Abstract
A health record database contains structured data fields that identify the patient, such as patient ID, patient name, e-mail and phone number. These data are fairly easy to de-identify, that is, replace with other identifiers. However, these data also occur in fields with doctors’ free-text notes written in an abbreviated style that cannot be analyzed grammatically. If we replace a word that looks like a name, but isn’t, we degrade readability and medical correctness. If we fail to replace it when we should, we degrade confidentiality. We de-identified an existing Danish electronic health record database, ending up with 323,122 patient health records. We had to invent many methods for de-identifying potential identifiers in the free-text notes. The de-identified health records should be used with caution for statistical purposes because we removed health records that were so special that they couldn’t be de-identified. Furthermore, we distorted geography by replacing zip codes with random zip codes.
Introduction
Electronic health record (EHR) systems store large amounts of data and are essential for all clinical work. According to ANSI, 1 important qualities of an EHR are confidentiality and accessibility only by authorized persons. An EHR system must ensure confidentiality since exposing health records are against law and ethical principles. In order to create data for testing EHR systems, for presenting them to others and for teaching, access is needed to large amounts of EHR data, but it is hard to get the necessary permissions. Access to de-identified (anonymized) health records would in many cases be sufficient. However, the de-identified data should meet certain quality criteria:
Medical correctness: Each health record must show a true medical picture of a real patient.
Anonymity: It must not be possible to see who the real patient is.
Readability: The health record must look real. As an example, patient names and addresses that have become F274, XXXX or **** don’t meet this criterion.
Consistency: The patient’s identifiers must be consistent with the medical picture, for instance, an age that is like the real person’s age. If the real patient’s name is Peter, but his de-identified name is Jens, then Peter must be replaced by Jens also in the clinician’s free-text notes. Furthermore, his wife’s health record may refer to him as Peter, and this Peter must also be changed to Jens.
A health record database contains fields with structured data that can identify the patient, such as patient ID, patient name and phone number. These data are fairly easy to replace with other identifiers, in that way ensuring anonymity. The database also contains fields with medical data such as diagnosis codes, blood pressure and other measured values. They have to be preserved to ensure medical correctness. The problem is the unstructured data fields with doctors’ free-text notes. They contain important medical information that has to be preserved, but may also contain phone numbers, patient names, name of the spouse and other identifying items. Furthermore, clinicians write notes in an abbreviated—often personal—style that cannot be analyzed grammatically.
Considerable work2–12 has been done in developing de-identification algorithms using various techniques such as natural language processing (NLP), named entity recognition and machine learning. These approaches de-identify database records (e.g. pathology documents) that do not relate to other records. We will refer to this as a record-oriented de-identification approach. Recent work13–15 has focused on utilizing a full database rather than records. The approach presented in this article was briefly presented by Pantazos et al. 16
Previous research has not looked at quality attributes for database-oriented de-identifications. In this article, we focus on the four quality attributes above: medical correctness, anonymity, readability and consistency.
Background
In 2010, we started work on an EHR system with a high degree of data visualization. We cooperated with a Danish software house that had delivered EHR systems to many clinics and small hospitals in Denmark. In order to get test data, we made a copy of the full database and de-identified it. The database consisted of 437,164 patient health records. The work took place on their premises since no real health records could go outside the company.
The idea was to make a mapping table that translated all patient identifiers into patient identifiers for other patients. In this way, patient B got patient C’s first name, patient D’s last name, patient E’s street name and so on. Patient B would get a randomized civil registration number (CPR) that preserved his year of birth and gender. In this way, we would ensure consistency across patients. Somebody looking at a full de-identified patient record would know that this was a real patient, but he or she was not called C, nor D and didn’t have address E. We had outlined the conversion program and expected the whole thing to take a couple of days, but—alas—unexpected problems turned up. We spent 3 months.
We had to invent many methods for locating and anonymizing potential identifiers in the free-text notes. To our surprise, 3–4 percent of the words in free-text fields were potential patient identifiers. Consistency across patients turned up to be more important than we had expected: around 90 percent of the patients had one or more relatives in the database.
We detected that many health records had been created for testing or were left uncompleted due to system errors (“corrupt” data). In 69,914 cases, we had to delete such patient records. In 43,119 cases, data couldn’t be safely de-identified without manual intervention, so we made the program delete these patient records. As an example, we deleted patients that had a rare Danish name that was also the name of a disease, for instance, Aaron, which is also a medical term. The program couldn’t tell whether Aaron in a free text was a medical term that shouldn’t be changed or the name of a related patient that should. We ended up with 323,122 patient health records.
We manually compared 369 random, anonymized patient records against the original records, checking for medical correctness, readability and anonymity. These quality factors were preserved on an acceptable level for our purpose. Consistency was ensured by the algorithm, so we checked it in a few places only.
Related work
A good de-identification system must replace all data that are personal identifiers in structured data, as well as in free text. 5 One of the first de-identification systems for patient records was Scrub. 2 It was evaluated against 275 English patient records and 3198 letters to physicians from the pediatric department. External sources, predefined templates and rules (e.g. the format of a phone number and address) were included in the algorithm. This algorithm had a 99–100 percent success rate for de-identifying personal identifiers. Another system was developed by Ruch et al. 3 It resembles Scrub, but added NLP. NLP tools use a medical semantic dictionary with word-sense and morph-syntactic labeling. This system located 98–99 percent of all personal identifiers. To anonymize the data, the authors replaced all identifiers with XXX’s, which had a negative impact on readability.
Several systems4–6,17 were developed in the last decade to de-identify pathology reports. Thomas et al. 4 developed an algorithm that scored 98.7 percent successful name replacements using English syntactic rules, prefixes, suffixes and names composed of first and last names. Gupta et al. 6 conducted an iterative evaluation of their system. At the end of the third iteration, the authors claimed that their method generated anonymized and readable reports. An algorithm designed by Berman 5 replaced words with codes from the Unified Medical Language System (UMLS) and asterisks. It produced hardly-readable documents. Beckwith et al. 17 evaluated an open source system which replaced identifiers with X’s in pathology reports.
Seven de-identification systems were evaluated in the “Challenge in NLP for clinical data” workshop, using medical discharge letters as input. 18 In this workshop, the systems were evaluated using three performance measures: precision, recall and f-measure. The highest f-measure was 99.7534 achieved by a novel approach based on Named Entity Recognition combined with iterative machine learning. 8 This application finds personal identifiers in the structured data and uses them to locate identifiers in free-text data.
Hanauer et al. 12 introduced the iterative tag-a-little, learn-a-little approach for a particular document type. The authors used the MITRE Identification Scrubber Toolkit 19 to integrate their approach. They obtained an f-measure of 95.
Susilo and Win 20 present a new approach for patient confidentiality that utilizes searching through encrypted data. Huang et al. 21 focused on portable EHRs for privacy preservation. The authors stress the feasibility of the approach, which can meet patient confidentiality requirements.
Even though most of the research has been in an English context, there are some studies on de-identifying in other contexts. Tveit et al. 22 present their approach to de-identify Norwegian general practitioner medical records. Their approach consists of six steps: create dictionaries, find exact match and tag, identify approximate match and tag, replace tags, tackle untagged words and generate the de-identified output. However, this approach was not evaluated empirically. A Swedish de-identification system was developed by Kokkinakis and Thurin, 7 using named entity recognition. This approach de-identified 200 Swedish discharge letters with a precision of 96.97 percent, recall of 89.35 percent and f-measure of 93. Velupillai et al. 10 adjusted an English de-identification system for Swedish medical records. This transformation did not produce the expected results (f-measure in total = 65, f-measure for names = 80). Consequently, the authors reported that building a Swedish system from scratch was more efficient. This phenomenon was also observed and confirmed by Grouin et al. 9 who adjusted a de-identification system from English to French and obtained poor results.
Meystre et al. 23 reviewed recent de-identification algorithms and found that the majority of the algorithms focus on de-identifying structured data and not free text. However, in accordance with Dalianis and Velupillai 24 and Hanauer et al., 12 there is immense valuable information in the free text. We found the same in our data.
Quality factors
An EHR contains database fields with structured data that can identify the patient, for instance, CPR, patient name and phone number. Other structured data fields contain medical data such as diagnosis codes and blood pressure. The EHR also contains free-text fields, for instance, doctor’s notes and discharge letters. It may also contain pictures of body parts, X-ray and so on, usually with a patient ID embedded in the picture. We have not dealt with pictures in this project.
Some data are quasi-identifiers because they can narrow down the set of patients that might have this health record. Examples are street name, zip code, birth date, hospital or clinician who treated the patient. Two or more quasi-identifiers in combination may identify the patient. 25
Anonymity
In order to ensure anonymity, all patient identifiers and quasi-identifiers must be de-identified, that is, replaced with something else. It is fairly easy to do this for structured data, but very hard for free-text data. Often the computer has no way to tell whether a free-text word is an everyday word, a medical term or part of a patient name. As an example, Aaron’s sign is a medical term, but it might also be the name of a person.
Readability
In order to ensure readability, we have to replace the patient name with a new name that looks real. Inside this patient’s record, we have to be consistent so that we replace with the same name for all occurrences.
Consistency
In the database we worked with, 90 percent of the patients had one or more relatives in the database. Most likely, the patient’s name and/or CPR will occur in one of these related health records in free-text fields. To ensure consistency, we have to replace also these identifiers with the same new identifier.
There are other aspects of consistency, for instance, that the distribution of names should remain much the same. If rare names suddenly turn up for a large number of patients, the health record database will not look real.
Medical correctness
If we replace a medical term that looks like a person name, with the new person name, the health record will look odd. We have lost medical correctness and readability. In many cases, a clinician can guess what the medical term was and in that way get to know the original name of all patients that have this new name.
Another aspect of medical correctness is age. If birth dates are transformed in a way that makes the patient have a very different age, it will not match the patient’s diagnosis pattern.
Solution
We will first give an overview of the solution and then explain the details and where the data came from.
Permutation tables
For some identifiers, we made a permutation table that mapped existing identifiers to new ones. We picked the new identifier at random from the same table, avoiding reuse of identifiers. Any occurrence of an identifier from this table would be translated into the new one.
As an example, we created a permutation table of all last names. The last name Jensen would be translated into Petersen wherever it occurred. Petersen was another last name in the table, with a similar frequency. This ensured readability and consistency across all patient records.
We made permutation tables for these identifiers and quasi-identifiers: first male names, first female names, last names, street names, zip codes, hospital and clinic names.
Distorted identifier table
For the CPR, we made a mapping table from existing CPR to a distorted CPR in this way: The Danish CPR format is: DDMMYY-CSSG where DDMMYY is the birth date. The day (DD) and month (MM) were changed to a random valid day and month. The year (YY) was kept. C indicates birth century (1900 or 2000). This was not changed. SS (serial number) was changed, while G indicates the gender and wasn’t changed.
This ensured readability (clinical users see lots of CPR numbers and can easily spot wrong ones) and medical correctness (because age and gender were kept).
Randomized identifier
For other identifiers, we randomized the identifier without caring about readability or consistency. This applied to phone numbers, e-mail addresses and URLs.
Ambiguous words
Ambiguous words could be part of a person’s name or something else, for instance, a medical term or a common word. Through many sources, we created a list of ambiguous words. When the de-identification program meets an ambiguous word B in a free-text field, it has three choices:
Replace the word B with its corresponding new name, C. If the word B actually is part of a person’s name, everything is fine. But if B actually is a medical term or a common word, the clinician can see from the context that C probably means B. If he knows the replacement rules, he now knows that everybody in the database with the name C is actually B. We lose not only medical correctness and readability but also some anonymity. For this reason, we never replace ambiguous words.
Keep the old word B. This ensures medical correctness, readability and consistency. If the word actually is a person’s name, the clinician can see it from the context. As an example, assume that the program finds Aaron in a free-text note. Since it is a medical term, it keeps it. However, a clinician can see that Aaron in this context is the name of a person. If he knows the rule of replacement, he now knows that the person referred to is really called Aaron, although this is not his name in the de-identified database. The clinician gets no clue to where Aaron’s health record is. If there are only a few Aarons in real life, he might guess whom it is. If there are many Aarons, he cannot know. We decided that 200 occurrences was a safe limit. If the ambiguous name occurs more than 200 times, we keep it in the database.
Delete all patient records with name B. We do this when the ambiguous name occurs less than 200 times. This ensures all four quality factors, but we lose data. If a free text for another patient refers to patient B, the reference will now be to a deleted patient. We have lost a bit of consistency, but such data could exist anyway in the database.
The database and the mapping tables
The EHR we de-identified is built on Microsoft Axapta, which is an ERP system that can be extended in many ways. It contained data from 79 clinics and hospitals (including a few in Greenland and the Faroe Islands) and contained 437,164 patient records in total. The entire database was 12 GB. There were 65 health-related tables:
43 tables had no fields that could expose the patient identity. They included reference tables of drug codes, treatment codes and diagnosis codes.
9 tables had fields that only contained personal identifiers in structured form, for instance, the patient table that contained patient ID, first name, last name, address, zip code, five phone numbers, birth date and date of death. Another example is a table of family relations, that is, relations between two patients. Clinicians, hospitals and clinics had their own tables with name, address and so on.
13 tables had fields with free text. The largest one was Medical Record Lines, which occupied 7 of the 12 GB in the database.
Mapping tables
To be able to replace existing identifiers with new identifiers, we created the following mapping tables.
CPR
We collected the CPR numbers from the patient table and gave each number a partially random new number according to the rules above. If the new number was already used as a new number, we randomized it once more.
Last names
We used three sources to collect last names: the database’s patient table, Danmarks Statistik’s website 26 and a study of Danish names at University of Copenhagen, 2005. 27 We merged these sources and obtained 56,339 last names. We counted how often each last name occurred in the patient table. Many names didn’t occur at all in the patient table, but might occur in free-text notes. It was important to catch them too and de-identify them.
For each name, we assigned a new name from the table with a frequency similar to the old name. We used this approach: we divided the names into groups according to frequency. The group of most frequent last names consisted of 20 names with frequencies from 14,712 to 5319. We rotated these names a random number of steps to obtain the new names (a cyclical permutation). We used the same approach for groups of 30 names with decreasing frequencies. Rare names (frequency < 200) were randomly replaced with another name in the frequent part of the list. This also took care of the names that didn’t occur at all in the patient table.
Male first names
We used the same approach to collect and de-identify male first names. For names occurring in the patient table, we got the gender from the CPR number. Our external sources had separate lists for male and female first names. In total, we got 11,415 male first names.
Female first names
We treated them in the same way as male names. In total, we got 13,044 female first names.
Street names
We collected street names from the patient table’s address field. The address field included also floor numbers and entrance letters. In Denmark, the street name is first, so we simply extracted the first real name from the address field. We also included street names from the CPR website. In total, we got 25,429 street names. We assigned a random street name as the new name without caring about frequencies or consistency with zip codes.
Zip codes
We collected zip codes and related city names from Post Danmark 28 and assigned a random zip code and city name as the new name. In total, we got 1396 zip codes.
Hospital names and clinic names
We collected hospital names from Region Hovedstaden, Region Sjælland, Region Syddanmark, Region Midtjylland, Region Nordjylland, Queen Ingrid’s Hospital in Greenland, Faroe Islands website and our own EHR Database. We used Sygehusvalg, 29 Brancheforeningen for Privathospitaler og Klinikker (the trade association) and our own EHR database to extract names of clinics. In total, we got a list of 219 clinic names and 93 hospital names. We did not randomly assign new names to the clinics and hospitals. This would reduce medical correctness because clinicians know which clinics do what. On the other hand, being treated in a specific clinic is a quasi-identifier. We manually selected 41 hospital names and 92 clinic names and used them as new names. In many cases, the new name was simply “Hospital” or “Clinic.” This was a reduction in readability and to some extent in medical correctness.
Ambiguous names
Ambiguous names in our context are first or last names of persons that happen to mean something else too. We need a table of them to decide how to treat such a name when it occurs in free text. As explained above, we have to delete patients with rare names if they appear in free text. If they are frequent names, we leave them as they are.
In healthcare, it is common that diseases, signs, symptoms and so forth are named after a person, most likely the one who discovered it. These names are called medical eponyms and may cause ambiguity. For example, according to Statistics Denmark in 2010, 26 there were 88 males using the name Aaron. At the same time, Aaron is part of a medical eponym (Aaron sign). The algorithm knows too little about the context to decide whether to de-identify this name or not.
A similar ambiguity exists also with common words in a language. Each language contains several words whose meaning depends on the context. For example, in Danish, the word “hans” can be a pronoun or a male name. Another ambiguous case is abbreviations used by clinicians. For instance, instead of writing “kirurgisk” (in English: “surgical”) they use the abbreviation “kir,” which can be a last name as well. As another example, it is common that a city, hospital, clinic or street name is used as a first or last name. For instance, Aalborg is a city in Denmark, but it is a last name as well.
We derived the table of ambiguous names from several sources. We checked our lists of first and last names against the Danish Dictionary from Microsoft Office Word 2010. This created a list of 3557 potentially ambiguous names. That a name exists in the dictionary doesn’t mean that it also has another meaning that can occur in health records. So, the medical specialist in our team (Lippert) scrutinized the list and came up with 1952 ambiguous names.
To the best of our knowledge, there is no official source that contains medical eponyms. So, we used the website “Who named it” 30 and extracted 3246 medical eponymous names from it. They too entered the list of ambiguous names.
Applying the mappings
The mappings must be applied to the structured fields as well as to the free-text fields. We applied the mappings to the structured fields according to Table 1. Notice the last rule: remove all patients above 90 years. It came from the US Health Insurance Portability and Accountability Act guidelines, HIPAA. 31 There are so few patients above 90 that their age exposes them.
Replacement rules for structured data fields.
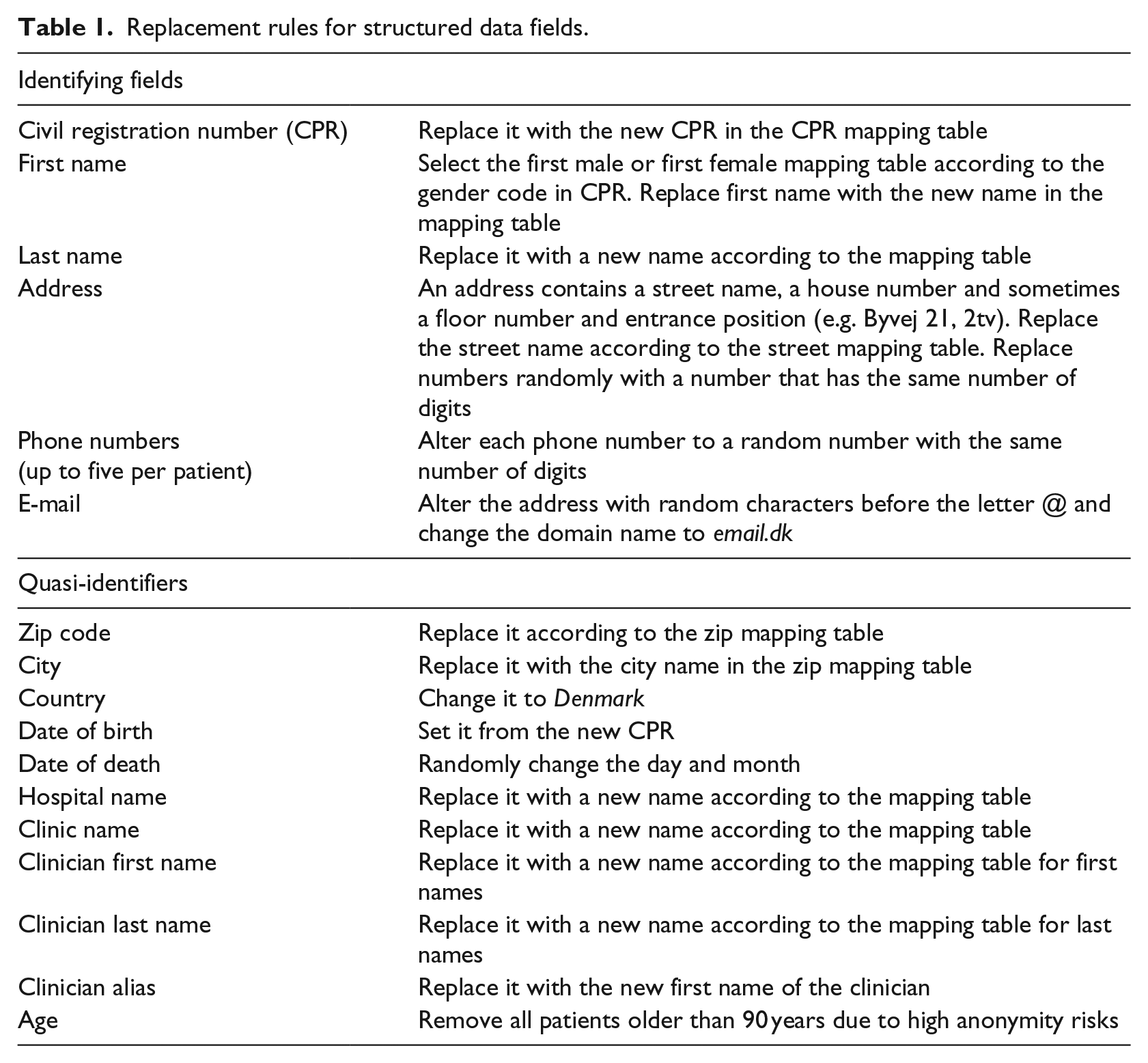
It was harder to apply the mappings in free-text fields because we don’t know whether an identifier is a first name, a street name and so on. The program analyzed the free-text token by token and applied these rules.
Name tokens
1. If the name is in one of the person name tables and also in the table of ambiguous words, do nothing or delete the related patient records depending on the name frequency.
2. If the name is in the last name table, replace it with the new name in the table.
3. If the name is in the first male name table, replace it with the new name.
4. If the name is in the first female name table, replace it with the new name.
5. If the name is in the table of ambiguous words, leave it as it is.
6. If the name is in the table of street names, replace it with the new name.
7. If the name is in the table of zip codes and city names, replace it with the new city name.
8. If the name is in the table of hospitals and clinics, replace it with the new name.
9. Otherwise, leave it as it is.
Number tokens
10. If the number is in the table of CPR numbers, replace it with the new CPR number.
11. If the number looks like a CPR number (10 digits starting with a date), randomize it as other CPR numbers.
12. If the number has eight digits and is next to a word like tlf, tel and fax, randomize the number.
13. If the number is in the table of zip codes and next to a city name, replace it with the new zip code.
14. Otherwise, leave it as it is. (It may be a measured value, a lab-test number (eight digits), a house number and so on.)
These rules give priority to anonymity rather than medical correctness. As an example, a lab-test number or a date-time that looks like a CPR number will be de-identified and thus reduce the medical correctness.
Evaluation of the quality factors
Anonymity, readability and medical correctness
In order to evaluate the actual anonymity, readability and medical correctness, we need to know how many words were replaced incorrectly.
We selected a random sample of 369 full patient records. A clinician manually compared all the free-text fields in the old and the new version, in total 73,150 words. The result is shown in Table 2.
Correct and incorrect replacements.

Seven words should have been de-identified but wasn’t. Only one of them was a person name. It was ambiguous and frequent (frequency >200) and consequently preserved according to our rules. Since it was frequent, we consider it a quasi-identifier. The other words were quasi-identifiers such as department names and misspelled street names that were not in our translation tables. In total, out of 73,150 words, we had seven anonymity leaks on quasi-identifiers and none on full identifiers.
A total of 109 words were replaced, but shouldn’t. They were ambiguous, but not in our table of ambiguous words. One example was the word “Uno,” which was the name of a drug, but also a male first name. These cases decreased medical correctness and readability. It also revealed a general weakness: also drug names should be considered a source of ambiguity.
Measured in the traditional way with recall and precision, the algorithm scored 99.5 percent for recall (the seven anonymity leaks) and 92.3 percent for precision (the 109 leaks in medical correctness). The f-measure was 95.7 percent. Our database-oriented approach compares favorably with previous work on record-based de-identification approaches.4–6,17
It would be interesting to compare how other de-identification approaches would handle our data. However, this is impossible because the approaches are very dependent on the language. Furthermore, we are not allowed to move our original data out of the company where it is hosted. We have not found publications about de-identification that discuss ambiguity. Most likely, they don’t pay attention to it. It will probably cause some leaks of confidentiality that isn’t detected.
Consistency
The database can record family relations and other relations between patients. Around 90 percent of the patients have one or more recorded relatives. When a person name is de-identified in the structured patient table, it is important that the same name is de-identified in the same way in the rest of the patient’s records and in records of related patients, also for free-text fields. This is solely a matter of correct programming. We checked it for a couple of patients in Table 3. Since the translation tables are used for all patients, consistency is also preserved for relatives who are not recorded as relatives.
Sample of 20 patients showing the variety of patient records.
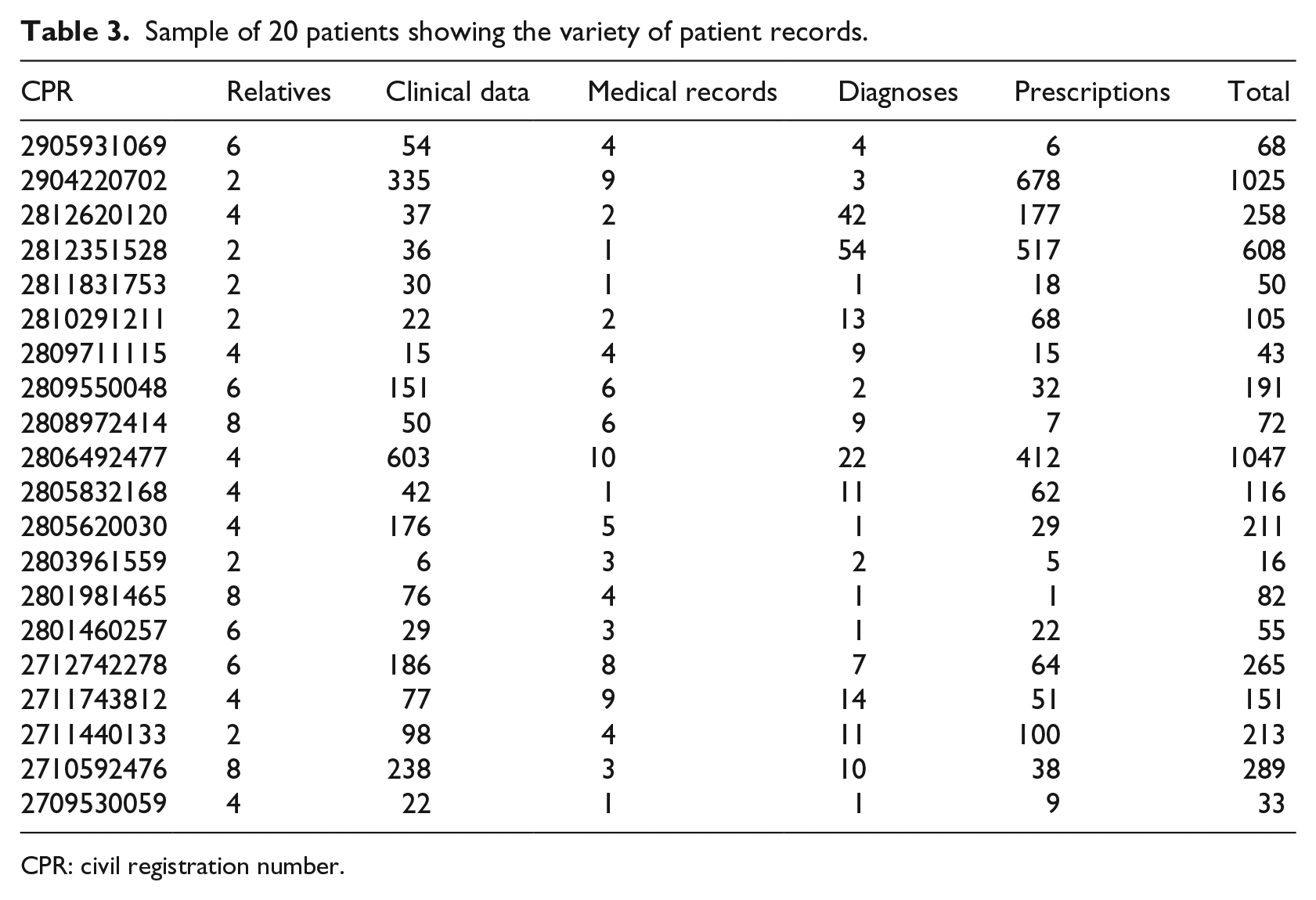
CPR: civil registration number.
Results
Table 3 shows a (non-random) sample of patients with two or more relatives. It gives an impression of the variety and complexity of patient records. Several patients have eight relatives in the database, many have more than 100 measurements with notes (Clinical Data), many have more than 10 diagnoses and several hundred prescriptions.
Table 4 shows the results for the Medical Record Line and Clinical Data tables, which contain most of the free text in the database. In total, 3–4 percent of the words are personal identifiers.
Number of identifiers in free text.
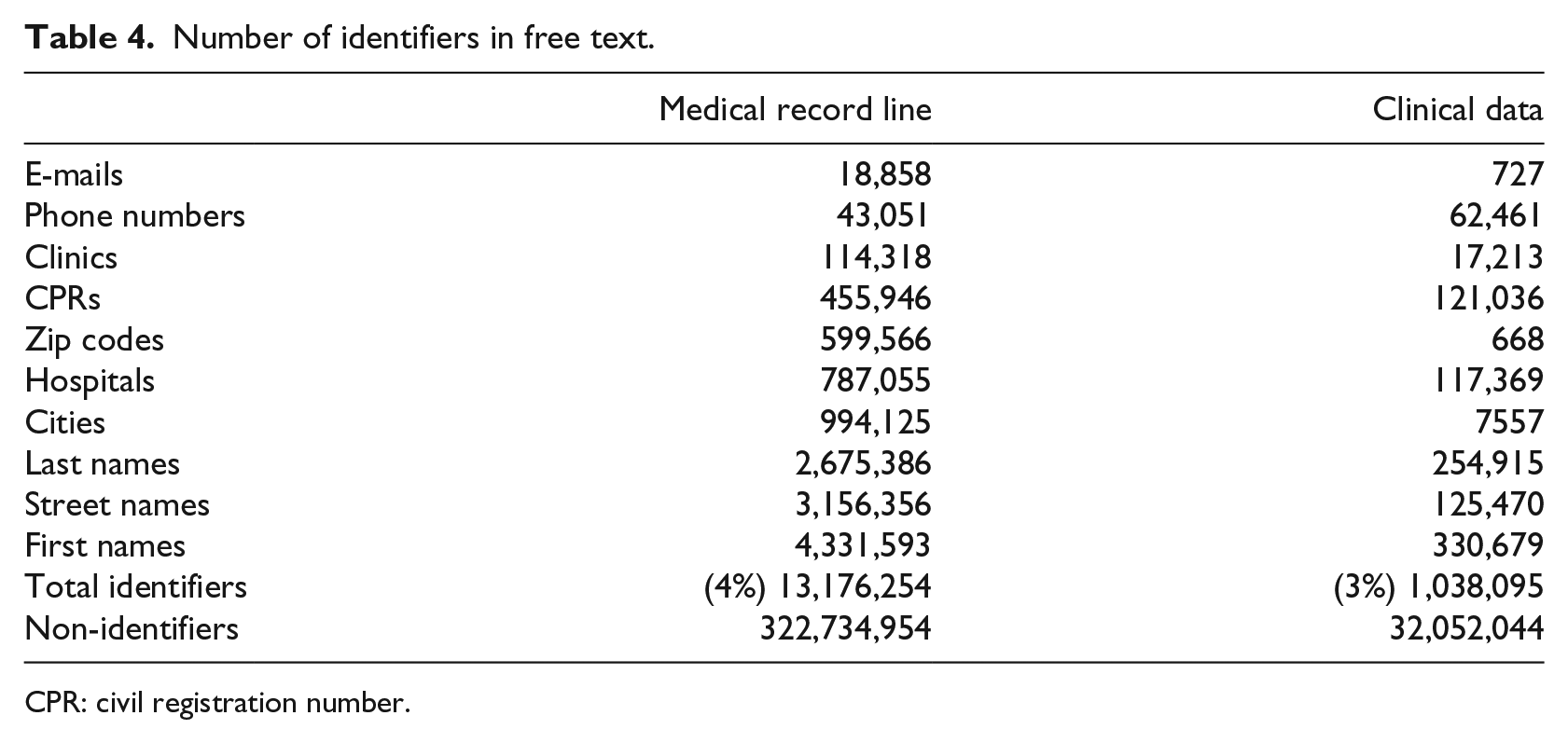
CPR: civil registration number.
This study is the first de-identification algorithm that focuses on anonymity, medical correctness, readability and consistency. Other approaches are limited to a few types of documents, while our approach deals with full EHR records from 79 hospitals and clinics. An important part of our approach was to collect ambiguous names from many sources.
We started out with 437,164 patient health records. We deleted 69,914 patient records because data were corrupted (old test data and records left after system failures). We deleted 43,119 patient records because of rare ambiguous names or because the patient was older than 90. We ended up with 323,122 patient health records.
The distinction between frequent and rare names (fewer than 200 occurrences) is somewhat arbitrary. The limit of 200 caused us to delete “only” 43,119 patient records because they had rare ambiguous names. If all names were considered rare, we would have lost another 55,000 patient records.
We made a manual review of 369 patient records with 71,721 free-text words. It revealed seven words where a quasi-identifier hadn’t been de-identified. It revealed 109 words where it was de-identified, but shouldn’t because the word wasn’t in our list of ambiguous names. This reduced medical correctness and readability.
Limitations and errors
An EHR database contains also binary files (e.g. X-rays), scanned documents and Word documents. They are not part of the database, but fields in the database contain the file names. Our approach is limited to structured data and free-text fields, and it doesn’t try to de-identify pictures and other files. The picture will usually contain patient identifiers such as CPR and name. De-identifying these would be a project of its own.
We have not tried to deal with spelling errors. It might have reduced the seven un-identified words above to around three. We could deal with spelling errors by looking at close matches of words instead of precise matches, but we don’t know how much it would have increased the number of false de-identifications (the 109 words above).
We forgot to put also pharmaceutical names in the list of ambiguous words. This could have removed some of the 109 false de-identifications above.
We missed several clinical abbreviations as potential ambiguous names. A language analysis of the free-text notes might have revealed them.
The de-identified data should be used with caution for statistical purposes because of the way we had to remove health records that couldn’t be de-identified and also because we deleted patients older than 90 and distorted geography by replacing zip codes with random zip codes.
For statistical purposes, the de-identification should have been different. We shouldn’t care about readability or consistency, but simply replace all potential identifiers in free text with asterisks or the like. We should only delete corrupted patient records. The mapping tables would still be needed, but only to detect what might be an identifier. We wouldn’t need to care about ambiguous words. The result would probably be similar to many other de-identification approaches.
Footnotes
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
