Abstract
Nucleos(t)ide analogues (NAs) are a mainstay of therapy for chronic hepatitis B (CHB) infections and have a profound effect on hepatitis B virus (HBV) suppression. We report a rare case of HBV reactivation in a CHB patient without cirrhosis following cessation of NA therapy that resulted in acute liver failure requiring liver transplantation. Investigation of the viral genetics and host immune responses suggest that viral mutations known to promote virus replication are associated with reactivation, whereas adaptive immunity to HBV remained defective in this patient. Viral sequencing may be useful for identifying mutations that are unfavorable for therapy withdrawal.
Introduction
Chronic hepatitis B (CHB) infection is a challenging public health issue which is currently uncurable. Most patients are untreated; those who are treated usually require lifelong treatment with nucleos(t)ide analogues (NAs), which inhibit hepatitis B virus (HBV) replication but do not eradicate the virus [1,2]. A functional cure—loss of hepatitis B surface antigen (HBsAg) with or without seroconversion of hepatitis B surface antigen antibody (HBsAb)—has been seen as a surrogate end point of treatment but is rarely achieved with NAs. Withdrawal of NA treatment is being considered as a possible path to functional cure by provoking HBV reactivation, followed by immune consolidation and subsequent HBsAg loss in some patients [2,3]. However, in rare cases, NA therapy withdrawal causes severe hepatitis flares, hepatic decompensation, or death. Predictors of hepatic decompensation or death with NA withdrawal have not been well established [3]. Here, we present a rare case of CHB infection reactivation in a patient without cirrhosis following cessation of NA therapy that resulted in acute liver failure (ALF) requiring liver transplantation. We performed detailed serological, histological, virological, and cellular analysis in this patient and compared the results to that of a group of CHB patients without ALF as well as normal blood donors (NBDs) in order to investigate the potential mechanisms leading to this unusual outcome.
Case report
A 63-year-old man with a history of hepatitis B e antigen (HBeAg) positive CHB infection was admitted at Scripps Clinic, La Jolla in October 2019 for ALF. He underwent liver biopsy in 2006 showing CHB with minimal activity and minimal fibrosis. In 2006, his pretreatment HBV DNA level was 316,000 IU/mL and was initiated on treatment with lamivudine initially in February 2006 but for unclear reasons was switched to entecavir 0.5 mg daily in June 2006 with subsequent undetectable HBV DNA levels. In April 2013, his HBV DNA increased from undetectable to 1500 IU/mL and he experienced HBeAg loss (HBeAg was positive from 2009 until 2013). He was continued on entecavir 0.5 mg, and in 2014, his HBV DNA levels further increased to 78,230 IU/mL and he was transitioned to tenofovir-alafenamide (VemlidyTM, Gilead Sciences) with subsequent negative HBV DNA levels. In July 2017, he had reverse seroconversion when a positive HBeAg was found along with a still undetectable HBV DNA level. In October 2019, following his undetectable HBV DNA level, he voluntarily discontinued all medications without informing his physicians, including tenofovir-alafenamide and subsequently developed jaundice, bloating, and fatigue.
On admission, the patient demonstrated hepatic encephalopathy. Laboratory testing (Figure 1A) revealed a rebound of HBV DNA (36,400,000 IU/mL) and alanine aminotransferase flare (5265 IU/L). He was positive for HBsAg and HBeAg but non-reactive for HBsAb. His international normalized ratio (INR) was 4.9. A computerized tomography of the abdomen showed a heterogeneous liver with enlarged portal veins but was otherwise normal. Further evaluation for the patient’s ALF showed that he was negative for hepatitis A virus, hepatitis C virus, hepatitis delta virus, hepatitis E virus, herpes simplex virus, Epstein–Barr virus, cytomegalovirus, serum markers for autoimmune hepatitis, and findings on urine drug screen. Despite resuming antiviral therapy, his INR increased to 5.8, his hepatic encephalopathy worsened, and he ultimately underwent deceased donor liver transplantation on 11/4/2019. The liver explant showed acute fulminant confluent (sub-massive, 70%) liver cell necrosis, consistent with HBV reactivation (Figure 1B). Clinical characteristics of the patient. (A) Selected laboratory data from prior to admission until post transplantation. (B) Histopathologic changes in the patient liver. Explant liver shows congested nutmeg appearing liver with multiple micronodular regenerative nodules (<0.3 cm) on the cut surface. Microscopically, there is confluent (submassive) liver cell necrosis that involves approximately 70% of the liver parenchyma. There is hemorrhage with exuberant small ductular proliferation in the areas of liver cell dropout. Portal tracts show mild chronic inflammation composed of lymphocytes and scattered plasma cells with focal interface activity. There is marked intracanalicular and intracytoplasmic cholestasis within the residual viable liver cells. Left, trichrome special stain shows portal with focal periportal fibrosis. Right, there are ground glass appearing hepatocytes and they are positive for hepatitis B surface antigen by immunohistochemistry. The findings are consistent with acute fulminant confluent (submassive) liver cell necrosis, due to hepatitis B viral reactivation.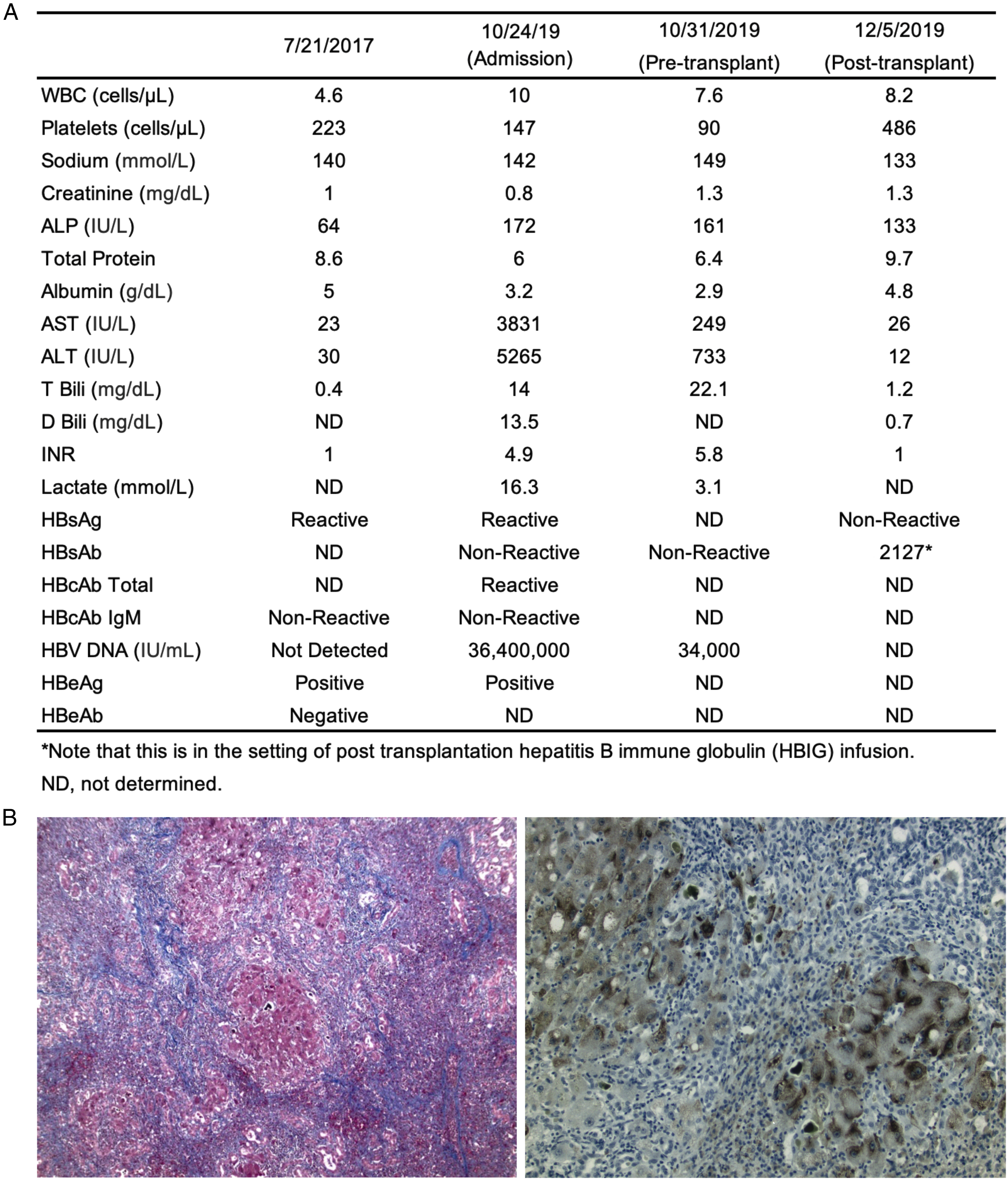
A phylogenetic analysis of full genome sequences of HBV in the patient’s liver biopsy and plasma collected on 10/31/2019 pre-transplant (GenBank No.: MT757436 - MT757457) confirmed that the patient was infected with a genotype B and serotype ayw virus, which is closest to the quasi-subgenotype B3 isolate AB219426 (98.7% nucleotide identity; Figure 2A) [4]. The virus in both the patient’s blood and liver had multiple mutations correlated with reactivation [5,6] (Figures 2B and 2C), including the basal core promoter (BCP) mutations A1762T/G1764A, T1753C, and T1768A that can reduce HBeAg expression while enhancing viral replication [7-9]. Intriguingly, a high prevalence of the A1752G mutation that may reduce HBV DNA replication [10] was found to coexist in the same genome, indicating distinct biological mechanisms involved in selection of these mutations during the long-term chronic infection. The pre-core stop codon mutation G1896A that can abrogate HBeAg production was detected in a small fraction of the virus in the liver (1 in 12 HBV genomic sequences). BCP overlaps with the X region, with the mutations T1753C, A1762T, G1764A, and T1768A causing I127T, K130M, V131I, and F132I amino acid substitutions in the HBx protein, respectively, which may promote HCC formation and contribute to ALF [11,12]. Several HBsAg mutations associated with reactivation, for example, P127S in the “a” determinant (the major target of HBsAb) within the major hydrophilic region, and S207N in a known T cell epitope, were also detected. These mutations are known to be responsible for vaccine- and diagnostic-escape [6,13]. Strikingly, we did not observe any previously reported drug resistance mutations within the reverse transcriptase domain [6,14], for example, I169T, T184G, S202I, M250V, M204I/V for entecavir, and A181V and N236T for tenofovir. HBV quasi-species and variations in the patient. (A) Phylogenetic analysis of complete HBV genomes for the patient and reference stains from nine genotypes A–I. *J is a putative genotype isolated from one individual [18,19]. Teal and orange dots indicate genomes from the patient liver (n = 12) and plasma (n = 10), respectively. Reference sequences (black dots) for genotypes A–H were obtained from the Hepatitis B Virus database (HBVdb) (accession numbers EU054331, X02763, AB219428, D00331, GQ924620, GQ358158, AF121240, FJ904433, AB106564, AY738145, AY090458, X75658, AB064313, AF160501, AY090454, and FJ356716), and genotypes I (AB562462.1 and FJ023671.1) and J (AB486012.1) were downloaded from GenBank. The red dot represents the genotype B ayw strain (AB219426). MEGA X was used to generate the maximum likelihood phylogenetic tree. (B–C) Frequency of major mutations in the HBV genomes from the patient liver (teal) and plasma (orange). (B) Nucleotide mutations in the base core promoter (BCP)/X gene and precore region. (C) Amino acid mutations in HBsAg.
Flow cytometry analysis of T and B cell subsets in the blood of the patient in comparison to four other CHB patients and three NBD revealed overall similar circulating immune profiles for the patient and other CHB patients (Figure 3A). The patients had a significantly lower frequency in IgG+ B cells but more CD27−CD21− atypical memory B cells (atMBC) in both global and HBsAg-specific B cells than NBD. The HBsAg-specific B cells in the patient displayed elevated expression of inhibitory and exhaustion markers (PD-1hiT-bethi). Compared to the CHB patients without ALF and NBDs, our patient had fewer circulating CD19+ total B cells, higher frequency of IgM+ B cells, and more HBsAg-specific B cells. We did not detect signatures of T and B cell activation in this patient (Data not shown). However, reduced intracellular IgG expression and HBsAg-specific IgG+ antibody secreting cell response of the patient in comparison to NBD were observed when the cells were stimulated in vitro (Figures 3B and 3C), suggesting that HBsAg-specific B cells in the patient have impaired capacity to produce HBsAb. Overall, cessation of NA therapy did not restore anti-HBV immune responses in this patient. Characterization of host immune responses in the patient. (A) Comparison of immune profiles in the peripheral blood mononuclear cell (PBMC) of CHB patients and NBDs. Our patient was shown in the CHB group as a red dot. cMBCs, classic memory B cells (CD27+CD21+); atMBCs, atypical memory B cells (CD27−CD21−). p values were calculated by two-tailed Mann–Whitney tests. *, p < 0.05. ns, not significant. (B and C) In vitro activation of virus-specific B cells by anti-human IgG/IgM and CD40L. The expression of intracellular IgG and HBsAg-specific IgG+ antibody secreting cells (ASCs) were measured by flow cytometry (B) and ELISPOT (C), respectively. The wells shown in the ELISPOT were plated with 0.6 × 105 PBMC.
Discussion
Reactivation of HBV, characterized by a rise in HBV DNA by at least 1-log compared to baseline, may be spontaneous, or triggered by chemotherapy, immunosuppression, solid organ transplantation, or discontinuation of antiviral medication [15]. Instances of ALF due to reactivation of HBV have been described in the setting of patients receiving anticancer therapy and cytotoxic therapy [15]. However, ALF from discontinuation of NA therapy is rare and usually occurs in patients with impaired hepatic reserve due to cirrhosis [1]. The case presented herein shows that liver failure can occur with the cessation of antiviral therapy alone in the setting of a relatively normal liver. Our patient did not have underlying cirrhosis or chronic liver disease based on laboratory parameters, prior liver biopsy, and relatively recent transient elastography. The possibility of reactivation causing liver failure is especially relevant because of recent efforts to achieve functional cure in HBeAg negative patients by cessation of NA therapy [3]. Notably, when our patient stopped his antiviral treatment, he would not have been eligible for cessation of NA therapy as his HBeAg was positive.
HBsAg loss is considered a functional cure for HBV infection but, unfortunately, is a rare occurrence following decades of therapy [2]. A recent large prospective trial of therapy cessation, investigating HBeAg negative chronic HBV infected patients who met the Asian Pacific Association for the Study of Liver Disease stopping criteria, found that 42 out of 691 patients (308 with cirrhosis) who stopped therapy had seroconversion of HBsAg during a median follow-up of 155 weeks. The estimated annual incidence of seroconversion was 1.78% in those who stopped therapy compared to 0.15% in those who did not. Seven out of 308 patients with cirrhosis who stopped therapy developed hepatic decompensation and three of them died [2]. In nine other NA therapy withdrawal trials, 11 (0.2%–1.6%) patients experienced liver-related deaths [3]. Despite risk of decompensation, the possibility of a functional cure remains alluring. The lack of reliable predictors of poor outcomes following NA therapy withdrawal makes the pursuit even more challenging and is a critical area of future research [3]. Unfortunately, large virologic studies of patients who develop ALF after NA withdrawal are unlikely. Thus, we believe reports of selected cases are important and enlightening.
Our investigation suggests both viral and host factors play a role in the ALF seen in a patient with HBeAg positive CHB infection, without cirrhosis, who developed liver failure after stopping tenofovir-alafenamide. While we did not detect a significant difference in B and T cells in comparison to CHB patients to explain the ALF, viral sequencing in the liver explant and plasma detected mutations that are associated with HBV reactivation. Similar to previous reports [16,17], the HBsAg-specific B cells in the patient showed phenotypic alterations and impaired antibody-producing potential which may cause insufficient serum HBsAb levels. Drug withdrawal did not improve antiviral immune responses in this patient. Further studies are required to understand the virus and host immune responses of patients with ALF participating in CHB therapy withdrawal to help predict the risk of ALF before the practice of therapy withdrawal becomes routine.
Footnotes
Acknowledgements
The authors thank Stefan Wieland, PhD for advice in HBV genome analysis, Jasneet Aneja at Massachusetts General Hospital (Boston, MA, USA), and Jamie Case, PhD at Scripps Clinic Bio-Repository (La Jolla, CA, USA) for assistance in sample coordination. ML is partly supported by the NIH grants AI079031 and AI123861.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: ML is partly supported by the NIH grants AI079031 and AI123861.
