Abstract
Genetic risk prediction for Alzheimer's disease (AD) has high potential impact, yet few studies have assessed the reliability of various polygenic risk score (PRS) methods at the individual level. Here, we evaluated the reliability of AD PRS estimates among 6338 participants from the Multi-Ethnic Study of Atherosclerosis. We compared four PRS models that have been previously associated with dementia risk. Despite similar population-level performance metrics, inter-model reliability of individual-level risk assessment was low, even among individuals classified in the top and bottom deciles. These findings raise serious concerns about the downstream application of PRS for guiding interventions for AD.
Introduction
Given that Alzheimer's disease (AD) has high heritability (58–79%), 1 genetic data shows promise for early identification of people at greatest risk.2,3 Genetic studies have identified dozens of loci associated with AD.4,5 While individual effects of each loci are modest and offer limited predictive value, by combining effects from across the genome, polygenic risk scores (PRS) capture the cumulative genetic risk for AD in a single score. The use of PRS has grown rapidly, with major efforts underway to develop new methods and to provide guidelines for clinical and commercial implementation.6–9
While PRS are conceptually simple—essentially a weighted sum of genetic risk variants—choices regarding which variants to include and how weights are assigned are complex and have led to an ever-increasing number of PRS approaches. Despite countless possibilities for constructing PRS, there are no obvious winners. Several studies, including our own, have shown that PRS constructed from thousands of variants do not necessarily outperform methods that include only the most significantly associated variants for predicting AD and dementia.10,11
According to population-level performance metrics including area under the curve (AUC), concordance index (Harrell's C), and variance explained (R2), differences in predictive performance across AD PRS methods are relatively small. As PRS begin to be used to guide decisions, however, critical questions about the choices of PRS methods emerge beyond summary-level performance: given that potentially life-altering screening, clinical trial enrollment, or other interventions are allocated differently to individuals with the highest and lowest genetic risk, how does the PRS method that is chosen affect who is identified for these interventions? In other words, what is the reliability of these various methods in identifying high-risk individuals?
Reliability, defined here as the extent to which a given measurement can be replicated, 12 is crucial to health screening, but not typically assessed when comparing the predictive performance across different PRS methods. Here, we explore how differences in PRS construction methods impact whether individuals classified as high or low risk for AD by one method are consistently identified as such by other methods. Unreliable risk assessments can have catastrophic downstream effects, undermining the efficacy of resulting interventions and perhaps causing more harm than benefit.
Methods
Study population
We used data from the Multi-Ethnic Study of Atherosclerosis (MESA), a longitudinal cohort study that recruited 6814 individuals across six sites at baseline in 2000–2002. 13 Participants were included in the analysis if they had genotyping data that passed quality control and were followed for hospitalizations or deaths due to dementia.
PRS construction
The PRS methods used have been previously described. 11 Briefly, we constructed PRS for AD using clumping and thresholding (C + T), 14 PRS-CS, 15 and PRS-CSx. 16 For the C + T models, we used an LD threshold of r2 = 0.01 and compared two p-value thresholds: 5e-08 (typical threshold for genome-wide significance) and 1e-05 (typical threshold for genome-wide suggestive association). Bayesian methods, PRS-CS and PRS-CSx, do not filter variants based on LD or p-value thresholds, instead tuning the effect sizes for each variant using continuous shrinkage. Therefore, although the variants selected using C + T are included in the Bayesian models, their effect sizes may be reduced while variants in strong LD retain larger weights. For PRS-CS and PRS-CSx, we used the -auto option and a global shrinkage parameter of 1e-02. All PRS were derived from GWAS summary statistics from the International Genomics of Alzheimer's Project (IGAP). 17 PRS-CSx allows for the integration of multiple summary statistics with differing ancestry backgrounds, so we also used summary statistics from the African Genome References Panel. 18 The APOE region was removed from all PRS, with APOE genotype included as an independent covariate in prediction models. All PRS were adjusted for the first three genetic principal components and subsequently standardized for comparison. 19
Assessing PRS reliability
To evaluate reliability, we calculated the intraclass correlation coefficient (ICC) using a two-way mixed effects approach, which measures the degree of correlation and agreement for a given construct between different approaches. 20 We also calculated Spearman's rank correlation coefficients to assess pairwise correlation of risk estimates. 21 ICC and Spearman's rank both offer measures of reliability at the population level, but do not differentiate between consistency of risk assessment at the extremes of PRS distributions compared to intermediate risk. It's possible that low to moderate correlation coefficients are driven primarily by inconsistencies in those at intermediate levels of genetic risk.
To address this limitation, participants were sorted into risk deciles based on scores from each model, and pairwise comparisons of decile rankings were conducted across models. Sankey plots visualize the reliability of risk decile classifications between models, with emphasis on the low and high extremes.
Sensitivity analysis
We conducted sensitivity analyses to assess whether inter-model variation could be driven by differences in genetic background. We restricted our analysis to participants with >80% 1000 Genome (1KG) non-Finnish European-like (NFE-like) ancestry, as previously defined by global ancestry inference.11,22 This subgroup represents a relatively homogeneous population that closely aligns with the ancestry of the GWAS summary statistics used to construct PRS. Additionally, we ran the analyses in groups stratified by APOE ε4 carrier status, as there is evidence that associations between PRS and dementia may differ across these groups.23,24
Results
Study population
6338 MESA participants were included in the analysis. The self-reported race/ethnicity of these participants are as follows: 2519 white, 774 Chinese, 1603 African American/Black, and 1442 Hispanic/Latino. 52% of participants are women and 26% are APOE ε4 carriers.
PRS models
Four PRS were calculated for each individual. The number of variants included in each model varied widely. PRS-CSx retains the largest number with 938,595 variants, followed by the PRS-CS model including 862,647 variants. The C + T with p < 5e-08 threshold results in a model with 15 variants, while the p < 1e-05 model includes 53 variants. In a previous study, we showed that all models are associated with incident dementia (Hazard Ratio (HR)5e−08 = 1.21, 95% CI:[1.11–1.31]; HR1e−05 = 1.12 [1.02–1.22]; HRCS = 1.13 [1.05–1.22]; HRCSx = 1.13 [1.04–1.23]). 11 Performance of PRS for predicting dementia was also similar across approaches in univariate models (AUC = 0.53–0.55) and models that included APOE, sex, and age (AUC = 0.8).
PRS reliability
The four different PRS had an ICC of 0.352 (95% CI: 0.338–0.365, Table 1). When comparing scores constructed using the same approach, either among C + T or Bayesian continuous shrinkage priors, the ICC improved (ICCC + T =0.65, 95% CI: 0.634–0.662; ICCCS−CSx = 0.584, 95% CI: 0.567–0.600). According to standard interpretations, an ICC value below 0.5 is considered to indicate poor reliability and ICC between 0.5 and 0.75 indicate moderate reliability. 20
Intraclass correlation coefficient (ICC) across polygenic risk score (PRS) approaches.
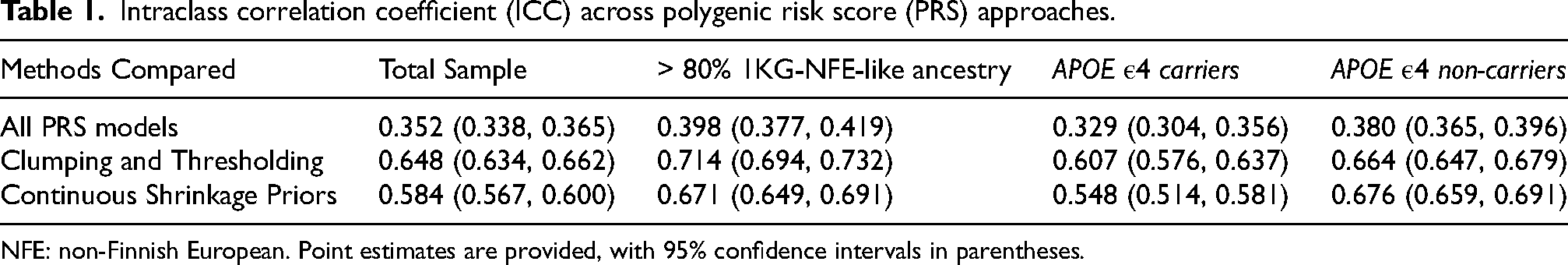

NFE: non-Finnish European. Point estimates are provided, with 95% confidence intervals in parentheses.
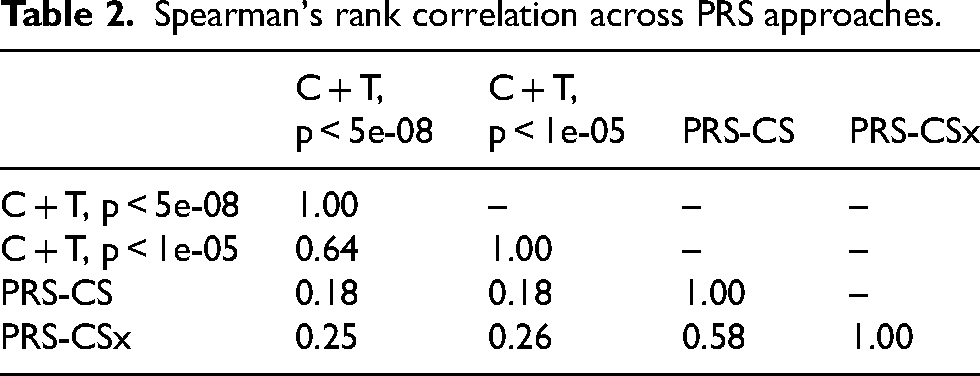
The C + T models that differ only in p-value threshold are the most strongly correlated, but the correlation is moderate (Spearman R = 0.64). The PRS-CSx and PRS-CS models are also moderately correlated (R = 0.58). The C + T and Bayesian approaches yield PRS that are weakly correlated (R = 0.11 to 0.25, Table 2).
Spearman's rank correlation across PRS approaches.
To directly examine the consistency in classification of high or low risk extremes across models, we conducted pairwise comparisons of inter-PRS model reliability (Figure 1-top). Of those who are in the top risk decile of PRS in the p < 5e-08 C + T model, 43% are considered in the top decile in the p < 1e-05 C + T model. 92% of those in the top risk decile in the p < 5e-08 model are in the top half of risk in the p < 1e-05 C + T. Comparisons between PRS-CS and PRS-CSx showed similar patterns, with 32% of those in the top risk decile in the PRS-CS model classified in the top decile in PRS-CSx and 83% remaining in the top half of risk. Results are similar when considering the reliability of a low-risk classification. Of those who are in the lowest risk decile of PRS in the p < 5e-08 C + T model, 43% are considered in the lowest decile and 90% are below median risk in the p < 1e-05 C + T model.
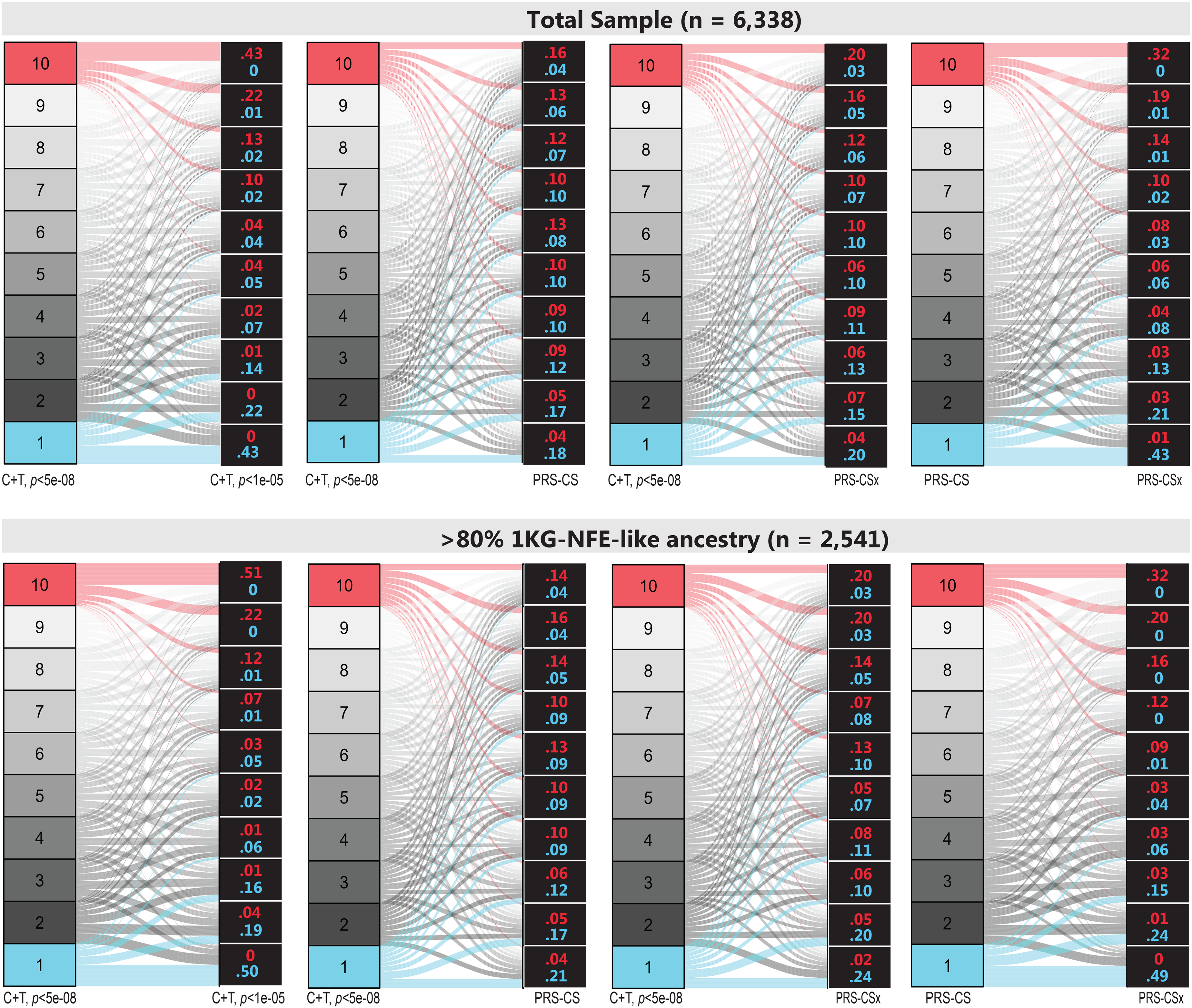
Sankey plot illustrating pairwise comparisons of PRS decile classifications. The top panels show the total sample (n = 6338), while the bottom panels focus on participants with >80% 1KG–NFE–like ancestry (n = 2541). In each panel, two PRS approaches are compared side by side: the left column indicates deciles of risk from the first approach (1 = lowest, 10 = highest), while the right column shows the deciles assigned by the second approach. The lines represent individuals flowing from one classification scheme to the other, and the numbers in the right column indicate the proportion of individuals from the first approach's top decile (top number) or bottom decile (bottom number) who are classified into each decile of the second approach.
In contrast, when comparing C + T and PRS-CS, only 16% of those who are in the top risk decile in the p < 5e-08 C + T model are considered in the top risk decile in the PRS-CS model. Alarmingly, 10% of participants classified as being in the lowest risk decile in the p < 5e-08 C + T model are classified as being in the top 20% of risk using the PRS-CS model.
Sensitivity analysis
Restricted analysis to those with >80% NFE-like ancestry. 2541 (40%) participants have >80% NFE-like ancestry. Reliability remains low within the NFE-like group, providing evidence that inconsistencies in risk assessment across models seen in the whole group analysis are not due to diverse genetic ancestry. While there is a modest improvement in reliability compared to the total sample, ICC = 0.398 (95% CI: 0.377–0.419) continues to indicate poor reliability among the NFE-like group (Table 1). Like the results from the total sample, there is higher concordance among PRS constructed using the same approaches, either using clumping and thresholding or continuous shrinkage priors (ICCNFE_C + T = 0.732, 95% CI: 0.694–0.732; ICCNFE_CS−CSx = 0.671, 95% CI: 0.649–0.691).
However, when comparing across PRS strategies, for example comparing C + T, p < 5e-08 and PRS-CS, those who are classified in the top or bottom deciles using C + T are nearly randomly distributed across the risk deciles when assessed using PRS-CS (Figure 1-bottom).
Stratified analysis by APOE ε4 carrier status. 1665 participants are APOE ε4 carriers. The inter-model reliability is low in both APOE ε4 carriers and non-carriers, with non-carriers having slight improvement in ICC (ICC ε 4 = 0.329, 95% CI: 0.304–0.356; ICCnon− ε4 = 0.380, 95% CI:0.365–0.396). The individual-level patterns in distribution of risk extremes are also aligned with those seen in the total sample (Supplemental Figure 1).
Discussion
Most evaluations of PRS performance focus on disease prediction or measures of association summarizing performance in the overall cohort. While these metrics provide insight into the overall accuracy of PRS or association between PRS and a trait of interest, they do not capture the reliability of individual-level risk assessment across different PRS methods. In this study, we found there is poor-moderate reliability across PRS methods for AD risk prediction. PRS method choice affects who is identified as high or low risk for AD.
All PRS compared here were derived from the same summary statistics; genotypes were called and imputed using the same arrays and references; and variants were filtered based on the same reference panels. In translational settings, there is no standard or guarantee that these parameters will be consistent. Additionally, PRS approaches are rapidly expanding, with methods that differ in SNP selection and effect weighting strategies. These different approaches may vary in performance across population or when applied to different summary statistics. It is highly likely that our comparisons underestimate the extent of variation that would be expected in translational settings.
Unreliable PRS estimates will have tangible consequences. PRS have already been proposed as a tool to improve the efficiency of clinical trials, enabling targeted enrollment to those at high or low risk extremes,25,26 with major pharmaceutical companies now investing in these approaches to potentially reduce trial participant sample sizes and shorten study times. However, unreliable PRS estimates could undermine these efforts by incorrectly excluding or including participants. The psychological impact of unreliable genetic risk prediction is another area of concern, as disclosure of high genetic risk has already been shown to cause anxiety among patients, which would be compounded when risk estimates lack reliability. 27 Furthermore, PRS unreliability has direct implications for emerging reproductive applications, and recent research has demonstrated that patients are highly interested in polygenic risk assessment for AD during preimplantation genetic testing. 28 Beyond clinical applications, genetic risk assessments have long been discussed in insurance underwriting, and PRS have already been shown to offer advantages to life insurance underwriting beyond clinical and demographic factors. 29 This use of PRS already raises ethical concerns about genetic discrimination, and it becomes more alarming when decisions are based on unstable predictions. 30
The lack of reliability in PRS is not restricted to AD. One study that examined the precision of PRS for coronary artery disease also found striking lack of consistency across models. 31 80% of participants classified as high risk (top quintile) for coronary artery disease using one score were also classified as low risk (bottom quintile) by another. Another study of 13 traits in the UK Biobank highlighted substantial individual-level uncertainty in PRS risk stratification, finding that 95% confidence intervals placed individuals anywhere between the 34th to 99th percentile. 32
Given the lack of reliability across methods, PRS for AD and other complex traits must be implemented in clinical and commercial settings with caution and transparency. While it is understandable that standard parameters for PRS will vary depending on the trait, application, and population, we strongly urge that the parameters selected for different models including variants and corresponding weights should be made available when used for downstream applications.
The PRS compared here are non-exhaustive. We focused on PRS that were constructed using the same GWAS summary statistics and found to be associated with dementia in a previous study. Future PRS may be improved by incorporating additional variants, particularly rare variants that often have larger effect sizes. Furthermore, while using PRS alone can be valuable for understanding genetic risk of endophenotypes or specific pathologies, their utility as predictive tools for disease development requires integration with non-genetic information, which we did not include here.
Conclusion
There remains a disconnect between PRS performance at the population level and translation into individual-level assessment. The lack of reliability across methods is a major challenge for translating PRS into a clinically meaningful tool for AD and dementia. Inconsistent individual-level risk assessments undermine confidence in the utility of PRS for guiding prevention, precision interventions, or stratification in clinical trials. We urge those implementing PRS for prediction in research and practice to proceed with caution.
Supplemental Material
sj-docx-1-alz-10.1177_13872877251375098 - Supplemental material for Risk score roulette: A cautionary tale of polygenic risk score reliability
Supplemental material, sj-docx-1-alz-10.1177_13872877251375098 for Risk score roulette: A cautionary tale of polygenic risk score reliability by Diane Xue, Elizabeth E Blue, Alexis C Wood, Jerome I Rotter and Alison E Fohner in Journal of Alzheimer's Disease
Footnotes
Acknowledgements
Genotyping was performed at Affymetrix (Santa Clara, California, USA) and the Broad Institute of Harvard and MIT (Boston, Massachusetts, USA) using the Affymetrix Genome-Wide Human SNP Array 6.0. The authors thank the other investigators, the staff, and the participants of the MESA study for their valuable contributions. A full list of participating MESA investigators and institutes can be found at ![]() .
.
Ethical considerations
Institutional Review Board approval was received from each of the six MESA study sites.
Consent to participate
All participants provided written informed consent.
Author contributions
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by NIA F99AG079792. MESA and the MESA SHARe projects are conducted and supported by the National Heart, Lung, and Blood Institute (NHLBI) in collaboration with MESA investigators. Support for MESA is provided by contracts 75N92020D00001, HHSN268201500003I, N01-HC-95159, 75N92020D00005, N01-HC-95160, 75N92020D00002, N01-HC-95161, 75N92020D00003, N01-HC-95162, 75N92020D00006, N01-HC-95163, 75N92020D00004, N01-HC-95164, 75N92020D00007, N01-HC-95165, N01-HC-95166, N01-HC-95167, N01-HC-95168, N01-HC-95169, UL1-TR-000040, UL1-TR-001079, and UL1-TR-001420, UL1TR001881, DK063491, R01HL105756, and R01AG058969 . Funding for SHARe genotyping was provided by NHLBI Contract N02-HL-64278.
National Institute on Aging, (grant number F99AG079792).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
Data used in this manuscript are from the Multi-Ethnic Study of Atherosclerosis and are available upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
