Abstract
The COVID-19 epidemic has demonstrated the important role that data plays in the response to and management of public health emergencies. It has also heightened awareness of the role that ontologies play in the design of semantically precise data models that improve data interoperability among stakeholders. This paper surveys vocabularies and ontologies relevant to the task of achieving epidemic-related data interoperability. The paper first reviews 16 vocabularies and ontologies with respect to the use cases. Next it identifies patterns of knowledge that are common across multiple vocabularies and ontologies, followed by an analysis of patterns that are missing, based on the use cases. Conclusions show that existing vocabularies and ontologies provide significant coverage of the concepts underlying epidemic use cases, but there remain gaps in the coverage. More work is required to cover missing but significant concepts.
Introduction
With the increasing occurrence of public health events, cities are faced with the dual problem of understanding the spread of disease and managing the resources necessary to reduce its impact. It has become obvious in the most recent COVID-19 pandemic that data plays a central role. From tracking the spread of disease, identifying the most vulnerable, distributing vaccines, determining vaccine efficacy to managing the provisioning of medical services, data has played a central, critical role in a city’s ability to respond.1,2 Nevertheless, cities, states, and countries have had to “go it alone” in developing the information systems to support these activities. They are often faced with variety of purpose-built systems that were not designed to interoperate. More importantly, they do not share a common data, making it difficult to integrate data from multiple data sources and systems.
In response to this challenge, ontologies have been proposed to establish a common data model that provides precision in the representation of medical and epidemiological knowledge, reduces the time to create the necessary information systems, and achieves interoperability amongst them.3–6 An ontology provides a unified and explicit representation of shared understanding.
7
It consists of a vocabulary comprised of classes and properties with explicit definitions in the form of logical axioms that restrict how terms in the vocabulary can be interpreted and used.
8
A class represents an abstract category of physical or abstract things. A class can have object properties to represent relations among classes and data properties that specify literal attributes. In Figure 1, Class “Vehicle” has an object property “ownedBy” and a data property “ID”. These properties represents that a vehicle is owned by an organization and its ID is a string. A simple example of a logical axiom states that a Vehicle is owned by exactly 1 Organization.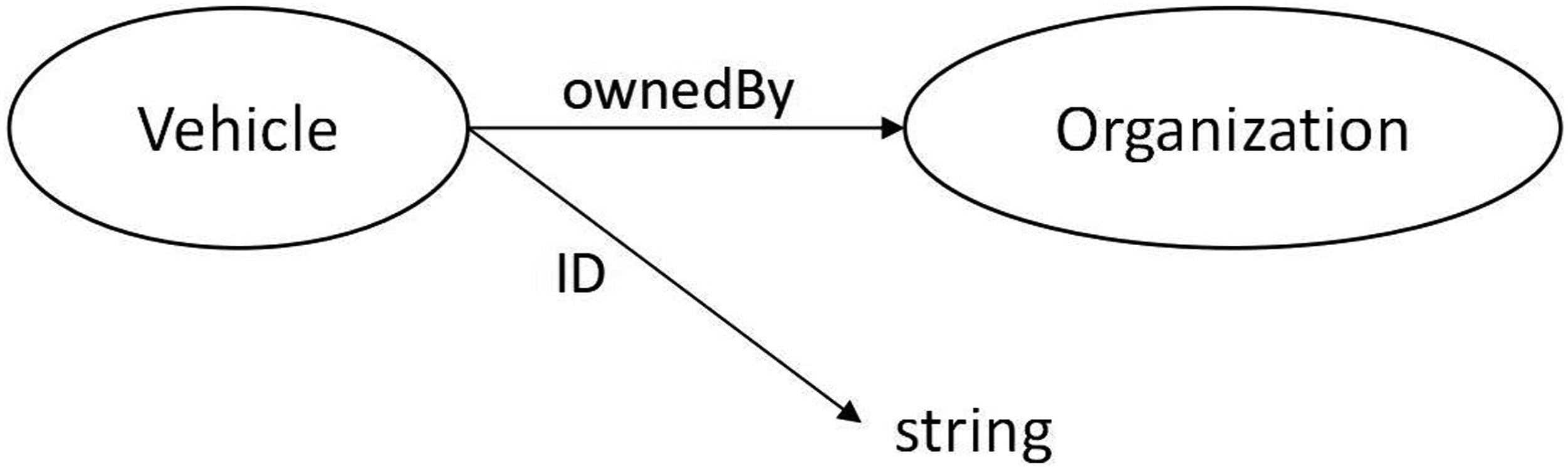
In this paper, we review and summarize existing epidemic vocabularies and ontologies (V/Os) and compare them from the perspectives of use cases, core classes and properties, patterns, etc. We define thirteen data-centred use cases for public health emergencies in a technical report and 11 of them are use cases for using data to manage epidemics. 9 These use cases provide a broader view of the activities needs for data. Use cases include Disease tracing, Contact tracing, Testing, Infection prevention for medical personnel, Vaccination management, Vaccine efficacy management, Vaccine distribution management, Transportation management during epidemics, Mental health management, Emergency medical resource management, and Stakeholder collaboration. 9 Use cases are drawn from different stakeholders such as public health department, medical organizations, medical personnel, citizens, etc. To evaluate the relevance of common patterns found in the reviewed V/Os, we identify these use cases in which they would be used. We also use the use cases to identify patterns that are missing in the reviewed V/Os. A pattern is a set of classes that are related by topic and inter-connected by properties, thereby forming a graph.
In the following, Section
Summary of Vocabularies/Ontologies
We summarize existing V/Os that overlap with Epidemic Management. We searched for related articles by keywords, and found new articles from the references of existing articles in an iterative manner. Some open-source V/Os related to epidemics in Github or BioPortal were also collected. At present, there are a limited number of V/Os proposed in the field of epidemic management. Reviewed V/Os meet the following requirements: • V/Os are published in English from 2006 to 2021; • V/Os are searched through Google scholars, Github or BioPortal; • Keywords are (“Covid-19” or “Epidemic” or “Infectious disease”) and (“Ontology” or “Vocabulary” or “Data model”); • V/Os that do not introduce application or uses are not included; • V/Os that mainly focus on the disease itself rather than the management of the disease are not included.
The remainder of this section summarizes seven V/Os. Additional nine related articles that are not directly related to epidemics or do not provide detailed V/Os are described in 2.10. For each V/O reviewed, the purpose, use case, core classes and properties are summarized.
EPO
Epidemiology Ontology (EPO) is designed for sharing epidemiology resources. 10 It reuses some existing ontologies and introduces new classes with the aim of providing unified semantic annotation for epidemiology-related data, such as demographic, geographic, social, and vaccination rates. It supports data integration, information retrieval, and knowledge discovery activities. EPO is accessible at https://code.google.com/p/epidemiology-ontology/.
Use case
A research team is investigating the model of incidence rates of an infectious disease as it relates to vaccination rates in different populations/locations. Data such as demographic, geographic, social, personal information, and vaccinations need to be collected to understand how herd immunity is related to vaccination rates. EPO would be used to provide a unifying data model so that the diverse sources of information can be integrated and used for analysis.
Core classes and properties
EPO has approximately 200 classes that cover transmission of infection, epidemiological and demographic parameters. EPO reuses Basic Formal Ontology (BFO) as an upper-level ontology.
11
An upper-level ontology defines and axiomatizes these most general categories. BFO contains generic classes such as “object” and “process.” Pathogen Transmission Ontology (TRANS) is imported as middle level that provides the taxonomy of transmission.
12
Then EPO extends it by defining specific modes of transmission such as sexual and biological transmission. Some classes from Infectious Disease Ontology (see Section
Figure 2 represents a portion of EPO and the relationship between classes in EPO and other ontologies. Unlabeled arrows represent subclass relations. The main class is transmission of infection from TRANS. A deeper taxonomy of transmission of infection is also defined. Transmission happens during communicable period. The focus of infection represents the location of transmission. Primary contact, infectious agent vector, and biological vector of infectious agent participate in different types of transmission. A representative portion of EPO. Reprinted from “The epidemiology ontology: an ontology for the semantic annotation of epidemiological resources” by Pesquita et al., 2014, 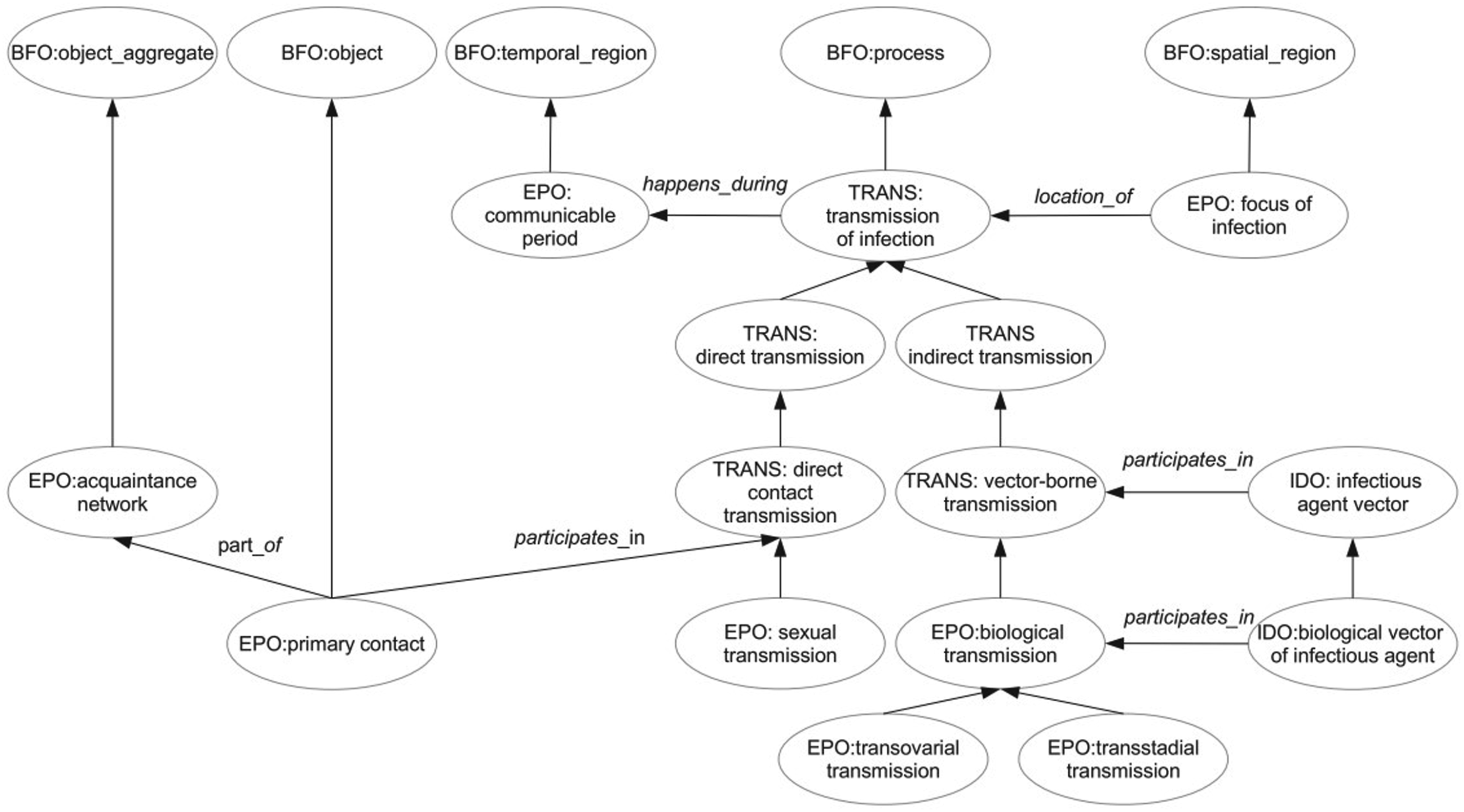
GeMInA
Genomic Metadata for Infectious Agents (GeMInA) is designed to provide a disease detection tool based on geospatial information. 12 GeMInA project has developed an ontological standards system that tracks the pathogen-related metadata by data mining. This is helpful to model the disease and comparatively analyze between strains and species. GeMInA is available at http://gemina.igs.umaryland.edu.
Use case
Some biomedical researchers are studying the interaction between different kinds of pathogens and their hosts and infectious diseases. They need to extract important information from the large amount of data of World Health Organization (WHO). GeMInA system provides researchers with quick access to a wide range of information, including the time and locations of outbreaks, the pathogens and hosts infected, and the cases and symptoms reported during a given time period.
Core classes and properties
GeMInA covers foundational components of infection to uniquely identify a pathogen. Some components are pathogen, host, disease, transmission method, and anatomy. Infectious incident data, such as gender and age of the infected people, date when incident occurs, are also included in GeMInA. Figure 3 shows an example of search results in GeMInA. Pathogen is the core class and it links to related disease, corresponding transmission method, host, anatomy, related incidents, etc. Example of search results for “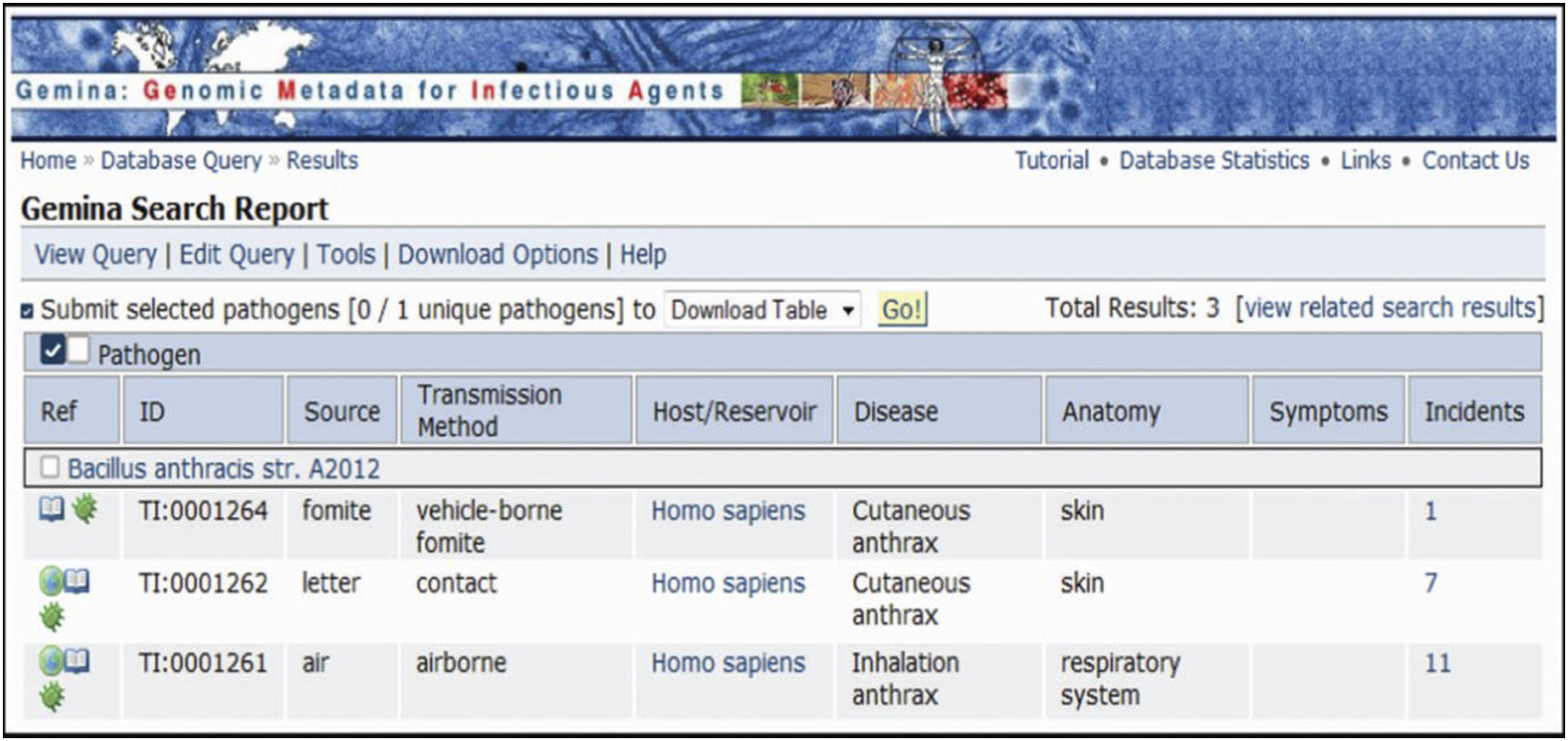
CIDO
Coronavirus Infectious Disease Ontology (CIDO) is a community-based, open-source biomedical ontology that facilitates the standardization, integration, sharing, and analysis of COVID-19 knowledge and data. 15 It is accessible at https://github.com/CIDO-ontology/cido.
Use case
To study the treatment and prevention of COVID-19, many clinical data and scientific research need to be studied to understand the etiology, transmission method, and pathogenesis of the disease. The amount of data has grown exponentially. CIDO helps to standardize COVID-19-related data and provide a unified representation that can be understood by humans and computers so that medical researchers can search and query data easily.
Core classes and properties
CIDO covers classes related to COVID-19, including its etiology, modes of transmission, epidemiology, pathogenesis, host-coronavirus interactions, diagnosis, prevention, and treatment.
15
In Figure 4, COVID-19 disease process realizes COVID-19 disease and occurs in animals’ lung. This disease process is caused by infection with a kind of virus: SARS-CoV-2. COVID-19 drug and vaccine are used to treat and immunize against such disease process. CIDO now contains over 4000 classes and instances, some of them are imported from existing ontologies. The design pattern of CIDO. Reprinted from “CIDO, a community-based ontology for coronavirus disease knowledge and data integration, sharing, and analysis” by He et al., 2020, 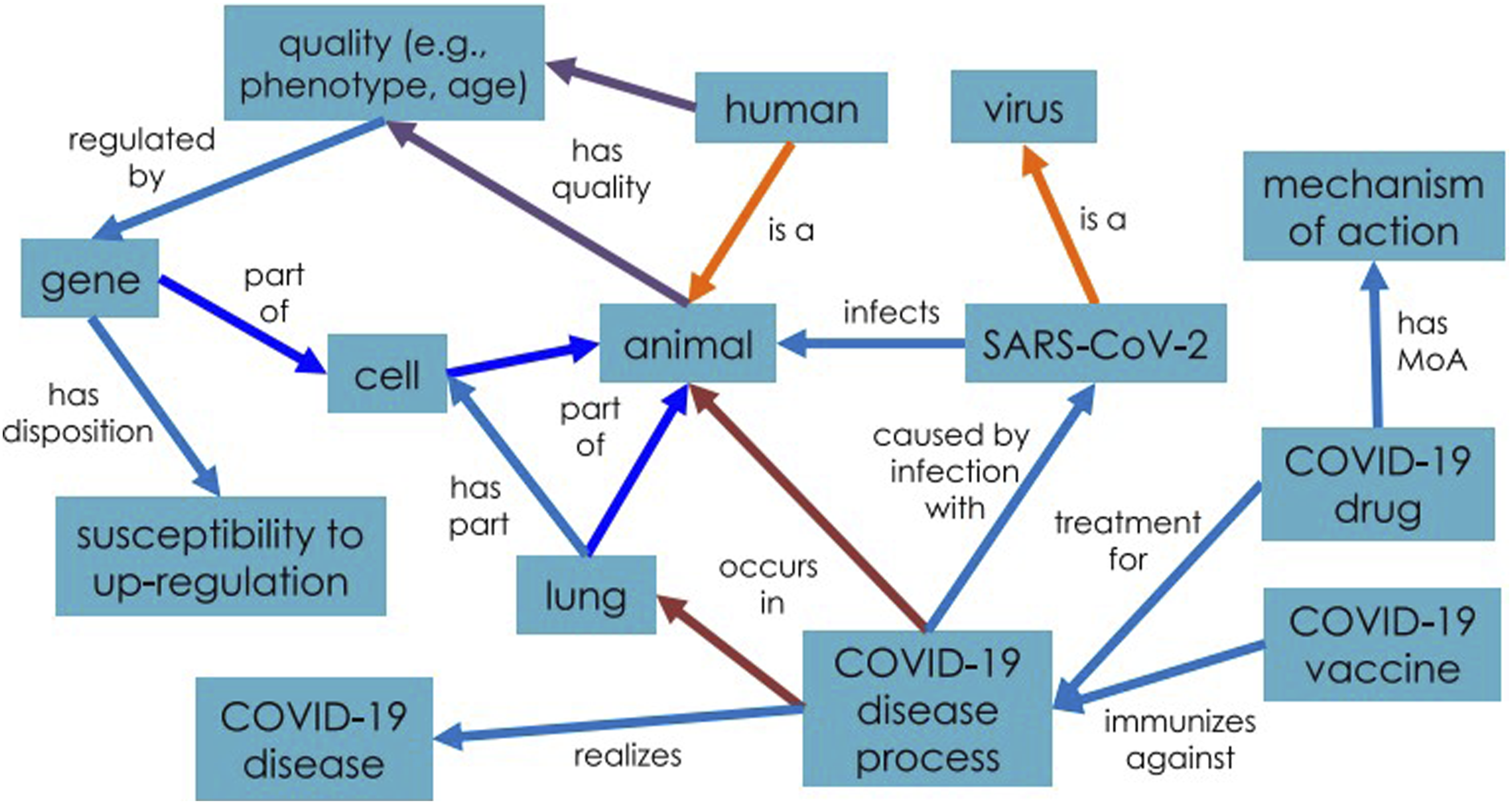
IDO Core
Infectious Disease Ontology (IDO) defines more than 500 classes and instances related to infectious diseases in general and serves as a foundation for more particular disease ontologies such as flu.13,14 The center of IDO is IDO Core, which extended by disease and pathogen-specific ontology modules. IDO and its datasets are available at https://github.com/infectious-disease-ontology/infectious-disease-ontology.
Use case
Researchers want to build a model to simulate the herd immunity of a population to an infectious disease. This requires the integration of biological and medical data and relevant public health, geographic and social science data, as well as data on the clinical presentation, diagnosis, and treatment of the patient with certain disease. Data is usually accessible locally since relevant information is acquired using various techniques and kept in geographically distributed and non-interoperable databases. This prevents researchers from comparing data across multiple dimensions and summarizing ways to prevent public health crises. IDO provides data integration and sharing methods to help them better access and use data.
Core classes and properties
BFO is used as the upper level and Ontology for General Medical Science (OGMS) is the middle level and provides definitions of disorder, disease, and disease course.11,16 OGMS covers relevant classes in clinical treatment. IDO Core is extended to define main classes including infectious disease, infectious disorder, and infectious disease course. Infectious disease course realizes infectious disease and infectious disorder is the material basis of infectious disease. In Figure 5, key classes are represented in boxes. Relationship between disease, disorder, and disease courses in IDO Core. Reprinted from “The Infectious Disease Ontology in the Age of COVID-19” by Cowell et al., 2020, 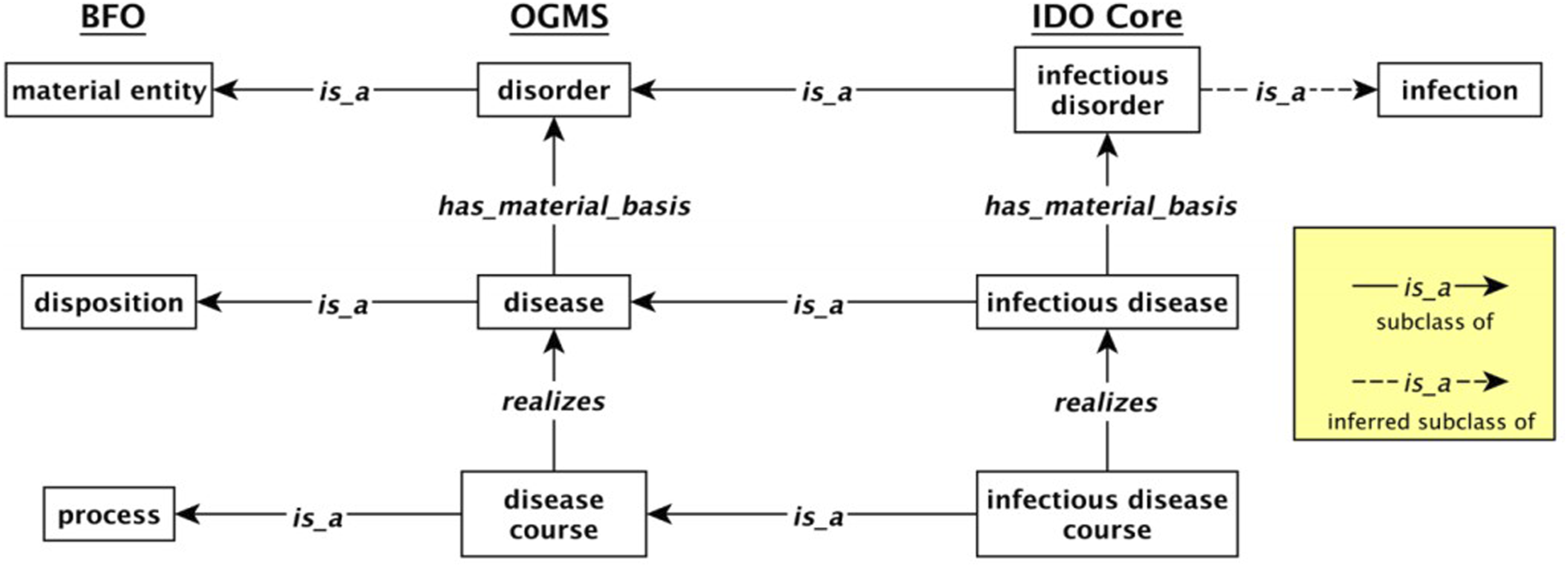
CODO
CoviD-19 Ontology for case and patient information (CODO) is designed to provide a knowledge graph (i.e., a graph database that uses an ontology for its data model) for COVID-19 data.
17
CODO also provides semantic services, applications, and vocabularies to help organizations annotate and interpret COVID-19 related data. Taking CODO as the schema and real pandemic data as instances, a knowledge graph is constructed to support data viewing in the form of graph and knowledge query and reasoning. In Figure 6, each node represents an instance and each edge represents a property. “P000003” is an instance of class “Diagnosed With Covid,” i.e. it represents a person who is infected. “P000011” is an instance of class “Person” and is the co-worker of “P000003.” CODO is mainly concerned with modeling the tracking of how patients are infected. It is available at https://github.com/biswanathdutta/CODO. Overview of CODO model. Reprinted from “CODO: an ontology for collection and analysis of COVID-19 data.” by Dutta et al., 2020, 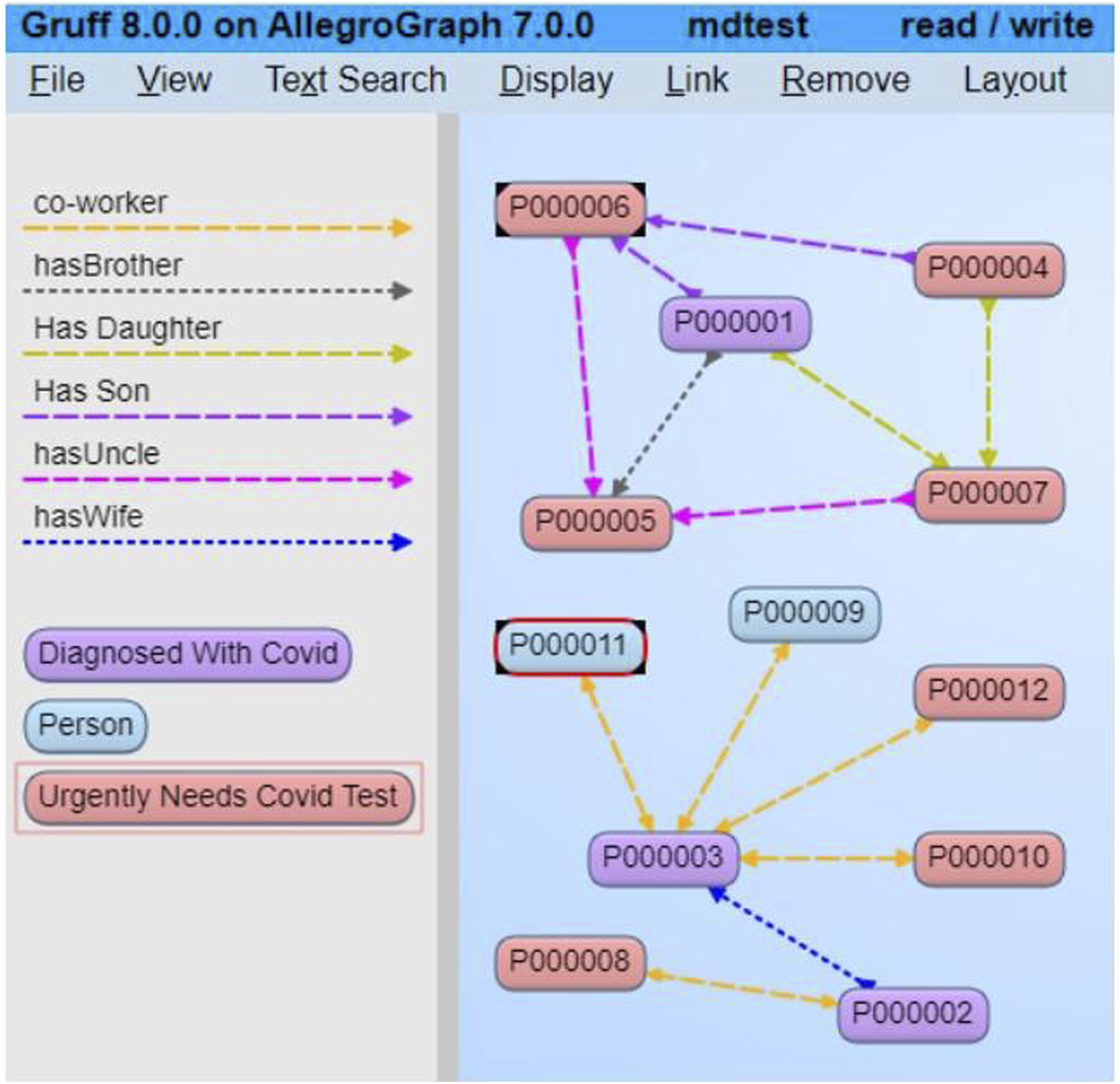
Use case
A project in Australia aims to develop an ontology-based COVID-19 risk detection system. CODO is used as a vocabulary to provide standardize COVID-19-related data and unified representations. These representations can be used as input for trend or growth study and help in analysing the behavior and transmission route of the disease.
Core classes and properties
CODO1.2 consists of 50 classes and more than 100 properties. Figure 7 depicts a high-level view of CODO. A person traveled from/to a place may become a patient and has symptom because of exposure to COVID-19. Therefore, person is admitted in a dedicated facility and get diagnosis. Participants, organizations, transmission modes, medical activities involved in this process as well as some statistics information are defined in CODO. Overview of the CODO model. Reprinted from “CODO: an ontology for collection and analysis of COVID-19 data.” by Dutta et al., 2020, 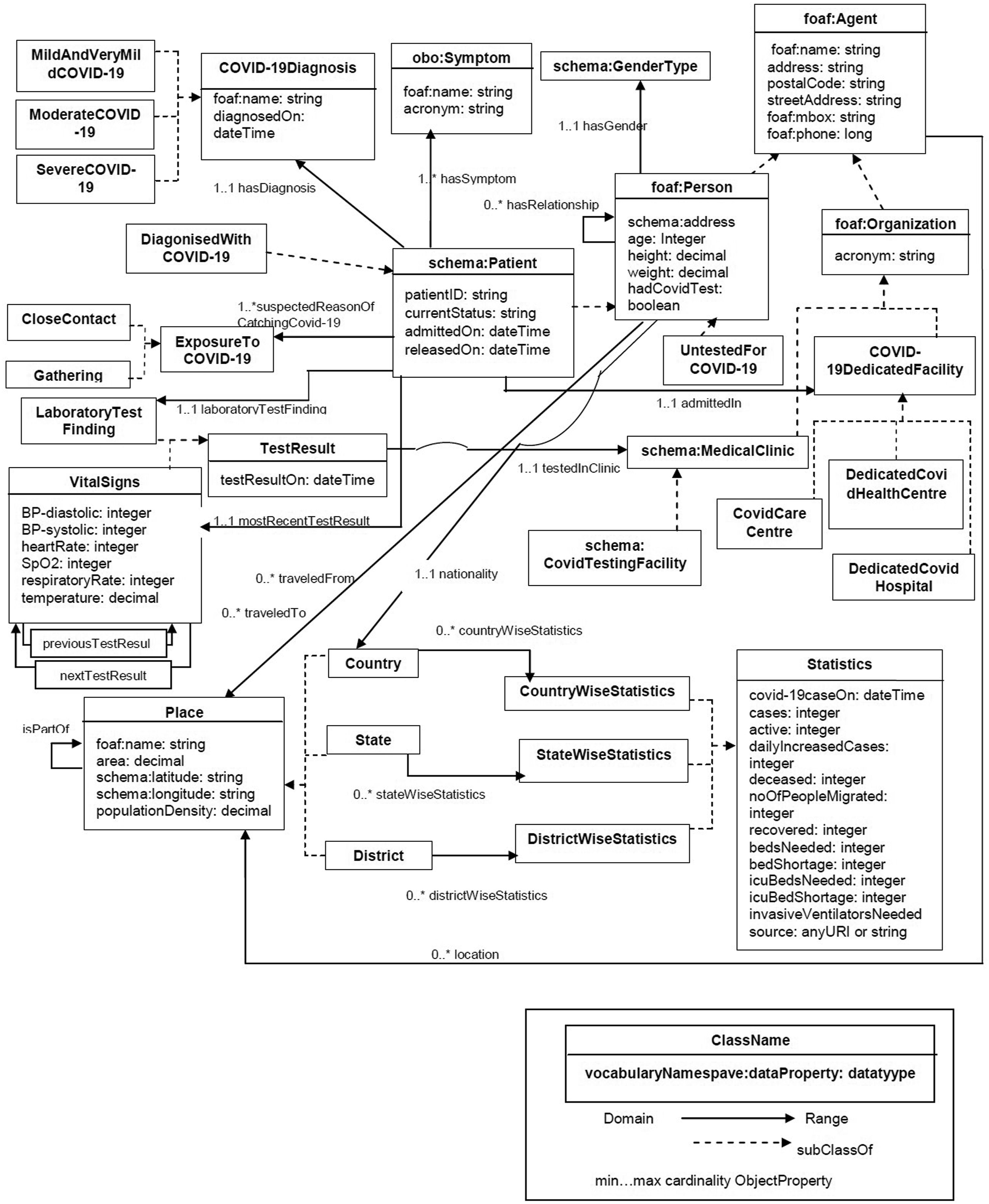
COVIDCRFRAPID
WHO COVID-19 Rapid Version Case Report Form (CRF) semantic data model (COVIDCRFRAPID) is designed for WHO’s COVID-19 case report form RAPID version. 18 It is intended to provide semantic references to questions and answers of the form. It is freely available at https://bioportal.bioontology.org/ontologies/COVIDCRFRAPID.
Use case
WHO collected information on COVID-19 from clinicians and patients in a systematic way and provide it to WHO Clinical Platform to help analyze and understand COVID-19. COVIDCRFRAPID was created based on this information. This ontology is used in a multilingual WHO’s Post COVID CRF representing standardized clinical data of patients after they have been discharged from the hospital or have had an acute illness. The standardized report supports online data entry of symptoms of COVID-19.
Core classes and properties
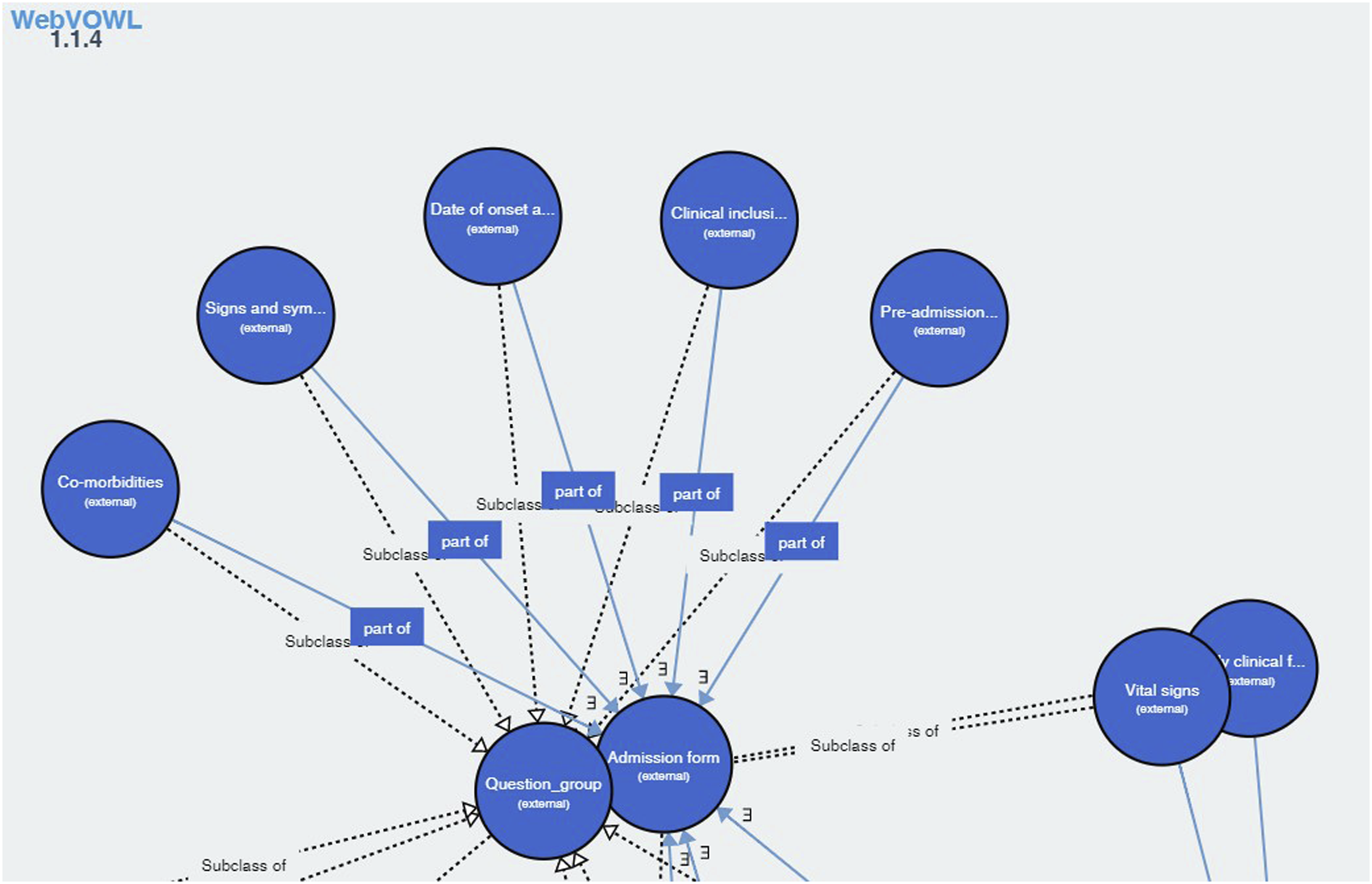
COVIDCRFRAPID 1.1.4 contains 398 classes and 13 properties. Most concepts describe clinical data, diseases and symptoms, occupations involved in medical activities. Figure 8 represents a portion of COVIDCRFRAPID and “Subclass of ” and “part of ” relationships between classes in COVIDCRFRAPID. Portion of COVIDCRFRAPID. Reprinted from “WHO COVID-19 Rapid Version CRF Semantic Data Model” https://vodan-ontology.github.io/.
18
OPM
An Ontology Proposal Model (OPM) is proposed as a semantic approach to analyze multi-sourced data and help to construct real-time statistics and reports on COVID-19. 19
Use case
A research project aims to identify and capture information about people who may have been infected with COVID-19 through localization technology. One approach is to use street cameras to catch person with symptoms that might be associated with COVID-19, such as coughing, trouble breathing, and bluish lips or face. Another way is to track the flight of people from high risk-level areas. The ontology combines the advantages of semantic web and big data, so that important concepts are defined and related information are acquired in real time.
Core classes and properties
In Figure 9, OPM contains classes extracted from Hospital, Airport and Camera Street Data Sources to capture and track potential infected person. Some significant class such as Person, Medical personnel, Patient, Passenger, and Flight are defined. Camera class is used to capture people in a Localisation way and detects their Behaviours and Symptoms. Disease class is also defined based on the categories of Viruses. The overview of ontology model. Reprinted from “Towards an ontology proposal model in data lake for real-time COVID-19 cases prevention.” by Kachaoui et al., 2020, 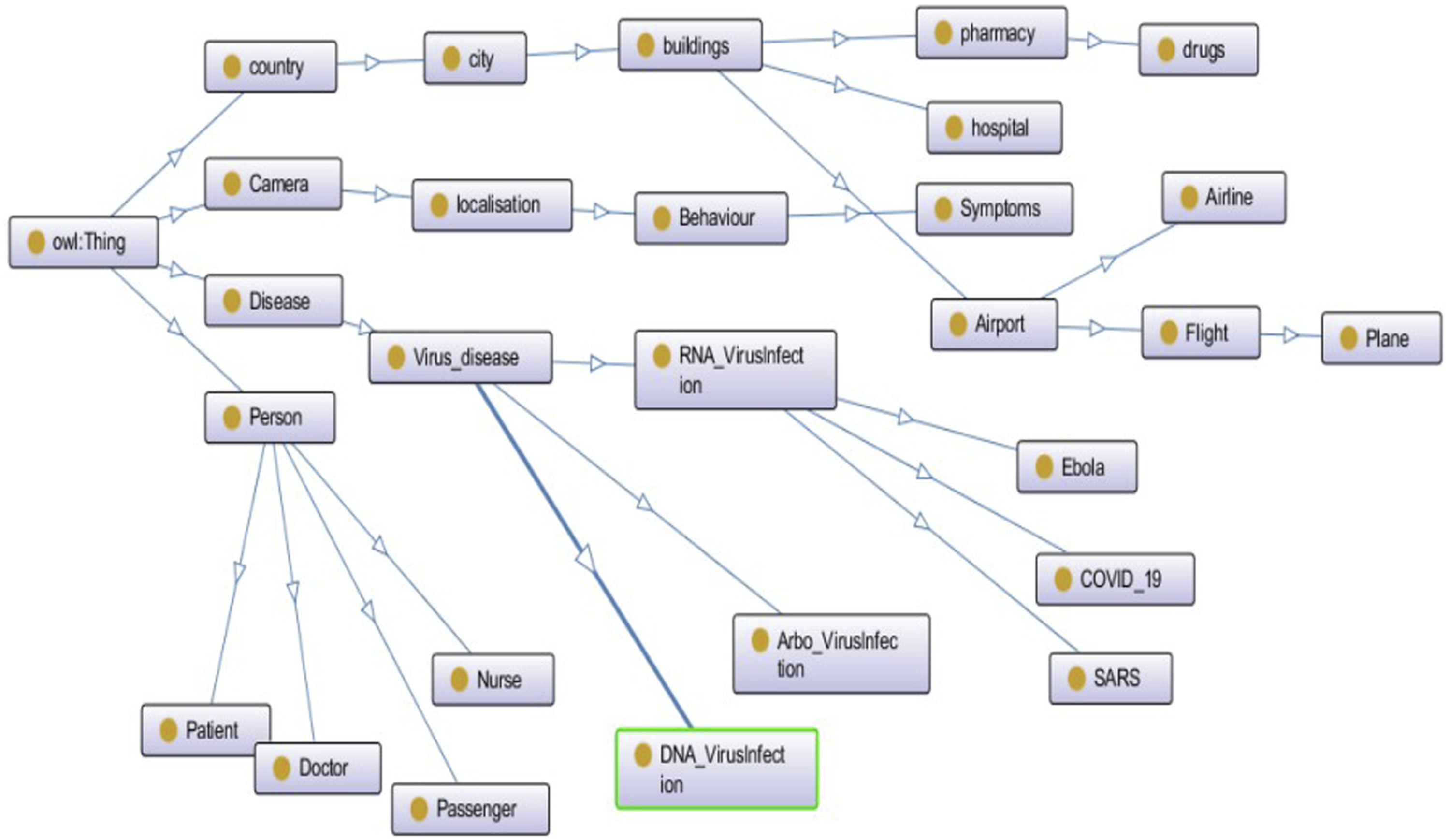
Other
The following briefly describes work that are not directly related to epidemics nor address the representations of epidemic related data, or do not provide sufficient information to be evaluated.
Fox et al. designed Global City Indicators Healthcare Ontology (GCIH) to represent definitions of healthcare indicators in ISO 37120:2018 standard.20,21 ISO 37120:2018 contains 19 themes and for each theme a set of indicators are defined to measure the quality and performance of that theme’s services. GCIH provides health domain concepts necessarily to represent a computationally precise definition for each health theme indicator using a machine-readable form, which means the definition of an indicator is deconstructed into its components, and each component is represented explicitly logically, and depicted as a graph. 22
Apollo Structured Vocabulary (Apollo-SV) provides a vocabulary for specifying input and output parameters for infectious disease simulators. 23 It helps to judge whether simulators are simulating same scenario and increases the availability of epidemiological simulators in public health events.
BioCaster Ontology (BCO) is a multilingual ontology constructed by Collier et al. for a text-based system that automatically monitors network information of different languages and detects and track outbreaks and distribution of epidemics.24–26 Based on BCO, BioCaster Event Ontology is designed to understand and analyze reported epidemic-related events. 27
Ferreira et al. 28 have developed a Network of Epidemiology-Related Ontologies. It constitutes the core of semantic annotation for data-intensive epidemiological information systems such as epidemic prediction infrastructure. They also developed a platform called Epidemic Marketplace, which aimed to the preservation of epidemiological resources.
Lopez et al. 29 developed a spatial data model to predict the epidemiological impact of influenza in India. They analyzed geographic information, environmental data, population data, and epidemic data and found the correlation between rainfall, wind speed, temperature, humidity, and H1N1 influenza prevalence.
Thiery et al. published a Linked COVID-19 Data Ontology to map COVID-19 related datasets from Johns Hopkins University, European Centre for Disease Prevention and Control and Robert-Koch-Institut. 30 Little information on its OWL implementation is provided.
KG-COVID-19 provides a knowledge graph for COVID-19 and SARS-COV-2. 31 It’s designed to support the translation of COVID-19 and SARS-COV-2 datasets into a knowledge graph for machine learning.
The ROC ontology represents COVID-19 pandemic data. 32 It mainly focuses on the government responses to the pandemic and evaluates the effectiveness and side effects of government responses to COVID-19 in different countries.
González-Eras et al. 33 constructed an ontological engineering method to build COVID-19-related ontologies. They integrated existing COVID-19 ontologies such as CODO and COVIDCRFRAPID and then constructed an ontology called Tepuy-COVID. This ontology construction method achieved high precision when aligning ontologies from different sources. Precision is used in the information retrieval sense, where it is defined as the percentage of correct mappings, relative to manual methods.
Common patterns
A goal of this survey is to identify commonalities among the V/Os. The following identifies ontology patterns that are common across at least two V/Os.
Disease pattern
It appears in all but one of the reviewed V/Os. In some it is simply an identifier for a disease and in others it provides a taxonomy of disease, the symptoms and mechanisms of the disease and host, and pathogens that cause them. The scope and scale of the disease taxonomy are also different. Some focus on the taxonomy of infectious diseases, while some focus on the categories of all diseases. In some of the V/Os, the anatomical terms are defined to help describe the disease’s mechanism. Disease pattern is an essential component of an epidemic data model. Epidemic incidents are associated with at least one infectious disease.
Person pattern
It appears in all but two of the reviewed V/Os. Person is the participants and undertakers of activities. It appears mostly as an identifier for a person. A person’s characteristics that are useful in epidemic management are included in this patten. The health status of a patient and the itinerary of a passenger are also included. Some V/Os only define residents to clarify the people being surveyed. This pattern is used in all use cases.
Organism pattern
It appears in three of these V/Os. It defines those organisms that can be infected, such as non-human animals, plants, fungi. Organism pattern is related to two use cases that describe the infection and transmission of non-human species and microorganisms.
Epidemiology pattern
It appears in three of these V/Os. The spatiotemporal information and information involved in clusters and outbreaks are included. This pattern is used in tracing-related use cases to help monitor infected people and close contacts.
Organization pattern
It appears in half of these V/Os. Medical organizations, social organizations, and government organizations are important participants and stakeholders in epidemic management. Hospital is a subclass of organization that is the most common medical organization in epidemic management. In some V/Os, they only focus on hospital. Organizations may contain medical services, medical resources, and medical personnel. The Organization pattern is used in all use cases.
Medical personnel pattern
It appears in four V/Os. Medical personnel are the main workers in the prevention and control of the epidemic and perform the services defined in the organization pattern. It covers personnel licensing and tasks. This pattern is used in some medical activity-related use cases because medical personnel are usually providers of medical activities.
Medical activity pattern
It appears in four of these V/Os. Testing, tracing, vaccination, and diagnosis are medical activities. This pattern covers patient treatment, the measures taken to control the epidemic, the tracing and surveillance for the epidemic.
Medical resource pattern
It appears in three V/Os. Medical and chemical products such as vaccines and drugs that are effective to treat infectious disease are defined in this pattern. It is directly applied to medical resource management use case, and appears in some use cases that need to consume medical resources.
Infection transmission pattern
It appears in half of the V/Os. It represents different types of transmission modes of infection. Related vectors, transmission participants and processes are often included in this pattern. It is used in several use cases to model the transmission route of infection.
Statistics pattern
It appears in two V/Os. It focuses on statistics and indicators that used to measure the severity of epidemic, health status of patients, etc. It is applied to several use cases related to statistical indicators.
City pattern
It appears in two V/Os. It represents different levels of administrative areas such as district, city, and country to represent the range of the epidemics. It is applied to all use cases because all they must take place in a city.
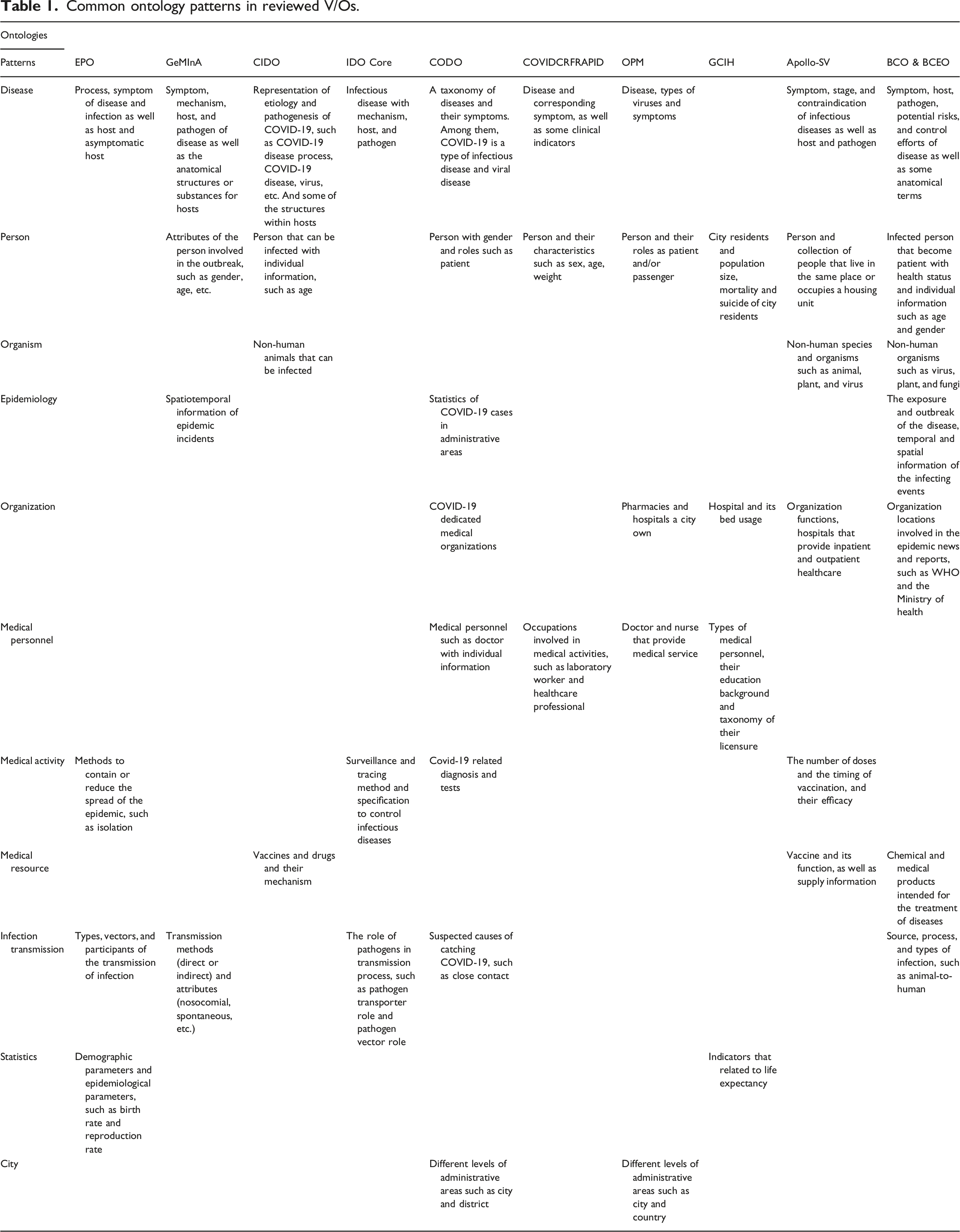
Common ontology patterns in reviewed V/Os.
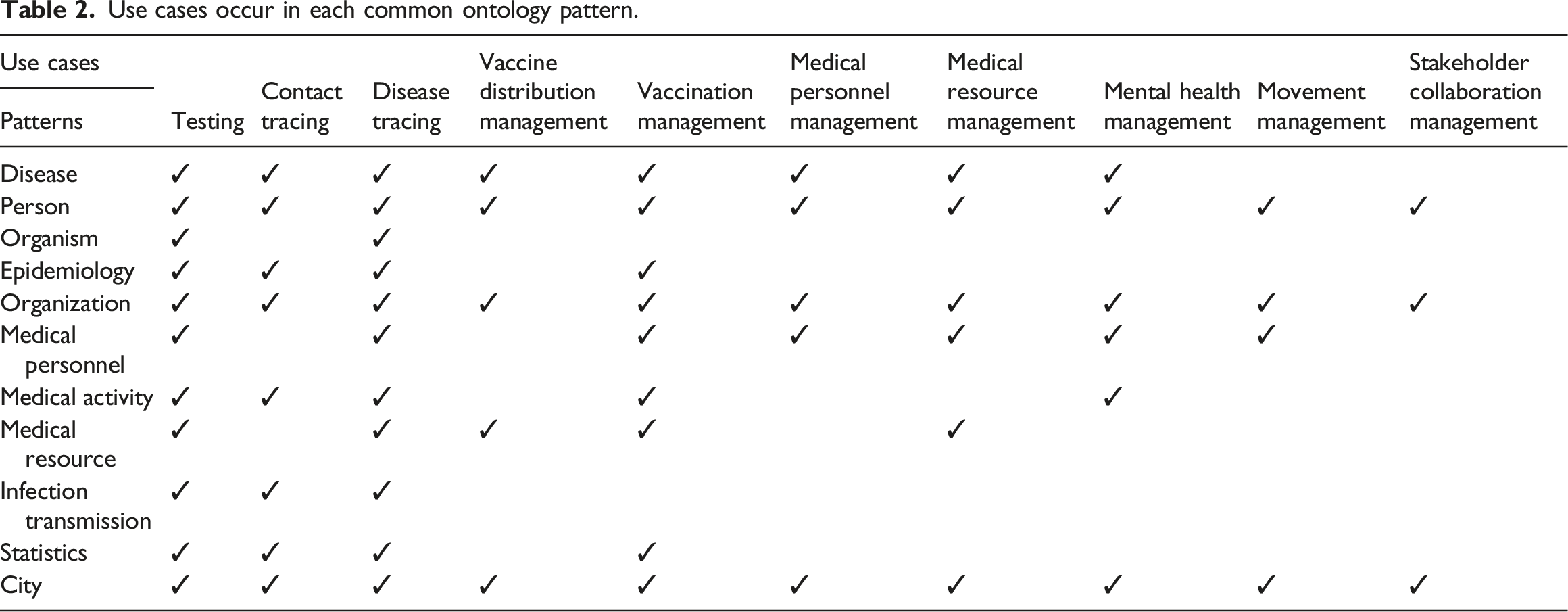
Use cases occur in each common ontology pattern.
What concepts are missing
In comparing the concept patterns described in Section
Conclusions
Over the last two decades there has been significant research into the development of vocabularies, and more recently ontologies, within the medical and health sciences. Much of this work has grown out of the need to share data within and across organizations. The COVID-19 pandemic has heightened the need for vocabularies and ontologies as the basis of data standards that enable sharing at national and international levels.
In our survey, we reviewed and summarized 16 existing epidemic V/Os. We compared them from the perspectives of use cases, core classes and properties, etc. The common patterns of each V/O and significant but missing concepts in epidemic use cases are also identified.
Our review demonstrates that existing V/Os provide significant coverage of the concepts underlying the epidemic use cases. Nevertheless, there remains gaps in the coverage. More work is required to represent concepts related to government organization; various types of people including visitors, volunteers, epidemic workers outside of the medical profession; medical infrastructure such as clinics; medical resources including tests, vials, and doses; and financial information. By expanding V/Os coverage of the concepts underlying public health emergencies use cases, knowledge related to epidemics can be more precisely and unambiguously defined and represented. It will enable city software applications to better share information, plan, coordinate, perform city tasks, and support decision making within and across city services.
Finally, in response to epidemics, Standards Development Organizations have developed guidance documents that define methods, processes, and checklists for how cities should respond. However, they also need to promote the development and publication of international standards for shared data models for epidemics to enhance interoperability of data generated in epidemics.
Footnotes
Acknowledgements
We thank Amanda Bell, who works for Toronto Public Health, for helpful comments. We also thank Zuohai Chen, Kun Yu, Wenqi Zhang and Hongli Lyu, for their great contribution.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by Ministry of Science and Technology of the People’s Republic of China under the High-End Foreign Expert Recruitment Plan [Project number G2022024004L] and the Department of Science and Technology of Shandong Province under the “Double-Hundred Talent Plan” on 100 Foreign Experts and 100 Expert Teams in 2020 [Project number WSG2020020].
