Abstract
Background
Little is known about the rare missense variants (RMVs) of APOE in East Asians, including the Japanese, and their association with Alzheimer's disease (AD) and lipid metabolism.
Objective
To identify APOE RMVs in the Japanese population and investigate their association with AD and lipid metabolism, including low-density lipoprotein cholesterol levels.
Methods
APOE RMVs were explored in the Niigata (NIG; 2589 subjects) and Tohoku (ToMMo; 3307 subjects) cohorts. A case-control study included 6261 AD cases and 16,331 controls, all of whom were aged 65 or older. Sanger sequencing, whole-exome sequencing, or a combination of both was performed on the NIG subjects. We used the genotype data from the ToMMo cohort. APOE RMV frequencies in the Japanese population were compared with various ethnic populations. Associations between APOE RMV genotypes, AD, and lipoproteins were examined.
Results
Fourteen RMVs were identified (minor allele frequency 0.02–0.73%), with 10 unique to East Asians. Five previously reported RMVs, such as the Christchurch RMV, were absent in Japanese individuals. Two RMVs (rs140808909 and rs190853081), which exhibit complete linkage disequilibrium, were found to have protective effects against AD: pBonferroni = 4.28E-02, OR (95% CI) = 0.70 (0.54–0.92). No significant differences in cholesterol levels were observed between RMV carriers and non-carriers.
Conclusions
The two APOE RMVs identified in Japanese individuals may have exhibited potential protective effects against AD. Further large-scale studies are needed to confirm these findings and to explore their roles in AD and lipid metabolism.
Introduction
Alzheimer's disease (AD) is the leading cause of dementia in the elderly. AD is a multifactorial disease resulting from interactions between environmental factors and individual genetic susceptibility. A large twin epidemiological study in Sweden estimated the heritability of AD to be 60–80%. 1 Multiple loci are thought to contribute to the onset of AD. 2 This indicates that genetic factors play substantial roles in the development and progression of AD. Large-scale genome-wide association studies (GWAS) have identified 90 susceptibility genes/loci for AD. 3 Recent studies have focused on the application of polygenic risk4–6 and resilience 7 score analyses, which integrate multiple AD susceptibility variants to assess individual risks, with potential clinical applications being explored.
Among the AD susceptibility genes/loci identified to date, APOE remains the most significant genetic risk factor across populations.8–11 A search for “Alzheimer” on the GWAS Catalog website 12 highlights the APOE genomic region as a multiple-hit locus with a remarkably strong association with AD, consistent with findings reported in our previous publications13–15 and those of others.3,16 Recently, rare missense variants (RMVs) of APOE were reported to be associated with AD, shedding light on new links between APOE and AD. For example, RMVs such as p.L46P (p.L28P; rs769452 [Pittsburgh]), 17 p.R154S (p.R136S; rs121918393 [Christchurch]), 18 p.R163C (p.R145C; rs769455), 19 p.V254E (p.V236E; rs199768005 [Jacksonville]), 20 and p.R269G (p.R251G; rs267606661) 21 have been reported in Western populations, suggesting potential resistance to amyloid-β (Aβ) accumulation, phosphorylated tau pathology, or the effects of the APOE ε4 allele. Rasmussen et al. 22 identified 20 APOE RMVs in an analysis involving more than 100,000 participants and demonstrated their association with plasma APOE levels and AD onset.
However, few reports have addressed the identification and significance of APOE RMVs in the Japanese population. Therefore, this study represents the first attempt to characterize APOE RMVs in this population. We examined the genetic association between the identified RMVs and AD, as well as differences in peripheral blood lipoprotein levels between carriers and non-carriers of these RMVs.
Methods
Subjects for identifying APOE RMVs
We explored APOE RMVs in two Japanese cohorts (Tables 1 and 2). The first cohort, referred to as the Niigata (NIG), consisted of 2589 participants (1150 females and 1439 males) from medical and research institutions across Japan. This cohort was primarily established for research-level genetic diagnoses, including AD and various other neurological disorders. Of these, 826 subjects were based on clinical diagnoses, and 1763 were based on pathological diagnoses. Additionally, 508 of the 2589 NIG subjects were derived from the Japanese Alzheimer's Disease Neuroimaging Initiative (J-ADNI). 23 Sanger sequencing (SS), whole-exome sequencing (WES), and a combination of both (SS + WES) were performed on 839, 1,509, and 241 subjects, respectively. The second cohort, referred to as Tohoku (ToMMo), consisted of 3307 community-dwelling healthy subjects (1810 females and 1497 males) recruited by the Tohoku Medical Megabank Organization. 24 In compliance with the data usage policy (Research ID: 2019-0074), individual genotype data for 176 variants within a genomic region of approximately 7.2 kb, including APOE (positioned at 45,407,254–45,414,449 bp in GRCh37/hg19), were obtained. The genotype data for each variant were previously obtained through whole-genome sequencing. 24 The total number of subjects from both cohorts was 5896 (2960 females and 2936 males), with participants aged ≥20 years (mean: 64.1 years; median: 66.0 years; range: 20–111 years).
The information on subjects used in the exploration of APOE RMVs.
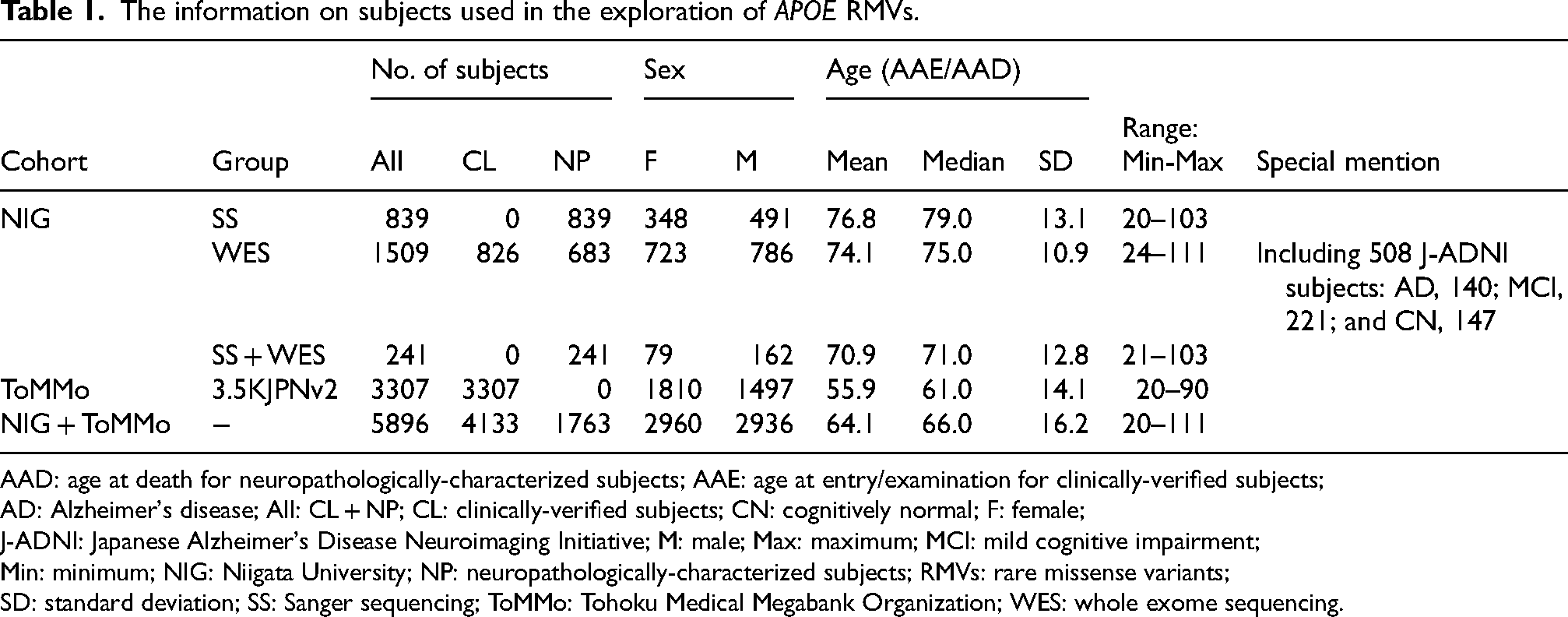
AAD: age at death for neuropathologically-characterized subjects; AAE: age at entry/examination for clinically-verified subjects;
AD: Alzheimer's disease; All: CL + NP; CL: clinically-verified subjects; CN: cognitively normal; F: female;
J-ADNI: Japanese Alzheimer's Disease Neuroimaging Initiative; M: male; Max: maximum; MCI: mild cognitive impairment;
Min: minimum; NIG: Niigata University; NP: neuropathologically-characterized subjects; RMVs: rare missense variants;
SD: standard deviation; SS: Sanger sequencing; ToMMo: Tohoku Medical Megabank Organization; WES: whole exome sequencing.
Genotypic and allelic distribution of APOE in the cohorts used for the exploration of APOE RMVs.
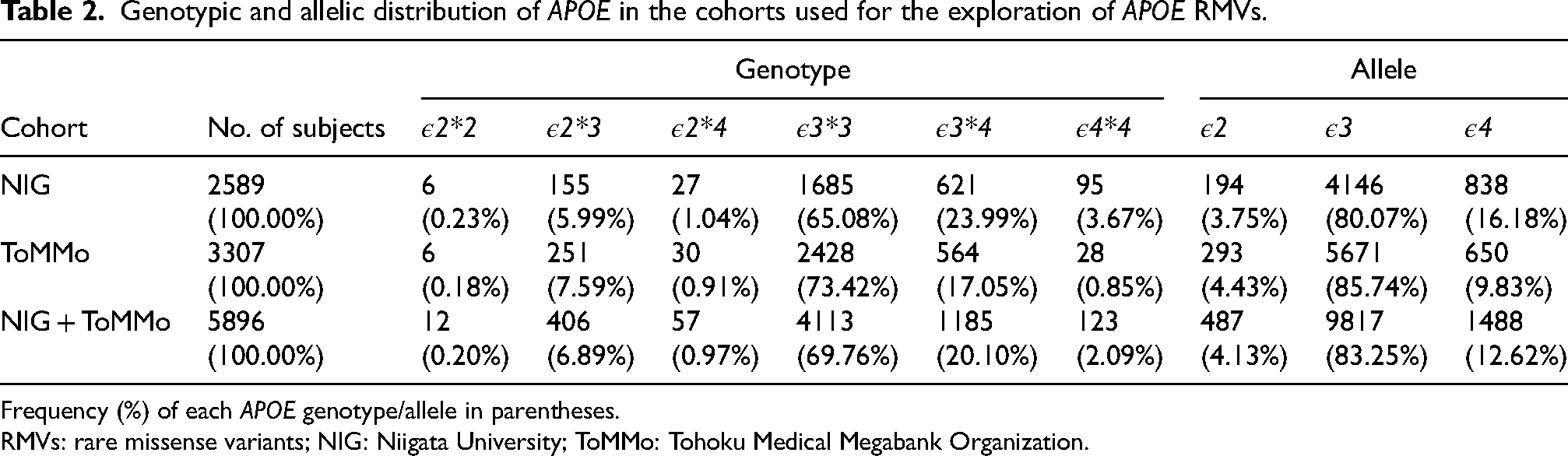
Frequency (%) of each APOE genotype/allele in parentheses.
RMVs: rare missense variants; NIG: Niigata University; ToMMo: Tohoku Medical Megabank Organization.
SS for APOE
SS was performed in 1080 subjects from the NIG cohort (839 for SS and 241 for SS + WES) (Table 1). Genomic DNA was extracted from peripheral blood or autopsied brain tissues and quantified using the NanoDrop spectrophotometer (Thermo Fisher Scientific). DNA quality was assessed by calculating the DNA integrity number (Agilent Technologies). The primers used to sequence the entire coding region of APOE is provided in Supplemental Table 1.
WES
WES was performed on 1750 subjects from the NIG (1509 for WES and 241 for SS + WES) (Table 1). Genomic DNA was used to prepare an exome library using the SureSelect Human All Exon V6 Kit (Agilent Technologies). The library was sequenced on a NovaSeq 6000 platform (Illumina) in 151-cycle paired-end mode. Sequence reads were processed with fastp (v.0.19.5) using the default settings for quality control and adapter trimming. The cleaned reads were then mapped to the human reference genome (hg38) using BWA-MEM (v.0.7.15-r1140) with default parameters. Subsequent analyses, including read processing, variant calling, and filtration, were performed following GATK4 Best Practices. The resulting variant call sets were annotated using snpEff (v.4.3t) and stored in an SQLite database using GEMINI (v.0.20.1).
Determination of six common APOE genotypes
We determined six common APOE genotypes (ε2*2, ε2*3, ε2*4, ε3*3, ε3*4, and ε4*4) in all 5896 subjects from the NIG and ToMMo cohorts (Table 1) based on nucleotide combinations (haplotypes) of the common missense variants rs429358 (T > C: p.C130R [p.C112R]) and rs7412 (C > T: p.R176C [p.R158C]) in APOE. For the NIG cohort, these genotypes were determined using both the SS and TaqMan methods. Information regarding the TaqMan assays used (TaqMan assay IDs: 19AyD0001 for rs429358 and 19AyD0002 for rs7412) is provided in Supplemental Table 1. For the ToMMo cohort, genotypes were determined based on digital data (genotype: 0/0, 0/1, and 1/1). The results are presented in Table 2. The effects of APOE genotypes and alleles on AD risk and protection are shown in Supplemental Figure 1 and Supplemental Table 6. Genotypic and allelic data for APOE were obtained from Asanomi et al. 25 and used in this analysis.
Case-control study
We conducted a case-control study to examine whether the identified APOE RMVs were genetically associated with AD. Information concerning the subjects used in this analysis is provided in Table 3, including sex, age (age at entry/examination/death), and APOE (genotype and allele). The diagnostic criteria used for subject recruitment are described in detail elsewhere.25,26
The information on subjects used in the case-control study.
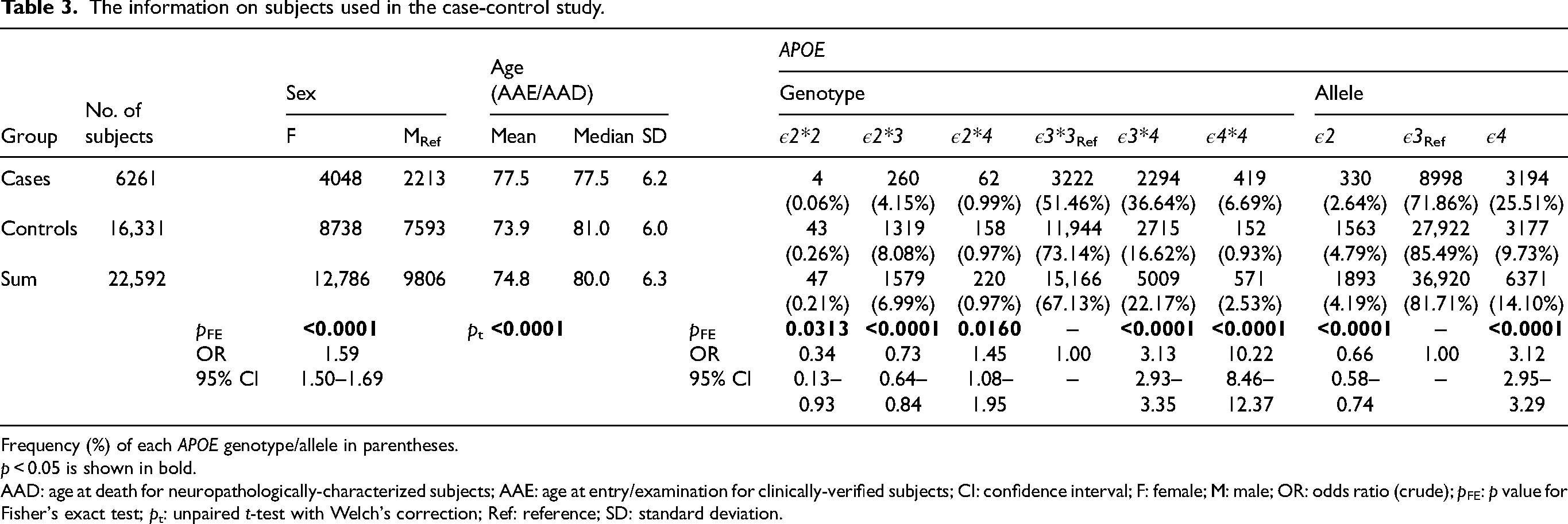
Frequency (%) of each APOE genotype/allele in parentheses.
p < 0.05 is shown in bold.
AAD: age at death for neuropathologically-characterized subjects; AAE: age at entry/examination for clinically-verified subjects; CI: confidence interval; F: female; M: male; OR: odds ratio (crude); pFE: p value for Fisher's exact test; pt: unpaired t-test with Welch's correction; Ref: reference; SD: standard deviation.
Genomic variants were genotyped using the Asian Screening Array (ASA) (Illumina). A total of 6261 AD cases and 16,331 controls (Table 3) were analyzed using the ASA.
None of the 14 APOE RMVs identified in the NIG and ToMMo cohorts were covered by the ASA platform. Therefore, genotypes were inferred through imputation. Imputation accuracy, reflecting the reliability of inferred genotypes, was evaluated using the INFO score. In SNP analysis, imputation is used to infer missing genetic variants from known genotype data and a reference panel. This approach increases the number of analyzed variants without the need for direct genotyping. SNP imputation was performed using Minimac4 with a Japanese reference panel constructed from 1000 Genomes Project Phase 3 (1KGP3 [May 2013, n = 2503]) and 3181 Japanese whole-genome sequences from the National Center for Geriatrics and Gerontology (NCGG) biobank database. To ensure high imputation accuracy, variants with an INFO score ≥ 0.7 were included in the association analysis. Among the 14 APOE RMVs, genotype data were obtained for four variants: rs121918392 (ε5), rs762461580, rs140808909 (ε7), and rs190853081 (ε7) (Table 4, Figures 1 and 2). The INFO scores for each imputed variant were as follows: rs121918392 (ε5), 0.93; rs762461580, 0.83; rs140808909 (ε7), 0.82; and rs190853081 (ε7), 0.82.
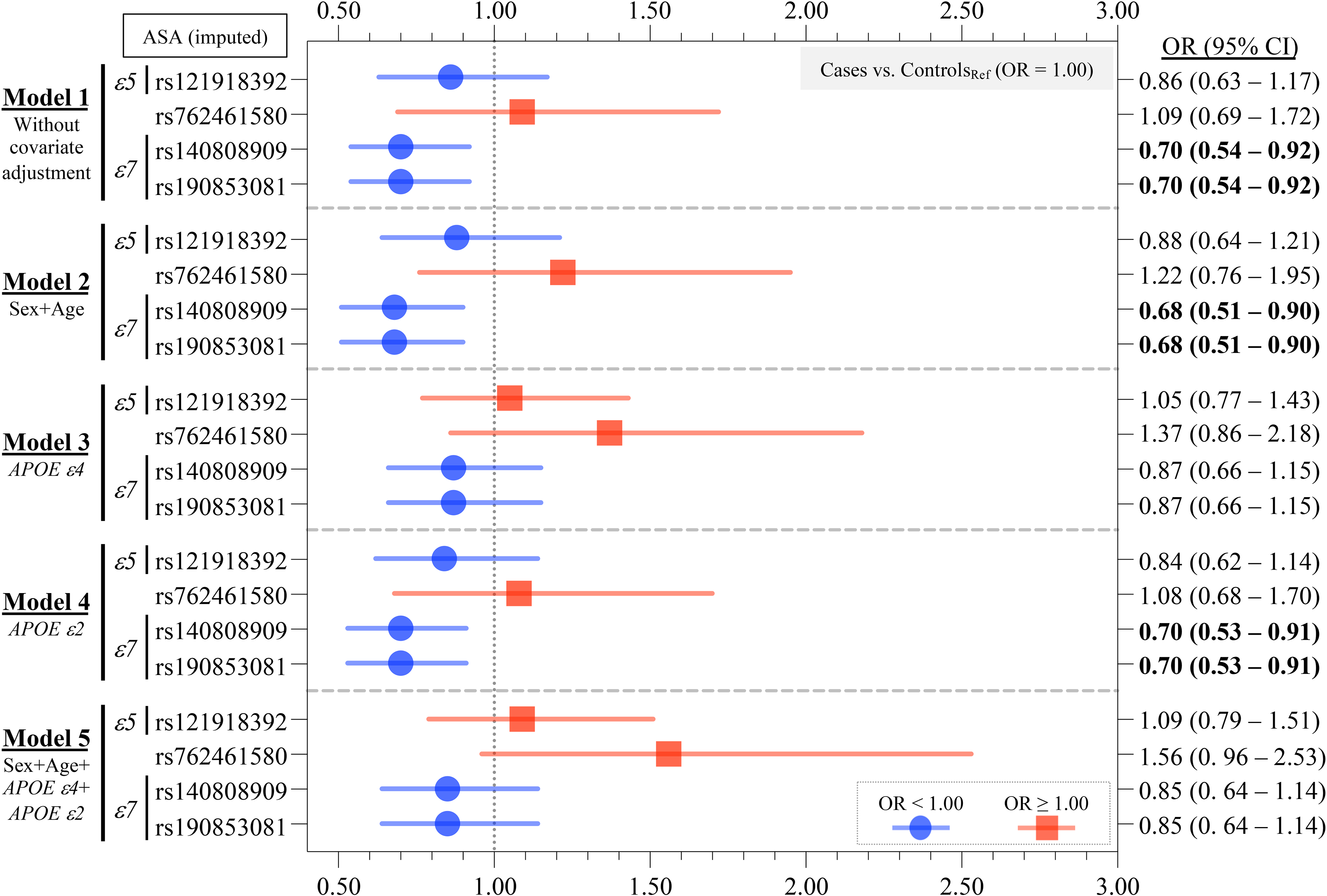
Case-control study on APOE RMVs identified. Of the 14 APOE RMVs identified in this study, four imputed variants were included in the analysis. Two RMVs, rs140808909 and rs190853081, which were in complete LD (Supplemental Figure 2) and define the ε7 allele, remained statistically significant after multiple comparison corrections in Models 1, 2, and 4. The OR was less than 1.0, suggesting a protective effect against AD. However, after adjusting for the ε4 allele in Models 3 and 5, the significant association of the ε7 allele disappeared. This suggests that the ε7 allele is not independent of the ε4 allele but rather is a dependent allele. The detailed numerical data are presented in Table 4. Bold text denotes that the 95% CI of the OR does not contain 1.00.
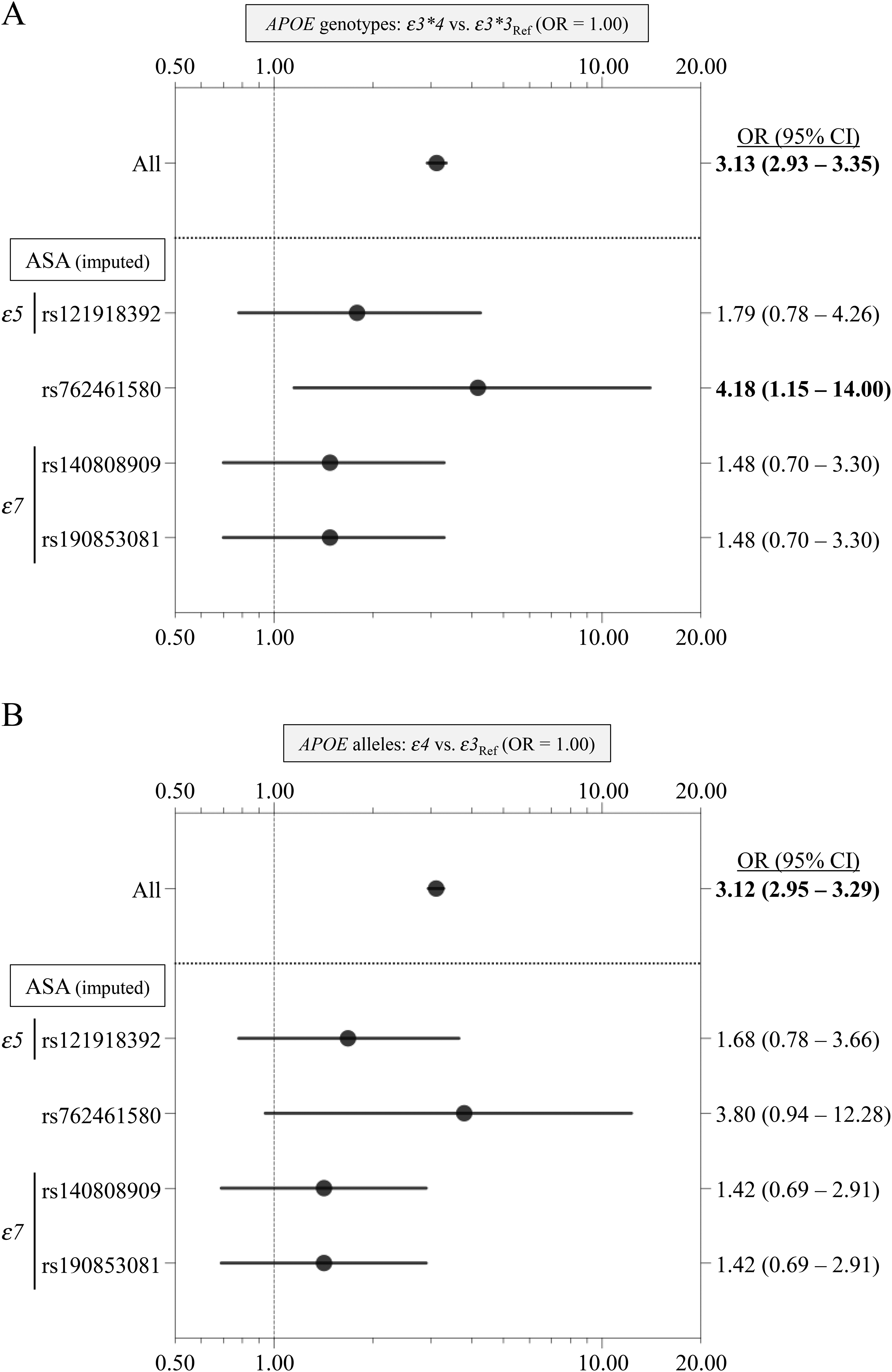
Case-control study on common APOE genotypes and alleles in carriers with APOE RMVs identified. For the four APOE RMVs analyzed in the case-control study (Table 4, Figure 1), we conducted a case-control analysis focusing on subjects carrying alternative alleles, examining common APOE genotypes (A) and alleles (B). Forest plots showing OR and 95% CI are provided. For genotypes (A), the OR (95% CI) of ε3*4 relative to ε3*3 was calculated. For alleles (B), the OR (95% CI) of the ε4 allele relative to the ε3 allele was calculated. Bold text denotes that the 95% CI of the OR does not contain 1.00. “All” indicates the OR and its 95% CI for all subjects (6261 AD cases and 16,331 controls) as presented in Table 3. Importantly, both the ε5 (rs121918392) and ε7 alleles (rs140808909-rs190853081) tended to attenuate the AD risk conferred by the ε4 allele slightly. Ref: reference.
Case-control study on APOE RMVs identified.
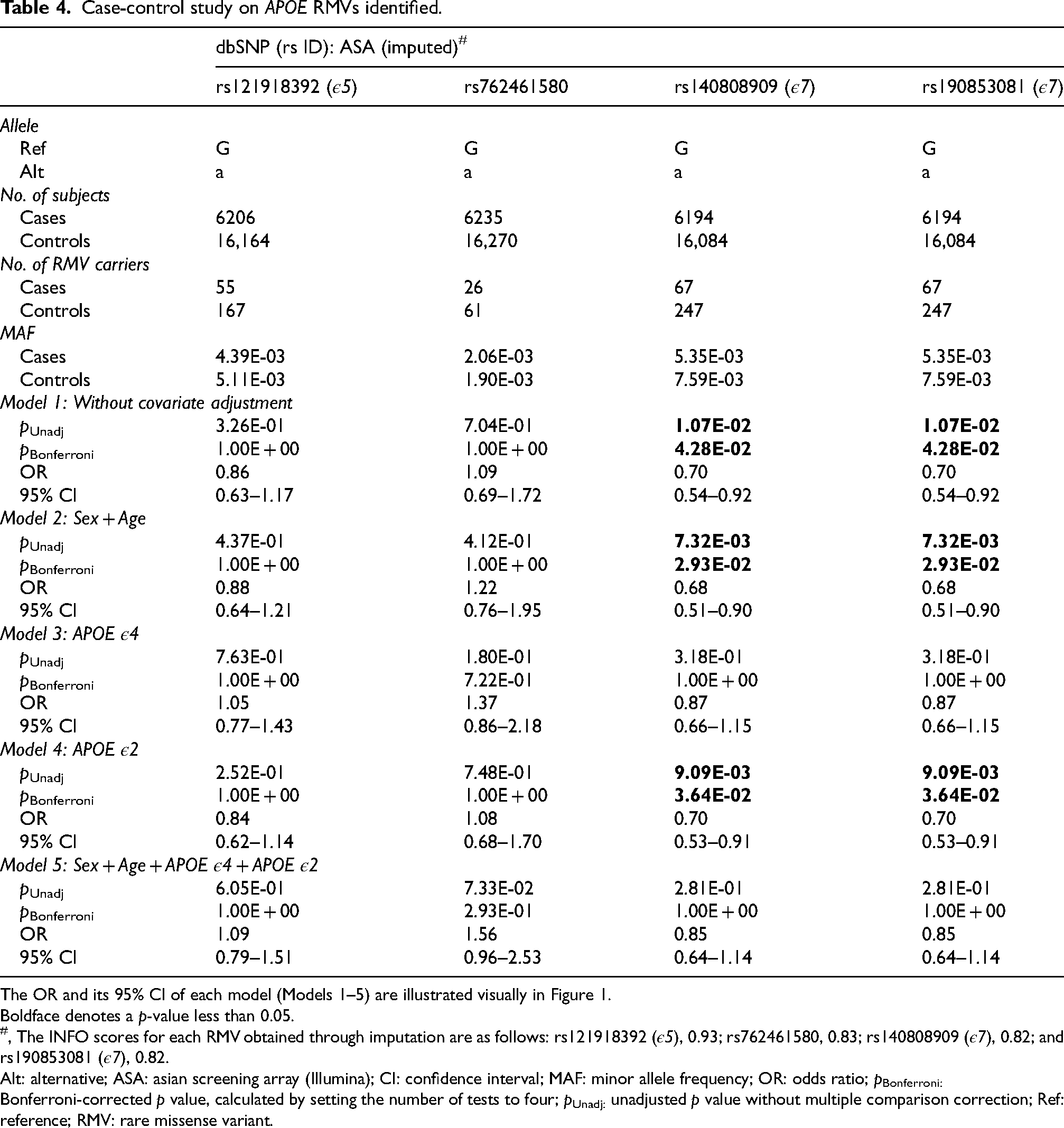
The OR and its 95% CI of each model (Models 1–5) are illustrated visually in Figure 1.
Boldface denotes a p-value less than 0.05.
, The INFO scores for each RMV obtained through imputation are as follows: rs121918392 (ε5), 0.93; rs762461580, 0.83; rs140808909 (ε7), 0.82; and rs190853081 (ε7), 0.82.
Alt: alternative; ASA: asian screening array (Illumina); CI: confidence interval; MAF: minor allele frequency; OR: odds ratio; pBonferroni: Bonferroni-corrected p value, calculated by setting the number of tests to four; pUnadj: unadjusted p value without multiple comparison correction; Ref: reference; RMV: rare missense variant.
APOE RMVs identified.
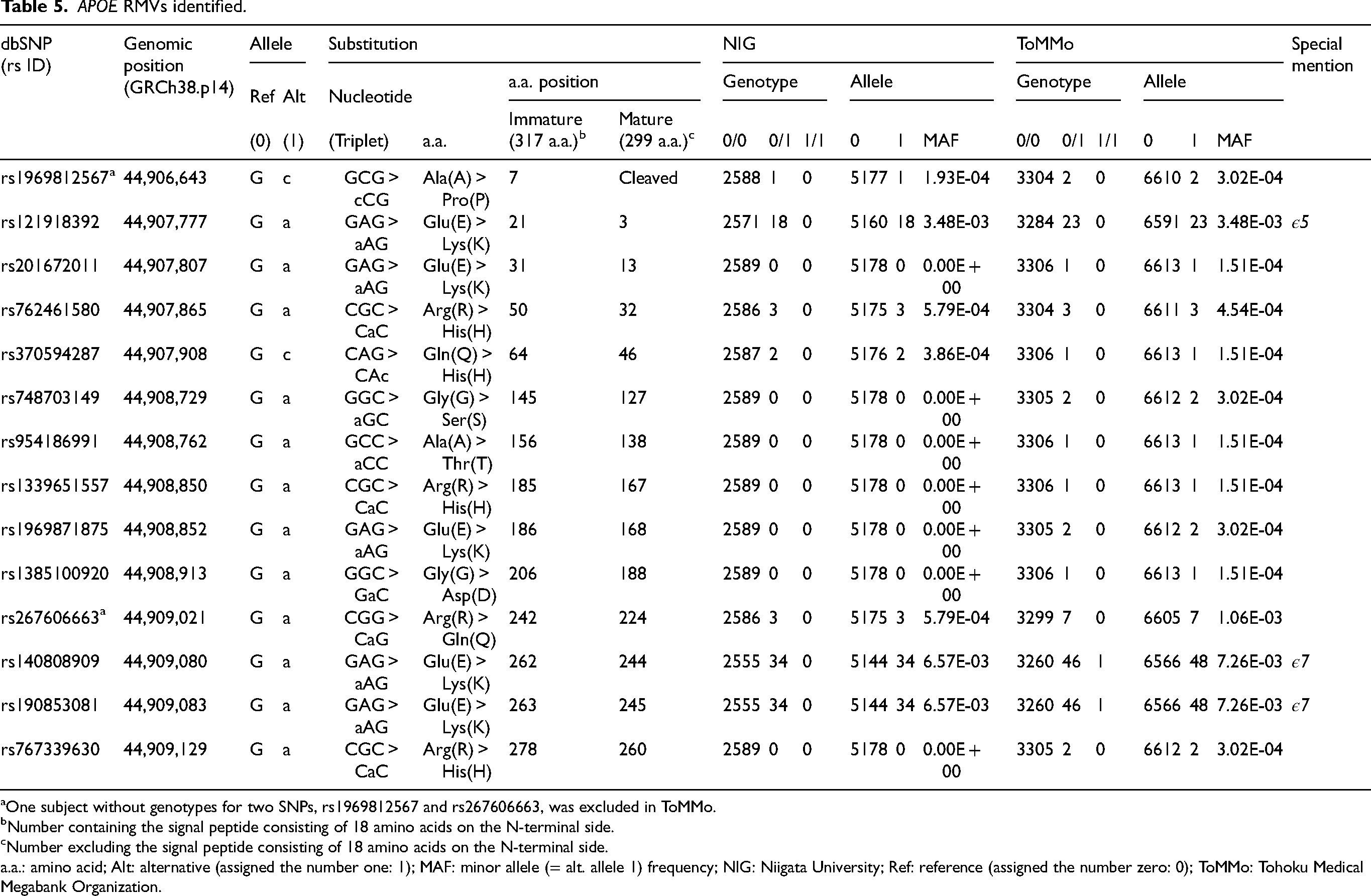
One subject without genotypes for two SNPs, rs1969812567 and rs267606663, was excluded in ToMMo.
Number containing the signal peptide consisting of 18 amino acids on the N-terminal side.
Number excluding the signal peptide consisting of 18 amino acids on the N-terminal side.
a.a.: amino acid; Alt: alternative (assigned the number one: 1); MAF: minor allele (= alt. allele 1) frequency; NIG: Niigata University; Ref: reference (assigned the number zero: 0); ToMMo: Tohoku Medical Megabank Organization.
TA cloning for confirmation of APOE RMV alleles
In one case (Subject ID #35: female, age at death 99, APOE ε3*3) harboring three APOE RMVs (all heterozygous: rs267606663-G/a, rs140808909 [ε7]-G/a, rs190853081 [ε7]-G/a) (Supplemental Table 5), we examined whether these RMVs were co-located on the same chromosome. All three RMVs mapped to exon 4 of APOE. PCR was performed using genomic DNA as a template, followed by TA cloning with the TaKaRa Mighty TA-cloning Kit. The PCR primers used were as follows: forward, 5’-TGCCGATGACCTGCAGAA-3’; reverse, 5’-GCTGCATGTCTTCCACCAG-3’.
Lipoproteins
Lipoprotein measurements, including total cholesterol (TC), low-density lipoprotein cholesterol (LDL-C), high-density lipoprotein cholesterol (HDL-C), and triglycerides (TG), were obtained from the J-ADNI subjects in the NIG (508 subjects: baseline) and ToMMo (TC: 234 subjects; non-TC: 1630 subjects) cohorts. The ToMMo cohort included subjects assessed under both fasting and non-fasting conditions, as well as those with missing fasting status data. In contrast, all blood samples from the J-ADNI cohort were collected under fasting conditions. We evaluated the quantitative differences in lipoprotein levels (TC, LDL-C, HDL-C, and TG) between APOE RMV carriers and non-carriers.
Statistical analysis
Differences in sex distribution and APOE genotypic and allelic distributions between cases (AD) and controls were assessed using Fisher's exact test based on 2 × 2 contingency tables (Table 3, Supplemental Tables 4 and 6). Additionally, unadjusted odds ratios (OR) and 95% confidence intervals (CI) were calculated (Table 3, Supplemental Tables 4 and 6). For age, an unpaired Student's t-test with Welch's correction was used (Table 3).
Logistic regression analysis was conducted in the case-control study to calculate p-values and OR (95% CI) (Table 4, Figure 1). Five models were used in the analysis: Model 1, no covariate adjustment; Model 2, adjusted for sex and age; Model 3, adjusted for the APOE ε4 allele; Model 4, adjusted for the APOE ε2 allele; and Model 5, adjusted for all four covariates used in Models 2, 3, and 4 (i.e., sex, age, the APOE ε4 allele, and the APOE ε2 allele). The Bonferroni method was applied to correct for multiple comparisons for the APOE RMVs analyzed.
A meta-analysis was conducted to integrate and assess the association between APOE genotypes/alleles and AD derived from the two cohorts, as described in the study by Asanomi et al.: 25 the NCGG and the Japanese Genetic Study Consortium for Alzheimer's Disease (JGSCAD) 27 (Supplemental Table 6, Supplemental Figure 1). Both fixed-effects and random-effects models were assumed, and Chi-square p-values and pooled OR (95% CI) were calculated. Cochran's Q and I² statistics were also computed to evaluate the heterogeneity between the two datasets.
Comparisons of TC, LDL-C, HDL-C, TG, and the LDL-C/HDL-C (LH) ratio between the two groups (APOE RMV carriers versus non-carriers) were conducted using the non-parametric Mann-Whitney U-test (Supplemental Figure 4).
A series of genetic statistical analyses related to the case-control study (Table 4, Figure 1) was conducted using PLINK (v.1.90; https://www.cog-genomics.org/plink/). 28 A meta-analysis (Supplemental Table 6, Supplemental Figure 1) was conducted using StatsDirect software (v.4.0.4; https://www.statsdirect.com). Other statistical analyses (Tables 1 and 3, Supplemental Tables 4 and 6, Figure 2, Supplemental Figure 4) were performed using GraphPad Prism software (v.10.3.1; https://www.graphpad.com). p-values of less than 0.05 were considered statistically significant.
Databases
Databases and analysis websites used in this study are listed below in alphabetical order.
Alzforum MUTATIONS APOE: https://www.alzforum.org/mutations/apoe Ensemble Human (GRCh38.p14): https://asia.ensembl.org/Homo_sapiens/Info/Index gnomAD (v4.1.0): https://gnomad.broadinstitute.org GWAS Catalog: https://www.ebi.ac.uk/gwas/home UCSC genome: https://genome.ucsc.edu
Results
Identification of APOE RMVs in the Japanese population
Using 2589 subjects from the NIG cohort (Table 1), we identified seven APOE RMVs (Table 5, Figure 3): rs1969812567 (p.A7P), rs121918392 (p.E21 K), rs762461580 (p.R50H), rs370594287 (p.Q64H), rs267606663 (p.R242Q), rs140808909 (p.E262K), and rs190853081 (p.E263K). All carriers were heterozygous, and no homozygotes were identified. A summary of the clinical and pathological diagnostic information of these patients is provided in Supplemental Table 5. Among these seven RMVs, the two variants with the highest alternative allele frequencies were rs140808909 (p.E262K) and rs190853081 (p.E263K), both with a minor allele frequency (MAF) of 0.00657 (Table 5). These two variants were in complete linkage disequilibrium (LD) (LD coefficient R² = 1), indicating that they were fully co-inherited (Supplemental Figure 2). In other words, knowing the genotype of one variant makes it possible to predict the genotype of the other with 100% accuracy. Both variants have been previously identified in Japanese patients with dyslipidemia and atherosclerosis and is defined as the APOE ε7 allele.29–31 rs121918392 (p.E21K) was defined as the APOE ε5 allele29,32–34 and was the third-most frequent variant after the APOE ε7-defining rs140808909 (p.E262K) and rs190853081 (p.E263K), with an MAF of 0.00348 (Table 5). rs121918392 (p.E21K) was also associated with dyslipidemia and atherosclerosis.29,32–34
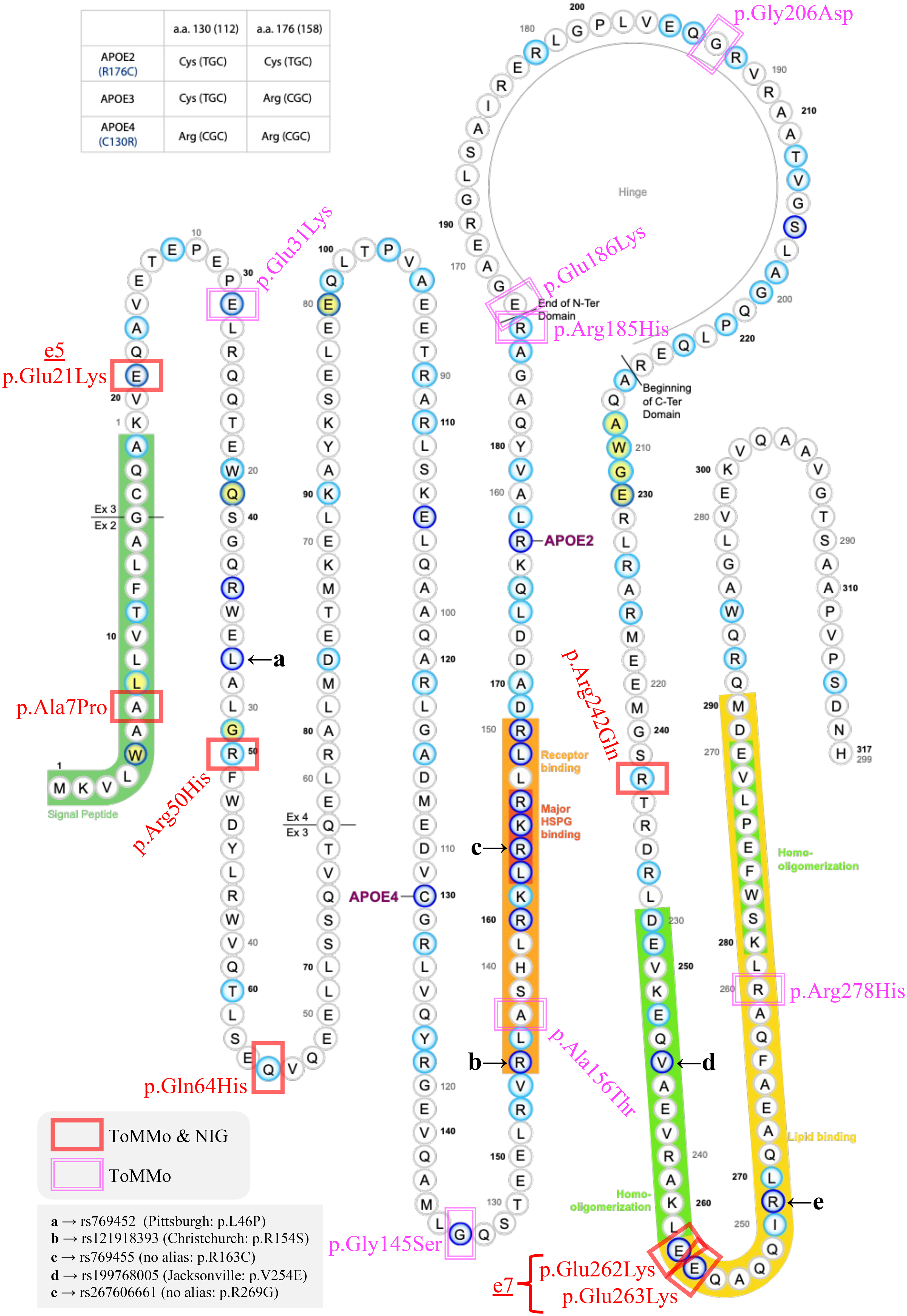
APOE RMVs identified in this study. The 14 APOE RMVs identified in this study were added to the Alzforum MUTATIONS APOE diagram (https://www.alzforum.org/mutations/apoe). RMVs identified in both ToMMo and NIG cohorts are marked in red, whereas those identified only in ToMMo are marked in magenta. The five RMVs previously reported are labeled with the letters “a” to “e”. The ε7 allele, which is the primary focus of this paper, is defined by two RMVs that replace two glutamic acid (Glu, E) residues with lysine (Lys, K) at the entrance of the lipid-binding domain (colors are visible in the online version).
We also examined the presence of loss-of-function variants in the NIG cohort (1750 subjects; Table 1) that underwent WES. No loss-of-function mutations were detected.
In the analysis of 3307 subjects from the ToMMo cohort (Table 1), we identified 14 APOE RMVs (Table 5, Figure 3). Seven variants were identified in both the NIG and ToMMo cohorts. The following seven other variants were found in the ToMMo cohort but not in the NIG cohort: rs201672011 (p.E31K), rs748703149 (p.G145S), rs954186991 (p.A156T), rs1339651557 (p.R185H), rs1969871875 (p.E186K), rs1385100920 (p.G206D), and rs767339630 (p.R278H). All carriers were heterozygous, and no homozygotes were identified. We identified one homozygous individual for the two variants in complete LD, rs140808909 (p.E262K) and rs190853081 (p.E263K): male sex; age at entry, 65 years; APOE genotype, ε3*3; and body mass index, 23.0. This individual exhibited LDL-C, HDL-C, and TG levels of 96, 49, and 67, respectively, all within the normal ranges (TC data were not available).
Five previously reported variants (Figure 3), rs769452 (p.L46P: Pittsburgh), 17 rs121918393 (p.R154S: Christchurch), 18 rs769455 (p.R163C), 19 rs199768005 (p.V254E: Jacksonville), 20 and rs267606661 (p.R269G), 21 were not detected in either the NIG or ToMMo cohorts. Furthermore, these variants were not observed in the Japanese multi-omics database jMorp (version 60KJPN) 35 (Supplemental Table 3). These variants were more frequently observed in non-East Asian populations (Supplemental Table 3).
Relationship between identified APOE RMVs and the common APOE alleles, ε2, ε3, and ε4
Clarifying the common APOE alleles (ε2, ε3, and ε4) on which each identified APOE RMV is located is crucial for understanding the potential genetic relationship between AD and APOE. Therefore, for each APOE RMV, we categorized subjects from the NIG and ToMMo cohorts (Tables 1 and 2) according to the common APOE genotypes: ε2*2, ε2*3, ε2*4, ε3*3, ε3*4, and ε4*4 (Supplemental Table 2). The most common APOE genotype among APOE RMV carriers was ε3*3. Several individuals were also identified with ε3*4 and ε2*3, and one individual with ε4*4. None of the APOE RMV carriers had ε2*2 or ε2*4. Variant rs121918392 (p.E21K) that defines the ε5 allele, and two variants, rs140808909 (p.E262K) and rs190853081 (p.E263K), that are in complete LD and define the ε7 allele, appeared to co-segregate with the ε3 allele (Supplemental Table 2). The rs767339630 (p. R278H) variant was observed in a subject with ε4*4, suggesting that this variant is likely to co-segregate with the ε4 allele (Supplemental Table 2).
Ethnicity dependent frequencies of APOE RMVs
To investigate whether the 14 APOE RMVs identified in the NIG and ToMMo cohorts were present in other populations and to assess their frequencies, we explored public databases (Supplemental Table 3). We initially examined the jMorp database. 35 Alternative alleles were observed in all 14 variants in this database. For two variants of complete LD, rs140808909 (p.E262K) and rs190853081 (p.E263K), 13 homozygous individuals were identified in the database (version 60KJPN). This finding was consistent with our analysis, which showed that these two APOE RMVs exhibited the highest alternative allele frequencies. The remaining 12 variants were exclusively observed in heterozygous individuals.
Four variants, rs201672011 (p.E31K), rs762461580 (p.R50H), rs370594287 (p.Q64H), and rs748703149 (p.G145S), were also observed in populations other than East Asian populations (Supplemental Table 3). In contrast, six variants, rs121918392 (p.E21K), rs954186991 (p.A156T), rs1339651557 (p.R185H), rs267606663 (p.R242Q), rs140808909 (p.E262K), and rs190853081 (p.E263K), were highly ethnicity-dependent and observed only in the East Asian population (Supplemental Table 3). Information regarding the frequencies of the four variants rs1969812567 (p.A7P), rs1969871875 (p.E186K), rs1385100920 (p.G206D), and rs767339630 (p.R278H) was not available for other populations (Supplemental Table 3).
Effect of APOE RMVs on AD
To investigate whether the 14 identified RMVs (Table 5, Figure 3) were genetically associated with AD, we conducted a case-control study involving 6261 AD cases and 16,331 controls (Table 3). Among the 14 RMVs examined, genotype data for four variants were obtained by imputation. Two variants, rs140808909 (p.E262K) and rs190853081 (p.E263K), which define the ε7 allele, exhibited a statistically significant association with AD in Models 1 (without covariate adjustment: pBonferroni = 0.0428, and OR [95% CI] = 0.70 [0.54–0.92]), 2 (adjusted for sex and age: pBonferroni = 0.0293, and OR [95% CI] = 0.68 [0.51–0.90]), and 4 (adjusted for the ε2 allele: pBonferroni = 0.0364, and OR [95% CI] = 0.70 [0.53–0.91]) (Table 4, Figure 1). These results suggest that the ε7 allele may confer protection against AD (OR < 1.00). However, in Models 3 and 5, where the ε4 allele was included as a covariate, the association was no longer significant: pBonferroni = 1.000 and OR (95% CI) = 0.87 (0.66–1.15) in Model 3; and pBonferroni = 1.000 and OR (95% CI) = 0.85 (0.64–1.14) in Model 5 (Table 4, Figure 1). Since the ε7 allele co-segregates with the ε3 allele and never with the AD risk-associated ε4 allele (Supplemental Table 2), its protective effect is likely attributable to the ε3 allele. Furthermore, since these two RMVs were in complete LD (Supplemental Figure 2), they exhibited identical p-values and OR (95% CI) across all models examined (Table 4, Figure 1).
Association between APOE genotypes/alleles and AD in APOE RMV carriers
By focusing on carriers of the four RMVs analyzed in the case-control study (Table 4, Figure 1), we investigated whether APOE genotypes and alleles were associated with AD. As no carriers of APOE RMVs with the ε2*2, ε2*4, or ε4*4 genotypes were identified, they were excluded from the analysis. Additionally, due to the low frequencies of the ε2*3 genotype and ε2 allele, they were excluded from the analysis. As a result, the comparison of genotypes focused on ε3*3 and ε3*4, and we compared ε3 and ε4 for alleles. The results of the analyses are presented in Figure 2 and Supplemental Table 4. For reference, the OR (95% CI) for the entire cohort, including both carriers and non-carriers of APOE RMVs and denoted as “All”, was also provided, derived from the data in Table 3. The OR (95% CI) for both APOE genotypes and alleles exhibited similar trends. Specifically, for the three RMVs defining the ε5 (rs121918392) and ε7 (rs140808909 and rs190853081) alleles, the risk associated with the ε4 allele showed a tendency to decrease. For example, in a comparison between the ε3*3 and ε3*4 genotypes, the OR (95% CI) for All was 3.13 (2.93–3.35), whereas it was 1.79 (0.78–4.26) for ε5 (rs121918392). In contrast, for rs762461580, there was a trend towards an increase in risk associated with the ε4 allele. For example, in the same genotype comparison, the OR (95% CI) for All was 3.13 (2.93–3.35), but was 4.18 (1.15–14.00) for rs762461580.
Comparison of lipoprotein levels between APOE RMV carriers and non-carriers
We compared blood lipoprotein levels between carriers and non-carriers of APOE RMVs in J-ADNI participants (508 subjects) from the NIG cohort and in participants from the ToMMo cohort (TC, 234 subjects; and non-TC, 1630 subjects). No statistically significant differences were observed between APOE RMV carriers and non-carriers in terms of TC, LDL-C, HDL-C, TG, or LH ratios (Supplemental Figure 4).
Discussion
In this study, we identified 14 APOE RMVs in 5896 Japanese individuals (Table 5, Figure 3). Some APOE RMVs were specific to East Asian populations, including the Japanese population (Supplemental Table 3). Notably, variants previously reported in Western populations (e.g., the Christchurch 18 and Jacksonville 20 variants) were absent in the Japanese population (Supplemental Table 3). These findings suggested that RMVs exhibit distinct ethnic and regional specificities. To investigate their genetic association with AD, we conducted a case-control study involving 22,592 individuals (Table 3). Two APOE RMVs, rs140808909 and rs190853081, which were in complete LD (Supplemental Figure 2) and define the ε7 allele,29–31 exhibited a statistically significant association with AD (Table 4, Figure 1). Although this effect may be influenced by the ε3 allele, it suggests a potential protective effect (Table 4, Figure 1). To the best of our knowledge, this is the first report to demonstrate an association between APOE RMVs and AD in East Asians. Future studies with larger sample sizes are strongly recommended to validate these findings as reproducibility is crucial. Given the genetic diversity and ethnic specificity of genomic variants, 36 detailed analyses for each ethnic group are essential. 37
In this exploratory study, we identified 14 APOE RMVs (Table 5, Figure 3), with the ε7 allele29–31 being the most intriguing due to its potential protective effects against AD (Table 4, Figure 1). The two RMVs that define the ε7 allele replace two glutamate (E) residues with lysine (K) at the entry point of the lipid-binding domain on the C-terminal side of APOE (Figure 3). This substitution reduces the binding affinity of the ε7 isoform to the LDL receptor (LDLR) and increases its affinity for heparin. 38 No previous studies have reported a genetic association between the ε7 allele and AD. However, Youn et al. 39 suggested that the ε7 allele may be involved in cognitive impairment based on a study of 344 amnestic patients and 345 controls in a Korean population. They identified three heterozygous carriers of the ε7 allele, all of whom belonged to the amnestic group. Two of these individuals were clinically diagnosed with AD and small vessel disease, and the remaining one was diagnosed with subjective cognitive impairment. They also predicted the three-dimensional protein structure of the ε7 isoform, indicating that its C-terminal structure resembled that of the ε4 isoform, despite having the ε3 isoform background. They concluded that the ε7 isoform may behave like the ε4 isoform and potentially exert adverse effects on cognitive function. Although our study suggests that the ε7 allele may have a protective effect against AD, the relationship between cognitive function and the ε7 allele remains an important topic for future investigation.
We compared the peripheral blood cholesterol levels (TC, LDL-C, HDL-C, TG, and LH ratios) between carriers and non-carriers of the identified APOE RMVs. Contrary to expectations, no significant differences were observed in cholesterol levels (Supplemental Figure 4). Due to the rarity of APOE RMVs, all carriers were grouped into a single category for analysis in this study. Different APOE RMVs may have varying effects on peripheral blood cholesterol levels; therefore, each APOE RMV should ideally be analyzed individually. For example, a study on the ε5 and ε7 alleles revealed a trend toward higher TC (>220 mg/dL) and TG (>150 mg/dL) levels compared with those in controls. 40 Considering the low population frequency of APOE RMVs (<1%), comparative analyses with larger sample sizes are required.
This study had several limitations. First, the number of identified RMVs (Table 5, Figure 3) was lower than expected. Therefore, further studies with larger Japanese cohorts are needed, particularly including individuals aged 65 and older for late-onset AD analysis. Second, we have not confirmed the genetic association between AD and the ε7 allele (rs140808909 and rs190853081) in a larger, independent East Asian sample set, highlighting the need for further investigation. Third, as this study was based on cross-sectional data, the longitudinal relationship between RMVs and AD risk remains unclear. Future studies should incorporate longitudinal data such as those from the ADNI project (https://adni.loni.usc.edu) to clarify the temporal relationships among RMVs, AD onset, and disease progression. Finally, no functional assays have been conducted for RMVs, including the ε7 allele, and their impact on AD pathology and the underlying molecular mechanisms remain unclear. Given APOE's role in lipid metabolism, elucidating its relationship with key AD pathologies like Aβ and tau is crucial. Notably, the ε7 allele has been linked to dyslipidemia and atherosclerosis,29–31 yet few studies have examined its binding to LDLR and heparin. 38 As these molecules play critical roles in Aβ clearance and tau aggregation,41,42 the ε7 isoform may influence AD pathology through these pathways.
APOE plays a crucial role in the pathophysiology of AD, particularly in relation to Aβ and tau.43,44 Previous research has largely focused on the major missense alleles of APOE, namely ε2, ε3, and ε4. However, interest in APOE RMVs, including the Christchurch variant (rs121918393: p.R154S [p.R136S]),18,45–49 is growing. Further research in this area is expected to provide valuable insights into their functional significance. Such studies will contribute to a deeper understanding of APOE's role in AD pathophysiology. To elucidate the biological implications of the 14 APOE RMVs identified in this study, additional investigations using in vitro (cultured cells) and in vivo (animal models such as mice) approaches are required.
Supplemental Material
sj-pdf-1-alz-10.1177_13872877251340710 - Supplemental material for Association of rare APOE missense variants with Alzheimer's disease in the Japanese population
Supplemental material, sj-pdf-1-alz-10.1177_13872877251340710 for Association of rare APOE missense variants with Alzheimer's disease in the Japanese population by Akinori Miyashita, Ai Obinata, Norikazu Hara, Risa Mitsumori, Daita Kaneda, Yoshio Hashizume, Terunori Sano, Masaki Takao, Ramil Gabdulkhaev, Mari Tada, Akiyoshi Kakita, Akira Arakawa, Maho Morishima, Shigeo Murayama, Yuko Saito, Hiroyuki Hatsuta, Tomoyasu Matsubara, Akio Akagi, Yuichi Riku, Hiroaki Miyahara, Jun Sone, Mari Yoshida, Haruyasu Yamaguchi, Tamao Tsukie, Mai Hasegawa, Kensaku Kasuga, Masataka Kikuchi, Hiroyasu Akatsu, Ryozo Kuwano, Takeshi Iwatsubo, , Shumpei Niida, Kouichi Ozaki and Takeshi Ikeuchi in Journal of Alzheimer's Disease
Supplemental Material
sj-xlsx-2-alz-10.1177_13872877251340710 - Supplemental material for Association of rare APOE missense variants with Alzheimer's disease in the Japanese population
Supplemental material, sj-xlsx-2-alz-10.1177_13872877251340710 for Association of rare APOE missense variants with Alzheimer's disease in the Japanese population by Akinori Miyashita, Ai Obinata, Norikazu Hara, Risa Mitsumori, Daita Kaneda, Yoshio Hashizume, Terunori Sano, Masaki Takao, Ramil Gabdulkhaev, Mari Tada, Akiyoshi Kakita, Akira Arakawa, Maho Morishima, Shigeo Murayama, Yuko Saito, Hiroyuki Hatsuta, Tomoyasu Matsubara, Akio Akagi, Yuichi Riku, Hiroaki Miyahara, Jun Sone, Mari Yoshida, Haruyasu Yamaguchi, Tamao Tsukie, Mai Hasegawa, Kensaku Kasuga, Masataka Kikuchi, Hiroyasu Akatsu, Ryozo Kuwano, Takeshi Iwatsubo, , Shumpei Niida, Kouichi Ozaki and Takeshi Ikeuchi in Journal of Alzheimer's Disease
Footnotes
Acknowledgments
We wish to thank all the participants for their involvement in this study. We also gratefully acknowledge the ToMMo cohort (Tables 1 and ![]() ) for providing individual genomic variant data regarding APOE and its surrounding regions as well as various datasets from the subjects (research ID: 2019-0074). In preparation for this manuscript, an initial proofreading in English was conducted using ChatGPT-4o and subsequently reviewed by native English speakers.
) for providing individual genomic variant data regarding APOE and its surrounding regions as well as various datasets from the subjects (research ID: 2019-0074). In preparation for this manuscript, an initial proofreading in English was conducted using ChatGPT-4o and subsequently reviewed by native English speakers.
ORCID iDs
Ethical considerations
This study was approved and conducted according to the ethical guidelines of the Ethics Committee of Niigata University and all participating medical and research institutions (approval number of the Genetic Ethics Review Committee, Niigata University: G2017-0012, G2018-0034, 2018-0409, and G2022-0009).
Consent to participate
Written informed consent was obtained from some participants. For the others, an opt-out method was employed with the approval of the Ethics Committee of Niigata University (approval numbers: G2017-0012, G2018-0034, 2018-0409, and G2022-0009).
Author contributions
Akinori Miyashita (Conceptualization; Formal analysis; Funding acquisition; Investigation; Methodology; Project administration; Resources; Supervision; Validation; Visualization; Writing – original draft; Writing – review & editing); Ai Obinata (Formal analysis; Investigation; Methodology; Writing – review & editing); Norikazu Hara (Formal analysis; Investigation; Methodology; Validation; Writing – review & editing); Risa Mitsumori (Formal analysis; Investigation; Methodology; Writing – review & editing); Daita Kaneda (Formal analysis; Funding acquisition; Investigation; Methodology; Resources; Writing – review & editing); Yoshio Hashizume (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Terunori Sano (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Masaki Takao (Formal analysis; Funding acquisition; Investigation; Methodology; Resources; Writing – review & editing); Ramil Gabdulkhaev (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Mari Tada (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Akiyoshi Kakita (Conceptualization; Formal analysis; Funding acquisition; Investigation; Methodology; Resources; Writing – review & editing); Akira Arakawa (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Maho Morishima (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Shigeo Murayama (Formal analysis; Funding acquisition; Investigation; Methodology; Resources; Writing – review & editing); Yuko Saito (Formal analysis; Funding acquisition; Investigation; Methodology; Resources; Writing – review & editing); Hiroyuki Hatsuta (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Tomoyasu Matsubara (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Akio Akagi (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Yuichi Riku (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Hiroaki Miyahara (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Jun Sone (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Mari Yoshida (Formal analysis; Funding acquisition; Investigation; Methodology; Resources; Writing – review & editing); Haruyasu Yamaguchi (Formal analysis; Investigation; Methodology; Resources; Writing – review & editing); Tamao Tsukie (Formal analysis; Investigation; Methodology; Writing – review & editing); Mai Hasegawa (Formal analysis; Investigation; Methodology; Writing – review & editing); Kensaku Kasuga (Formal analysis; Investigation; Methodology; Writing – review & editing); Masataka Kikuchi (Formal analysis; Funding acquisition; Investigation; Methodology; Writing – review & editing); Hiroyasu Akatsu (Resources; Writing – review & editing); Ryozo Kuwano (Resources; Writing – review & editing); Takeshi Iwatsubo (Funding acquisition; Resources; Writing – review & editing); Shumpei Niida (Funding acquisition; Resources; Writing – review & editing); Kouichi Ozaki (Formal analysis; Funding acquisition; Investigation; Methodology; Resources; Writing – review & editing); Takeshi Ikeuchi (Conceptualization; Funding acquisition; Investigation; Methodology; Project administration; Resources; Supervision; Writing – original draft; Writing – review & editing).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by: (1) JSPS Grant-in-Aid for Scientific Research (KAKENHI) (C), 21K07271 (AM); (2) JSPS Grant-in-Aid for Scientific Research (KAKENHI) (B), 21H03537 (AM); (3) AMED, JP21dk0207055 (AM); (4) JSPS Grant-in-Aid for Scientific Research (KAKENHI), 22H04923 (DK); (5) AMED, JP21wm0425019 (MT); (6) Intramural fund from NCNP, Grant Number 6-8 (MT); (7) JSPS Grant-in-Aid for Scientific Research (KAKENHI) (C), 18K06506 (MT); (8) JSPS Grant-in-Aid for Scientific Research (KAKENHI) (C), 21K06417 (MT); (9) JSPS Grant-in-Aid for Transformative Research Areas, 22H04923 (MT); (10) the Research Committee of Prion Disease and Slow Virus Infection, the Ministry of Health, Labour and Welfare Health and Labour Sciences Research Grants, Japan (MT); (11) AMED, JP24wm0425019 (AK) ; (12) AMED, JP21wm0425019 (YS, SM); (13) JSPS Grant-in-Aid for Scientific Research (KAKENHI), 22H04923 (CoBiA) (YS); (14) JSPS Grant-in-Aid for Scientific Research (KAKENHI) (C), 22K07550 (YS); (15) MHLW Research on rare and intractable diseases Program Grant Number JPMH23FC1008 (YS); (16) AMED, JP21wm0425019 (MY); (17) AMED, JP18kk0205009 (SN); (18) AMED, JP21dk0207045 (SN); (19) AMED, JP22dk0207060 (SN); (20) AMED, JP21dk0207045 (KO); (21) AMED, JP22dk0207060 (KO); (22) Research Funding for Longevity Sciences from NCGG (24-15) (KO); (23) a grant from the Japanese Ministry of Health, Labour, and Welfare for Research on Dementia (KO); (24) AMED, JP23wm0525019 (MK); (25) AMED, JP23dk0207060 (MK); (26) JSPS Grant-in-Aid for Scientific Research (KAKENHI) (B), 23K27515 (TIwatsubo); (27) AMED, JP22dk0207060 (TIkeuchi)
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The data supporting the findings of this study are available upon request from the corresponding authors (AM and TIkeuchi). However, the WES data used in this study are not publicly available due to privacy or ethical restrictions.
Supplemental material
Supplemental material for this article is available online.
Correction (June 2025):
The originally published supplemental material lacked figure numbers. Some figure legends were also presented in Japanese rather than English. Additionally, the file was initially provided in PowerPoint format, which is not ideal for archiving or referencing. These issues have been corrected, and the supplemental material has been updated to a properly formatted PDF version.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
