Abstract
Keywords
Introduction
Problem
Comprehensive Cancer centers need to quickly integrate clinical genomics data from disparate vendor systems for oncology operations and to facilitate research use of the data with minimal additional cost.
What is already known
Clinical data warehouse and molecular data warehouse architectures are costly to construct and brittle, not readily amenable to the rapid changes in oncology research.
Significance
The hybrid cloud Clinical Omics Data Lake architecture described in this paper solves the two main problems of conventional clinical and molecular data warehouses while maintaining two metadata sources that allow for file inventory and cohort search building within the Data Lake. The architecture presented is low-cost and can be constructed rapidly. Multiple test vendors and clinical sites are supported by the architecture. A software engineer with a cloud computing background is the only expertise required to implement this Data Lake architecture.
The wide adoption of electronic health records (EHR) by healthcare providers led to expectations among researchers that it would become easier to access patients’ digitized clinical data. The realization of this expectation has turned out to be much more difficult, as several problems have arisen. First, EHR adoption has made it easier to consolidate a patient’s clinical data within a provider or provider network sharing the same EHR framework. However, having a holistic view of a patient’s clinical data is still difficult. Advancements toward achieving precision medicine, especially for treating diseases such as cancer, are genome-driven. 1 It is generally understood that new progress in precision medicine will be driven mainly by discovering actionable variations in a person’s omics (genome, transcriptome, proteome, epigenome, etc.) data. Integrating clinical and omics data is necessary for any oncology research seeking precision medicine-based therapies. 2 Omics data are big data, and as genomic tests (both somatic and germline) are becoming more affordable and a routine part of patient care, the problem of data size increases storage and egress costs. In addition, somatic tests are becoming serial (since somatic genomes are not invariant 3 ), so more data are being generated per patient. 3 Raw sequencing data is stored long-term for future re-analysis. The abundance of ‘omics data adds to existing challenges in the storage, integration, accessibility, and privacy of data for oncology research. Another challenge with omics data is the diversity of platforms that are commonly used to obtain data and the rapid evolution of technologies.
There have been attempts to solve this clinical and omics data integration problem, and many suggested architectures are based on Clinical Data Warehouse (CDW) concepts. These solutions fall into four categories: translational platforms, Informatics for Integrating Biology and the Bedside (i2b2) based CDW solutions, custom platforms, and cloud-native solutions. Translational platforms are designed to be turn-key, meaning they are useful out of the box with minimal configurations. Canuel and coauthors reviewed seven such translational research products. 4 The primary design consideration of these platforms is their ability to fit into researchers' workflow. These include platforms such as cBioPortal, 5 tranSMART, 6 and caTRIP,4,7 among others. Canuel and coauthors concluded that these platforms lack privacy, data exchange capabilities, and interoperability. Moreover, these platforms do not provide ready modifications by institutions to support new ‘omics technologies as these evolve.
I2b2-based integrations are based on CDW implementations and are seen as quick additions leveraging data from an existing CDW. These implementations are popular with academic researchers, but the integration process can be complex, and often, the help of an i2b2 integration expert is required. 3 Murphy and others reported on three sample integration methods applying various techniques: Sequence Ontology-based, tranSMART, and a NoSQL database (CouchDB) based solution. 3 These authors reported that all three methods led to the development of tools that could answer a researcher’s queries from the Molecular Data Warehouse (MDW) application. However, they also report that all three solutions suffered various usability problems.
Custom integration solutions vary from those built from the ground up to solutions leveraging Enterprise Data Warehouse (EDW) applications provided by software vendors such as IBM Inc. And Oracle Inc. Moffitt Cancer Center (MCC) developed an MDW solution based on the Oracle Healthcare Translational Research product. 8 The Dana-Farber Research Center developed its MDW, PPIP, based on IBM’s Netezza EDW. 9 These MDWs have processing, analytic, and visualization capabilities, but both articles describing these MDWs noted the difficulties in getting these MDWs operational in production. For example, Dana-Farber Center’s PPIP took 10 years to develop. 9 The MCC solution development is reported to have been challenging, 8 and the developers noted that even after development, during use, a complex query “requires a collaborative team with dedicated expertise in oncology, translational research, bioinformatics, and database querying”. 8
Because of these difficulties, the three major cloud service providers (CSP), Google Inc., Microsoft Inc., and Amazon Inc.’s Amazon Web Services (AWS), offer tools enabling genomic data storage, analysis, and visualization on the cloud. 1 These companies have created platforms that are compliant to the Health Insurance Portability and Accountability Act (HIPAA) and offer out-of-the-box processing, analysis, and visualization of genomic data. These solutions have low start-up costs and enable processing at scales that individual academic institutions cannot easily match. Their cost model is ‘pay-on-use’, meaning that initially, the costs are low compared to traditional MDW development efforts. Some researchers consider the use of cloud services as one solution for the management of healthcare data complexity problems. 10 However, anecdotal evidence suggests that researchers have been slow in embracing these offerings and are concerned by perceived patient confidentiality and privacy problems. 11
Developing a comprehensive MDW that serves both clinical and research needs and is flexible enough to accommodate undefined omics data is complex and costly. Additionally, smaller organizations with limited resources, such as a lack of dedicated expertise in oncology, bioinformatics, or software development, cannot develop a comprehensive MDW. Therefore, we designed and implemented a Data Lake architecture that takes advantage of the low cost of cloud storage and is straightforward to implement. Various definitions of a Data Lake have been offered.12,13 For this research, we consider a Data Lake as a scalable data storage system that ingests raw data, maintains metadata on the data, places governance restrictions on access to the data, and can support egress mechanisms that allow flexible data egress for other services.
The ‘omics data lake architecture’
Our Data Lake architecture builds upon the zone architecture preferred in healthcare and demonstrated in previous reports.14–16 This Data Lake architecture includes the four zones commonly found in healthcare Data Lakes: an ingestion layer, storage layer, transformation layer and interaction layer
15
as shown in the generalized framework presented in Figure 1. The contributions of the Data Lake presented here are specifically designed for the setting, functional needs and constraints of an academic cancer center and expand upon the zone architecture for these settings. First, the ingestion process splits the metadata into two layers. To support lowering the overall storage cost of the raw sequencing files, they are immediately placed in archival cloud storage, necessitating a process for inventorying the sequencing files and establishing an easily accessed link between test orders, patients, and ordering physicians without retrieving the files. These file annotation metadata power the data request module in the application layer and facilitate the retrieval of files from archival storage. The second metadata set, extracted by the transformation layer, annotates the test results in the on-premises application layer tools. It is important to define the interaction layer in the cloud environment which enables utilization of cloud tools for machine learning or data science separately from the on-premises application layer. Additionally, matching the patient consent to the resulting tests occurs in the pipeline after the metadata is pulled into the on-premises environment. Tools to support data governance are included in the architecture. These tools include query tools enabling an honest broker to create consented study cohorts and to submit requests to the system for extracting raw sequencing files related to those cohorts for distribution on approved research projects. Conceptual diagram of a zone-based architecture. The framework of the hybrid cloud architecture builds upon a zone architecture by expanding previously established layers (Ingestion, Storage, Transformation, Interaction) to facilitate the use of the hybrid framework in a learning health system environment where clinical data informs research, and research informs the clinical decision-making process. Additional layers which were added to support this environment include a Consent Layer, Data governance layer, Application layer and Egress layer. The Transformation layer in the cloud generates metadata which is utilized by the application layer to provide role appropriate views into the data lake. The Egress layer leverages the Consent, Data governance, and Application layers to link consented patients to available tests and associated files then initiate a request to pull the files from archive storage in the cloud. Tools to facilitate clinical and clinical research roles such as patient screening for trials are built out in the Application layer.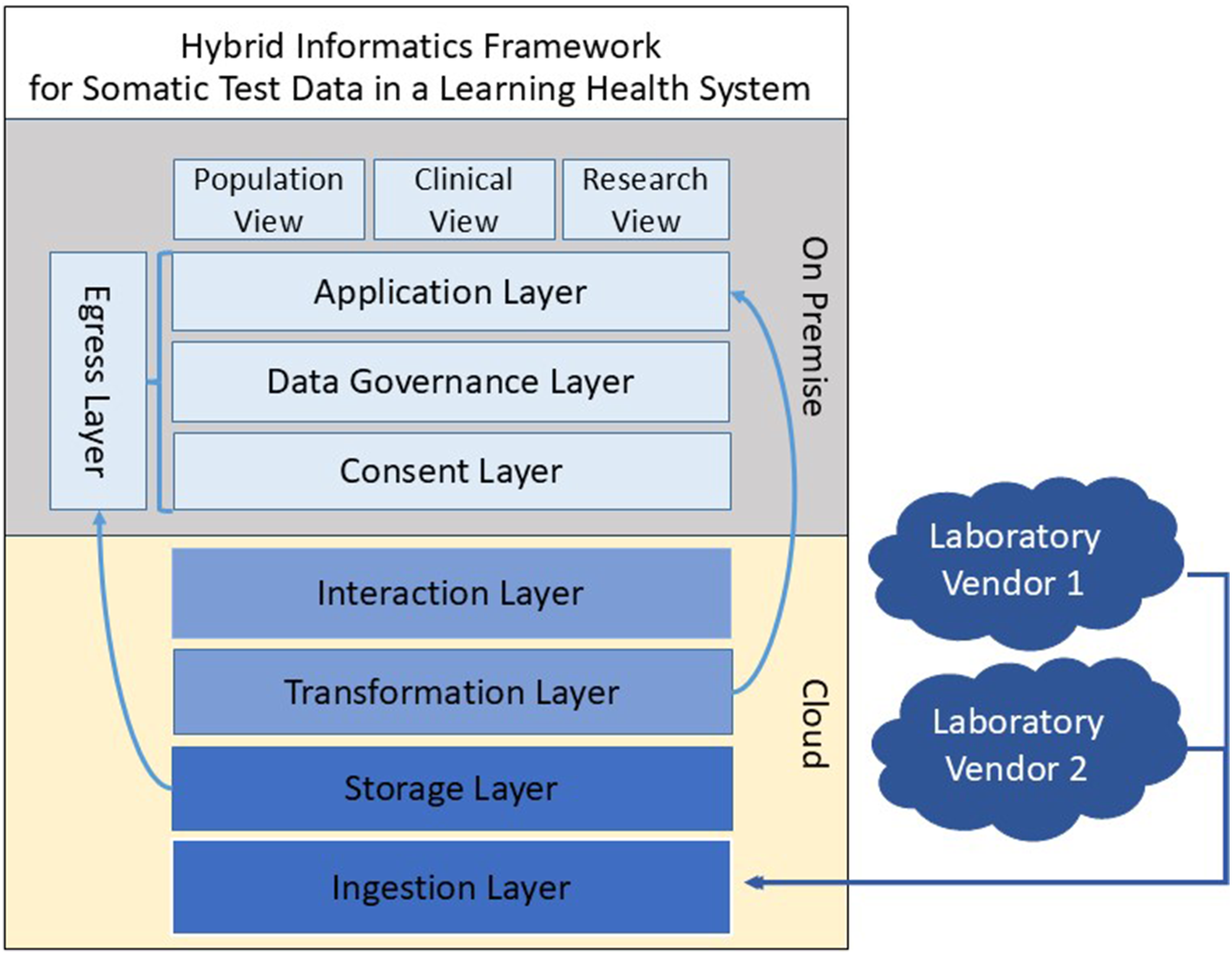
This work presents a framework that builds upon previous work by other researchers14,15,17 by including additional layers to facilitate research use of the data in the form of a consent layer, data governance layer, application layer and egress layer. The on-premises application layer facilitates user interface tools for supporting clinical and clinical research functions, and is the foundation for clinical research support. The egress layer allows for the distribution of large raw sequencing files such as FASTQ and bam to research projects. Additional provisions to lower the start-up costs and storage costs with a hybrid cloud and on-premises solution are included in this framework while also expanding metadata creation in the Transformation layer, to improve data utility downstream for separate functions of molecular cohort building and data sharing. This Data Lake, with its expanded architecture layers, has been evaluated at an academic cancer center and shown to be flexible to accommodate the ingestion of data from multiple sequencing vendors and to offer services effectively to a diverse set of clinicians, researchers, and bioinformaticians.
Methods
The following section will provide a detailed implementation of the Clinical Omics Data Lake architecture shown in Figure 1. We will describe the data ingestion steps, metadata creation, security and confidentiality, and data governance. After describing the implementation, we will describe one application by our Institution that relies on the Data Lake. As shown in Figure 2, oncologists order genomic tests with a test vendor via a Health Level Seven (HL-7) Fast Healthcare Interoperable Resources (FHIR) module integrated with an EHR or some other test ordering system. The genomic sequencing objects are deposited in the vendor’s storage on the cloud, while the summary results are returned to the clinician, for example, as PDF reports. These PDF reports contain patient data (text and graphics) important to the treating clinician. The PDF report is generated based on the genomic test reporting standard established by the American College of Medical Genetics and Genomics (ACMG)
18
and HL-7. The PDF files are stored in AWS and linked through the application layer to the patient record in the metadata. The patient metadata allows the clinician to perform searches based on gene mutation, PDL-1 status, MMR status, and so forth. Once the clinician has targeted the patient population they are interested in, then they can open individual PDF files to view the annotations of clinical trials and treatment options for each individual patient. The genomic objects are then copied from the vendor’s storage to the Data Lake’s storage on the cloud, and metadata are generated for these objects. Whereas the clinician may download genomic test files for their patient, any requests for research purposes are made through a data governance committee. If the governance committee grants access to the data of consented patients, the omics data are downloaded via Globus or directly from the cloud provider. Below, we describe the methods used for the implementation of the Clinical Omics Data Lake architecture. The implementation descriptions provide the details required to implement the architecture. The descriptions are based on AWS, but other implementations may be used by other cloud providers. Data flow diagram showing the process from genomic test ordering to genomic file access by a researcher. Abbreviations: AWS- Amazon Web Services; ICTR- The Institute for Clinical and Translational Research; EPIC- EHR Vendor; EHR- Electronic Health Record; FHIR- Fast Healthcare Interoperability Resources; ODL- Omics Data Lake; S3-Simple Storage Service.
Data integration
The Apache Hadoop framework is the most common Data Lake development approach. We considered four factors in deciding whether to have a custom Hadoop based implementation versus a cloud-based implementation. These factors include scalability, cost of maintenance, integration with analytic services and security and compliance. The most critical of these factors were security and compliance. Cloud-based offerings (e.g., from AWS, GCP and Azure) provide built-in security features (such as data encryption and access control) and compliance certifications (e.g., HIPAA). The second key factor for consideration is the cost of maintenance. Managed cloud services reduce administrative overheads, as opposed to custom implementations. An in-depth look at these factors can be found in a recent review. 19 We, therefore, chose not to use the Hadoop-based implementation due to these two main considerations: security and compliance requirements and the cost of maintenance.
Instead, we created a Data Lake architecture based on AWS Simple Storage Service (S3). We designed this Data Lake following the hybrid cloud integration paradigm, with two Virtual Machines (VM) on-premises and the raw sequencing files stored on AWS. The result of genomic testing has well-defined file types (FASTA, BAM, VCF, etc.), but there is still a need to integrate these into the rest of the patient data in other systems. The data integration is performed via record linkages. This is possible since the metadata maintains an identifier that resolves the patient. At our academic medical center, we support distributed clinical environments with patients whose records are recorded in multiple EHR systems with different patient IDs. Patients are also transient and frequently seen at more than one of our clinical affiliates resulting in multiple medical record numbers. To limit duplication of patients in the Omics Data Lake, the integration layer solves this issue by resolving the incoming patient MRN and MRN site matches. This initial step provides a blocking technique. Running the resulting patient records through a custom weighted pairwise matching algorithm using first name, last name, and date of birth has resolved all but a handful of records that are screened manually. The Data Lake stores the large raw data (FASTQ and BAM) files under the S3 archive storage class, and the Variant Call Files (VCF) and clinical reports in PDF format are under the S3 Standard storage class. We utilized Globus 20 file transfer to transfer files to a user’s file system.
Metadata
To prevent the Data Lake from becoming a data swamp,
21
we attach metadata to every file in the Data Lake. There are several potential metadata management applications. The first is the AWS Data Catalog which requires each table to be associated with a single database. To seamlessly integrate the metadata across test vendor platforms, we did not think extending the AWS Data Catalog would offer any advantages. AWS Data Catalog tracks basic metadata (name, size, location, update dates) for genomic file objects, but is not capable of managing, versioning or querying binary objects. With AWS Catalog, adding rich metadata content is possible with the use of other AWS-based services ( such as AWS Glue crawlers), and with the use of classifier services, but these have no capability to infer meaning from this binary file, and pre-processing with custom classifiers would be required to infer richer metadata. This process involves creating custom metadata tables within the AWS Glue services, and these tables must reside in the same database. While AWS Catalog can be utilized for managing metadata for genomic files objects, from an efficient standpoint, it offers no advantages over a custom metadata service. The second, Apache Atlas, is a metadata and governance application. Apache Atlas could not appropriately manage our Data Lake’s metadata, for two main reasons. First, being optimized for a Hadoop-based system, a quick analysis showed that Apache Atlas would not be flexible for cloud environments without Apache’s stack of tools. Second, the three main cloud providers (AWS, GCP, and Azure) do not offer Apache Atlas. We could not find any other suitable metadata management model, so we developed a file metadata model that links the patient identifier and test identifier to the files while annotating the file type (BAM, FASTA), size, and nucleic acid materials (DNA, RNA) as shown in Figure 3. The file metadata, along with the PDF test result files, are persisted in an on-premises NoSQL database (MongoDB). Metadata generation process. Two sets of metadata are generated for different functional purposes. Shown in blue, the file meta data is used to link ordering clinicians to tests, patients, and files and is stored in a NoSQL document store database. The test metadata shown in green is utilized to identify cohorts of patients for clinical research and governance approved research requests and resides in a relational database. Both metadata sets reside on premise and are utilized by the consent, governance, and application layers of the architecture.
The transformation layer generates a second set of metadata encapsulating the minimal information needed to identify cohorts from the test results shown in Figure 3. This metadata includes minimal patient, test, mutation, RNA, and therapy data. The patient data consists of a unique patient ID, patient demographics, and an anonymized patient ID. The test data contains information such as ordering physician, patient diagnosis, test type, specimen type, and accession ID, enabling the capture of serial testing data. The mutation and RNA findings contain information about the gene variants and clinical significance. The metadata (file metadata and test metadata) are both heterogeneous. The metadata fields can grow depending on the vendor test and genomic files being deposited. Implementations of the metadata need to be consistent with established genomic data standards such as the Human Genome Variant Guidelines (HGVS) 22 to enable data sharing. Compliance with clinical and genomic data standards was made easier due to the vendor’s adherence to the ACMG and HL-7 FHIR standards for clinical genomic reporting.18,23
Security and confidentiality
HIPAA requires that covered entities that rely on third parties to manage Protected Health Information (PHI) must have a Business Associate Agreement (BAA). Our institution has a BAA with test vendors and AWS. AWS’s S3 stores encrypted data and downloads from S3 are also encrypted. Patients are consented to participation in future research projects at the time of specimen collection according to institutional procedures, and the consent information is stored outside this Data Lake framework. Consent information is integrated nightly with the metadata to identify clinical sequencing files available for the data governance committee to consider for research distribution.
Data governance
Data governance can introduce complexity into a distributed data project, and we sought to minimize complexity in data governance by articulating its steps. We established a formal data governance process to address research use of the data once use cases were clearly defined, Figure 4. Data request and the governance process. A researcher’s request for data starts the workflow process. The researcher initially interacts with the Honest Broker to locate the presence of tests from patients of interest, then submits a request for consideration by the governance committee. A researcher needs IRB approval before accessing data de-identified by the Honest Broker before distribution.
The Data Lake is made available for clinicians in the operations of their clinical practice and research use by researchers with Institutional Review Board (IRB) approval. The Data Governance Committee membership is comprised of clinical stakeholders, and the committee’s membership changes regularly, with the chairperson rotating biennially. While the genomic analyses are performed as a clinical service, consenting patients to research applications can burden the clinical team. Therefore, the system has become a trusted resource by empowering clinicians with governance over access to the research data. The governance process begins when a researcher approaches the Honest Broker to identify data of interest or submit a data request application. The Honest Broker is an informatician who assists in preparing and de-identifying any clinical data to be taken out of the Data Lake. The committee can deny, amend, or approve the request and communicate its decision to the researcher and the Honest Broker.
Results & Discussion
The methods described above were utilized to implement the Omics Data Lake architecture. During implementation, each zone may utilize a wide variety of technologies. The choices range from commercial offerings from cloud providers to custom implementations based on both commercial and open-source components. In the implementation we have described above the choices of components to use were driven by the cost of maintenance, security, and compliance considerations. We did not perform an analysis to ensure that the General Data Protection Regulations (GDPR) compliance is met by the implementation, but the architecture implementations can utilize different compliance components, as necessary. In the healthcare setting, patient consent is a critical consideration when patient data is to be shared. The implementation places governance (including patient consent) as part of the workflow, making it possible for researchers to access data in a way that meets compliance regulations. The implementation currently enables genomic test results from two commercial test vendors, Tempus, and Caris, but it is vendor-agnostic. The Data Lake currently stores approximately 240 TB of data on 5800 patients from two independent clinical sites and supports 31 disease groups. A total of 149 clinicians have utilized the system. In the following discussion, we will describe the choices made during the implementation, the advantages and limitations of the Data Lake architecture, and discuss some of the publications that benefited from the Data Lake. Because a Data Lake does not have a schema, it was necessary to create an application that summarizes the state of the Data Lake. We, therefore, developed an in-house data summary dashboard utilizing the Data Lake’s metadata. The dashboard application also demonstrates the ease of accessing data from the Data Lake.
Example application: Population view summary dashboard
A dashboard application with a summary of the represented patient population was implemented as a web application showing the contents of the Data Lake, Figure 5. This dashboard is not part of the Data Lake but is a sample application built on the application layer that consumes data from the Data Lake. The dashboard allows physicians, clinical research coordinators, and researchers to view aggregated data and make de-identified queries on the Data Lake. The metadata information enables the integration of data between the Data Lake with clinical applications. By the first quarter of 2024, two institutional providers representing 16 clinics had participated. The total number of physicians participating was 149, with 31 disease groups represented. Possible uses of the summary query dashboard include data exploration for hypothesis generation, cohort identification, scanning for patients eligible for clinical trials, and presentation of patients for molecular tumor boards. An example dashboard of an intranet application utilized to identify the contents of the Omics Data Lake. For ease of visibility, additional graphics are not shown.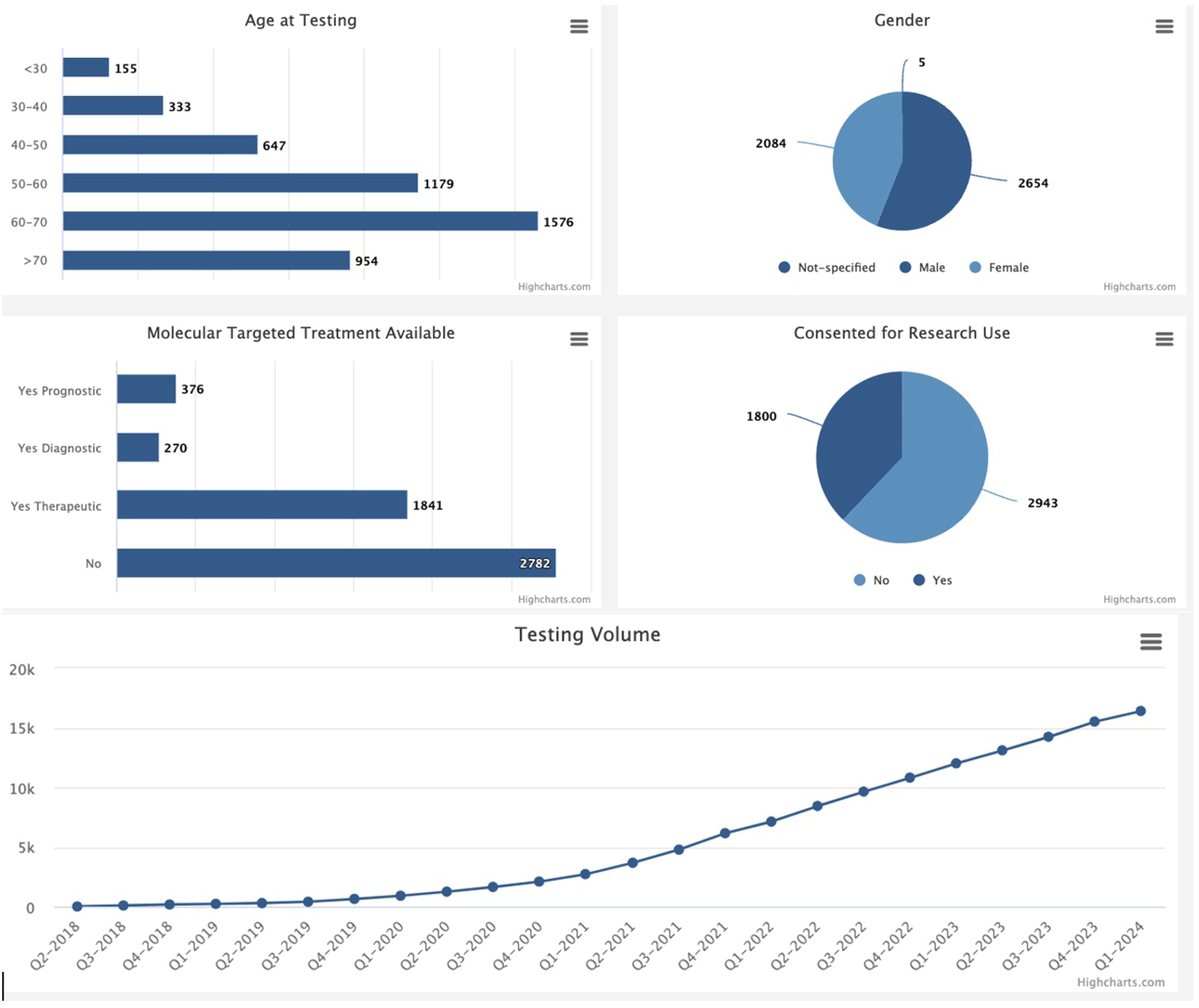
Adaptability and applied use cases
Since deploying the Data Lake in 2021, it has addressed several use cases for clinical and research purposes. A selection of the clinical use cases addressed by the Data Lake implementation includes support for the molecular tumor board, oncologists querying for expected mutations during serial testing, mutation levels below the clinical reporting threshold, and false negatives due to areas of low sequencing coverage. Translational use cases addressed include molecular screening of existing patients for a newly established clinical trial and clinical trial feasibility assessment based on patients seen in the previous 5 years to predict the prevalence of patients eligible to enroll in a future clinical trial if established at our clinical sites. Research data requests have presented the most diverse use cases, including requests for sequencing test files identifying and reporting gene fusions in Glioma,24,25 RNAseq files for 200 prostate cancer patients stratified by race and ethnicity for gene expression analysis, case report files for patients diagnosed with the simultaneous occurrence of two independent primary cancers, 26 and serial liquid biopsy test results for breast cancer patients while on treatment. 27 One recently developed application of omics data lake has been to support the opening of a Clonal Hematopoiesis of Indeterminate Potential (CHIP) clinic at our academic medical center by referring existing patients. CHIP refers to the clonal expansion of white blood cells that carry acquired mutations. It is more common in older individuals. It carries an increased risk of developing certain blood cancers and/or cardiovascular disease and, therefore, warrants monitoring. Patients receiving NGS testing at our clinical facilities are consented on an excess tissue institutional biorepository protocol. The metadata in the Omics Data Lake is being used to annotate biospecimens by filtering consented patients by molecular mutations. The results are then cross-referenced against the biorepository inventory. The Hybrid Data Lake model has enabled the successful distribution of these data without modification to the architecture framework. Through addressing these requests, the implementation team has taken note of the functionalities that would further enable discovery and self-service data access for future development.
Discussion
Our institution needed a system to integrate genomic and clinical data from disparate systems to serve clinicians in our Comprehensive Cancer Center across multiple affiliate clinical sites. The institution’s existing clinical data warehouse (CDW) was neither flexible nor efficient at handling molecular data, so it did not serve the needs of our oncologists or researchers. Additionally, the institution serves multiple affiliated hospitals and clinical environments, some of which do not have CDWs. The Institution needed a system enabling clinicians to gather clinical genomic test data from genomic testing vendors that were not tied to any specific testing vendor and allowed clinicians to change vendors in the future without losing historical testing data. Additionally, institutional researchers needed to be able to access raw sequencing data for research. With limited resources, it was impossible to construct a traditional MDW to serve this need. In addition, the institution’s physicians have patients from multiple clinical affiliate sites with varying levels of technical integration needs. Building an MDW spanning different organizations is a difficult task that takes years. 9 With those constraints, a Data Lake architecture was a more viable architecture option.
There are two general approaches to integrating clinical data from various sources: CDW, Clinical Data Lake, 28 and service-oriented architectures. 29 The most popular CDW offers out-of-the-box analytics and ease of use, but its model presupposes that the creators of the CDW know all the truth at the time of construction. Due to the specificity of the data requirements, the construction process is costly. However, the resulting DataMart model is brittle, not readily amenable to different use cases, and is slow to change. Such a CDW is unsuitable for genomics research, where rapid change in data and methodologies is standard. On the other hand, a Data Lake imposes no model at design time, so the genomic data are stored in their raw format. The lack of model-on-write makes setting up a Data Lake quick and just as rapid to change it. Whereas the center bears the operational costs of operating the Data Lake, the downstream analyses and model creation costs are delegated to the end user, that is, the associated monetary and time costs are pushed to the end user. These costs include egress charges for requesting the genomic files out of the Data Lake, and any other costs the user may incur when creating models from the data, which might include performing data analytics such as applying machine learning applications.
Strengths & weaknesses
The architecture has three main strengths. First, the cost of implementation and operation is comparatively minimal: It took one software engineer 9 months at 0.5FTE to initially implement the architecture. The monthly costs of operation are approximately $350. The Data Lake currently includes approximately 5800 patients, with about 20,000 tests. The stated cost includes persisting 240 TB of data and egress charges in 2024. Egress charges can easily be separated from the ongoing data operational costs and passed onto researchers. There are other minimal costs, such as those for maintaining the on-premises VMs that host the applications managing the Data Lake, and modest maintenance personnel costs. Second, putting the Data Lake in operation took less time than implementing a traditional MDW. Third, whereas architecture development requires domain knowledge from biomedical informatics, bioinformatics, and oncology, the architecture implementation requires expertise from software engineering. MDW implementation using traditional CDW approaches requires expertise from many domains. 8
The Data Lake architecture has three noted weaknesses. First, a Data Lake does not have out-of-the-box analytics capabilities. For any use case, expertise would be needed to develop a model from various domains. We think the schema-on-read approach of a Data Lake is an advantage; only the model demanded by a use case is created, as opposed to the schema-on-write approach of an MDW, where the DataMart attempts to meet future use cases that may never be utilized. Second, a Data Lake can become a data swamp if metadata management fails. Third, while the initial start-up cost may be low, cloud storage and egress from the Glacial storage can be expensive. To address this burden, the design of the Data Lake pushes the cost of the egress charges onto the data requestor, in effect transferring the cost from the institution to the beneficiary of the data. Finally, whereas a summary information dashboard has been generated to enable some insights into the Data Lake, to realize the full utility of this data in a learning health system, more data science applications would be required. These future applications will leverage the data lake to create tools that provide insights into clinic practices, support precision medicine clinical decision-making, and provide analytics to facilitate improved population health.
Conclusion
We succeeded in creating a cost-effective Data Lake for storing clinical sequencing results ordered by oncologists at our comprehensive cancer center, which can retrieve data on-demand and is vendor-agnostic. The combination of the cloud-based Data Lake in conjunction with a local document database for metadata enables enterprise applications to easily utilize the Data Lake. Establishing a data governance process and leveraging the institutional patient consent mechanism has allowed these data to be used to both move our comprehensive cancer center closer to a learning health system and to stimulate research on the molecular drivers of cancer in our patient population. Finally, creating a governance environment that empowers clinicians to participate in the distribution of data that they have generated fosters collaborations between clinicians and researchers while also engendering trust within the system and use of the data environment.
Footnotes
Ethics approval
Any use of data for research from this data repository requires that an IRB-approved protocol has been obtained. Data governance and data release by the honest broker system only occur once a review of an IRB-approved protocol has been processed to ensure that the proposed research lies within the scope of the IRB-approved protocol. The purpose of our data lake environment is to store and manage data that can be used for research studies as well as for clinical management of patients. A primary purpose of the data lake is to provide access by clinicians to data from patients they have supported and ethical approval is not required for that aspect of our design. We do manage informed consent tracking in this data lake so that research can be conducted for research studies for individuals who have consented to them. Patients who would like to consent to research were offered opportunities to participate in several studies, including H-18245, H-14435, and H-49185, which were approved by the Baylor College of Medicine Institutional Review Board.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is supported by funding from the Dan L Duncan Comprehensive Cancer Center P30 Cancer Center Support Grant NCI-CA125123; This work is funded by the Institute for Clinical and Translational Research at Baylor College of Medicine, support Number: 9351200101/BA 7210.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
