Abstract
Background
In Alzheimer's disease (AD), soluble tau oligomers are central to neurodegeneration and cognitive decline. Resilient individuals, such as those with non-demented Alzheimer's neuropathology (NDAN) or primary age-related tauopathy (PART), offer critical insights into protective mechanisms against tau-mediated neurodegeneration. NDAN individuals exhibit AD neuropathology without cognitive impairment or neurodegeneration, while PART, characterized by hippocampal- and entorhinal-restricted tau pathology, manifests with minimal-to-no amnestic changes. Brain-derived tau oligomers (BDTO) from these cohorts provide a unique platform to explore molecular pathways underlying both vulnerability and resilience to tau pathology.
Objective
To identify vulnerability- and resilience-associated pathways by comparing BDTO interactomes across AD, NDAN, and PART.
Methods
BDTO were isolated from AD (n = 4; 2M, 2F), NDAN (n = 4; 2M, 2F), and PART (n = 4; 1M, 3F) hippocampal autopsy specimens using co-immunoprecipitation. Proteins were identified via liquid chromatography-tandem mass spectrometry, and non-specific interactors were filtered using SAINTq. Interactome networks and enrichment analyses were performed using Metascape. Findings were cross-referenced with the Neuropro database and existing literature on tangle-associated proteins. Key interactors were validated through reverse co-immunoprecipitation.
Results
A total of 203 proteins were identified, including eight novel interactors not previously linked to AD. All interactomes were enriched in proteins related to tau physiology and lysosomal degradation. NDAN and PART interactomes showed unique enrichment in proteins involved in cellular responses to reactive oxygen species.
Conclusions
One vulnerability-associated and 18 resilience-associated pathways that may mitigate tau-mediated neurodegeneration were identified, laying the groundwork for novel diagnostic and therapeutic strategies targeting pathological tau oligomers.
Introduction
The A/T/N framework for diagnosing neurodegenerative diseases encompasses biomarker reactivity for three critical hallmarks: amyloid-β (A), tau (T), and neuroinflammation and neurodegeneration (N). 1 Alzheimer's disease (AD; A+T+N+) is the most prevalent neurodegenerative disorder, afflicting nearly seven million Americans. 2 AD is characterized by the progressive accumulation of extracellular amyloid-β (Aβ) plaques and intracellular tau neurofibrillary tangles (NFT) in nearly all brain regions; these neuropathological features are accompanied by neurodegeneration and cognitive decline (Figure 1(a)). 2 Non-demented with Alzheimer's neuropathology (NDAN) individuals represent approximately one-third of the aging population (NDAN; A+T+N−).3–5 These individuals, while exhibiting similar Aβ and NFT burden in cortical and sub-cortical brain regions to AD patients, are distinct from AD patients in that NDAN individuals lack substantial neurodegeneration or cognitive decline (Figure 1(a)).3,5,6 The intact cognitive functioning and absence of neurodegeneration, which occur in the face of advanced AD neuropathology, in NDAN individuals further differentiate them from preclinical AD patients and normally-aging individuals. 4 Primary age-related tauopathy (PART; A−T+N±) is a neurodegenerative condition that is distinct from both AD and NDAN. PART is a broadly encompassing term used to describe patients with complete absence (i.e., definite PART) or sparse presence (i.e., probable PART) of senile Aβ plaques, mild to moderate NFT burden localized to the entorhinal cortex and limbic areas, and minimal neurodegeneration with varying cognitive impairment (Figure 1(a)).7–9 AD, NDAN, and PART thus create a continuum of vulnerability and resilience (Figure 1(b)) centered around tau pathology: while AD patients are vulnerable to tau spreading and neurodegeneration, PART patients are resilient to tau spreading, and NDAN individuals are resilient to neurodegeneration.
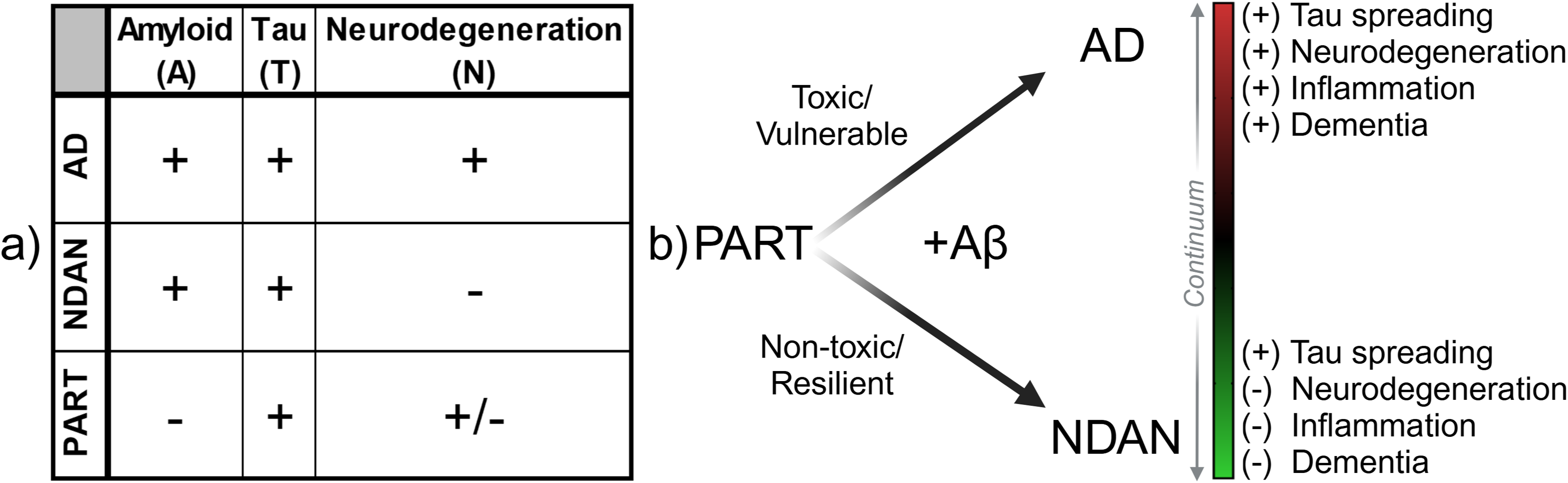
Comparative clinical and histopathological framework of AD, NDAN, and PART. (a) Clinical and histopathological features of Alzheimer's disease (AD), non-demented with Alzheimer's neuropathology (NDAN), and primary age-related tauopathy (PART), as classified by the established A/T/N (Amyloid/Tau/Neurodegeneration) biomarker framework. (b) Conceptual diagram illustrating the continuum of tau pathology extending from PART (intermediate resilience, mixed features) toward either AD (vulnerable/toxic) or NDAN (resilient/non-toxic) states, based on the presence or absence of amyloid-beta (Aβ). Progression along this continuum is associated with differential degrees of tau spreading, neurodegeneration, inflammation, and dementia.
Braak staging quantifies the propagation and regional spread of NFT burden.10,11 This staging remains the strongest pathological correlate for cognitive decline in AD and higher Braak stages are associated with greater cognitive impairment in PART.8,12–15 Preceding NFT formation, the microtubule-associated protein tau aggregates into tau oligomers (tauO) and further aggregates to form paired helical filaments (PHF). The detachment of tau monomers and aggregation into soluble tauO are associated with microtubule instability and disrupted synaptic homeostatic processes, which can ultimately contribute to neurodegeneration and cognitive dysfunction.16–18 Soluble tauO are recognized as one of the most toxic species across multiple neurodegenerative diseases.16–21 Prior studies have characterized tau interactomes of various tau conformations, including total, PHF, and NFT tau, and compared protein enrichment profiles between AD patients and healthy controls.22–28 Total tau interactomes encompass physiological (i.e., monomeric) and pathological (i.e., misfolded, aggregated) tau, making it difficult to discern proteins associated with non-toxic and toxic tau species. PHF and NFT interactomes capture late-stage AD pathology, raising questions regarding the diagnostic and therapeutic utility of the identified proteins. As putative, early drivers of neurodegeneration and cognitive dysfunction, the protein interactome of soluble tauO remains a crucial yet unexplored facet of tau interactome studies. An additional critically lacking dimension of tau interactomes is an understanding of tauO protein interactions in resilient individuals. The existence of individuals, such as those in the NDAN and PART cohorts, suggests that complete removal of toxic neuropathology may not be necessary for effective resilience against neurodegeneration and cognitive dysfunction. Therefore, identifying tauO protein interactions in resilient conditions can provide innovative targets for diagnosing, modulating, or treating AD. This dual focus on both pathological and resilient tauO interactomes can offer a comprehensive understanding of tau pathology and resilience mechanisms to tau pathology.
To address this critical gap, brain-derived tau oligomers (BDTO) were isolated from hippocampal autopsy specimens of individuals with AD, NDAN, and PART via co-immunoprecipitation and subjected to high-resolution protein identification using liquid chromatography-tandem mass spectrometry (LC-MS/MS). Non-specific interactors were identified using IgG isotype controls and rigorously filtered using Significance Analysis of INTeractome (SAINTq). Condition-specific BDTO interactomes were constructed and functional enrichment analyses were performed on the Metascape web platform. These findings were systematically cross-referenced with the Neuropro database and existing literature on tangle-associated proteins, providing a robust framework for comparative analysis. To foster further discovery, we have made our raw proteomic data publicly accessible herein, enabling the scientific community to build upon the molecular network of tau. Key interactors identified in these networks were subsequently validated through reverse co-immunoprecipitation, ensuring the reliability of our results. This integrative approach (Figure 2) enabled the delineation of distinct molecular pathways underlying both vulnerability and resilience to tau-mediated neurodegeneration.
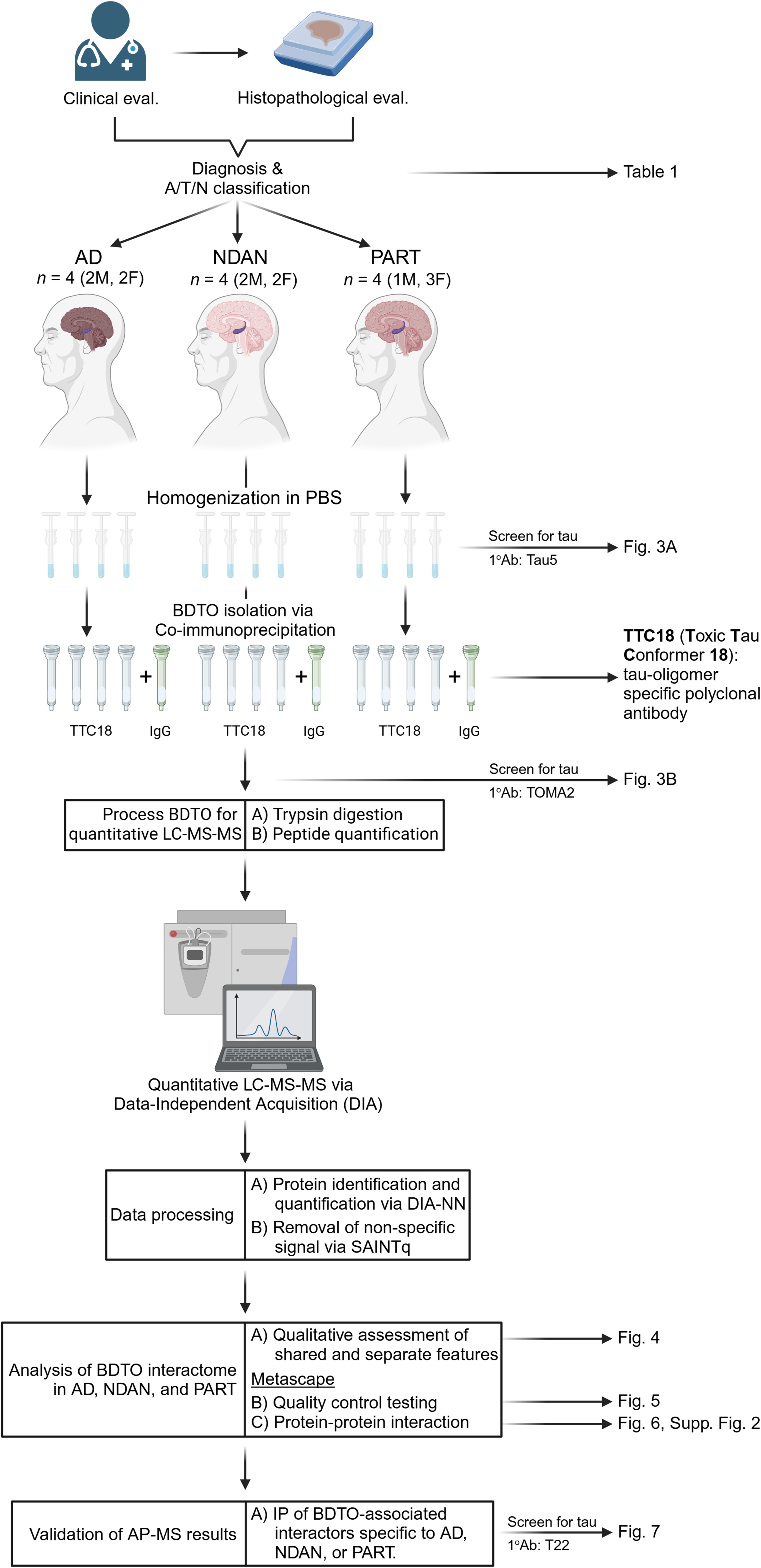
Experimental workflow schematic. Overview of sample selection, brain-derived tau oligomer (BDTO) isolation, and subsequent proteomic analysis, culminating in a multi-faceted, data-driven evaluation of disease-specific protein signatures. Created with BioRender.com.
Methods
Human subjects and autopsy brain tissue
All specimens used in the current study were evaluated by expert neuropathologists who utilized objective and standardized criteria to reach their diagnoses. Male and female AD patients and NDAN individuals provided informed consent and enrolled at the Layton Aging and AD Center (ADC) at Oregon Health and Science University (OHSU). PART patients were enrolled at the Alzheimer's Disease Research Center (ADRC) at the University of Kansas (UK). All protocols were approved by the Institutional Review Board (IRB) at OHSU and UK, respectively. ADC and ADRC de-identified all donor subject samples before providing them to the University of Texas Medical Branch (UTMB); therefore, no approval was required from the UTMB IRB per CFR §46.101(a). In sum, there are no human participants in this article and informed consent is not required. The presence or absence of Aβ pathology differentiates possible and definite PART, respectively.7,29 Prior work from our laboratory did not detect pathological Aβ in synaptosomes isolated from the UK PART cohort, thus suggesting a definite PART classification. 30 Each subject received annual neurological and neuropsychological evaluations by experienced clinicians. Patient diagnosis, age at death, sex, postmortem index (PMI), brain region, Braak stage, and mini-mental state exam (MMSE) scores (when available) are provided in Table 1. CERAD scores and Thal stages were requested and unavailable for AD patients, NDAN individuals, and PART patients.
Pathological classification and patient information of brain autopsy specimen utilized.
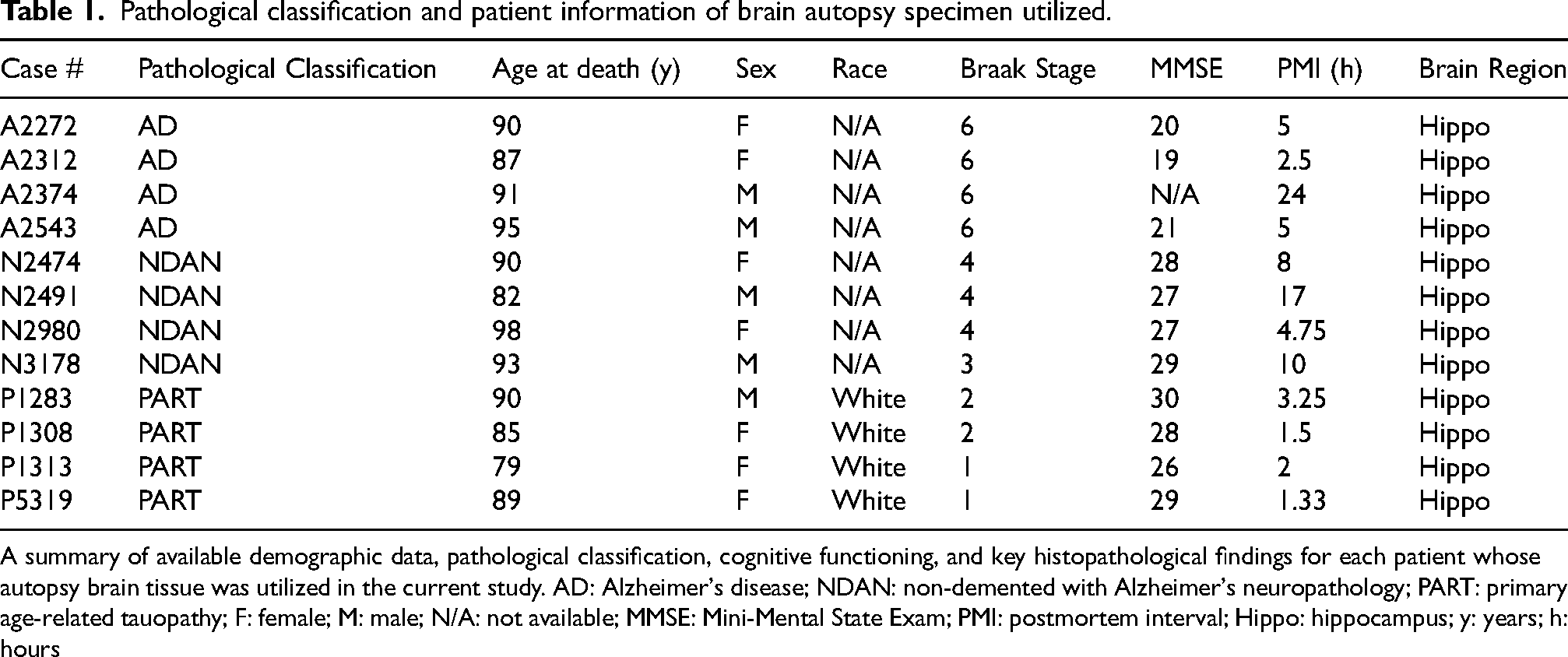
A summary of available demographic data, pathological classification, cognitive functioning, and key histopathological findings for each patient whose autopsy brain tissue was utilized in the current study. AD: Alzheimer's disease; NDAN: non-demented with Alzheimer's neuropathology; PART: primary age-related tauopathy; F: female; M: male; N/A: not available; MMSE: Mini-Mental State Exam; PMI: postmortem interval; Hippo: hippocampus; y: years; h: hours
Isolation of brain-derived tau oligomers
Preparation of PBS-soluble brain homogenates. BDTO were prepared from fresh frozen hippocampal autopsy brain specimens of AD patients (n = 4; 2M, 2F), NDAN individuals (n = 4; 2M, 2F), and PART patients (n = 4; 1M, 3F). Low retention plasticware, including tips and tubes, was used to minimize sample loss. All steps for BDTO isolation were performed on ice or at 4°C unless otherwise specified.
Fresh frozen hippocampal specimen (75 mg) from each patient or individual was gently homogenized in a Dounce homogenizer (10–15 gentle strokes until no visible tissue chunks) in ice-cold Dulbecco's phosphate-buffered saline (DPBS; 2.68 mmol KCl, 1.47 mmol KH2PO4, 136.89 mmol NaCl, 8.10 mmol Na2HPO4 (anhydrous); Corning, cat. 21–031-CV), supplemented with protease (Roche, cat. 11836153001) and phosphatase inhibitors (Roche, cat. 490683700), at a 1:5 ratio (w/v) of brain:PBS+protease+phosphatase buffer (henceforward referred to as PBS lysis buffer). Brain homogenates were transferred to a 1.5 mL microcentrifuge tube and centrifuged at 9,168x g for 10 min at 4°C. The supernatant (i.e., PBS-soluble homogenate) was carefully removed and either immediately used for subsequent co-immunoprecipitation or stored at −80°C.
Co-immunoprecipitation. BDTO were isolated via co-immunoprecipitation (co-IP) using the Pierce™ Co-Immunoprecipitation Kit (ThermoFisher, cat. 26149). The primary modification was substitution the of IP lysis/wash buffer with 1X PBS lysis buffer to minimize protein complex disruptions by detergents, thereby preserving the integrity of the isolated interactomes. Briefly, spin columns were prepared by coupling 75.0 µg of TTC18 (toxic tau conformation 18; prepared in-house by Dr Rakez Kayed's laboratory) antibody with the AminoLink Plus Coupling Resin, as directed by the manufacturer's protocol. Four tau antibodies were screened for tauO immunoprecipitation efficiency, and TTC18 provided the greatest tauO enrichment (Supplemental Figure 1). Spin columns for isotype controls were prepared in parallel using rabbit IgG (Millipore, cat. 12-370). Prior to resin quenching, the antibody coupling efficiency for each column was verified by measuring IgG A280 (Nanodrop 2000c; ThermoFisher Scientific, Wilmington, DE, USA).
The resin-antibody mixtures were equilibrated with PBS lysis buffer and incubated with equal amounts of individual brain homogenates overnight at 4°C. Following incubation, the resin-antibody-homogenate mixtures were centrifuged, and the flowthroughs were collected and stored at −80°C for subsequent analyses. Spin columns were then washed three times with PBS to reduce non-specific interactors. BDTO were eluted twice in 50 µL of elution buffer and immediately neutralized with 1.0 M Tris (pH 8.0), as per protocol recommendations. Post-elution, BDTO samples were placed on ice for 30 min to promote meta-stability of BDTO conformers. Resin-antibody mixtures were regenerated by washing twice and stored in diluted coupling buffer at 4°C.
Western blotting
PBS-soluble brain homogenates and BDTOs were analyzed using conventional western blotting techniques, as previously described.5,31–34 Key technical steps for optimized detection of each sample type are outlined below.
Proteins in the PBS-soluble homogenates were quantified in duplicate using the Bicinchoninic Acid assay (BCA) (ThermoFisher, cat. 23227). Samples were prepared in 1X NuPAGE™ LDS Sample Buffer (ThermoFisher, cat. NP0007) and 1X NuPAGE™ Sample Reducing Agent (ThermoFisher, cat. NP0004), then denatured at 75°C for 10 min. Pre-cast NuPAGE 4–12% Bis-Tris SDS-PAGE gels (ThermoFisher, cat. NP0323BOX) were loaded with 10.0 µg protein per well and underwent conventional wet-tank membrane transfer (100V for 1 h at 4°C). For total homogenates, fluorescence-based immunodetection was employed due to the flexibility of multi-channel detection. Proteins were transferred onto a nitrocellulose membrane (Cytiva, cat. 10600001). Prior to blocking, membranes were stained with Revert™ 700 Total Protein Stain (LI-COR Biotech, cat. 926-11011) and imaged for normalization. The membrane was blocked in Intercept® (TBS) Blocking Buffer (LI-COR, cat. 927-60001) for 1 h at room temperature. Primary and IRDye-conjugated secondary antibodies were prepared in Intercept® T20 (TBS) Antibody Diluent (LI-COR, cat. 927-65001). Membranes were incubated with the primary antibody overnight at 4°C and the secondary antibody for 1 h at room temperature, both with gentle rocking. Post-incubation washes were performed four times for 7 min each at room temperature in TBS-T (0.1% Tween-20 in 1X TBS). Membranes were imaged using the Odyssey LI-COR Classic Infrared imaging system (LI-COR Biosciences, Lincoln, NE, USA).
To assess Aβ pathology, RIPA-soluble hippocampal homogenates were also analyzed. Fresh frozen hippocampal tissue was mechanically lysed as described for PBS-soluble homogenates, except RIPA buffer (ThermoFisher, cat. 89900) was used in place of PBS. To extract insoluble Aβ species, samples were sonicated on ice for five 15-s bursts (30% amplitude), with 1-min rest intervals. This was followed by vortexing on ice every 5 min for a total of 30 min. Homogenates were then centrifuged at 14,000g for 10 min at 4°C, and the supernatant (i.e., RIPA-soluble fraction) was collected for western blot analysis or stored at −80°C. For detection of Aβ monomer and oligomers, samples were prepared under reducing and denaturing conditions in 1X Tricine SDS Sample Buffer (ThermoFisher, cat. NP0007), loaded onto pre-cast Novex 16% Tricine SDS-PAGE gels (ThermoFisher, cat. EC6695BOX) and transferred to membrane using wet-tank transfer at 80 V for 45 min at 4°C. Immunoblotting was performed using the 6E10 monoclonal antibody (1:1000, BioLegend, cat. 803002) as the primary probe.
Due to limited sample availability, BDTO protein concentration was measured using the fluorometric peptide assay (ThermoFisher, cat. 23290) and confirmed by relative A280 absorbance measures (Nandrop 2000c, ThermoFisher Scientific, Wilmington, DE, USA). BDTO samples were prepared in 1X NuPAGE™ LDS Sample Buffer (ThermoFisher, cat. NP0007) without reduction or denaturation to preserve conformational stability. Pre-cast NuPAGE 4–12% Bis-Tris SDS-PAGE gels were loaded with 10.0 µg BDTO per well and transferred using conventional wet-tank membrane transfer (100V for 1 h at 4°C). For co-immunoprecipitated samples (i.e., BDTO and IgG eluates), enhanced chemiluminescence (ECL) detection was utilized to detect low-abundance proteins. Proteins were transferred onto a polyvinylidene fluoride (PVDF) membrane (Millipore Sigma, cat. ISEQ00010) and blocked in 3% BSA in TBS-T for 1 h at room temperature. Primary and HRP-linked secondary antibodies were prepared in 1% BSA in TBS-T. Membranes were incubated with the primary antibody overnight at 4°C and the secondary antibody for 1 h at room temperature, both with gentle rocking. Following antibody incubation, membranes were washed four times for 7 min each at room temperature in TBS-T. Membranes were exposed to regular (Cytiva, Amersham™ ECL Regular, cat. RPN2209), enhanced (ThermoFisher, Pierce™ ECL Plus, cat. 32132), or super-enhanced (ThermoFisher, SuperSignal™ West Atto, cat. A38554) HRP substrates, and developed on Amersham hyperfilms (Cytiva, cat. 28906842). Substrate selection, exposure, and development times were optimized for each membrane to achieve the best signal-to-noise ratio.
Primary antibodies and their respective working concentrations: 6E10 (1:1000, BioLegend, cat. 803002), TOMA2 (1:1000, courtesy of Dr Rakez Kayed); T22 (1:2000, courtesy of Dr Rakez Kayed); Tau5 (1:1000, Abcam, cat. ab80579); CCT7 (1:5000, ThermoFisher, cat. A304-730A); S100β (1:5000, Abcam, cat. ab52642).
Secondary antibodies and their respective working concentrations: HRP-linked IgG anti-mouse (1:5000, Cell Signaling, cat. 7076S); HRP-linked IgG anti-rabbit (1:5000, Cell Signaling, cat. 7074P2); IRDye® 680RD IgG anti-mouse (1:10000, LI-COR, cat. 926-68070); IRDye® 800CW IgG anti-rabbit (1:10000, LI-COR, cat. 926-3221).
Protein abundance was quantified using ImageJ FIJI software (https://imagej.nih.gov/ij, NIH, Bethesda, MD, USA), with individual values normalized to the Total Protein signal.
Liquid chromatography-tandem mass spectrometry (LC-MS/MS) of BDTO interactomes
Proteomic sample digestion. Individual samples were prepared similarly as previously described. 35 Briefly, protein samples were adjusted to a volume of 50 µL with a composition of 5% SDS and 50 mM TEAB (pH 7.1) and were reduced by 10 mM Tris (2-carboxyethyl)phosphine (TCEP) (ThermoFisher, cat. 77720); the samples were incubated at 65°C for 30 min. The sample was then cooled to room temperature and 1 μL of 0.5 M iodoacetamide acid was added and allowed to react for 30 min in the dark. Then, 2.7 μl of 12% phosphoric acid was added to the 52 μL protein solution followed by 165 μL of binding buffer (90% Methanol, 100 mM TEAB (pH 8.5)). The resulting solution was administered to an S-Trap spin column (Protifi, Farmingdale, NY) and passed through the column via centrifugation. The spin column was washed twice with 150 μL of 50% methyl-tert-butyl ether with 50% methanol and then twice with 150 µL of the binding buffer; each solution addition was followed by centrifugation. Six hundred nanograms of trypsin (Promega, cat. V5280) were added to the protein mixture and incubated overnight at 37°C. Peptides were eluted twice with 75 μL of 50% acetonitrile, 0.1% formic acid. The resultant peptides were quantified by a fluorometric peptide assay (ThermoFisher, cat. 23290), and peptide quantities corresponding to 230 ng were then dried in a speed vacuum at room temperature for 1.5 h and resuspended in 10 μL of 2% acetonitrile, 0.1% formic acid, 97.9% water. The samples were then aliquoted into autosampler vials, and a pooled sample was created from equal volumes of peptides from all samples.
NanoLC MS/MS analysis. Peptide mixtures were analyzed by nanoflow liquid chromatography-tandem mass spectrometry (nanoLC-MS/MS) using a nano-LC chromatography system (UltiMate 3000 RSLCnano, Dionex), coupled on-line to a Thermo Orbitrap Fusion mass spectrometer (ThermoFisher, San Jose, CA) through a nanospray ion source. A trap-and-elute method was used with an Acclaim PepMap (100 µm×2 cm, 5 µm; ThermoFisher, cat. 164750) as the trap column and an Acclaim PepMap (75 µm×2 cm, 3 µm; ThermoFisher, cat. 164569) equipped with a stainless-steel nano-bore emitter (ThermoFisher, cat. ES542) as the analytical column. After equilibrating the column in 98% solvent A (0.1% formic acid in water) and 2% solvent B (0.1% formic acid in acetonitrile (ACN)), 4 µL of the samples (92 ng) were injected and eluted by gradient elution as follows (300 nl/min unless stated otherwise): isocratic at 2% B, 0 to 5 min; 2 to 4% B, 5 to 6 min; 4 to 24% B, 6 to 86 min; 24 to 44% B, 86 to 93 min; 44 to 90% B, 93 to 95 min; isocratic at 90% B, 95 to 96 min; 90 to 10% B, 96 to 98 min; isocratic at 10% B, 98 to 99 min (increase flow to 450 nl/min); 10 to 90% B, 99 to 102 min (increase flow to 600 nl/min); isocratic at 90% B, 102 to 105 min; 90 to 2% B, 105 to 107 min; isocratic at 2% B, 107 to 117 min; isocratic at 2% B, 117 to 118 min (decrease flow to 450 nl/min); isocratic at 2% B, 118 to 119 min, (decrease flow to 300 nl/min); and isocratic at 2% B until 120 min.
The Thermo Orbitrap Fusion acquired six gas-phase fractions of the pooled sample using 4 Da fully staggered windows in 100 m/z increments from 400–1000 m/z as previously described. 36 Individual sample data were acquired using 16 Da fully staggered windows from 400–1000 m/z. All LC-MS/MS data were acquired in positive ion mode using data-independent acquisition (DIA). The survey scans (m/z 390–1010) were acquired in the Orbitrap at 60,000 resolution (at m/z = 400) in centroid mode, with the maximum injection time set to auto and an AGC target of 100,000 ions. The S-les RF level was set to 60. Isolation was performed in the quadrupole, and HCD MS/MS acquisition was performed in profile mode using the Orbitrap at a resolution of 30,000 using the following settings: collision energy = 33%, max injection time set to auto, and AGC target of 500,000 ions.
Bioinformatics. Raw data from each sample was demultiplexed to mzML with 10 ppm accuracy after peak picking in MSConvert. 37 The resulting mzML files were searched in MSFragger and quantified via DIA-NN (https://github.com/vdemichev/DiaNN) using the following settings: peptide length range 7–50, protease set to Trypsin, two missed cleavages, three variable modifications, clip N-term M on, fixed C carbamidomethylation, variable modifications of methionine oxidation and N-terminal acetylation, MS1 and MS2 accuracy set to 20 ppm, 1% FDR, and DIA-NN quantification strategy set to Robust LC (high accuracy). The files were searched against a human database acquired from UniProt (18 December 2023). The gas-phase fractions were used to generate the spectral library, which was used to analyze the individual samples.
Statistical analysis was performed using Fragpipe-Analyst using an R script based on the ProteinGroup.txt file produced by DIA-NN. 38 First, contaminant proteins, reverse sequences, and proteins identified ‘only by site’ were filtered out. In addition, proteins identified by a single peptide and not identified or quantified consistently in the same condition were removed. The DIA data was converted to a log2 scale, grouped by samples and conditions, and missing values were not imputed. Protein-wise linear models combined with empirical Bayesian statistics were used for the differential expression analyses. The limma package from R Bioconductor generated a list of differentially expressed proteins for each pair-wise comparison. A cutoff of 0.05 for the adjusted p-value (Benjamini-Hochberg method) along with an absolute log2 fold change of 1 was applied to determine significantly regulated proteins in each pairwise comparison. The raw proteomic data for each sample is provided as Supplemental Material (Supplemental File 1).
Significance analysis of INTeractome (SAINTq)
Removal of nonspecific protein interactors is critical towards accurate interpretation of BDTO protein interactomes. To achieve this, we employed the SAINT software,39,40 a probabilistic model that identifies high-confidence protein interactors from affinity purification–mass spectrometry experiments by incorporating spectral counts or protein intensities from negative controls. Specifically, we utilized SAINTq, the latest iteration of the SAINT software package, which supports DIA analysis and allows for data compression. 40 In this study, protein intensity DIA data were analyzed, and protein interactions were deemed significant with a false discovery rate (FDR) of less than 0.05. Control intensities were not normalized, and the compression parameter was set to two for both controls and replicates. 40
Survey of prior human tau proteomic studies
To contextualize our findings within the existing knowledge of tau-associated proteins, we compared our BDTO interactome with previously published studies through a systematic literature survey and data mining of earlier proteomics research. A systematic search was performed using the PubMed Advanced Search Builder with the following keywords and Boolean operator combination: ((tau interactome) OR (MAPT interactome)). The filter ‘Humans’ was applied, returning 56 studies. After a thorough evaluation, the work by Kavanagh and colleagues (2022) 41 was particularly informative due to their detailed compilation of tau interactome and proteomic studies. In addition to the studies identified through PubMed, individual studies from this compilation were extracted for further screening and analysis.
To ensure translational relevance and reliable interpretation, we screened the results hierarchically. First, only studies directly interrogating human biological specimens were included; those using humanized cell or animal tau models were excluded. Second, only studies examining brain tissue from human adults were included; studies involving non-brain tissues (e.g., blood, plasma, CSF) or fetal samples were excluded. Third, studies focusing on specific cellular sub-compartments (e.g., nuclear, mitochondrial, synaptic) were also excluded.
Our survey yielded outcomes in three categories: a) total tau interactome, b) PHF tau interactome, and c) NFT tau localized proteomics. No prior proteomic studies have been conducted in the context of NDAN or PART, limiting the interpretation of our findings to AD tau pathology. Investigations focusing on the AD total tau interactome were excluded for two reasons: a) total tau includes both physiological/non-toxic and pathological/toxic conformations, complicating the interpretation of interacting proteins; and b) studies on total tau interactomes define differential protein enrichment associated with AD tau compared to non-demented controls, differing from our enrichment parameters. The final list of studies (n = 5) fulfilling these criteria is summarized in Table 2.
Summary of prior PHF interactome and NFT proteomic studies.
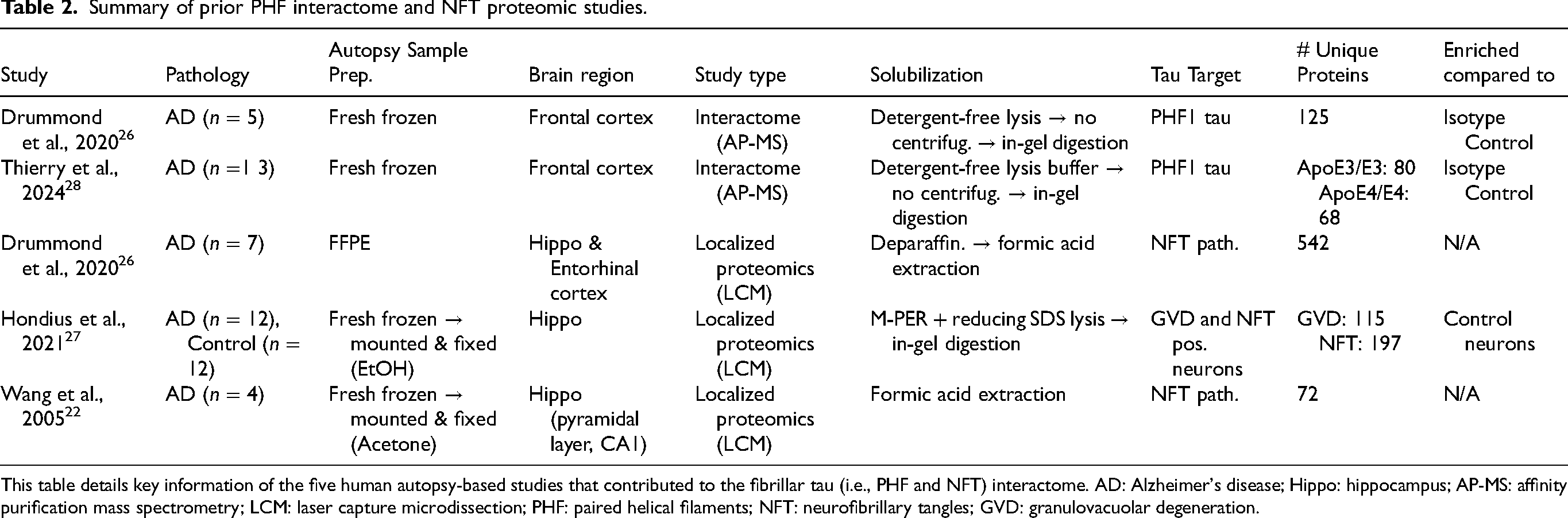
This table details key information of the five human autopsy-based studies that contributed to the fibrillar tau (i.e., PHF and NFT) interactome. AD: Alzheimer's disease; Hippo: hippocampus; AP-MS: affinity purification mass spectrometry; LCM: laser capture microdissection; PHF: paired helical filaments; NFT: neurofibrillary tangles; GVD: granulovacuolar degeneration.
We adopted the vote-count system used by Kavanagh and colleagues (2022) 41 to evaluate and integrate bona fide tau protein interactors across independent investigations. We merged findings from the PHF interactome (two studies)26,28 and NFT localized proteomics (three studies)22,26,27 for two reasons. First, PHFs are the primary constituents of NFTs, 42 collectively reflecting late-stage ‘fibrillar’ tau species. Second, merging studies increases the sampling pool, enhancing the reliability of identified proteins. Proteins present in the majority (>60%), or at least three of the five reports, were considered bona fide tau interactors. Proteins within the BDTO interactome not previously associated with the human AD proteome were classified as novel tau interactors, identified using the 'Multi-Protein Query' function in NeuroPro, 43 an extensive online repository for human AD proteomic studies.
Metascape Custom Analysis
The Custom Analysis feature was utilized to control the CAME (i.e., Conversion, Annotation, Membership, and Enrichment) process in Metascape. 44 No changes were made in the Conversion or Membership tabs; all terms were unselected in the Annotation tab, and one GO annotation (i.e., cellular component, biological processes, and molecular function) was selected at a time in the Enrichment tab. The human brain was set as the background protein set to compare significantly enriched terms and pathways in AD BDTO versus PHF and NFT (hereafter referred to as fibrillar tau) interactomes. When comparing the three BDTO interactomes (AD versus NDAN versus PART), the human hippocampal formation was set as the background protein set.
Enrichment analyses. Hairball networks (Figure 6) were created by providing all protein interactors within the fibrillar tau, and AD, NDAN, and PART BDTO interactomes. Protein-protein interaction (PPI) networks were isolated from the hairball network and underwent enrichment analysis using Metascape's Molecular Complex Detection (MCODE) algorithm; the MCODE algorithm allowed the assignment of specific biological roles to each PPI network. 44
Validation studies: reverse co-immunoprecipitation
In order to validate specific protein interactions identified in the BDTO interactomes, reverse co-IPs of CCT7 from NDAN and PART hippocampal PBS-soluble homogenate and S100β from AD hippocampal PBS-soluble homogenate were performed. Reverse co-IPs were performed in the same manner as described above (see Co-immunoprecipitation), using the hippocampal PBS-soluble homogenates pooled in equal amounts from the AD (n = 4), NDAN (n = 4), and PART (n = 4) samples used for BDTO isolation. Resins were prepared with 20 μg of anti-CCT7 (ThermoFisher, cat. A304-730A), anti-S100β (Abcam, cat. ab52642), or rabbit IgG isotype control (Millipore, cat. 12-370) antibodies. To confirm successful immunoprecipitation, the reverse co-IP samples were probed for CCT7 (1:5000, ThermoFisher, cat. A304-730A) and S100β (1:5000, Abcam, cat. ab52642) via western blots, as detailed in the Western blotting section. Confirmation of BDTO co-IP was completed by probing the reverse co-IP samples with T22 (1:2000, courtesy of Dr Rakez Kayed's laboratory).
Software for general analysis, figures, and illustrations
GraphPad Prism 10.0 was used to depict the MAPT abundances in AD, NDAN, and PART BDTO relative to their IgG isotype control samples. Adobe Illustrator and BioRender were used to visualize the PPI networks better and create the experimental workflow schematic, respectively.
Results
Isolation and confirmation of brain-derived tau oligomers (BDTO)
The neuropathological characteristics and classification, as determined by expert neuropathologists, and key demographic data of the corresponding patient and individual specimens were carefully assessed prior to inclusion in this study (Table 1). The absence of pathological Aβ in synaptosomes isolated from the same PART specimens analyzed in this study 30 supports their classification as definite PART cases.7,29 To further confirm the Aβ status of each group, western blotting was performed on pooled RIPA-soluble hippocampal homogenates, where a clear band at the expected molecular wright of Aβ (∼4 kD) was detected in AD and NDAN but not in PART samples (Supplemental Figure 2), providing additional biochemical confirmation of the absence of overt Aβ pathology in PART. The average Braak stage of each condition aligns well with previously reported NFT burden (AD: M = 6.00, SD = 0.00; NDAN: M = 3.75, SD = 0.50; PART: M = 1.50, SD = 0.58).2,3,6,30–32 AD patient MMSE scores (M = 18.75, SD = 2.63) were consistent with a moderate AD diagnosis (i.e., cognitive decline, executive dysfunction, behavioral changes, etc.), whereas the MMSE scores of NDAN individuals (M = 27.75, SD = 0.96) and PART patients (M = 28.25, SD = 1.71) indicated intact cognitive function. 45 No significant differences were observed in age at death (AD: M = 91.75, SD = 3.95; NDAN: M = 90.75, SD = 6.70; PART: M = 85.75, SD = 4.99) or postmortem interval (AD: M = 6.63, SD = 5.06; NDAN: M = 9.94, SD = 5.18; PART: M = 2.02, SD = 0.87).
BDTO were isolated via co-IP from PBS-soluble homogenates of AD (n = 4; 2M, 2F), NDAN (n = 4; 2M, 2F), and PART (n = 4; 1M, 3F) hippocampal autopsy specimens (Table 1). TTC18, an anti-tauO antibody, was utilized for the co-IP as it was the most efficient at tauO immunoprecipitation (Supplemental Figure 1). To determine non-specific components of respective BDTO interactomes, eluates from IgG isotype control IP were assayed for each condition. The presence of BDTO were confirmed at two critical stages in the experimental workflow: following a) homogenization (Figure 3(a)) and b) co-immunoprecipitation (Figure 3(b)). Initial screening of individual PBS-soluble total homogenate fractions probed for total tau (Ab: Tau5) revealed oligomeric, high molecular weight (HMW) tau (i.e., 75 to 250+ kD) in all three conditions (Figure 3(a)). Following co-immunoprecipitation, BDTO eluates exhibited significant enrichment of oligomeric tau species (Figure 3(b); Ab: TOMA2), with only minimal detection of non-specific proteins observed in IgG isotype controls (Supplemental Figure 3b: Total Protein Stain). Due to limited sample availability, BDTO preparations with the highest protein concentrations from each condition (Supplemental Table 1) were selected for this query.
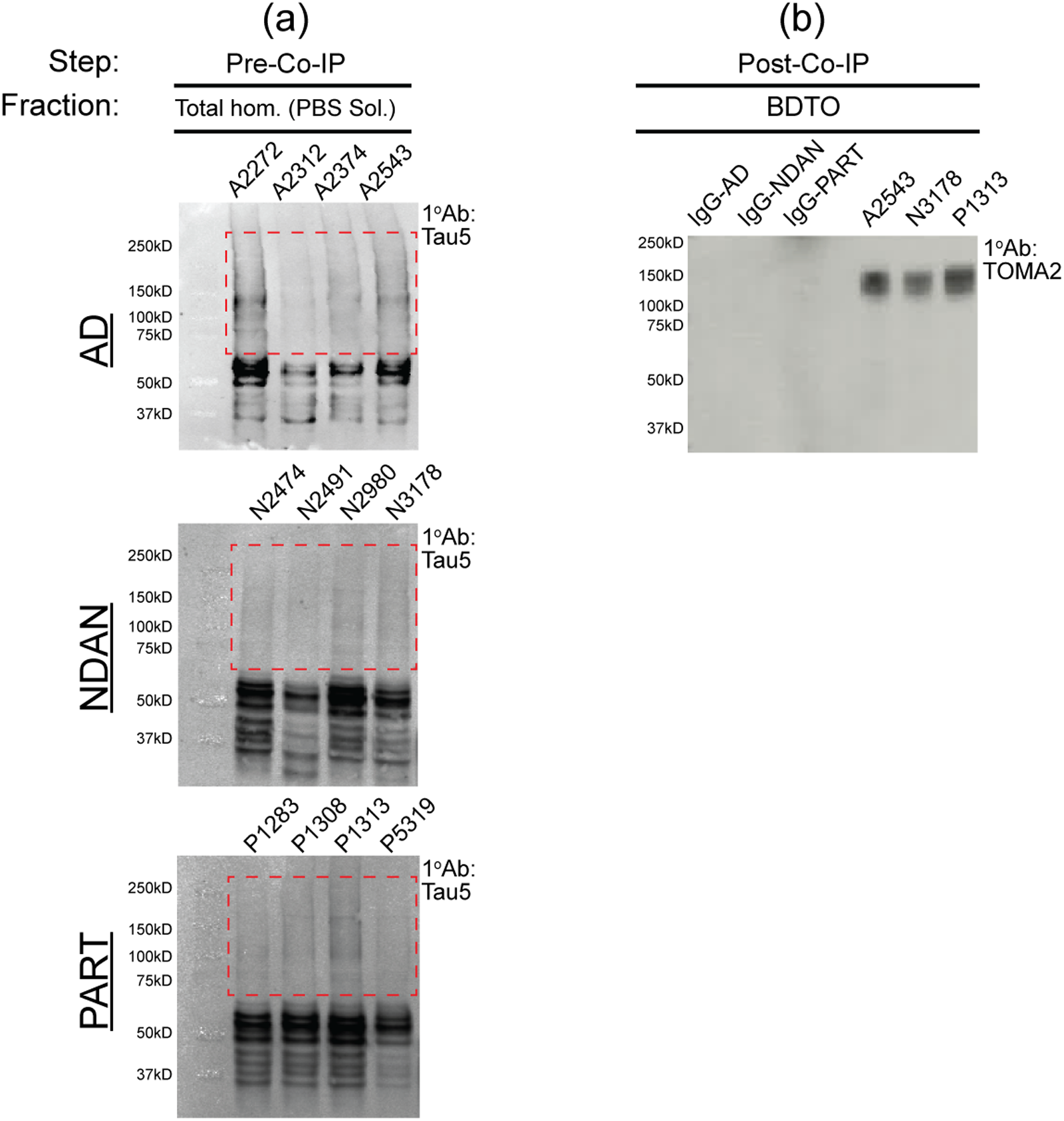
Sequential tau screening across stages of BDTO isolation. The experimental step and corresponding brain tissue fraction are specified above each panel. (a) PBS-soluble homogenates from AD, NDAN, and PART hippocampal specimens (n = 4 each) were screened for total tau (Ab: Tau5) via western blot analysis, with lanes labeled by sample ID as detailed in Table 1. (b) Subsequent analysis on co-immunoprecipitated BDTO fractions were performed using the TOMA2 antibody to probe tauO, focusing on the sample with the highest protein concentration (Supplemental Table 1). Total protein staining for the respective immunoblots are shown in Supplemental Figure 3.
SAINTq analysis and distribution of BDTO interactors
Following liquid chromatography-tandem mass spectrometry, or LC-MS/MS, with data-independent (DIA) acquisition for each sample (12 BDTO and 3 IgG isotype controls), 789 unique proteins were identified across all the samples. Control eluates were used to quantify and remove non-specific features of the BDTO interactome by SAINTq. 40 After implementing SAINTq, 203 specific proteins were identified. The AD, NDAN, and PART BDTO interactomes contained 69, 102, and 154 proteins, unique and shared, respectively (Figure 4). The number of unique proteins was nearly three times greater in the PART BDTO interactome (npr = 64), as compared to the AD (npr = 20) and NDAN (npr = 22) BDTO interactomes. NDAN and PART BDTO interactomes shared 47 proteins, AD and PART BDTO interactomes shared 16 proteins, and AD and NDAN BDTO interactomes shared 6 proteins; all three interactomes shared 27 proteins. Specific BDTO interactors were cross-referenced with published AD brain proteomics studies using the NeuroPro online database;43,46 195/203 proteins (96%) have been reported previously. Eight proteins identified across the three interactomes have not been previously reported in AD proteomics studies (yellow highlighted proteins in Figure 4). The raw data generated from the LC-MS/MS and used for analysis is available as supplemental data (Supplemental File 1).
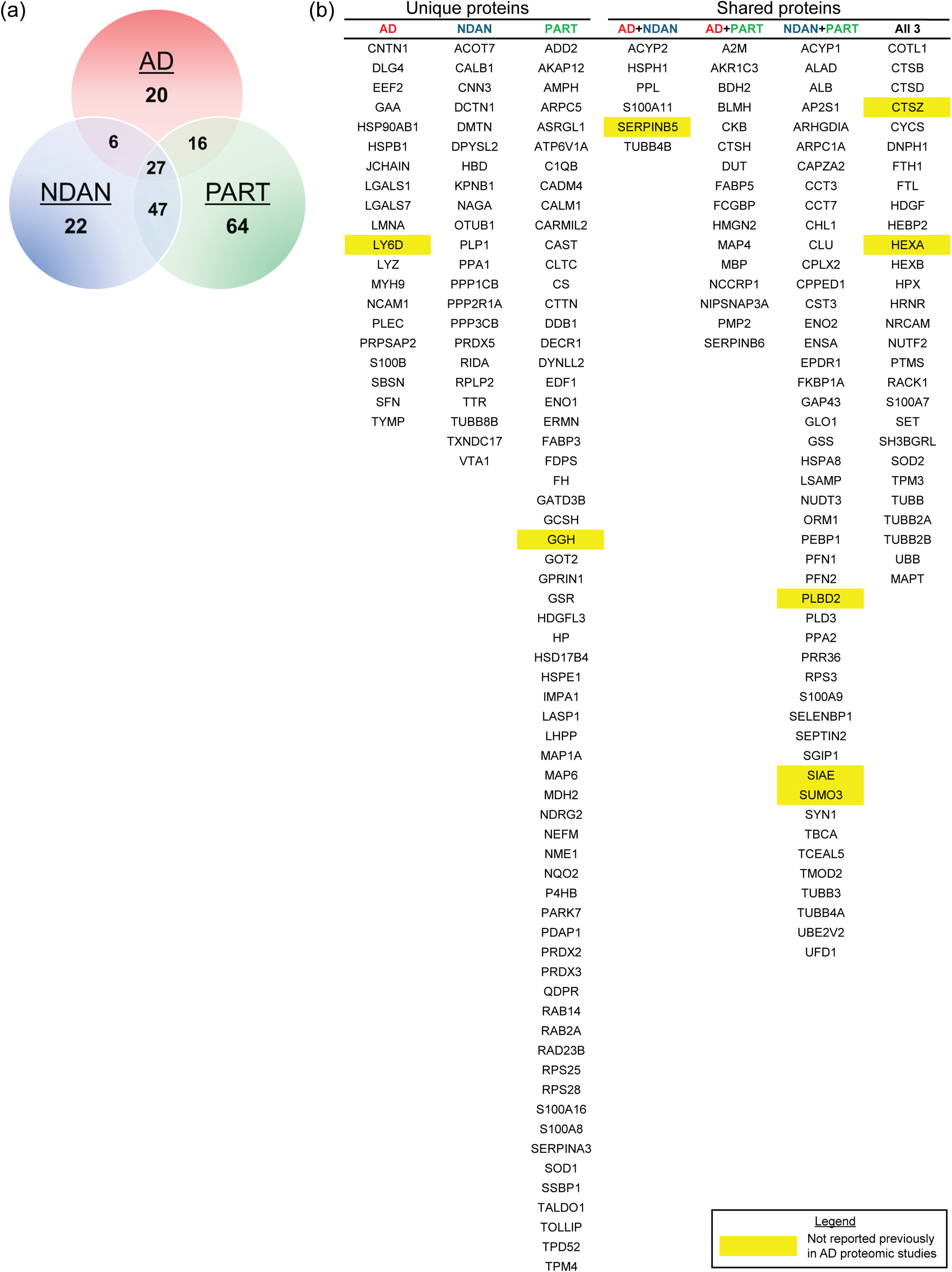
Specific proteins inform differential BDTO interactomes in AD, NDAN, and PART. (a) Venn diagram depicting the unique and shared proteins across BDTO interactomes, with AD (red), NDAN (blue), and PART (green), illustrating the proteomic diversity inherent to each condition. (b) Accompanying lists detail the specific proteins enumerated in the Venn diagram. All proteins were cross-referenced with the NeuroPro database and individual AP-MS experiments related to Alzheimer's disease. Proteins highlighted in yellow represent novel tau interactors not previously identified in any referenced sources.
Following SAINTq, BDTO interactomes were evaluated for quality control via DisGeNET pathway enrichment analysis, the largest database for known human gene-disease associations. The three BDTO interactomes were enriched in many putative neurodegenerative disease phenotypes (e.g., Lewy body disease, senile plaques, familial and sporadic amyotrophic lateral sclerosis, Down syndrome, motor neuron diseases, etc.) (red asterisks in Figure 5). Interestingly, this analysis clustered NDAN and PART BDTO interactomes separate from the AD BDTO interactome. Furthermore, of the three primary clusters created by DisGeNET, one cluster was enriched only in proteins from NDAN and PART interactomes. All three parent clusters contained neurodegenerative disease phenotypes, including tauopathies (first cluster), presenile dementia (second cluster), and Pick disease of the brain (third cluster).
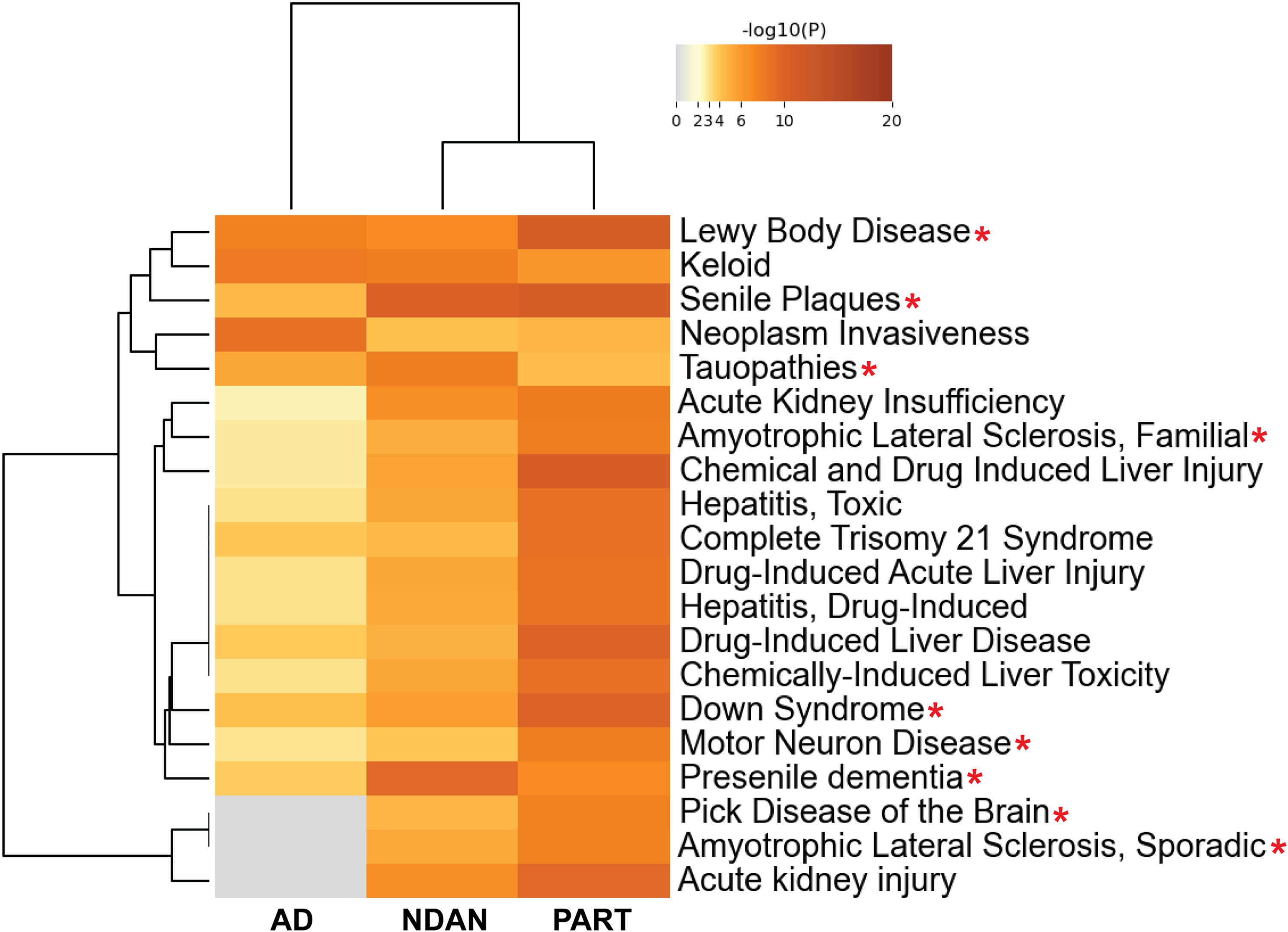
DisGeNET pathway analysis of BDTO-associated proteins reveals enrichment of relevant neurodegenerative disease phenotypes. Cluster heatmap generated from DisGeNET ontologies showcases enriched neurodegenerative disease pathways across AD, NDAN, and PART BDTO. Tau pathology-associated terms/conditions/pathways are indicated with a red asterisk. Selected terms meet defined significance thresholds (p-value < 0.01, minimum count ≥ 3, enrichment factor > 1.5) and are organized into clusters based on kappa similarity scores (> 0.3), with each row representing the most significant term of a cluster. The color-coded p-value scale at the top guides the interpretation of enrichment levels, with gray indicating non-significance. Created in Metascape. 44
In sum, the presence of BDTO was confirmed at key stages of the experimental workflow. The SAINTq algorithm was successfully applied to generate a list of statistically significant, specific proteins within AD, NDAN, and PART BDTO interactomes. Subsequent analysis revealed that proteins within the respective interactomes were enriched in neurodegenerative disease phenotypes and displayed unique distributions across the three conditions. Notably, the PART BDTO interactome had the highest number of unique proteins, while NDAN and PART BDTO interactomes showed the greatest overlap.
Overview of the BDTO proteome
As this is the first interactome analysis of its kind, we aimed to identify proteins that interact with BDTO irrespective of clinical condition. Twenty-seven of the 203 specific proteins were shared across all three interactomes. Twenty-five of the 27 proteins (∼92%) were previously reported to be altered in AD brain proteomics studies.43,46 Shared proteins underwent enrichment analysis using the three GO annotations: cellular component, biological processes, and molecular function. Proteins enriched in all three BDTO interactomes were associated with microtubules (red MCODE1) and lysosomes (blue MCODE2) and involved in microtubule dynamics (MCODE1) (e.g., microtubule organization; GTP phosphatase and binding activity) (Supplemental Figure 4 and Supplemental Table 2). These results are consistent with tau's primary function as a microtubule stabilizer and suggest BDTO engage one of the two primary cellular degradation systems.
Enrichment analysis comparison of tau interactomes based on different structural conformations
A key goal of our study was to understand how the structural conformation of pathological tau (oligomeric versus fibrillar) influences its protein interactome. We compared the AD BDTO interactome (evaluated here) to those of fibrillar (i.e., PHF and NFT) tau. Using a systematic literature survey, we identified five comparable studies that used similar methodologies, including postmortem brain specimens and co-immunoprecipitation with isotype controls. Unlike our study, which is the first to examine the BDTO interactome, two studies investigated PHF tau and three examined NFT tau interactomes (Table 2).22,26–28 Bona fide interactors were defined as proteins appearing in more than 60% of these interactomes. The complete list of compiled proteins, along with their rankings, is available in Supplemental File 2. Surprisingly, minimal protein overlap was observed: out of 114 proteins, only ubiquitin (UBB) and tau (MAPT) were shared.
The fibrillar tau (Figure 6(a)) and AD BDTO (Figure 6(b)) interactomes were converted into hairball and PPI networks. The hairball networks (left) contain all proteins identified within individual interactomes, whereas the MCODE-identified PPI networks (right) only contain proteins with known direct, physical interactions extracted from the hairball network; a discussion of the MCODE algorithm is provided in the Metascape Custom Analysis section within the Methods. Enrichment analysis of the hairball and PPI networks was conducted using the three GO annotations, as summarized in Table 3.
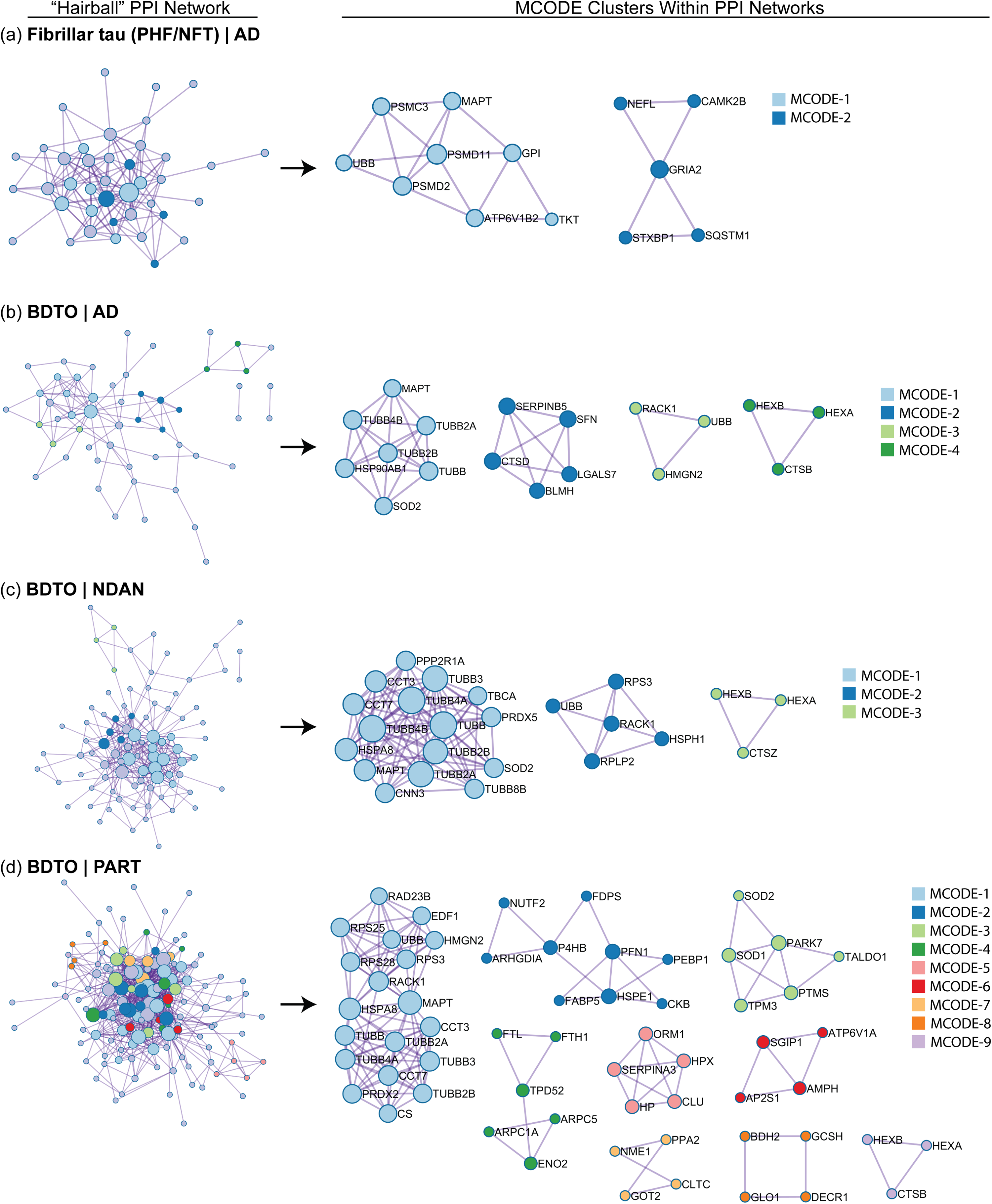
Identifying hubs within the fibrillar tau and BDTO interactomes through PPI network analysis. Protein-protein interaction (PPI) networks, derived from high-confidence physical interaction data, reveal functional protein networks within fibrillar tau from AD (a) and BDTOs from AD (b), NDAN (c), and PART (d). Each panel represents the PPI network filtered to include only physically interacting proteins, thereby enhancing the biological relevance, and reducing noise. The left side of each panel displays the full fibrillar tau or BDTO PPI (‘hairball’) network for each condition; the right illustrates sub-networks of densely connected proteins, indicative of functional modules, identified using the MCODE algorithm. Created in Metascape. 44
Enrichment of MCODE clusters derived from respective interactome proteins.
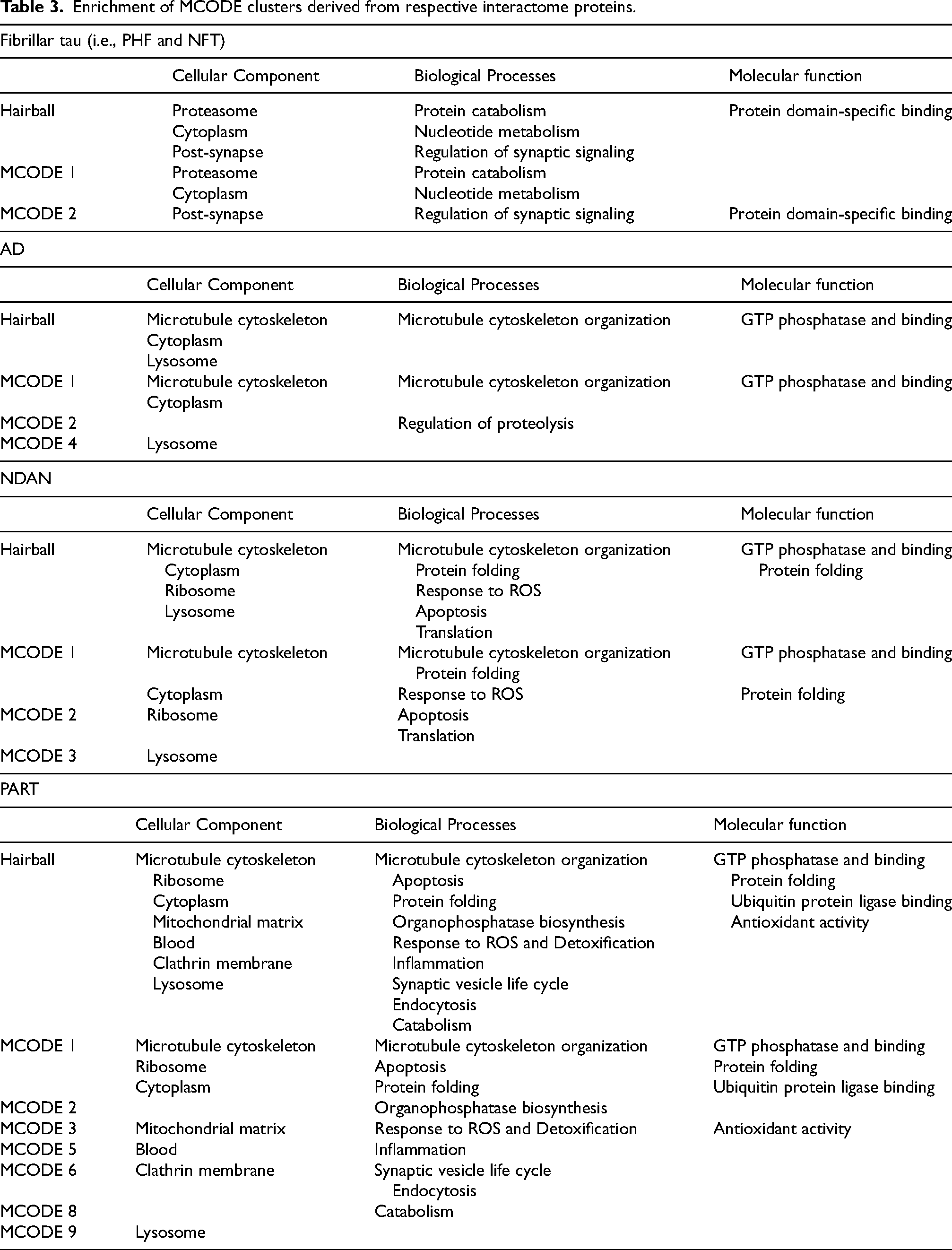
The fibrillar tau interactome contained two enriched protein clusters. The first cluster (light blue MCODE1) was enriched in proteins associated with the proteasome and cytoplasm and involved in protein catabolism and nucleotide metabolism, whereas the second cluster (dark blue MCODE2) was enriched in proteins associated with the post-synapse and involved in the regulation of synaptic signaling. The primary molecular function of proteins found in the fibrillar tau interactome was protein domain-specific binding (MCODE2).
Conversely, enriched terms in the AD BDTO interactome differed significantly. Regarding cellular components, the AD BDTO interactome was enriched in proteins that localize to the microtubule cytoskeleton (light blue MCODE1) and lysosome (dark green MCODE4). In terms of biological processes and molecular function, proteins associated with AD BDTO were involved in microtubule cytoskeleton organization and GTP phosphatase and binding activity (MCODE1) and regulation of proteolysis (dark blue MCODE2).
In sum, a comparison between the fibrillar tau and AD BDTO interactomes demonstrates minimal shared proteins and differential enrichments in cellular components, biological processes, and molecular functions.
Enrichment analysis comparison unveils unique features of AD, NDAN, and PART BDTO interactomes
After examining the overlap between interactomes of different tau structures, we evaluated BDTO interactomes across three distinct clinical conditions. The BDTO interactomes for AD (Figure 6(b)), NDAN (Figure 6(c)), and PART (Figure 6(d)) were converted into hairball and PPI networks and underwent enrichment analysis as previously described. The AD BDTO interactome was primarily enriched in proteins associated with the microtubule cytoskeleton and its dynamics, including microtubule cytoskeleton organization and GTP phosphatase and binding activity. NDAN and PART BDTO interactomes were also enriched in the same terms (light blue MCODE1 for both NDAN and PART). Notably, all three BDTO interactomes contained proteins associated with the lysosome (MCODE4 in AD; light green MCODE3 in NDAN; light purple MCODE 9 in PART).
Proteins within the NDAN and PART BDTO interactomes demonstrated additional enrichments, which aligns with their increased complexity compared to the AD BDTO interactome. NDAN and PART BDTO interactomes shared proteins associated with the ribosome (dark blue MCODE2 in NDAN; MCODE1 in PART) and were involved in apoptosis (MCODE2 in NDAN; MCODE1 in PART), protein folding (MCODE1 for NDAN and PART), and response to reactive oxygen species (ROS), or ROS (MCODE1 for NDAN; light green MCODE3 for PART). Interestingly, PART interactome proteins were specifically enriched in detoxification and antioxidant activity in addition to ROS response (MCODE3), which was lacking in the NDAN interactome. While NDAN and PART interactomes were enriched in ribosome-associated proteins, only the NDAN interactome contained proteins associated with translation (MCODE2).
Altogether, all three BDTO interactomes were enriched in proteins associated with microtubules and lysosomes and implicated in the former's organization and dynamics. NDAN and PART BDTO interactomes shared additional unique enrichments.
Validation of AP-MS results
CCT7, a member of the chaperonin containing TCP1 complex (CCT) (i.e., the TCP1 ring complex (TRiC) family), and S100β, part of the S100 protein family, were chosen as validation targets for three main reasons. First, both CCT7 and S100β were previously reported in AD brain proteomic studies, providing confidence these proteins were indeed bona fide interactors, yet an interaction with tauO has not been reported.43,46 As a molecular chaperone, CCT7 binds to misfolded proteins and can regulate protein oligomerization and oligomer toxicity.47–51 Interestingly, CCT subunits are reported to have neuroprotective roles in other neurodegenerative diseases, specifically Parkinson's disease and Huntington's disease.49–51 S100β levels are significantly elevated in AD patients compared to non-demented controls and mechanistic studies suggest that S100β levels may increase tau hyperphosphorylation, representing one avenue of progression to NFT formation.52,53 Further, S100β levels positively correlate with NFT levels and S100β was identified physically clustered within NFT.52,54,55 Second, despite no significant differences in CCT7 or S100β protein levels in AD, NDAN, and PART PBS-soluble homogenates (Figure 7(a)), CCT7 and S100β were identified by the SAINTq algorithm as significant protein interactors exclusively in NDAN and PART BDTO interactomes and AD BDTO interactome, respectively (Figure 4). Third, commercial, IP-verified antibodies were available for both targets. To this end, we performed reverse co-immunoprecipitation using S100β in AD lysates and CCT7 in NDAN and PART lysates, based on the diagnostic groups in which each interactor was specifically identified. Following confirmation of successful target immunoprecipitation in the appropriate groups (Figure 7(b), top and middle), pooled reverse co-IP samples were probed for tauO (Ab: T22). As predicted by our interactome analysis, tauO was detected in the CCT7 IP samples from NDAN and PART, and in the S100β IP sample from AD, supporting the presence of disease-specific tauO–protein interactions.
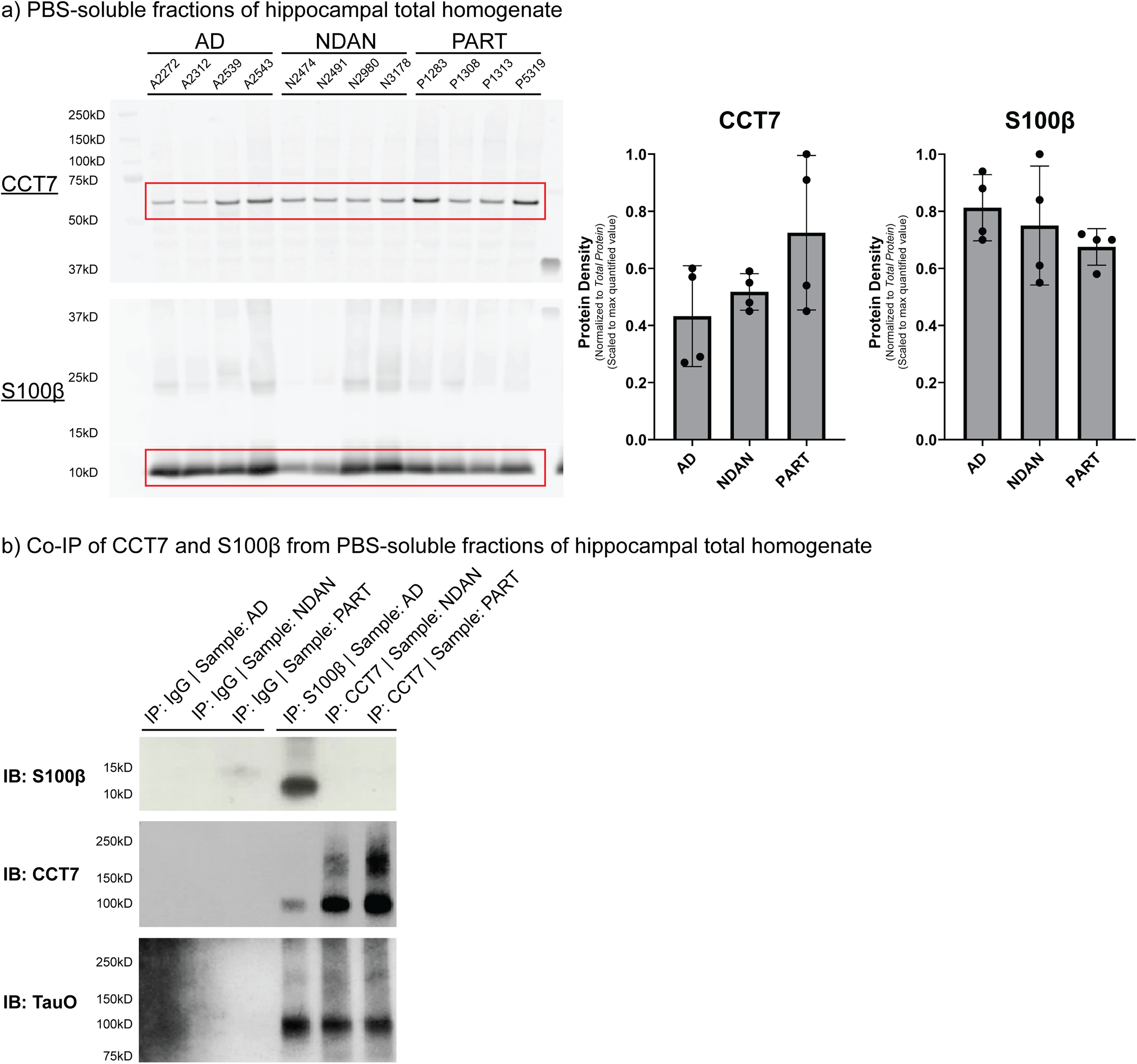
Validation of BDTO interactome findings via reverse co-immunoprecipitation. (a) PBS-soluble fractions of hippocampal total homogenates from AD, NDAN, and PART specimens (n = 4 each) were analyzed for the presence of CCT7 and S100β via Western blotting. Lanes are labeled by sample ID. Bar graphs represent quantified protein density normalized to the total protein signal (Supplemental Figure 5), showing no significant differences in CCT7 (p = 0.14) and S100β (p = 0.37) levels between groups. Statistical analysis was performed using one-way ANOVA (α = 0.05) with Tukey HSD post-hoc test. (b) Co-IP of CCT7 and S100β was performed to validate their selective associations found with BDTO. S100β was co-immunoprecipitated from pooled AD brain homogenates, and CCT7 was co-immunoprecipitated from pooled NDAN and PART brain homogenates. Western blot analysis confirms successful co-IP of these proteins, as well as the presence of tau oligomers (Ab: T22) associated with both CCT7 and S100β across all three groups.
Discussion
This work aimed to elucidate shared and unique features of three BDTO interactomes across the tauopathy spectrum. To this end, we characterized the first-of-its-kind hippocampal BDTO interactomes from AD patients, NDAN individuals, and PART patients. We further contextualized our findings within published pathological tau protein interactomes by comparing the interactome of soluble AD BDTO, likely mediators of early AD progression, to a late-stage fibrillar tau (i.e., PHF and NFT) interactome. This comparison identified eight previously unreported proteins. Additionally, analyzing our three BDTO interactomes revealed potential pathways conferring resilience to tau pathology, offering novel therapeutic targets. We have made our raw data publicly accessible, enabling the scientific community to expand on these molecular networks and accelerate research into tau-mediated neurodegeneration.
Comparison of BDTO interactomes with previous interactomes, facilitating identification of eight novel proteins of interest
Previous studies have laid the groundwork for defining the AD tau interactome, primarily focusing on total tau (both toxic and physiological) or late-stage fibrillar, tau.22–28 We investigated BDTO, representing an intermediate stage in tau pathology, believed to be the most toxic species in AD and an ideal target for early therapeutic intervention. The AD BDTO interactome was enriched in proteins associated with microtubules, microtubule dynamics, and the lysosome, while the fibrillar tau interactome was enriched in synaptic and proteasomal degradation proteins. Only tau and ubiquitin overlapped between the 114 combined proteins. The enrichment of multiple tubulin isoforms in all three BDTO interactomes, absent in the fibrillar tau interactome, suggests that these tau species may retain some physiological function, likely represent an intermediate stage of tau pathology.22,26–28 These findings highlight significant differences between intermediate and late-stage pathological AD tau interactomes.
The AD BDTO and fibrillar tau interactomes shared proteasomal enrichment. However, fibrillar tau proteins were more directly associated with the proteasome, whereas AD BDTO interactome proteins were involved in the regulation of proteolysis. The proteasome, a key degradation system in the brain, is responsible for breaking down short-lived, misfolded, oxidized, and damaged proteins.56–59 Prior research indicates that tauO are not initially heavily ubiquitinated but become increasingly so as they progress to PHF and NFT. 56 The AD interactome was uniquely enriched in proteolysis-related terms, suggesting that AD BDTO may follow a pathological progression, unlike NDAN and PART BDTO, which may diverge from typical toxic progression.
There may be numerous reasons why the AD BDTO and fibrillar tau interactomes do not significantly overlap. It is plausible and supported by our data that the stage of disease progression, and thus the degree of dysfunction, affects tau protein interactions. Our BDTO samples are derived from the PBS-soluble tissue fraction, while a majority of PHF and NFT tau remain in the insoluble fraction.22,26–28 Different tau structural conformations may mask or reveal different sites for interaction. Lastly, it is reasonable to argue that differences between the studies in data collection, analysis, and stringency may impact overlap. Altogether, the AD BDTO interactome overlapped minimally with the fibrillar tau interactome, suggesting notable differences in protein interactions based on tau structural conformation.
We compared the proteins in our BDTO interactomes with those identified in other mass-spectrometry proteomic studies using NeuroPro. This analysis revealed eight novel proteins not previously associated with AD brain changes: Lymphocyte Antigen 6 Family D (LY6D) in the AD interactome; gamma-glutamyl hydrolase (GGH) in the PART interactome; serpin family B member 5 (SERPINB5) in AD and NDAN interactomes; phospholipase B domain containing 2 (PLBD2), sialic acid acetylesterase (SIAE), and small ubiquitin-like modifier 3 (SUMO3) in NDAN and PART interactomes; and cathepsin Z (CTSZ) and hexosaminidase subunit alpha (HEXA) in all three interactomes.
LY6D is a member of the LY6 family, proteins involved in TGFβ signaling, AKT pathways, and immune regulation. 60 Through modulation of α7 nicotinic acetylcholine receptor signaling, LY6 members may affect hippocampal glutamatergic signaling; however, this potential cross-talk and its relevance to neurodegenerative diseases require further investigation. 60 GGH is a key enzyme in folate metabolism and maintains intracellular folate homeostasis. 61 Folate deficiency is observed in AD patients and reduced folate levels have been associated with a two-fold greater risk of developing AD.62,63 Beyond folate homeostasis, changes in GGH can influence DNA methylation and synthesis, gene expression, and cellular repair.61,64,65 SERPINB5 is part of the serine protease (i.e., serpin) superfamily, proteins with various physiological functions. 66 Serpin dysregulation has been implicated across a range of neurodegenerative diseases. 66 Distinct from other serpin proteins, SERPINB5 is non-inhibitory and increases tumor cells’ sensitivity to death.67,68 Previous research suggests pathological situations may influence SERPINB5's post-translational modifications and cellular localization.69,70 PLBD2 is a lysosomal phospholipase that preferentially acts on inositol-containing phospholipid substrates.71,72 PLBD2 inhibition by glycerophosphodiesters leads to the accumulation of toxic lysophospholipid species and lysosomal dysfunction, a commonly observed feature in AD neuropathology.58,72,73 Additionally, PLBD2 acts as a senescence molecule, further implicating its role in aging and potentially, neurodegenerative phenotypes. 74 SIAE is an important enzyme in sialic acid metabolism. 75 Sialic acids are enriched in brain tissue and play a critical role in axon myelination, synaptic development, synaptic transmission, and microglial homeostasis; therefore, perturbations in sialic acid metabolism can affect many cellular and molecular events. 76 For example, patients with mild cognitive impairment (MCI) have been shown to exhibit increased sialic acid metabolism, and this increase has been associated with elevated levels of phosphorylated and total tau and reduced cognitive performance and hippocampal volume.76,77 SUMO3 is part of the small ubiquitin-like modifier protein family and has emerged as a key modulator in conferring neuroprotection. 78 SUMO3 overexpression in the hippocampus of aged C57BL/6 mice correlated with better spatial learning ability. 79 However, SUMO3 overexpression significantly increased APP and β-secretase (BACE) levels and Aβ40 and Aβ42 peptide secretion.80,81 CTSZ is an abundant lysosomal cysteine protease involved in proteostasis.82,83 The cathepsin protease family, particularly CTSB and CTSD, has been implicated in AD progression and displays abnormal distributions in the AD patients’ hippocampi.84,85 The unique features of CTSZ (e.g., differing carboxypeptidase activity and limited similarity with other cathepsins) may make it a valuable target for further investigation.82,86 HEXA is a lysosomal hydrolase, and HEXA deficiency is related to the neurodegenerative progression observed in the lysosomal storage disease, Tay-Sachs. 87 MCI and AD patients display decreased HEXA activity in intracellular lysosomes compared to control subjects. 88
The eight novel proteins identified herein warrant further investigation. While further replication and validation are necessary, their identification in BDTO interactomes suggests that the proteins may be important targets that have not been previously considered for diagnostic and therapeutic development.
Comparison of BDTO interactomes across clinical conditions, highlighting potential pathways that may confer resilience to tau pathology
To better understand similarities and differences between BDTO interactomes across the tauopathy spectrum, we compared the AD, NDAN, and PART BDTO interactomes. All interactomes were enriched in proteins associated with microtubules and microtubule dynamics, suggesting maintenance of some physiological function and further supporting that our BDTO represent an intermediate stage in tau pathology progression. Tau's primary physiological function is to promote the polymerization and stabilization of microtubules.89,90 Tubulin isoforms were enriched in all three interactomes, with different enrichments across interactomes. βI-tubulin and βIIb-tubulin were found in all three interactomes; βIVb-tubulin was a significant interactor with AD and NDAN BDTO whereas βIII-tubulin and βIVa-tubulin were specific to NDAN and PART BDTO. βIVb-tubulin was previously shown to be selectively enriched in oligodendrocytes, and recent evidence implicates oligodendrocytes as a key modulator of pathological tau seeding and spreading.91,92 βIII-tubulin is suggested to protect neurons against free radical damage and ROS, aligning closely with what we observe in NDAN and PART BDTO interactomes. 93 Interestingly, only β-tubulin was enriched across our interactomes, not α-tubulin. Both α-tubulin and β-tubulin bind GTP, but it is the GTP bound to β-tubulin that undergoes hydrolysis to GDP during or following polymerization, likely contributing to the predominance of GO terms relating to GTP binding and activity. 94 Tau has not been previously reported to exhibit preference in its interaction with β-tubulin, as compared to α-tubulin; however, it is suggested that physiological tau has a greater binding affinity for GDP- than GTP-containing tubulin.89,95,96
Lysosomal proteins were enriched in all BDTO interactomes. In contrast to the proteasome, the lysosome primarily degrades long-lived proteins and damaged organelles.58,73 It is not surprising that we observe lysosomal enrichment in all interactomes as BDTO are likely long-lived misfolded proteins that would engage the lysosome, irrespective of clinical condition. Evidence suggests that the lysosome, particularly due to its ability to handle large molecules like tauO, may be the primary mechanism for degrading pathological tau.73,97,98 Lastly, we identified two novel lysosomal proteins, CTSZ and HEXA, in all three interactomes, further implicating the lysosome in tau pathology. Whether this reflects an attempt to maintain proteostasis or a disturbance in lysosomal signaling is unclear.
NDAN and PART BDTO interactomes were both enriched in proteins associated with the ribosome; however, the NDAN interactome displayed further enrichment in translation-involved proteins. Ribosomal dysfunction has been associated with reduced translation and is strongly implicated in AD memory loss. 99 It is also well-established that protein synthesis is crucial for memory formation.100,101 Physiological and pathological tau interact with ribosomes, but the latter interaction is enhanced in tauopathic brains, thus reducing nascent proteins, including those critical to synaptic plasticity. 99 BDTO are pathological tau aggregates; however, our data and prior literature suggest that compared to AD, NDAN does not experience comparable neuropathology-associated toxicity.5,33,34 It is tempting to speculate that the enrichment in ribosome-associated and translation-involved proteins in the NDAN interactome may reflect interactions that maintain or increase the propensity for translation, resulting in preserved protein synthesis and cognition.
One of the most salient features within our data was enrichment in the cellular response to ROS observed in NDAN individuals and PART patients. This phenotype is likely contributed to by peroxiredoxin 2 (Prx2) and peroxiredoxin 3 (Prx3) from the PART interactome and peroxiredoxin 5 (Prx5) from the NDAN interactome. Peroxiredoxins are thought to function as redox sensors, transmitting signals in response to oxidative stress. 102 A recent proteomic study identified elevated levels of Prx2 in hippocampi of AD patients compared to healthy control; however, another study found that the inactivated form was the predominant species in AD patients.103,104 Prx5 is well-documented to be neuroprotective and anti-inflammatory and has been shown to protect mouse hippocampal neurons from glutamate-induced toxicity (i.e., excitotoxicity), and thus may represent a possible mechanism of resilience in NDAN.105–107 βIII-tubulin, enriched in NDAN and PART BDTO interactomes, has been shown to protect neurons against free radical damage and ROS. 93 Glutathione (GSH) is an endogenous antioxidant critical for numerous cellular processes. Decreased levels of GSH and dysfunction in GSH synthesis have been observed in AD brains and correlate with neurodegeneration.108,109 Glutathione synthetase (GSS), the enzyme that catalyzes the production of GSH, was enriched in NDAN and PART interactomes, suggesting possible preservation of GSH synthesis.
While NDAN and PART BDTO interactomes were enriched in the cellular ROS response, the PART interactome was particularly enriched in antioxidant activity. Of note, our group has previously reported increased antioxidant activity in the frontal cortex of NDAN individuals. 31 The PART interactome was also enriched in superoxide dismutase 1 (SOD1) and Parkinsonism Associated Deglycase (Park7). Substantial prior research links SOD1 to AD neuropathology. For example, it has been shown that SOD1 levels are significantly reduced in AD patients, and higher SOD1 levels are associated with better global cognitive function.110,111 Park7 mutations are associated with early-onset Parkinson's disease, and Park7 interacts with pathological tau in the AD brain. 112 However, Park7 also exhibits antioxidant properties; through interactions with other proteins, Park7 renders cells resistant to oxidative stress-induced cell death.113,114 Taken together, our data suggests that multiple proteins and pathways in NDAN and PART BDTO interactomes converge, possibly resulting in a more efficient cellular response to ROS.
We find a significant overlap between the interactomes of NDAN and PART BDTO, as well as a distinct separation of both interactomes from AD BDTO, supporting our conceptualization of these conditions along a continuum of vulnerability and resilience. We postulate that NDAN and PART BDTO interactomes overlap the most because resilient pathways and processes likely oppose those occurring in the AD brain. An example of such events, as suggested by our data, is response to oxidative stress. Prior literature highlights the importance of an efficient antioxidant response in conferring protection to many cell types, including neurons. 115 While these responses are compromised in AD brains, it appears, from our results as well as other independent investigations, that resilience is associated with improved antioxidant activity and protection against free radical damage.31,116
We further note that the PART BDTO interactome contained the highest number of unique proteins, as compared to AD and NDAN. While the exact reason for this difference is unclear, it may be linked to the preserved cognitive function observed in PART subjects. Prior mass spectrometry-based biomarker studies in neurodegeneration emphasize the importance of sample quality for accurate protein identification. 117 Consistent with intact cognition indicated by high MMSE scores, 45 PART subjects likely experienced minimal neurodegeneration, thereby maintaining neuronal integrity and yielding higher-quality tissue for LC-MS/MS analysis. Such preservation may enhance the detection of low-abundance or unique protein species.
Collectively, AD, NDAN, and PART BDTO interactomes contain a complex set of unique and shared features. All three interactomes share proteins that localize to the microtubule and are involved in its dynamic regulation and proteins that engage the lysosomal degradation system. Both NDAN and PART interactomes contain proteins that contribute to the cellular response to ROS, and the PART interactome is further enriched in antioxidant activity. These results suggest that while a similar tau structural conformation may contribute to shared features, other factors likely drive unique protein interactions and functional outcomes.
Limitations
To the best of our knowledge, this is the first interactome characterization of BDTO from three distinct clinical conditions with varying vulnerability and resilience to tau pathology. We isolated BDTO directly from hippocampal autopsy specimens, enhancing the translational relevance of our results. The hippocampus was chosen because it is one of the earliest sites of tau pathology formation and tau pathology-associated neurodegeneration, facilitating comparison across the three clinical conditions.10,11,30,118 However, the limited availability of these high-value specimens restricted our sampling to four per condition. To compensate for the small sample size, we applied a rigorous methodological approach, including control IP samples and the implementation of SAINTq to remove nonspecific proteins quantitatively. This resulted in interactomes containing only statistically significant and specific proteins. All samples utilized in this study were rigorously assessed by expert neuropathologists, who provided a pathological classification. However, the potential incidence of co-pathologies within the overall pathogenesis of these conditions cannot be ignored; α-synuclein inclusions, cerebral amyloid angiopathy, and TDP-43 are frequently observed in AD patient brains.119–121 In our samples, α-synuclein was detected in 3 out of the 4 AD BDTO, all 4 NDAN BDTO, and all 4 PART BDTO, as well as the IgG isotype eluates across all conditions in comparable quantities; for this reason, α-synuclein was not identified (per SAINTq) as a significant interactor in any BDTO interactome. TDP-43 was not detected in any samples.
Tau-related mechanisms of toxicity represent a promising avenue of investigation, and therefore, the goal of our study was to generate insight into the tau interactome across three clinical conditions spanning vulnerability and toxicity. While all assertions mentioned within the discussion are supported by prior literature, validating the findings remains an essential next step. Expanding our work to consider other brain regions and the interactomes of other common AD co-pathologies represents a possible future direction.
Conclusions
BDTO and their associated interactomes were isolated from three clinical conditions across the tauopathy spectrum, from wholly vulnerable neurodegeneration with tau spreading (AD) to wholly resilient to neurodegeneration with tau spreading (NDAN) to resilient to tau spreading with localized neurodegeneration (PART). BDTO presence was confirmed at key steps in the experimental workflow. Using SAINTq, nonspecific proteins were quantitatively removed, resulting in 203 unique and shared proteins across the three BDTO interactomes. Enrichment analysis highlighted known tau physiological functions in all three interactomes and the predominance of cellular response to ROS in NDAN individuals and PART patients. While further investigation is needed, these proteins and pathways could be valuable therapeutic targets due to their association with BDTO. We hope our work serves as a foundation for future studies to elucidate the roles these proteins may play in conferring resilience to tau pathology.
Supplemental Material
sj-docx-1-alz-10.1177_13872877251352382 - Supplemental material for Comparative analysis of brain-derived tau oligomer interactomes in Alzheimer's disease, non-demented with Alzheimer's neuropathology, and primary age-related tauopathy: Implications for neurodegeneration and cognitive resilience
Supplemental material, sj-docx-1-alz-10.1177_13872877251352382 for Comparative analysis of brain-derived tau oligomer interactomes in Alzheimer's disease, non-demented with Alzheimer's neuropathology, and primary age-related tauopathy: Implications for neurodegeneration and cognitive resilience by Danielle Jamison, Shrinath Kadamangudi, Batbayar Tumurbaatar, Wen-Ru Zhang, Lee Palmer, Steve Kunkel, Rakez Kayed, Agenor Limon and Giulio Taglialatela in Journal of Alzheimer's Disease
Supplemental Material
sj-xlsx-2-alz-10.1177_13872877251352382 - Supplemental material for Comparative analysis of brain-derived tau oligomer interactomes in Alzheimer's disease, non-demented with Alzheimer's neuropathology, and primary age-related tauopathy: Implications for neurodegeneration and cognitive resilience
Supplemental material, sj-xlsx-2-alz-10.1177_13872877251352382 for Comparative analysis of brain-derived tau oligomer interactomes in Alzheimer's disease, non-demented with Alzheimer's neuropathology, and primary age-related tauopathy: Implications for neurodegeneration and cognitive resilience by Danielle Jamison, Shrinath Kadamangudi, Batbayar Tumurbaatar, Wen-Ru Zhang, Lee Palmer, Steve Kunkel, Rakez Kayed, Agenor Limon and Giulio Taglialatela in Journal of Alzheimer's Disease
Supplemental Material
sj-xlsx-3-alz-10.1177_13872877251352382 - Supplemental material for Comparative analysis of brain-derived tau oligomer interactomes in Alzheimer's disease, non-demented with Alzheimer's neuropathology, and primary age-related tauopathy: Implications for neurodegeneration and cognitive resilience
Supplemental material, sj-xlsx-3-alz-10.1177_13872877251352382 for Comparative analysis of brain-derived tau oligomer interactomes in Alzheimer's disease, non-demented with Alzheimer's neuropathology, and primary age-related tauopathy: Implications for neurodegeneration and cognitive resilience by Danielle Jamison, Shrinath Kadamangudi, Batbayar Tumurbaatar, Wen-Ru Zhang, Lee Palmer, Steve Kunkel, Rakez Kayed, Agenor Limon and Giulio Taglialatela in Journal of Alzheimer's Disease
Footnotes
Acknowledgements
The authors thank Dr William Russell for his technical mass spectrometry assistance.
Ethical considerations
Not applicable
Consent to participate
Not applicable
Consent for publication
Not applicable
Author contributions
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This manuscript was supported by NIH grants F30AG085974 (to SK) and 1R01AG073133 (to GT).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The data supporting the findings of this study are available within the article and its supplemental material.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
