Abstract
Background
Biomarkers of amyloidopathy/tauopathy/neurodegeneration (A/T/N) are being invoked as evidence of Alzheimer's disease (AD). However, some individuals are resilient against their effects. We have developed a psychometric algorithm to distinguish resilient from afflicted persons, and have demonstrated it in relation to inflammation, adipokines and amyloidopathy.
Objective
This analysis addresses neurodegeneration.
Methods
1737 participants in the Alzheimer's Disease Neuroimaging Initiative (ADNI) were assigned to neurodegeneration affliction classes through a “Line of Identity (LOI)” approach. A neurodegeneration “N Factor” indicated by three biomarkers was constructed. A previously validated measure of dementia severity, "dTEL″, was regressed onto the N Factor. Affliction classes were defined by each subject's deviation from the LOI obtained from the correlation of dTEL's N-Factor-adjusted residual (CR) versus unadjusted dTEL scores. Moderation effects on the N Factor's association with dTEL were tested by Chi Square difference. Class effects on prospective conversion to clinical “AD” from non-demented baseline diagnoses (NC + MCI) were tested by survival analysis.
Results
49.4% of ADNI subjects were afflicted by neurodegeneration. The Afflicted class had greater dementia severity, lower (adverse) N factor composite scores and higher observed levels of CNS neurodegeneration. These differences reflected the unique effect of neurodegeneration. Affliction class was not associated with other AD-related biomarkers. Afflicted cases were more likely to convert to clinical “AD” over 48 months [by Cox's F: F (164, 356) = 3.65, p < 0.001.
Conclusions
Our approach could allow for more accurate prediction of biomarker effects and guide precision interventions.
Introduction
Efforts are being made to diagnose Alzheimer's disease (AD) by biomarkers alone. The recently proposed “A/T/N” scheme 1 aims to make that diagnosis on the basis of amyloidopathy (A), tauopathy (T), and neurodegeneration (N). While these are generally present at autopsy in “AD” cases, obstacles remain to the scheme's implementation in vivo. First, central nervous system (CNS) pathology is not being measured, but is being estimated via more easily acquired measures and biomarkers. Second, while multiple biomarkers are considered acceptable proxies for A, T and N domains, 2 it is not clear that they have convergent validity and they are often discordant. One might expect that valid indicators of those domains would share variance with each other. That could be tested empirically through factor analysis, but such studies are rare. 3 Instead, individual biomarkers are often used interchangeably as proxies for A/T/N domains, either as individual measures 4 or as indices constructed from several. 5 This can lead to wide variations in diagnostic accuracy depending on the exact set of features selected. 4
A more significant limitation may be the existence of cases that are resilient against the purported adverse effect of a biomarker of interest. 6 The mere demonstration of a biomarker's altered level may not be sufficient for prognosis. Finally, “resilience” is often determined by a “residuals” approach, which may have methodological weaknesses. 7 In that approach, a measure of cognitive function is regressed onto a biomarker. If a case is found to have better cognition than their biomarker value would predict, they are considered to be “resilient”. One major limitation to that interpretation is that dementia may be “overdetermined”, i.e., determined by the summed effects of multiple competing and relatively independent dementia-related processes. The variability in observed outcomes may have more to do with unmodeled comorbid processes than the biomarker of interest.
This situation is depicted in Figure 1. The totality of dementia's variance may be explained by the additive effects of multiple modeled and unmodeled (e.g., “et alia”) biomarkers and /or risk factors. The challenge is to isolate the dementing impact of any desired single biomarker and not to conflate “affliction” or “resilience” by that specific biomarker with the influence of other modeled and /or unmodeled processes. A further challenge is make this determination in an individual. Then, interventions targeting a biomarker of interest could be directed towards the subset individuals who are afflicted by it (i.e., personalized medicine).
Dementia severity is determined by multiple biomarkers. APOE4: the number of apolipoprotein ε4 alleles; Abeta: amyloid beta regional standardized uptake values (SUVr) by positron emission tomography: N Factor: neurodegeneration. This cartoon is not generated from data. The relative contribution of each biomarker is arbitrarily assigned.
This undertaking has many potential advantages. In addition to selecting individuals for treatment, we could more accurately interpret the significance of a biomarker's detection, say a positive amyloid-β (Aβ) positron emission tomography (PET) scan. 8 We could divide patients into afflicted and resilient subsets and model the outcomes of each class. The mechanisms of resilience could be investigated via cross-group differences in the afflicted and resilient classes. Treatments could then be proposed to change class membership to a more favorable group, rather than bluntly trying to alter the biomarker of interest.
Figure 1 also points to a potential weakness of the diagnosis of resilience by residuals. It shows that the variance in dementia severity residual to the effect of any one biomarker is likely to be determined by many competing modeled and unmodeled processes. What we should be seeking is a determination of resilience to the specific effect of the biomarker of interest. Because individual biomarkers are likely to explain only a fraction of dementia severity, the true impact of resilience may be more circumspect than is widely reported. The current approach risks to conflate biomarker-specific resilience with variations in competing dementia risks, and could easily overestimate a biomarker's impact, and therefore the true risks and benefits of intervention on it.
We have developed an algorithm that may achieve some of these goals. Instead of defining “resilience” relative to a point estimate, we define it relative to the line of identity (LOI) in a scatterplot of biomarker adjusted- versus unadjusted dementia severity.8–10 If a biomarker has no effect, then a case will present along the LOI. When cases present off the LOI, the biomarker has demonstrably impacted dementia severity. If the biomarker-adjusted “cognitive residual” (CR) value is better than observed, the biomarker has adversely impacted the subject and they are “afflicted” by it. If the CR is found to be worse than observed, the same biomarker has somehow imparted a beneficial effect and they are “resilient”. Since only the difference between observed and CR scores is used to assign cases, cross-class differences are specific to the biomarker of interest. Affliction class categorizes the effect, but the difference between CR and observed performance provides a continuous measure of the biomarker's impact. By defining “resilience” relative to the LOI instead of a point estimate derived by regression, we can be confident that we are not conflating biomarker-independent with biomarker-specific effects.
As a further refinement, we have been using the dementia-specific phenotype, “δ” as the outcome of interest. δ is a latent variable representing the shared variance between a cognitive battery and a measure of Instrumental Activities of Daily Living (IADL). It is derived from Spearman's general intelligence factor “g” 11 via bifactor analysis in a Structural Equation Model (SEM) framework. 12 δ is strongly associated with clinician-rated dementia severity and consistently achieves high areas under the receiver operating characteristic curve (AUC /ROC) for dementia's diagnosis. Regardless, δ is agnostic to etiology and cannot distinguish any two dementing conditions.13,14 This suggests δ is dementia's essential phenotype whereas disease-specific cognitive changes cannot be.
As it is derived from g, δ is “indifferent” to its indicators and can be calculated from an almost unlimited variety of psychometric measures. So many potential batteries are available that we must distinguish each instance as a specific δ “homolog”. In genetics, a homolog is a gene descended from an ancestral gene in the same species which retains the original's function. All δ homologs are strongly associated with the CDR “Sum of Boxes” 15 and achieve high AUCs for AD's diagnosis, e.g.,.16,17
Two latent biomarker constructs, INFLAMMATION 18 and Adipokines, 19 have been submitted to our algorithm.9,10 Adipokines are thought to engender adverse effects in multiple organs, including brain. 20 Our construct (i.e., “Adipokines”) was indicated by eight plasma proteins. It exhibited a dose-dependent effect on clinical diagnoses and was significantly associated with dementia severity. 19 Similarly, inflammation was estimated by a latent variable (i.e., INFLAMMATION) indicated by multiple biomarkers. The reified INFLAMMATION score was associated with clinician-rated dementia severity, across cohorts and biofluids. 18
We have also submitted positron emission tomography (PET) imaged amyloidopathy to our algorithm. 8 39.3% of participants with (+) PET results in the Alzheimer's Disease Neuroimaging Initiative (ADNI) were assigned to the afflicted class. Afflicted PET (+) subjects had greater dementia severity, higher amyloid burdens, and were more likely to convert to clinical AD over 48 months. Moreover, the association between amyloid burden and dementia severity was moderated by affliction class. PET assessed amyloid burden was not related to dementia severity in resilient cases, i.e., the majority of PET (+) cases.
Those findings suggest significant limitations to the A/T/N project. The present analysis extends our approach to a second A/T/N domain [i.e., neurodegeneration (N)]. We hypothesize that in contrast to the neurodegeneration-afflicted group, resilient cases will have less adverse levels of neurodegeneration, lesser levels of dementia severity and a lower risk of prospective conversion to clinical AD from non-demented states. If so, then this would have significant implications for the A/T/N project as it would confirm cases in a second A/T/N domain who are resilient to the effects of a proposed AD-specific biomarker.
Methods
Subjects
ADNI. Briefly, this is a secondary analysis of ADNI data. 21 ADNI is a well-characterized longitudinal convenience sample developed to validate the magnetic resonance imaging (MRI), positron emission tomography (PET), cerebrospinal fluid (CSF), and genetic biomarkers of AD. It was launched in 2003 as a public-private partnership. Its original goal was to test whether serial MRI, PET, other biological markers, and clinical and neuropsychological assessment could be combined to measure the progression of mild cognitive impairment (MCI) and early AD. ADNI's current goals include validating biomarkers for clinical trials, improving the generalizability of ADNI data, and to provide data concerning the diagnosis and progression of AD to the scientific community.
Clinical variables
CDR-SB. The Clinical Dementia Rating Scale (CDR) 22 assesses a patient's abilities in six cognitive domains. Domain-specific scores are summed to provide the CDR “Sum-of-Boxes” (CDR-SB) with improved psychometric characteristics. 15
MMSE. The Mini-Mental State Exam (MMSE) 23 is a well-known screen for cognitive impairment.
dTEL. We used the “dTEL” δ homolog. 24 dTEL is indicated by Logical Memory I (LMI) and II (LMII) from the Wechsler Memory Scale 25 and Category Fluency (Animals) from the Consortium to Establish a Registry for Alzheimer's Disease (CERAD) battery. 26 The Functional Assessment Questionnaire (FAQ) 27 was used as dTEL's functional status indicator. It has been successfully incorporated into other δ homologs.15,16,28
N factor biomarker indicators
CSF total tau (CSF tau), fluorodeoxyglucose (FDG)-PET, and whole brain volumes by MRI have been associated with neurodegeneration and advocated as proxies for neurodegeneration in the A/T/N classification scheme.1,29 Each are available in ADNI. We used them as indicators of a formative “N Factor”.
CSF tau. CSF tau levels were obtained from ADNI. The data were acquired by the ADNI Biomarker Core Laboratory (University of Pennsylvania) using the multiplex xMAP Luminex platform (Luminex Corp, Austin, TX) with Innogenetics (INNO-BIA AlzBio3; Ghent, Belgium) immunoassay kits.
Whole brain volume. Whole brain volumes were obtained from the ADNI website. The method has been described in detail. 30
FDG PET. PET data was obtained from the ADNI website. 31 A detailed description of PET image acquisition and processing is available at. 32
Competing dementia risks
Age, an APOE ε4 (+) genotype, central nervous system (CNS) Aβ in CSF or by PET, depressive symptoms, and gender can be considered as competing determinants of dementia severity as measured by δ, and contribute independently to its variance.
Age. Age was calculated in years from date of birth.
APOE genotyping. APOE genotyping was conducted in both datasets using standard polymerase chain reaction (PCR) methods. APOE ε4 burden (APOE) was coded dichotomously by possession of one or more ε4 alleles.
Aβ. CNS amyloidosis was estimated by florbetapir/Amyvid PET (AV45). Regional Standardized Uptake Values (SUVr) were calculated as the mean AV45 uptake in four neocortical regions (temporal, frontal, parietal and cingulate) divided by a reference region (cerebellum). SUVr >1.2 have been associated with dementia. 33 CSF Aβ levels were obtained from the ADNI database.
Depression. Depressive symptoms were assessed in both studies by the Geriatric Depression Scale (GDS) 34 which has been validated in persons with dementia. 35 Higher scores are worse.
Gender. Self-reported gender was scored dichotomously with female = 0 and male = 1.
Statistical analyses
Approach. Our LOI algorithm has been previously described.8–10 We again use the “dTEL” homolog. dTEL was initially developed to facilitate assessment by telephone, but in ADNI, its indicators were obtained face to face. 24
Next, we constructed a latent measurement model to provide a latent “N Factor” representing the variance shared by several putative biomarkers of neurodegeneration. Then, we regressed dTEL scores onto the N Factor. This produces a cognitive residual, “CR”, representing all variance in dTEL (i.e., in dementia severity) not attributable to neurodegeneration. Any and all competing influences on dTEL are captured in that residual.
Next, we drew a LOI through a CR by dTEL scatterplot. Cases off the LOI have been demonstrably impacted by the N Factor. Cases can be said to be “afflicted” if their standardized CR-scores improve relative to those of the unadjusted dTEL. Otherwise, they are “resilient”.
We validated the affliction class by testing its effect on baseline clinician-rated dementia severity (CDR-SB), N Factor scores, and observed values of each of the N Factor's indicators by Analysis of Variance (ANOVA). The specificity of affliction class to neurodegeneration was confirmed by multivariate regression and by Factorial ANOVA testing the direct effects of affliction class, baseline clinical diagnosis and their interaction on AD-related biomarkers. Clinical diagnoses were coded categorically, i.e., Normal Control (NC) = 0, MCI = 1, and AD = 2. We estimated time to AD conversion from non-demented baseline diagnoses over 48 months. The effect of affliction class on conversion was tested by survival analysis (i.e., Kaplan-Meier and Cox proportional hazards models). 36
SEM. The analysis was performed using Analysis of Moment Structures (AMOS) software. 37 The maximum likelihood estimator was selected. Full information Maximum Likelihood (FIML) methods were used to address missing data. 38
Fit indices. The validity of structural models can be assessed by certain statistical tests. A non-significant chi-square signifies good fit to the data but is difficult to achieve in large samples. 39 However, the ratio of the chi-square to the degrees of freedom in the model (i.e., the CMIN/DF ratio) is also of interest. A CMIN/DF ratio < 5.0 suggests an adequate fit. 40 The comparative fit index (CFI), compares the specified model with a model of no change. 41 CFI values range from 0 to 1. Values of 0.95 or greater indicate excellent fit. A root mean square error of approximation (RMSEA) of 0.05 or less indicate good fit. 42
Results
Descriptive statistics are presented in Table 1. dTEL had excellent fit (CHISQ = 2.76, df = 2; CFI = 1.00, RMSEA = 0.02) and exhibited a strong association with CDR-SB (r = -0.78, p < 0.001), as previously reported. 24 That inverse association, and dTEL's inverse association with FAQ, suggest that higher dTEL scores are beneficial. dTEL was reified as a composite “dTEL score”.
Descriptive statistics by neurodegeneration-affliction class.
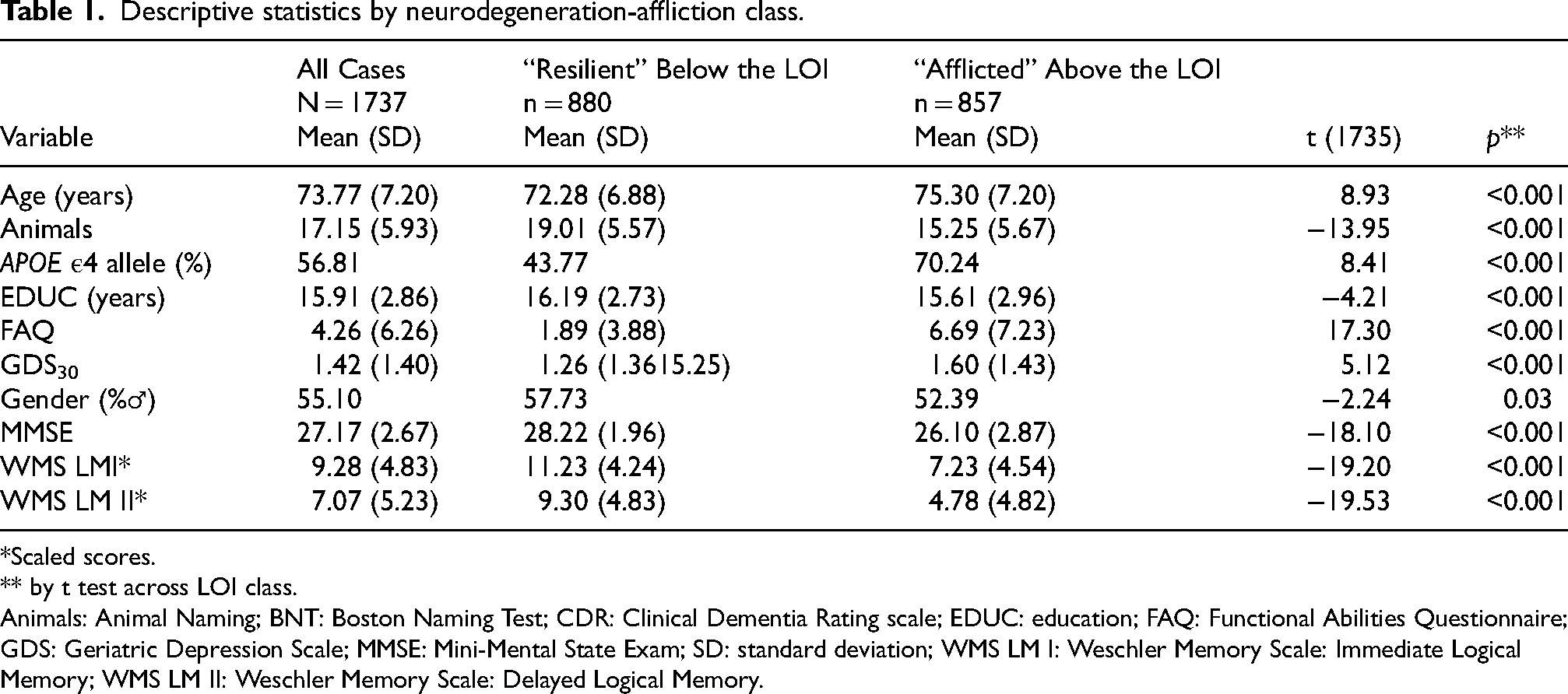
*Scaled scores.
** by t test across LOI class.
Animals: Animal Naming; BNT: Boston Naming Test; CDR: Clinical Dementia Rating scale; EDUC: education; FAQ: Functional Abilities Questionnaire; GDS: Geriatric Depression Scale; MMSE: Mini-Mental State Exam; SD: standard deviation; WMS LM I: Weschler Memory Scale: Immediate Logical Memory; WMS LM II: Weschler Memory Scale: Delayed Logical Memory.
As an expected artifact of its formative construction, the N Factor had perfect fit (CHISQ = 0.00, df = 0; CFI = 1.0). Regardless, all indicators loaded significantly, ranging from total brain volume (r = 0.15, p < 0.001) to FDG PET (r = 0.70, p < 0.001) (Figure 2). The N factor's parameterization was such that higher scores were associated with beneficial effects on dementia severity. It was strongly positively correlated with dTEL (r = 0.77, p < 0.001), and inversely with CDR-SB (r = -0.67, p < 0.001).
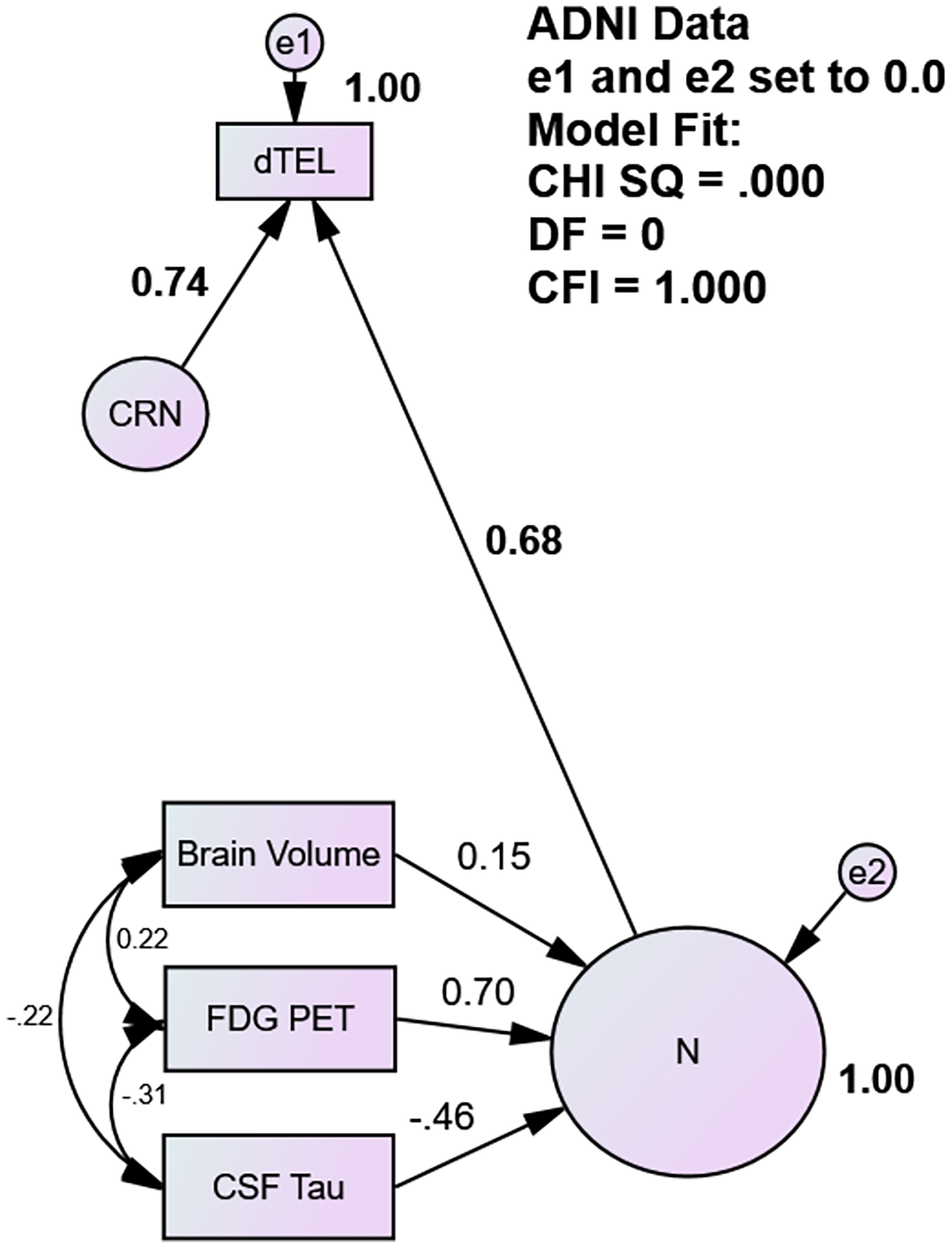
Cognitive residual factor (CR) (ADNI). ADNI: Alzheimer's Disease Neuroimaging Initiative; CFI: comparative fit index; CR: N Factor cognitive residual; CSF: cerebrospinal fluid; DF: degrees of freedom; FDG: fluorodeoxyglucose; PET: positron emission tomography; RMSEA: root mean squared error of approximation.
Figure 2 also illustrates the CR factor's derivation. The residual e1 is constrained to zero variance to force dTEL's residual variance into CR, i.e., the variance residual to neurodegeneration's effect. The N Factor is strongly positively associated with dTEL (r = 0.68, p < 0.001), a beneficial effect. Appropriately, total brain volume and FDG uptake load positively on the N Factor while CSF tau is inversely associated with it. The N factor explains variance in CDR-SB independently of competing dementia risks (Table 2). As expected, CR and the N Factor together explain 100% of dTEL's variance (by multivariate regression: F (2, 1928) = 496E, 15 p < 0.001. R²=1.00.
N factor adds variance to CDR-SB independently of competing dementia risks.
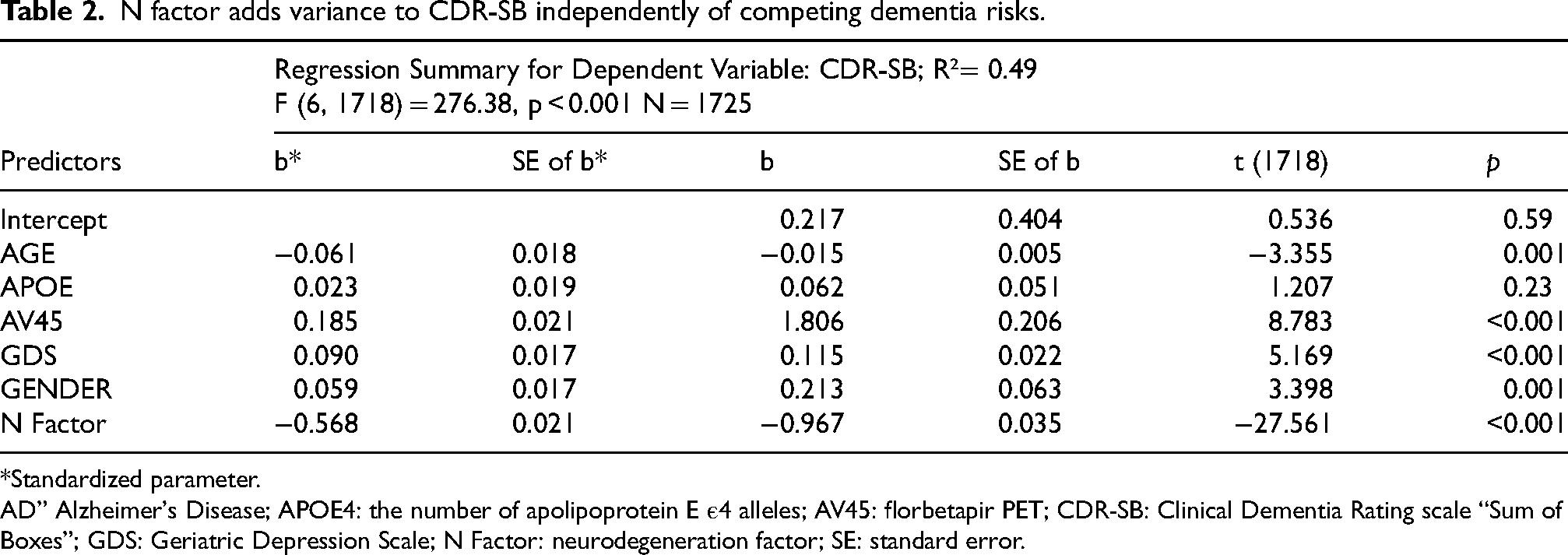
*Standardized parameter.
AD” Alzheimer's Disease; APOE4: the number of apolipoprotein E ε4 alleles; AV45: florbetapir PET; CDR-SB: Clinical Dementia Rating scale “Sum of Boxes”; GDS: Geriatric Depression Scale; N Factor: neurodegeneration factor; SE: standard error.
Figure 3 presents the LOI analysis. The standardized adjusted- (i.e., CR) and unadjusted- (i.e., dTEL) composites were strongly correlated (r = 0.82, p < 0.001). Higher dTEL, N Factor, and CR scores are beneficial. All three are inversely associated with CDRSB (dTEL r = -0.80, N Factor r = -0.67, CR r = -0.61, all p < 0.001). Therefore, cases with CR scores above the LOI have relatively improved dementia severity after the N Factor's effect is accounted for. They are “Afflicted”. The remaining cases are “Resilient” to the effect of the N Factor (i.e., neurodegeneration). LOI class is positively associated with CDRSB when afflicted cases are assigned a value of “1” (r = 0.41, p < 0.001).
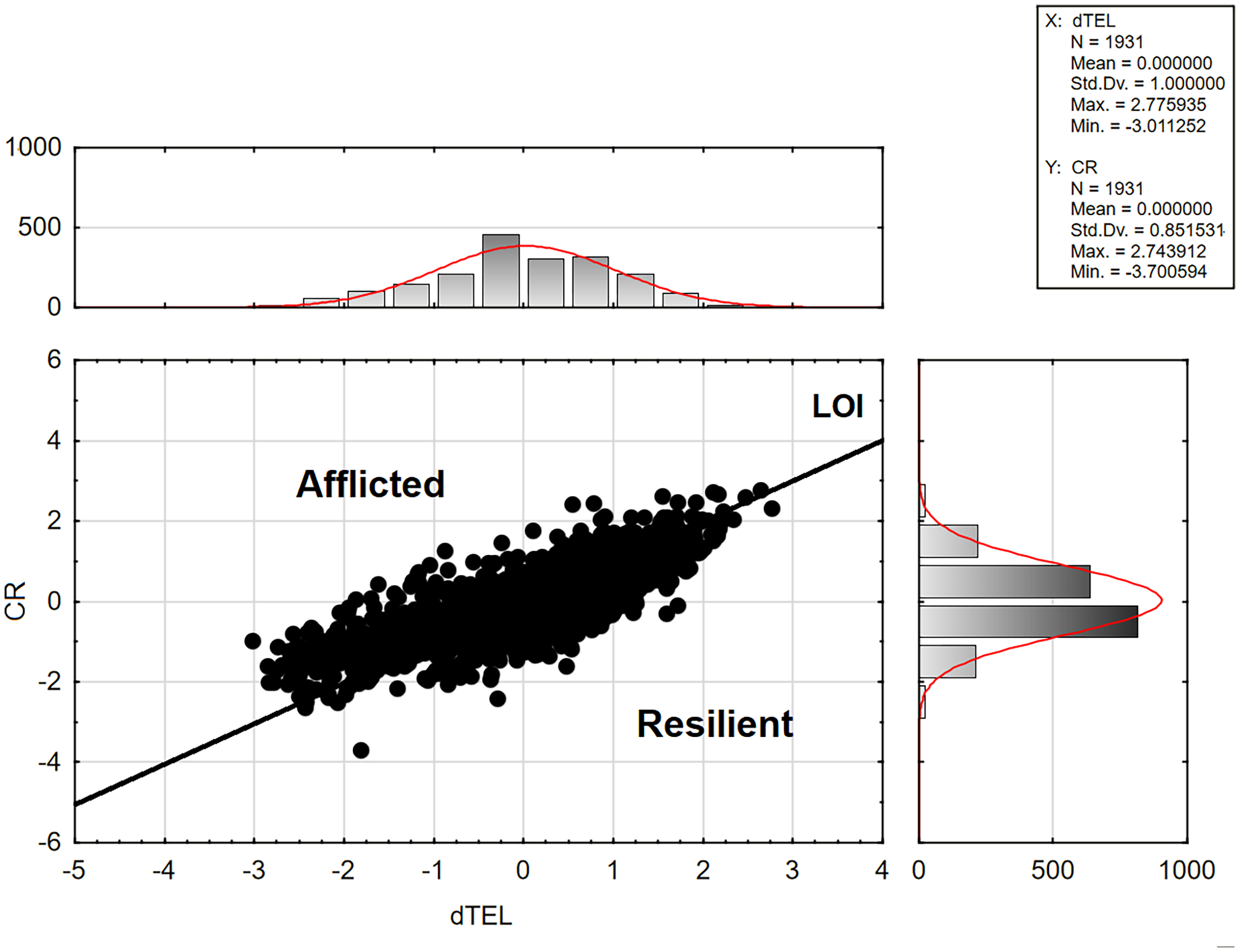
Scatterplot of N factor-adjusted (CR) x unadjusted (dTEL) composite scores with line of identity (LOI) (ADNI) r = 0.82, p < 0.001. ADNI: Alzheimer's Disease Neuroimaging Initiative; CR: N Factor cognitive residual; LOI: line of identity; Std Dv: standard deviation; Max: maximum; Min: minimum.
N = 857/1737 (49.40%) of ADNI's subjects were adversely impacted by neurodegeneration. Afflicted cases presented across dTEL's entire range (Figure 3). Table 1 presents descriptive statistics for each affliction class. Adversely impacted cases were more likely to be ε4 carriers, more impaired on multiple cognitive measures and were less well educated. They had higher levels of informant-reported disability, were more likely to be depressed and were more likely to endorse female gender. CDR-SB scores were significantly higher in the afflicted class [by ANOVA: F (1, 1735) = 354.88, p < 0.001.
Affliction class explained variance in dementia severity independently of the N Factor's observed indicators (Table 3). This suggests both that affliction by the N Factor, a systems level biomarker, contributes to dementia independently of its indicators and also that those indicators impact dementia severity independently of both each other and a subject's affliction class.
Affliction class explains variance in dementia severity independently of observed biomarkers of neurodegeneration.
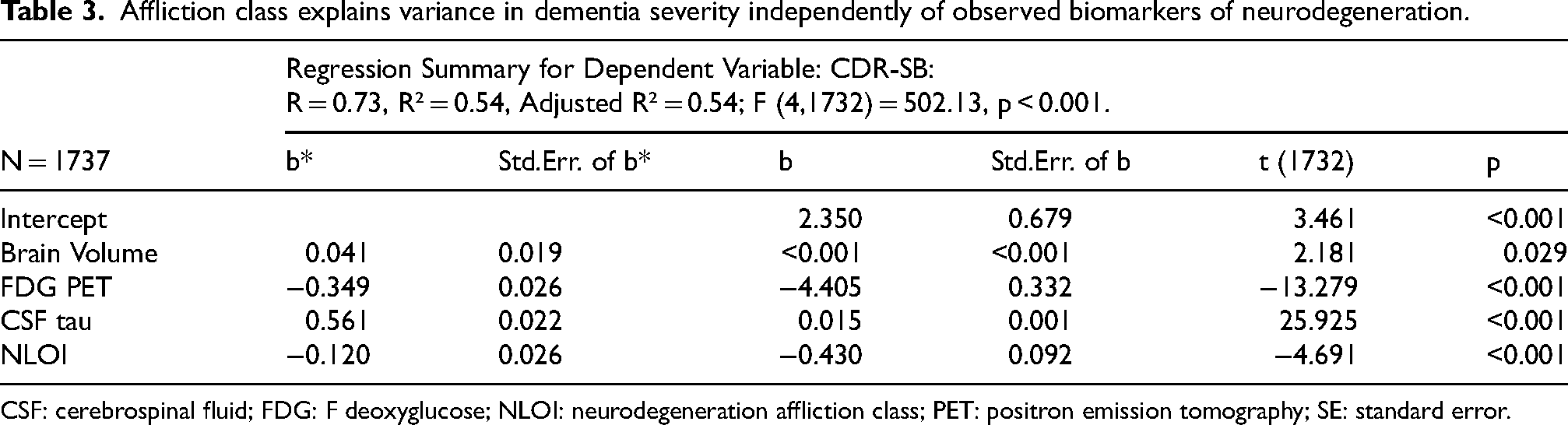

CSF: cerebrospinal fluid; FDG: F deoxyglucose; NLOI: neurodegeneration affliction class; PET: positron emission tomography; SE: standard error.
N Factor scores were significantly lower (adverse) in afflicted subjects [by ANOVA: F (1, 1735) 2049.50, p < 0.001], as were observed levels of each of the construct's indicators (i.e., by Tukey's HSD, all p < 0.001). The frequency of afflicted cases rose as a function of clinical diagnosis. 29.5% of NC, 45.40% of MCI cases, and 84.80% of AD cases were afflicted. To confirm that NLOI is not merely selecting more severe stages of AD, we tested its effect on putative AD biomarkers (APOE4, CSF Aβ, Aβ PET and the N Factor itself) and for interactive effects with baseline clinical diagnosis on those biomarkers (Table 4).
Clinical diagnosis has significant effects on the N factor, its indicators, and other AD-related biomarkers. Affliction by neurodegeneration (NLOI) impacts only the N Factor and its indicators. It has no direct or interactive effects on other AD-related biomarkers (by factorial ANOVA; ADNI).
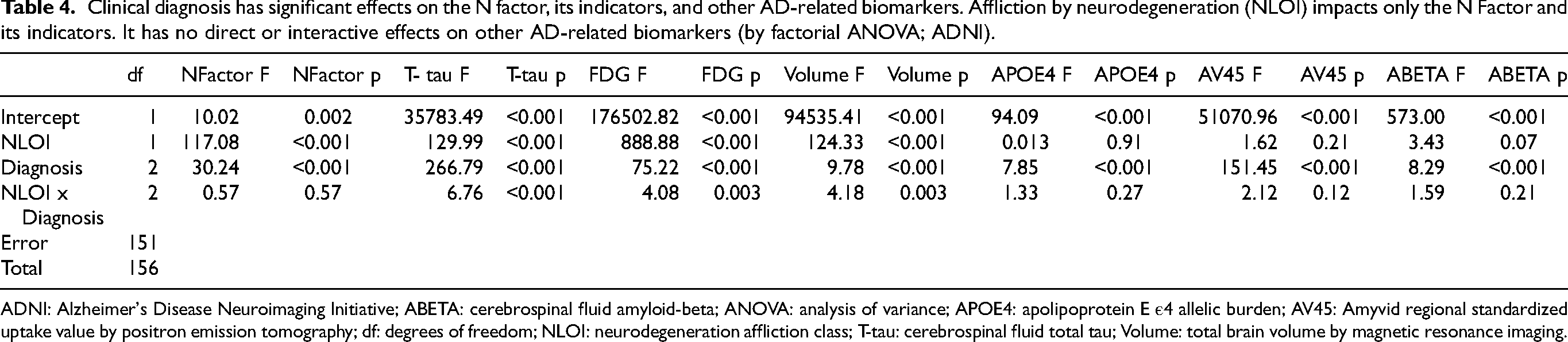
ADNI: Alzheimer's Disease Neuroimaging Initiative; ABETA: cerebrospinal fluid amyloid-beta; ANOVA: analysis of variance; APOE4: apolipoprotein E ε4 allelic burden; AV45: Amyvid regional standardized uptake value by positron emission tomography; df: degrees of freedom; NLOI: neurodegeneration affliction class; T-tau: cerebrospinal fluid total tau; Volume: total brain volume by magnetic resonance imaging.
Clinical diagnosis had significant effects on the N Factor, its indicators, and other AD-related Biomarkers (Table 4). However, the NLOI impacted only the N Factor and its indicators. It had no direct or interactive effects on other AD-related biomarkers. So, the association between affliction class and clinical diagnosis cannot be explained by a more “advanced” stage of AD, as that would entail differences in other AD-related biomarkers than neurodegeneration.
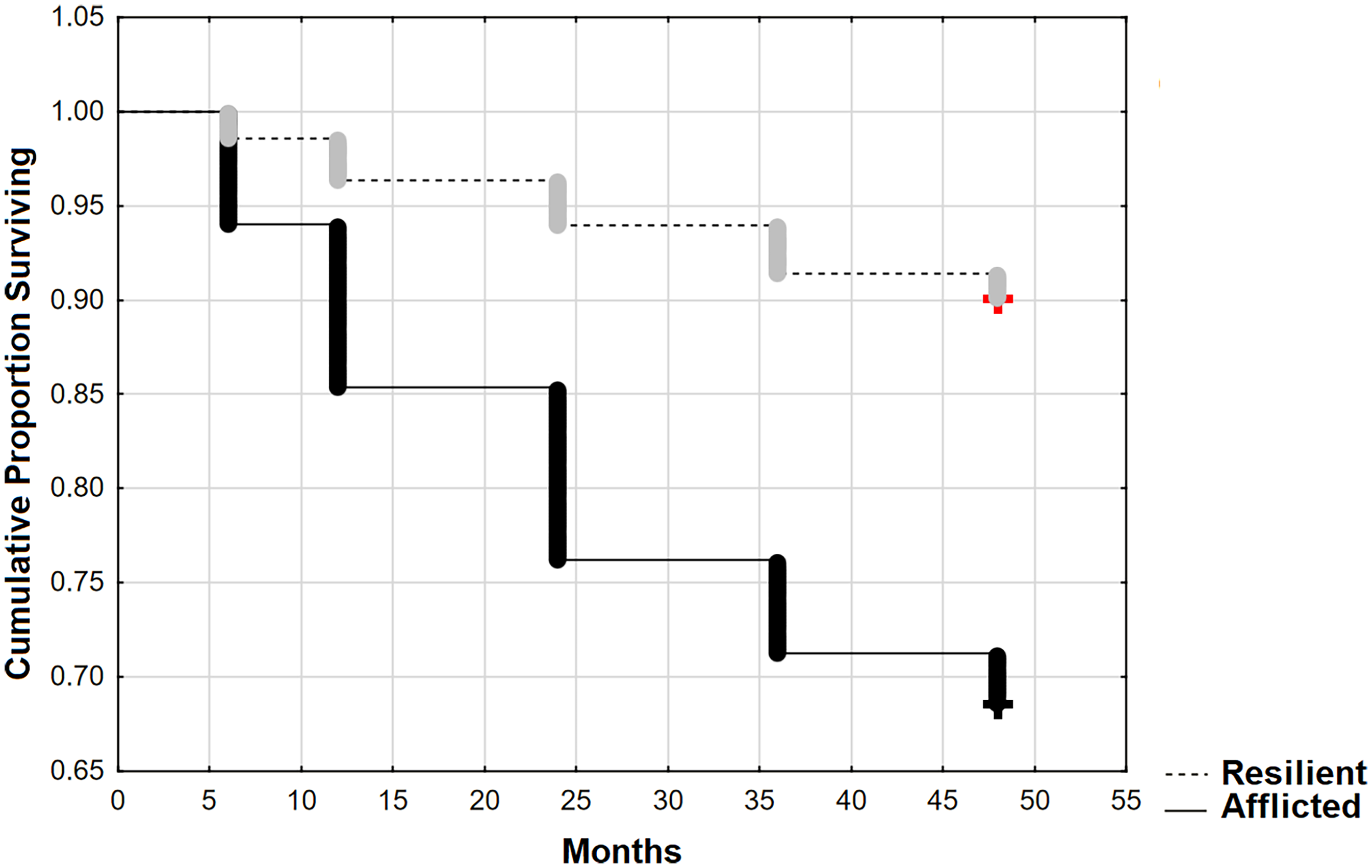
Baseline affliction class impacted prospective CDR-SB at 48 months (by ANOVA: F (1614) = 97.84, p < 0.001). It also had a significant effect on four year (48 month) prospective conversion to clinical “AD” from non-demented states [by Cox's F: F (164, 356) = 3.65, p < 0.001; Figure 4.
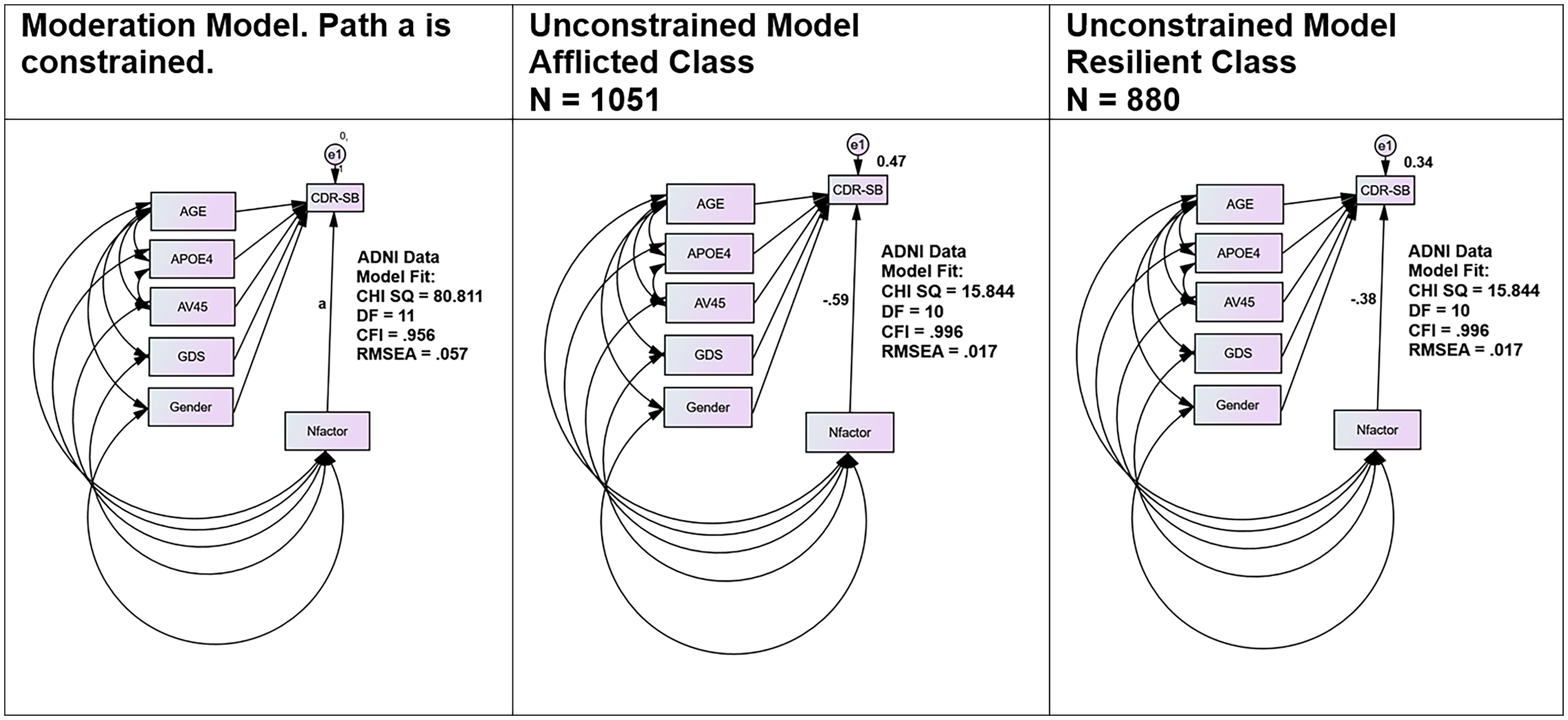
Moderation effects. ADNI: Alzheimer's Disease Neuroimaging Initiative; APOE4: the presence or absence of apolipoprotein ε4 alleles; AV45: florbetapir PET SUVr; CDR: Clinical Dementia Rating Scale “Sum of Boxes”; CFI: comparative fit index; CR: N Factor cognitive residual; DF: degrees of freedom; GDS: Geriatric Depression Scale; MMSE: Mini-Mental State Examination; PET: positron emission tomography; RMSEA: root mean squared error of approximation; SUVr: regional standardized uptake value.
We formally tested the moderation effect of Affliction class on the N Factor's association with clinician-rated dementia severity (i.e., CDR-SB). The effect was tested by change in ChiSQ fit (ΔCHISQ) in constrained versus unconstrained regression models, stratified by affliction class. Their association was adjusted for Age, APOE4, AV45, GDS and GENDER (Figure 5(a)). The moderation effect was significant [ΔCHISQ = 64.97 (ΔDF = 1), p < 0.001. In the afflicted class, the N Factor explained more than twice (241%) the variance in CDR-SB scores (Figure 5(b)) it could explain in the resilient class (Figure 5(c)).
Conversion to clinical “
Discussion
These results confirm our earlier findings that biomarker effects on dementia severity are moderated by LOI-derived Affliction Classes.8–10 They highlight potential weaknesses in the proposed A/T/N diagnostic scheme. First, we have distinguished the N Factor's shared variance from the unique effects of its indicators. Observed variables conflate the two sources of variation but it is the N Factor that is most strongly associated with dementia severity. The same was noted for blood-based proteins.9,10 Second, we have shown the N Factor's association with dementia to be independent of multiple other dementia-related biomarkers and risk factors. The same was noted for other biomarkers.8–10 This finding speaks to dementia's overdetermined nature and undermines its attribution to a single cause (e.g., “AD”) except in very specific and possibly infrequent circumstances. Instead, the dementia observed in so called “AD” cases is likely somewhat less determined by A/T/N biomarkers and somewhat more determined by unnamed comorbidities. The variance in dementia severity explained by those comorbidities should also be targets for intervention. For example, more ADNI participants are reported in this analysis to be afflicted by the N Factor than we found to be afflicted by Aβ PET, 8 and the two contribute to CDR-SB independently of each other. The N Factor's effect is stronger (Table 2). Third, we have qualified the N Factor's unique effect by affliction class, which moderates its association with dementia. The same was noted for blood-based proteins9,10 and for PET imaged amyloidopathy. 8
Our factor analytic approach distinguishes covaried and unique effects. We have used it to distinguish the N Factor's shared variance from the unique effects of its indicators. In contrast, by using observed biomarkers only, A/T/N classification risks to conflate the two.
Any one of our neurodegeneration-related biomarkers might plausibly stand in for N in an A/T/N analysis, which is the approach taken by many studies. However, the latent N Factor is qualitatively and psychometrically distinct. It is a system level biomarker, capturing the coordinated variation in its indicators. The residual variances of each biomarker (i.e., their N Factor independent effects) explain relatively little variance in dementia severity. The N factor's indicators were most strongly associated with dementia through that latent construct (Table 2). This suggests that studies that invoke any one of these observed biomarkers as a proxy for the N domain conflate dementia-related and unrelated effects, and may offer an inadequate view of the complexity of biomarker interactions.
The distinction is important. Treatment outcomes may depend on whether an intervention is acting at a systems level versus individual biomarkers. It may prove easier to intervene on a single indicator (e.g., CSF tau), than on the N Factor. However, any change effected in a single biomarker will be orthogonal (unrelated) to the N Factor score and may be less effective in changing a patient's d-score.
Similarly, A/T/N nomenclature risks to conflate the unique effects of individual biomarker domains with systems level integration of their effects. A/T/N-related biomarkers would have more validity if they could be shown to relate to δ through their shared variance. Their unique effects are unrelated to each other (by definition) and cannot then be invoked as the biomarkers of a single disease process, even if the observed pathologies can be demonstrated together in cases diagnosed with “AD”. For now, whether A/T/N effects are mediated through their covaried or unique residual effects remains an empirical question.
Our analysis suggests they may not be covaried. The N Factor's association with CDR-SB was shown to be independent of AV45 PET (Table 2, Figure 5), arguably a more AD-specific biomarker and itself a proxy for “A” in the A/T/N scheme. Moreover, FDG PET, which loaded most strongly on the N Factor, is arguably least specific for AD per se. CSF tau might be more AD-specific, but it loaded statistically weakly on the N Factor and had the strongest unique effect on CDR-SB, independent of N factor Affliction Class (Table 5). N Factor-derived affliction class had no impact on CSF Aβ, APOE, or AV45 PET (Table 4). Which of these then, are AD-specific biomarkers?
Affliction class explains variance in dementia severity independently of observed biomarkers of neurodegeneration.
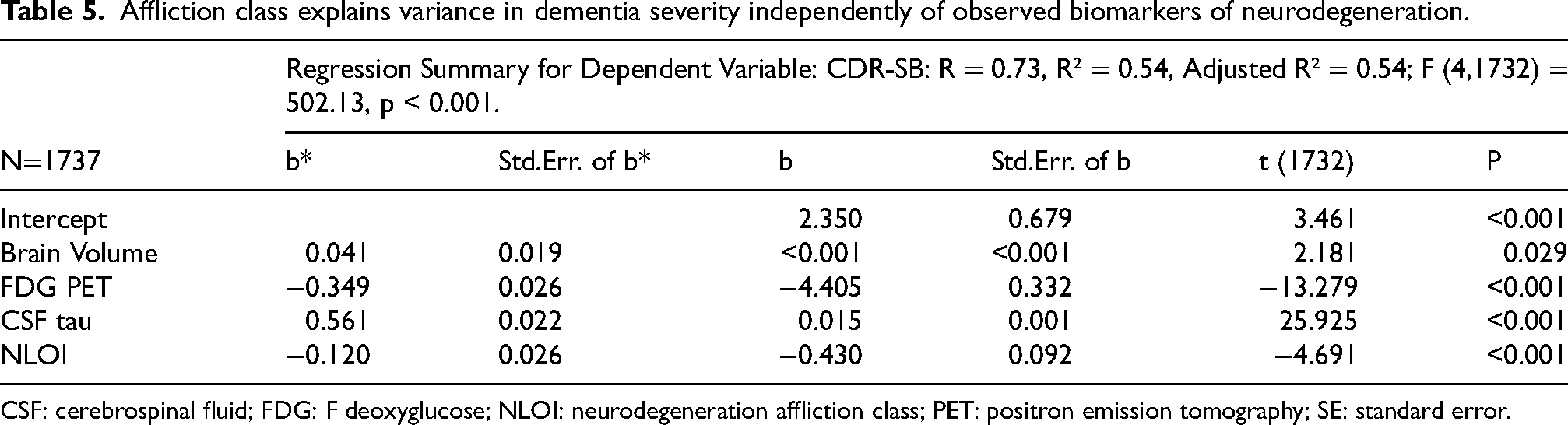

CSF: cerebrospinal fluid; FDG: F deoxyglucose; NLOI: neurodegeneration affliction class; PET: positron emission tomography; SE: standard error.
Similarly, the effect of affliction class is independent of comorbid processes (Table 5) and specific to the biomarkers of neurodegeneration (i.e., the N Factor and its indicators) (Table 4). This specificity is an advantage of our approach, but it undermines N's contribution to a monolithic “AD” process co-labeled by A, T and N pathologies.
On the other hand, we have already shown A and N-related biomarkers to be associated with the d-score. It seems likely that T will be associated with δ as well. We have not modeled T yet because its indicators are unavailable to us in ADNI. If A, T, and N can be associated with the d-score though a latent A/T/N construct, then AD has more validity. Were that construct then submitted to LOI, we could, for the first time, identify individuals in vivo who are resilient to AD and assess the true frequency and impact of A/T/N interactions.
Second, the traditional residuals approach to resilience risks to conflate biomarker-specific variation with differences in unrelated dementia processes (represented by CR in this analysis which is unrelated to the N Factor's effect). Our LOI approach results in biomarker-specific effects. Membership in the N Factor affliction class impacts only the N Factor and its indicators. It has no direct or interactive effects on other AD-related biomarkers (Table 4). Other nominal “AD” biomarkers have been shown to be associated with CDR-SB independently of the N Factor (Table 2; Figure 5). Similarly, clinicians’ diagnoses in ADNI non-specifically impacted both the N Factor and competing AD-related biomarkers (Table 4). The residual N-factor adjusted variance in clinical diagnosis, i.e., “CR” in Figure 2, is likely to reflect the contributions of similar unmodeled comorbidities. However, it is the variance in dementia severity residual to a point estimate, i.e., the very fraction from which “Resilience” is attributed by a traditional residuals approach. Our CR is better understood as a measure of “cognitive reserve”, 43 i.e., the variance in dementia-salient cognitive performance that is unrelated to the biomarker of interest and subject to competing dementia-related processes (if any) (Figure 1).
Finally, the affliction class qualifies the N Factor's effect on dementia severity. The resilient classification selects cases whose dementia severity worsens when the effect of the N Factor is extracted from their d-score. This suggests that neurodegeneration in resilient cases offsets the impact of competing dementia risks, i.e., it has a beneficial effect on dementia severity. However, our moderation models do not appear to confirm a direct beneficial effect. Instead, they show a significantly weakened adverse association between the N Factor and CDRSB. A beneficial effect would have changed in the inflection of their association. So, the resilient class's improved survival curve results from a weakened, yet still adverse association. One way these might be reconciled would be if neurodegeneration is acting antagonistically to one or more other, unmodeled, dementia-related biomarkers or risk factors. Then, the weakened effect of neurodegeneration in resilient cases would accentuate those other biomarkers’ effects. This implies the possibility of mediation effects among competing dementia-related biomarkers and goes beyond the scope of this analysis.
Our qualification of biomarker associations by affliction class suggests that A/T/N classifications may need adjustment. For example, the N Factor explained significantly more variance in CDR-SB in afflicted relative to resilient subjects, resulting in a relatively large difference in their survival curves (Figure 4). That suggests, 1) prognostication of prospective outcomes from biomarkers becomes problematical and potentially misleading without knowledge of their LOI class assignment, and 2) the risk /benefit ratio of interventions directed at a biomarker may vary by an individual's NLOI class assignment. So, the significance of a biomarker could be better assessed if qualified by class membership. This would increase the number of possible A/T/N designations from eight (currently) to 27. A more nuanced assessment of A/T/N impacts could then be determined, as could the relative benefits of intervention against each biomarker domain as a function of a case's affliction class.
Our approach is limited by some significant caveats. First, we did not model all possible indicators of neurodegeneration. CSF neurofilament light chain (NfL), glial fibrillary acidic protein (GFAP), and neurogranin (Ng) are also associated with neurodegeneration but were not available to us in our cut of the ADNI dataset. It will be an empirical question whether they contribute to δ. Not all neurodegeneration may do so. Regional sulcal variability is strongly associated with observed cognitive performance in AD, 44 but whole brain volume contributes independently of local atrophy 45 and is associated with intelligence. 46 It remains an empirical question whether CSF NfL, GFAP, or Ng can be associated with δ and whether they do so independently of our N Factor's indicators or in covariation with them (i.e., as additional indicators of the N Factor construct).
Figure 1 implies that major dementia-related biomarkers act independently of each other. This assumption may not be broadly true. We have shown that the N Factor's effect is independent of several other biomarkers, and that they themselves are contributing to the d-score independently of each other (Table 3). Regardless, the N Factor itself reflects the integrated effects of multiple biomarkers. Future analyses might address potential mediation and moderation effects among δ-related biomarkers or the impact of integrated biomarker effects, e.g., by LOI-affliction classification relative to a factor integrating A, T and N-related biomarkers.
Furthermore, we do not know the mechanisms of resilience against any of these biomarkers. However, our approach opens a way forward. We can now assign individuals to LOI-affliction classes relative to any pre-specified δ-related biomarker. We can look at the stability of class membership over time and across stages of dementia's evolution. We can examine either cross-sectional or longitudinal cross-class differences for clues as to the mechanisms that explain them.
Finally, it remains to be seen if our approach can be generalized to demographics other than non-Hispanic Whites (NHW) as ADNI is neither ethnically nor racially diverse. However, our INFLAMMATION classifier, also from ADNI, was replicated in a more diverse sample with a high fraction of Mexican American (MA) participants. 19
In conclusion, our LOI algorithm can select individuals with biomarker-specific profiles and predict prospective conversion to “AD” from non-demented states. Sizable fractions of non-demented subjects may be “resilient” to a biomarker's effect. These findings have implications for both the A/T/N enterprise and for AD's treatment by biomarker-specific interventions.
Footnotes
Acknowledgements
Data collection and sharing for the ADNI is funded by the National Institute on Aging (National Institutes of Health Grant U19AG024904). The grantee organization is the Northern California Institute for Research and Education. In the past, ADNI has also received funding from the National Institute of Biomedical Imaging and Bioengineering, the Canadian Institutes of Health Research, and private sector contributions through the Foundation for the National Institutes of Health (FNIH) including generous contributions from the following: AbbVie, Alzheimer's Association; Alzheimer's Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. ADNI had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. However, ADNI investigators contributed to the design and implementation of ADNI and/or provided data. A complete listing of ADNI investigators can be found at: ![]() .
.
Ethical considerations
ADNI's protocol was approved by institutional review boards of all participating institutions. It was conducted in accordance with the Declaration of Helsinki. Written informed consent was provided by all the participants or their guardians.
Consent to participate
This is a secondary analysis of deidentified data provided by the Alzheimer's Disease Neuroimaging Initiative (ADNI). ADNI is a multi-site observational study. Each site obtained approval for the ADNI protocol from its local Institutional Review Board (IRB). No data was collected from participants without their informed consent.
Consent for publication
Not applicable.
Author contribution(s)
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors are supported by NIA 1R01AG080548-01 (Royall PI) “Novel Methods for Clinical Trials in Dementia and Cognitive Decline”. Dr. Royall is also supported by the Julia and Van Buren Parr Endowment to the University of Texas Health Science Center at San Antonio.
National Institute on Aging, (grant number 1R01AG080548-01).
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The authors have disclosed their invention of δ, its homologs and orthologs to the University of Texas Health Science Center at San Antonio, which in turn holds a patent relating to the use of a non-overlapping set of δ-related blood-based biomarkers as predictors of Mild Cognitive Impairment (MCI) in non-Hispanic whites (U.S. Patent 13/943,654: “A Serum Biomarker Screen for the Diagnosis of Clinical and Pre-clinical Alzheimer’s Disease in Non-Hispanic Populations”). There are no other patents, products in development or marketed products to declare.
