipeline was primarily implemented using open-source tools, including LibreOffice and PyMuPDF, and was supported by the external GPT-4o API. All processing steps are reproducible based on the procedures described in this paper, and the workflow and program configurations have been thoroughly documented. Upon authorization, sample data and execution scripts may also be provided to support reproducibility and external validation.
Evaluation metrics and experimental results
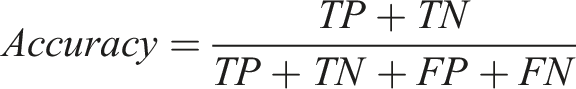
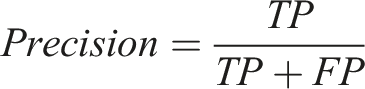
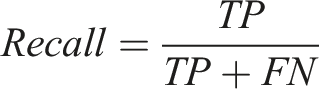
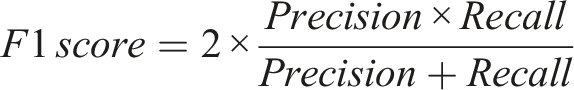
Definition of true positive (TP), true negative (TN), false positive (FP), and false negative (FN).
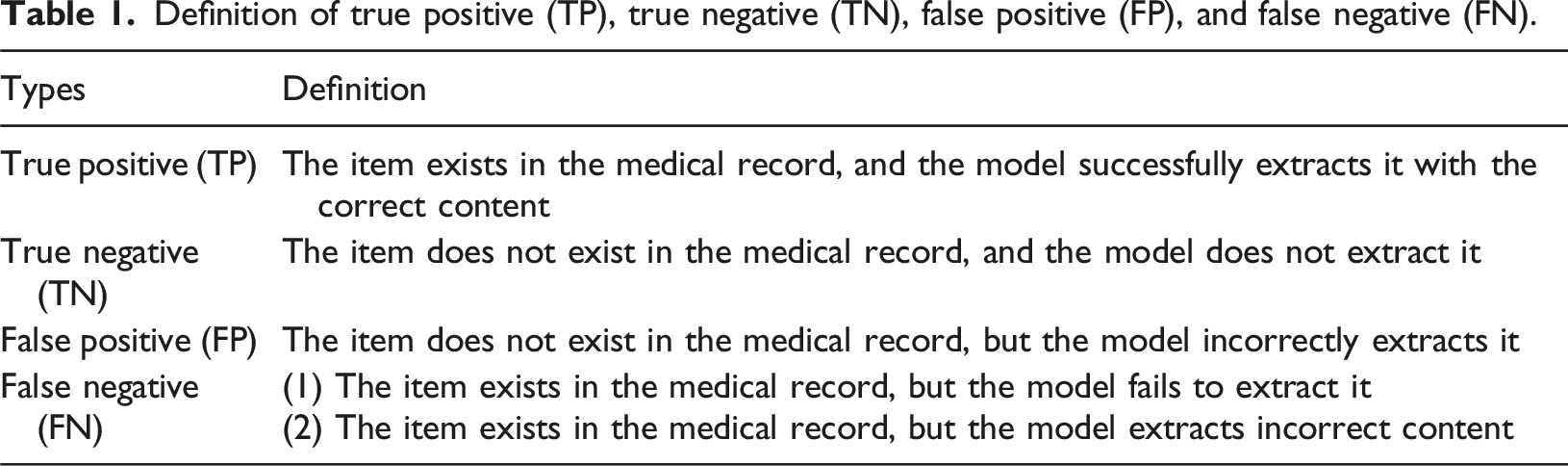

The comparative results of X-RAG and the other seven methods.
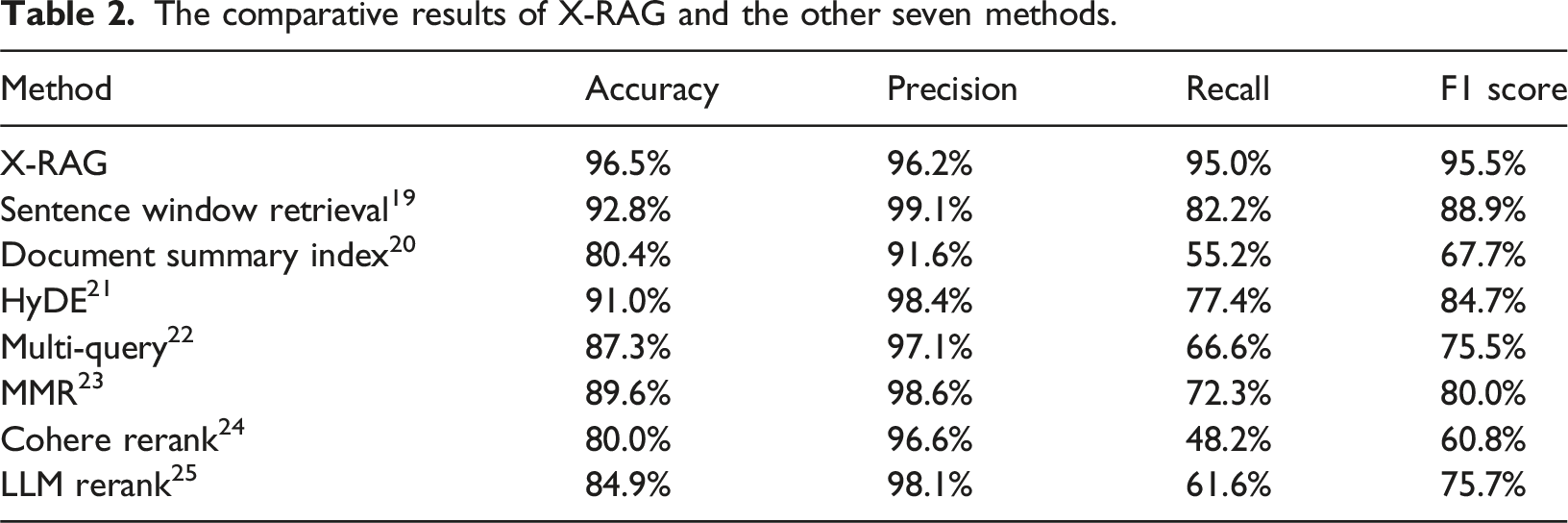
In clinical decision-making, it is crucial to avoid both incorrect extraction and omission of critical information. Therefore, the F1 score, as a comprehensive metric that balances Precision and Recall, is suitable for evaluating the overall performance of medical information extraction tasks and is widely adopted in related studies.10–12
Statistical significance testing
To evaluate the performance advantage of X-RAG over seven other RAG methods in medical information extraction, we conducted pairwise statistical tests on the accuracy and F1 score of each medical record. Specifically, the Wilcoxon signed-rank test was used to compare the performance differences between X-RAG and each of the baseline RAG methods. A two-sided test was used with a significance level of
Given that seven baseline RAG methods were compared, involving multiple hypothesis tests, we further applied the Holm–Bonferroni correction to adjust the p-values and control the family-wise error rate (FWER). This correction method sequentially compares p-values in ascending order against decreasing thresholds, thereby balancing error control with statistical power and ensuring the reliability of statistical inference.
In addition to reporting p-values, we calculated the effect sizes for each model comparison using the rank-biserial correlation, which corresponds to the Wilcoxon signed-rank test. This method ranks the absolute values of all non-zero paired differences and calculates the final value based on the sum of the ranks of positive and negative differences. The resulting effect sizes range from −1 to 1, with larger values indicating a greater performance advantage of X-RAG. This method more accurately reflects both the direction and magnitude of performance differences.
Wilcoxon signed-rank test results comparing X-RAG and seven baseline RAG methods on accuracy (Holm-Bonferroni adjusted).
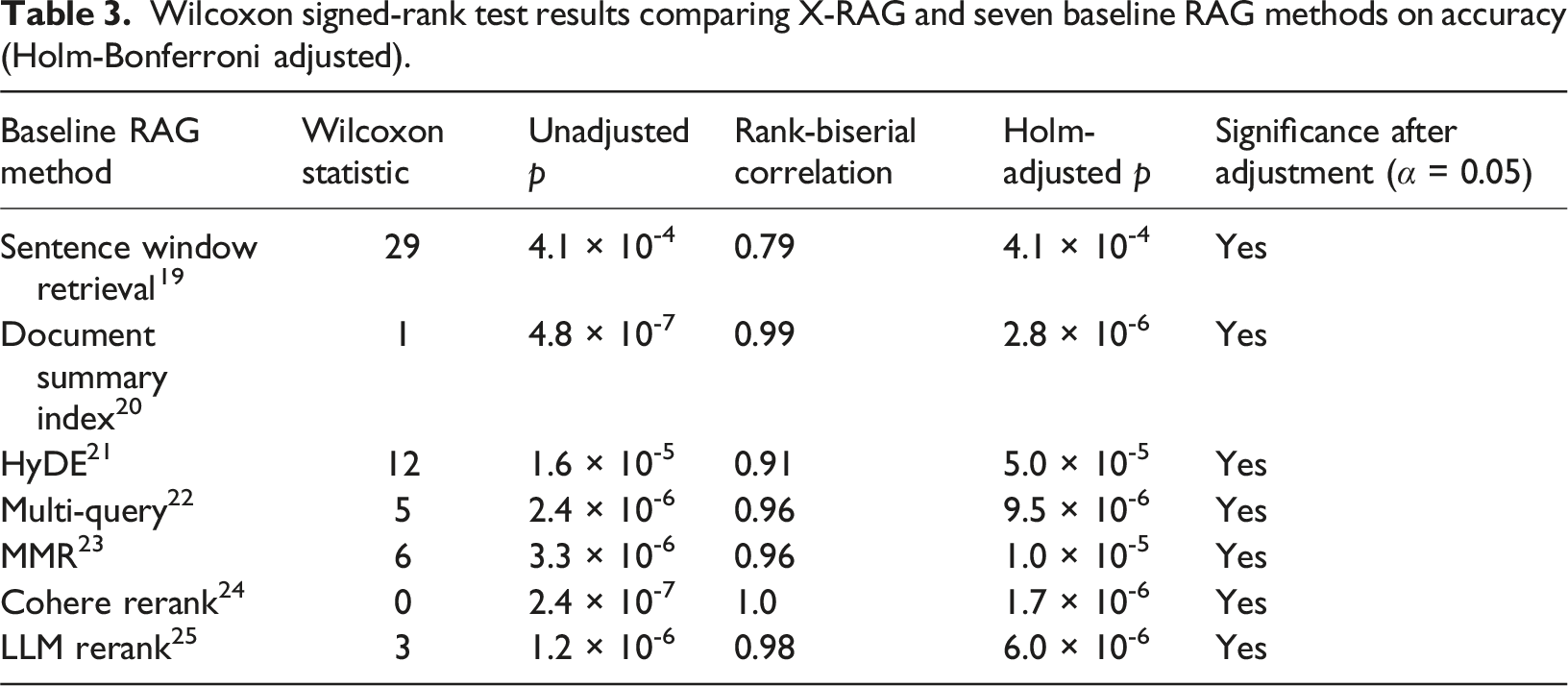
Wilcoxon signed-rank test results comparing X-RAG and seven baseline RAG methods on F1 score (Holm-Bonferroni adjusted).
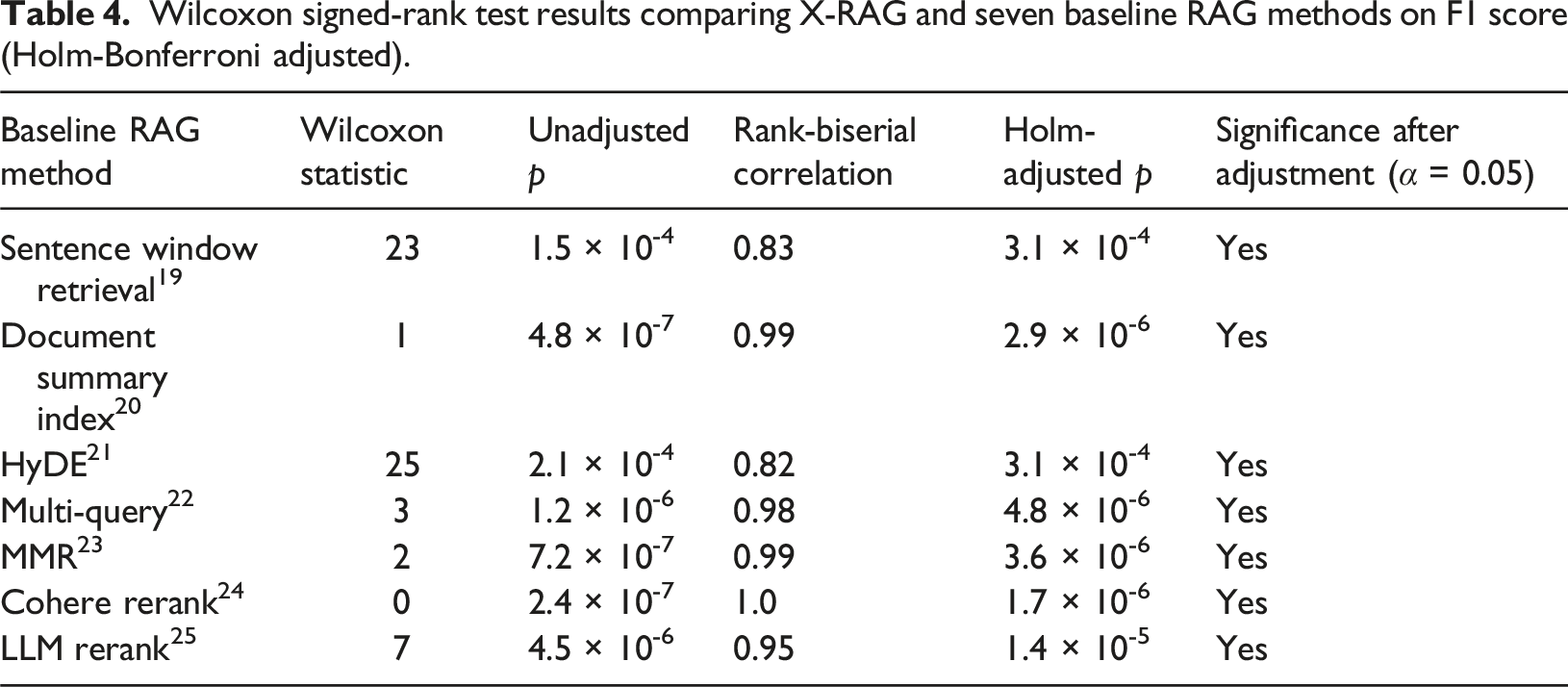
Comparison with existing models for medical information extraction
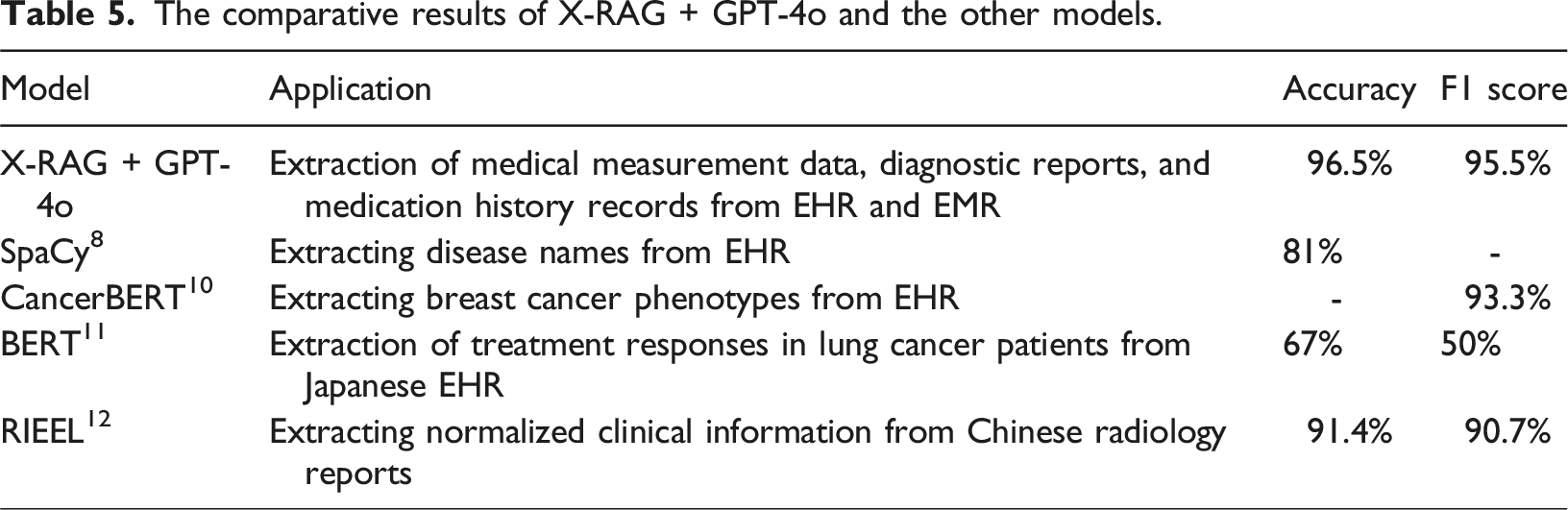
The comparative results of X-RAG + GPT-4o and the other models.

Ablation study
The ablation study.
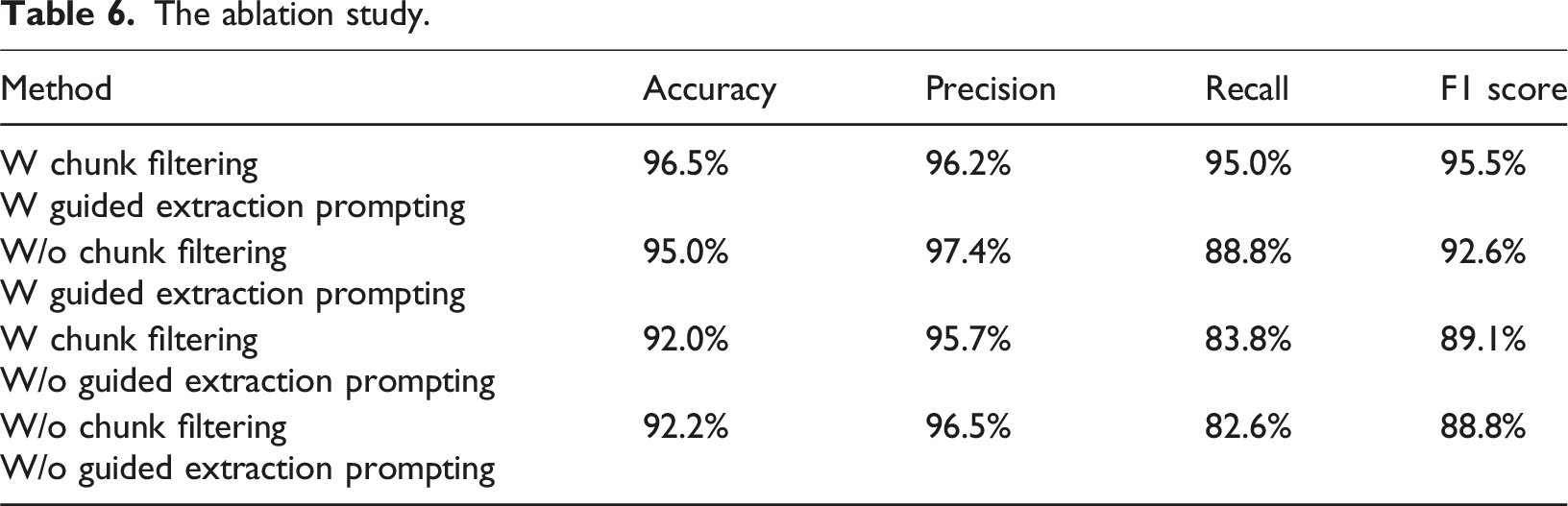
Overall, the ablation study confirms the complementarity between Chunk Filtering and Guided Extraction Prompting. Chunk Filtering enhances recall, while Guided Extraction Prompting improves overall accuracy and the F1 score. Therefore, the optimal strategy is to integrate both techniques to achieve the best performance in medical information extraction.
Field testing and doctors’ feedback
Doctor selection
To ensure that the system is applicable across various levels of healthcare institutions and suitable for different medical specialties, we invited eight physicians from a medical center and a primary care clinic to evaluate the system. The participants included three doctors from a medical center (one general surgeon, one radiologist, and one neurosurgeon) and five from a primary care clinic (two family medicine doctors, two cardiologists, and one pulmonologist). They used the system during daily consultations and provided feedback after the test.
System integration into outpatient workflow
Figure 4 illustrates how the Medical Record Summarization System developed in this study is integrated into the outpatient visit workflow. During patient registration, front desk staff or nurses upload the patient’s previous EHR or EMR files to the system. The system is automatically activated and generates a patient summary report that includes medical measurement data, diagnostic reports, and medication history records, assisting doctors in quickly understanding the patient’s condition during consultations. The outpatient visit workflow (in blue) integrated with our Medical Record Summarization System (in red).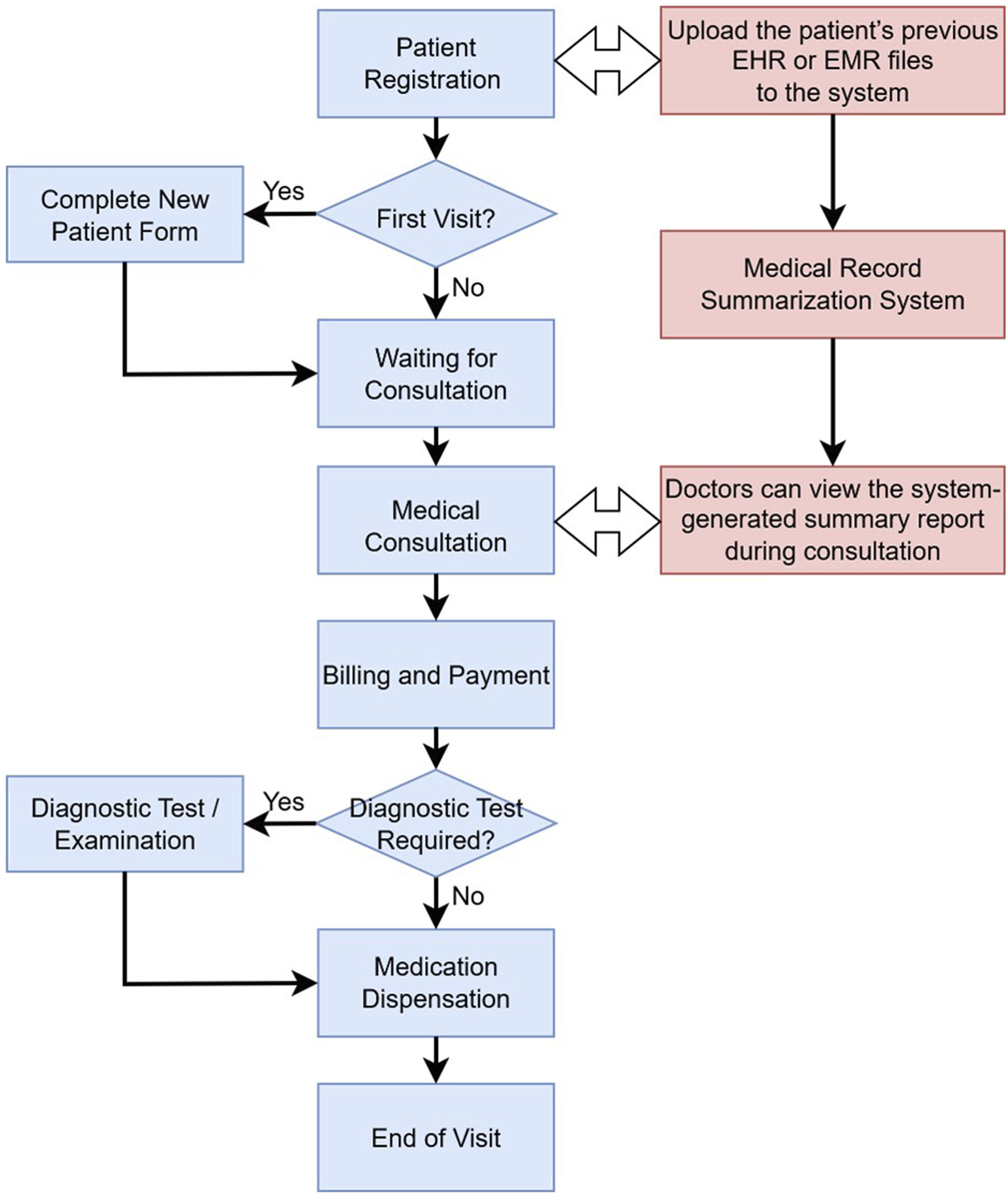
The system can be seamlessly embedded into the existing outpatient workflow without altering the operational steps of medical personnel. It only introduces two additional steps: “medical record upload” and “summary report review”, resulting in minimal disruption while ensuring both feasibility and practicality.
Doctors’ feedback
Doctors’ feedback indicated that, compared to the existing workflow, the system significantly reduced the time required for organizing medical records and retrieving information, saving approximately 40% of the time spent on patient data processing. They also reported that the summary reports generated by the system were clear and easy to read, facilitating the rapid identification of key information and enhancing clinical decision-making.
Discussion
Data security and ethical considerations
To ensure compliance with medical data privacy and ethical standards throughout system development and field testing, rigorous protective measures were implemented at every stage of this study. All 23 medical records used in this study were obtained with informed consent from the patients and were de-identified prior to use. Personally identifiable information, such as names and national identification numbers, was removed to effectively minimize the risk of data disclosure.
This study utilized the OpenAI API, including the natural language processing and OCR capabilities of GPT-4o, to assist with text conversion and semantic analysis. To ensure data security, all content transmitted to the API was fully de-identified and contained no personally identifiable information. All API calls were executed within a locally controlled environment, with data logging and training feedback functionalities disabled to ensure that no data records were stored or used for model learning during the processing.
During the field testing, clinical doctors were invited to use the system as part of their routine consultations. All test data were medical records with prior informed consent from patients, and the system served solely as a supportive tool without directly intervening in the clinical decision-making process. The overall research process was also reviewed and approved by the Human Research Ethics Committee to ensure that all aspects of data usage, informed consent, and ethical compliance adhered to current regulations and professional standards.
Limitations
Although the proposed system demonstrated strong performance in medical information extraction, certain limitations remain regarding the scale and diversity of the dataset. The current validation dataset includes only 23 medical records. Although these records were collected from 11 medical institutions of various levels, including medical centers, regional hospitals, district hospitals, and primary care clinics, and thus offer a certain degree of representativeness, the overall scale and structural complexity remain limited, which may affect the model’s generalizability in real-world clinical settings.
Furthermore, the medical records used in this study were primarily written in a mix of Traditional Chinese and English, and the formats were limited to Word documents, PDF files, and scanned images. Other languages and more diverse data types were not included, which may limit the model’s applicability and robustness in diverse healthcare environments. Additionally, the system currently focuses on processing specific categories of medical information, such as medical measurement data, diagnostic reports, and medication history records. It has yet to include other critical clinical information, such as surgical records and nursing records, which are also highly valuable for gaining a comprehensive understanding of a patient’s condition.
Future research will focus on the following directions: (1) Continuously expanding the medical record dataset to include a wider range of medical institutions, languages, and file formats; (2) Collaborating with more clinical units to establish a cross-specialty, multi-language, and multi-institutional validation platform; (3) Extending the scope of information processed by the system to incorporate a broader range of clinical data, with the goal of comprehensive medical record structuring and multi-category information summarization. These measures are expected to further enhance the system’s practicality and applicability across diverse clinical settings.
Practical application
In the outpatient workflow, the only manual step required is for front desk staff or nurses to upload the EHR or EMR files after the patient has registered. All subsequent processing is handled automatically by the system, resulting in minimal disruption to clinical operations. Regarding staff training, only basic operational instructions, such as uploading files and viewing summary reports, are required to complete initial user training, without the need for additional IT personnel or long-term training programs. The adoption of this system not only improves the efficiency of medical information processing but also has the potential to reduce the administrative burden on clinical staff, thereby promoting the implementation of smart healthcare.
Conclusion
This study developed a medical record summarization system that integrates the innovative X-RAG technology with GPT-4o to achieve more accurate medical information extraction and summarization from EHR and EMR. Experimental results show that X-RAG has an accuracy of 96.5% in medical measurement data, diagnostic reports, and medication history records, outperforming existing RAG technologies in terms of accuracy, recall, and F1 score. Through easy-to-read patient summary reports, doctors can quickly obtain patient information, reducing the time for reviewing and organizing medical records by approximately 40%. In summary, the proposed X-RAG technique and medical record summarization system offer an innovative solution for medical information processing and are expected to enhance the effectiveness and efficiency of clinical medical work.
Footnotes
Acknowledgement
We sincerely acknowledge Doctors’ Doctor Clinic and National Cheng Kung University Hospital for providing a valuable testing environment, which enabled us to validate the feasibility and practicality of our system in real-world clinical workflows.
Ethical consideration
This study was approved by the National Cheng Kung University Human Research Ethics Committee (approval no. NCKU HREC-E-114-0044-2) on April 01, 2025.
Author contributions
Conceptualization: Jhing-Fa Wang, Eric Cheng and Yuan-Teh Lee.
Investigation: Te-Ming Chiang, Hong-I Chen and Tzu-Chun Yeh.
Methodology: Che-Chuan Chang, Te-Ming Chiang and Tzu-Chun Yeh.
Supervision: Jhing-Fa Wang and Eric Cheng.
Validation: Te-Ming Chiang, Hong-I Chen and Yuan-Teh Lee.
Writing – original draft: Jhing-Fa Wang and Che-Chuan Chang.
Writing – review & editing: Jhing-Fa Wang and Che-Chuan Chang.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
