Abstract
Objective:
Examine if the gut microbiota composition changes across repeated samples in paediatric-onset multiple sclerosis (MS) or monophasic-acquired demyelinating syndromes (monoADS).
Methods:
A total of 36 individuals (18 MS/18 monoADS) with ⩾2 stool samples were included. Stool sample-derived DNA was sequenced. Alpha/beta diversities and genus-level taxa were analysed.
Results:
Mean ages at first sample procurement (MS/monoADS) = 18.0/13.8 years. Median time (months) between first/second samples = 11.2 and second/third = 10.3. Alpha/beta diversities did not differ between stool samples (p > 0.09), while one genus – Solobacterium did (p = 0.001).
Conclusions:
The gut microbiota composition in paediatric-onset MS and monoADS exhibited stability, suggesting that single stool sample procurement is a reasonable first approach.
Keywords
Introduction
Many gut microbiota-related studies in multiple sclerosis (MS) have typically relied on procuring a single sample from participants. 1 Little is known about whether this is sufficient.
Individuals with paediatric-onset MS and related diseases represent unique opportunities to explore the gut microbiota early on in the disease process, such as the recently demonstrated relationship between the gut microbiome (using metagenomics) and disease activity. 2 We assessed if the gut microbiota composition remained stable across repeated stool samples (⩾2) procured over the short-term in individuals with paediatric-onset MS and monophasic-acquired demyelinating-syndromes (monoADS).
Method
Individuals with symptom onset <18 years and no antibiotic use ⩽30 days prior, diagnosed with MS (McDonald criteria, 2017) or monoADS who provided ⩾2 stool samples were eligible. See Supplementary Data (Methods) for details. Included individuals were enrolled from six sites across Canada and one in the United States (November/2015-March/2018).
Cohort characteristics were captured by trained coordinators using standardized forms and questionnaires were completed at home by parents/participants.
Stool samples were shipped on ice, stored at −80oC, then DNA was extracted, amplified and sequenced (V4 hypervariable region, 16S rRNA gene, via the Illumina MiSeq platform), and clustered into amplicon sequence variants (ASVs) using Deblur via QIIME2 (Quantitative Insights Into Microbial Ecology; v.2019.1). 3 The gut microbiota’s composition was assessed using alpha (Shannon) and beta (unweighted UniFrac) diversity metrics.
Characteristics of the MS and monoADS participants were described. Alpha diversity was compared between the first and second stool samples using the pairwise Wilcoxon signed-rank test and between all three samples using the Friedman test. Unweighted UniFrac was displayed using the first two components from principal coordinates (PC) analysis and confidence ellipsoids. Permutational multivariate analysis of variance (PERMANOVA) was used to compare between stool samples. Phylum-level relative abundances were summarized. The most important genus-level taxa were identified using feature volatility analysis. 4 The top five genera were analysed for variability between stool samples using linear mixed effects models with sample number (first, second, third) as the response variable, including a participant-specific random intercept, adjusted for age at stool sample, sex and diagnosis (MS/monoADS). 4 Longitudinal similarity of all samples was compared between MS/monoADS participants using the non-parametric microbial interdependence test (NMIT). 4 Complementary analyses (MS cases only) explored whether any change in disease-modifying therapy (DMT) use associated with microbiome changes (alpha diversity). QIIME2 v2020.11, R v4.0.1 and SPSS v23.0 were used for analyses.3,5,6 Informed consent/assent and ethics were obtained (Supplementary file).
Results
A total of 36 individuals provided 77 stool samples (all participants provided 2, 5 of them provided 3) over a 26-month period. The median time (months) between the first/second sample = 11.2 (range = 2.0–25.0) and second/third = 10.3 (range = 1.8–13.6). Participant characteristics, including the Bristol stool scale, body mass index (BMI) and diet, did not differ significantly between timepoints (Table 1, Wilcoxon signed-rank/Friedman p > 0.05; second/third sample-related characteristics not shown). Alpha and beta-diversity did not differ significantly between samples for the entire cohort or MS cases alone (Wilcoxon signed-rank/Friedman p > 0.09; PERMANOVA p > 0.9, Figures 1, Supplementary Figure E1–E2). The most abundant phyla were the Firmicutes and Bacteroidetes, followed by Actinobacteria, Proteobacteria, Verrucomicrobia and Lentisphaerae (Supplementary Figure E3). From the feature volatility analyses and linear mixed models, the only genus that changed (increased) significantly over time was Solobacterium (adjusted p = 0.001). Genus-level NMIT results indicated that longitudinally, the MS and monoADS participants’ gut microbial composition did not differ statistically from each other (p = 0.289, Supplementary Figure E4). Alpha diversity did not differ significantly among the four MS participants who stopped/started/switched DMT between their first/second stool samples (Wilcoxon signed-rank p > 0.14, Supplementary Figure E5).
Cohort characteristics for the paediatric-onset multiple sclerosis (MS) and monophasic acquired demyelinating syndromes (monoADS) participants, primarily at the first stool sample (with select characteristics shown between the first and second stool samples).
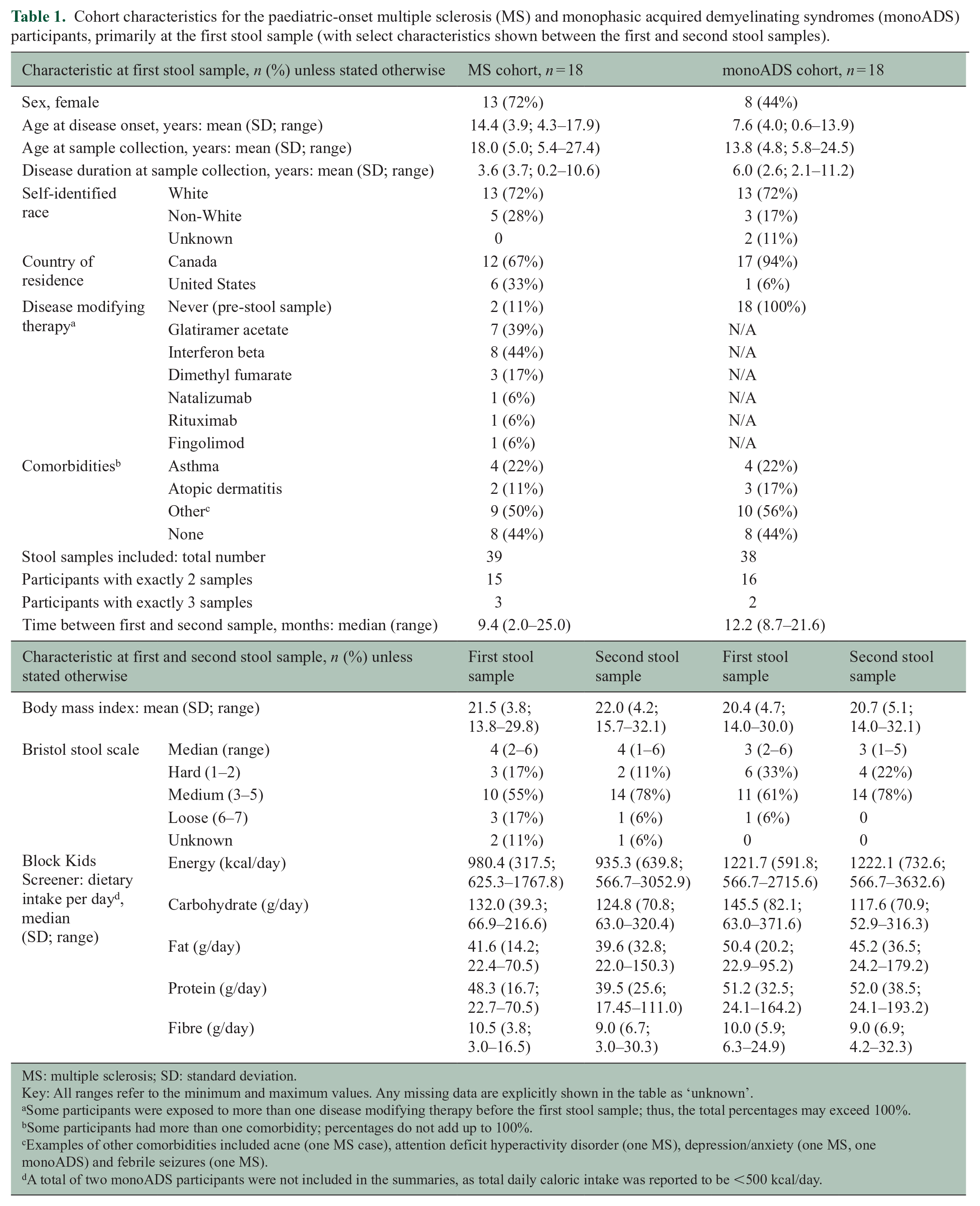
MS: multiple sclerosis; SD: standard deviation.
Key: All ranges refer to the minimum and maximum values. Any missing data are explicitly shown in the table as ‘unknown’.
Some participants were exposed to more than one disease modifying therapy before the first stool sample; thus, the total percentages may exceed 100%.
Some participants had more than one comorbidity; percentages do not add up to 100%.
Examples of other comorbidities included acne (one MS case), attention deficit hyperactivity disorder (one MS), depression/anxiety (one MS, one monoADS) and febrile seizures (one MS).
A total of two monoADS participants were not included in the summaries, as total daily caloric intake was reported to be <500 kcal/day.
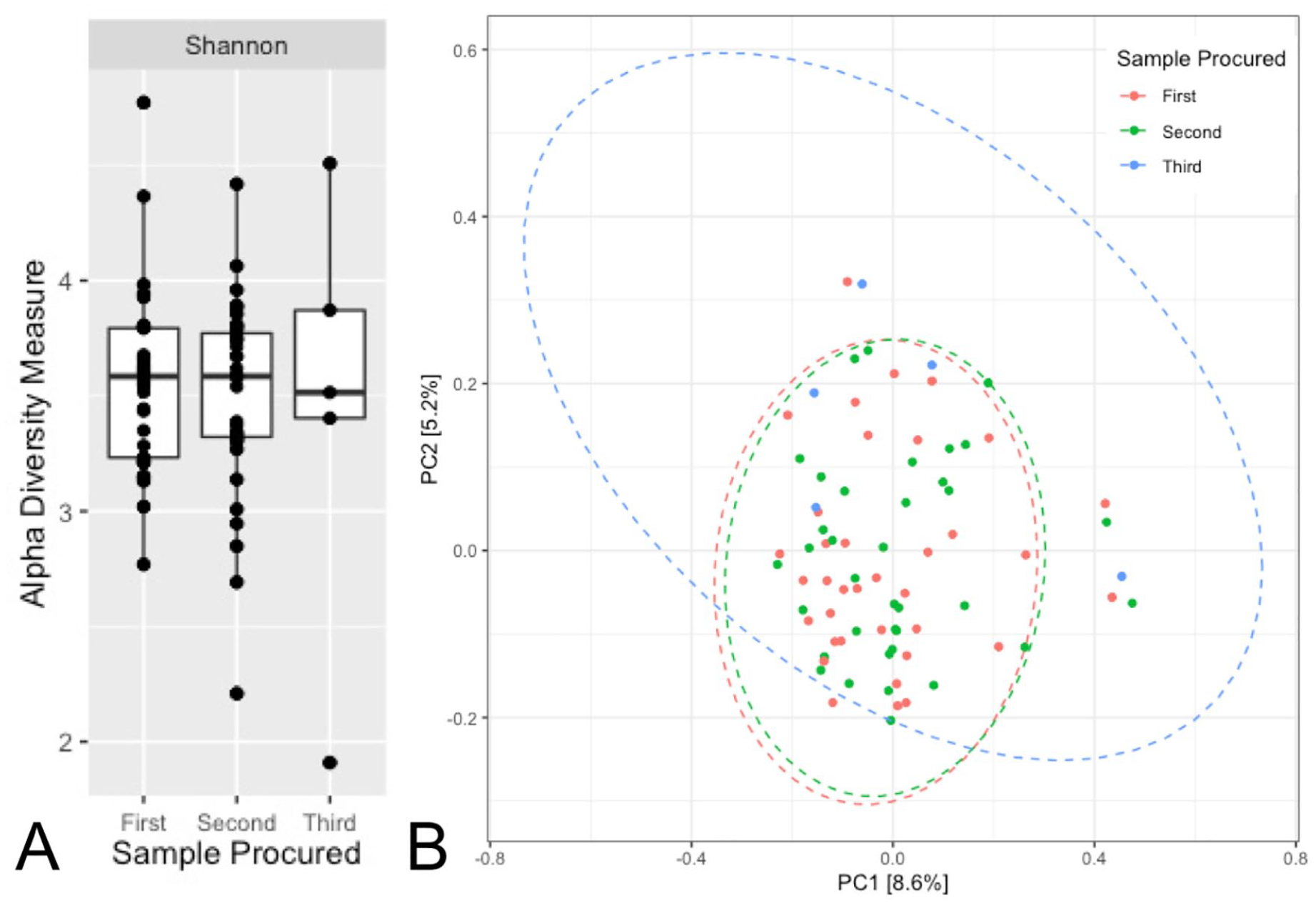
Alpha (Shannon) and beta (principal coordinates analysis of unweighted UniFrac) diversities of the gut microbiome over time for the entire cohort (multiple sclerosis and monophasic acquired demyelinating syndrome participants combined) for all stool samples procured.
Discussion
We examined the longitudinal stability of the gut microbiota in a cohort of individuals with paediatric-onset MS and monoADS across at least two, and up to three stool samples procured over 26 months. Neither alpha nor beta diversities differed significantly between stool samples procured from participants at up to three different points in time, with most samples procured 9–12 months apart. Furthermore, longitudinally, the overall genus-level composition of the gut microbiota samples (procured from the same individual) did not differ between the individuals with MS compared with those with monoADS. We can infer that, at least in the short-term, the gut microbiota composition in paediatric-onset MS and monoADS participants was relatively stable.
We also examined the five most important genera, as selected by feature analysis, and found that the only genus that changed (increased) significantly over time was Solobacterium. This genus belongs to the Clostridium cluster XVI, which has been associated with MS.1,7 Solobacterium moorei, the only species identified in the Solobacterium genus, in concert with other microbes, has been implicated as a potential driver of a proinflammatory gut microenvironment. 7
We were unable to find another published study that examined whether the gut microbiota is stable or fluctuates over time in a paediatric-onset MS or monoADS population. One study did report (primarily descriptively) stability in the gut microbiota in 102 adult participants with/without various immune-mediated diseases (19 had MS), at least across 2 samples procured over 2 months. 8
Others have also reported relative stability of the gut microbiota over time in healthy adults. The first study procured up to 4 stool samples from 9 participants over 3 months; authors reported minimal within-person variability in gut microbiota composition. 9 A second study spanned 68 months, with 2–13 stool samples procured from 37 participants; authors reported overall stability with >70% strain similarity between samples. 10
To the best of our knowledge, our study is the first to explore the stability of the gut microbiota in paediatric MS and related demyelinating diseases. Paediatric-onset MS remains relatively rare such that our sample size was modest, and we had access to a limited number of samples per participant. It would be of value for future studies to assess larger cohorts of individuals, with more samples collected at different time intervals and over longer periods of time. Trends observed in our study could potentially become significant in larger studies.
Nonetheless, our study findings are reassuring and help inform the design and analyses of future gut microbiota studies in MS and monoADS. Particularly when resources are finite and participant burden is prohibitive, our findings provide preliminary evidence to suggest that analysis of single samples from participants is a reasonable approach.
Supplemental Material
sj-docx-1-msj-10.1177_13524585221079533 – Supplemental material for Stability of the gut microbiota in persons with paediatric-onset multiple sclerosis and related demyelinating diseases
Supplemental material, sj-docx-1-msj-10.1177_13524585221079533 for Stability of the gut microbiota in persons with paediatric-onset multiple sclerosis and related demyelinating diseases by Geoffrey Liang, Feng Zhu, Ali I Mirza, Amit Bar-Or, Charles N Bernstein, Christine Bonner, Jessica D Forbes, Morag Graham, Janace Hart, Natalie C Knox, Ruth Ann Marrie, Julia O’Mahony, Gary Van Domselaar, E Ann Yeh, Yinshan Zhao, Brenda Banwell, Emmanuelle Waubant and Helen Tremlett in Multiple Sclerosis Journal
Footnotes
Acknowledgements
The authors are grateful for the involvement of the children and youth with multiple sclerosis and their parents, as well as all the investigators and their institutions involved, without whom this study would not have been possible. They also acknowledge the important contribution of the Tremlett team (University of British Columbia); Thomas Duggan in facilitating the study set-up, coordination and data collection; Bonnie Leung for additional study coordination; and Michael Sargent (Department of Internal Medicine, and the University of Manitoba IBD Clinical and Research Centre laboratory, Winnipeg, Canada) for managing the biobank. We are also grateful to the investigators and study teams at each site who participated in the Canadian Paediatric Demyelinating Disease Network study, including Douglas Arnold.
Author contributions
H.T., Y.Z., C.N.B., G.V.D., M.G., J.H. and E.W. contributed to the original funded gut microbiota grant proposal. A.B.-O., R.A.M., J.O.M., E.A.Y. and B.B. were part of the original Canadian Paediatric Demyelinating Disease Network study and facilitated collection of the cohort characteristics. J.H. facilitated training of study coordinators in the collection of stool samples. C.N.B. oversaw the biobanking. M.G., N.C.K. and G.V.D. oversaw the 16S rRNA sequencing and bioinformatics; C.B. performed the stool extractions and 16S rRNA sequencing; G.L., F.Z. and A.I.M. performed the bioinformatics, and G.L. the statistical analyses and creation of figures. All the authors contributed to the interpretation of the data. G.L. and H.T. drafted the manuscript. All the authors revised the manuscript and approved the final version to be published.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship and/or publication of this article: G.L., C.B., M.G., J.H., N.C.K. and J.O.M. have no conflict of interest to report. F.Z. and Y.Z. were funded through research grants held by H.T., including The Multiple Sclerosis Scientific and Research Foundation (PI: Tremlett, EGID: 2636). A.I.M. is funded through the MS Society of Canada endMS Doctoral Studentship (EGID: 3246) and was funded through The Multiple Sclerosis Scientific and Research Foundation (PI: Tremlett, EGID: 2636). A.B.-O. is funded by the NIH, ITN, NMSS and MSSOC. A.B.-O. has participated as a speaker in meetings sponsored by and received consulting fees and/or grant support from Janssen/Actelion; Atara Biotherapeutics, Biogen Idec, Celgene/Receptos, Roche/Genentech, Medimmune, Merck/EMD Serono, Novartis and Sanofi-Genzyme. C.N.B. has served on advisory boards for Abbvie Canada, Amgen Canada, Bristol Myers Squibb Canada, Roche Canada, Janssen Canada, Takeda Canada, Pfizer Canada Sandoz Canada, consulted to Mylan Pharmaceuticals and Takeda, has received educational grants from Abbvie Canada, Pfizer Canada, Takeda Canada, Janssen Canada and has been on the speaker’s panel for Janssen Canada, Takeda Canada, Pfizer Canada and Abbvie Canada. J.D.F. was part-funded through research grants held by H.T., including The Multiple Sclerosis Scientific and Research Foundation (PI: Tremlett, EGID: 2636). G.V.D. is the Chief Bioinformatics Scientist with the National Microbiology Laboratory – Public Health Agency of Canada and has received research support in the past 3 years from the National MS Society, the Canadian Institute of Health Research and Genome Canada. R.A.M. received research funding from CIHR, Research Manitoba, Multiple Sclerosis Society of Canada, Multiple Sclerosis Scientific Foundation, Crohn’s and Colitis Canada, National Multiple Sclerosis Society, CMSC and the US Department of Defense. She is supported by the Waugh Family Chair in Multiple Sclerosis. She is a co-investigator on a study funded by Biogen Idec and Roche. E.A.Y. has received research support in the past 3 years from the National MS Society, Canadian Institutes of Health Research, National Institutes of Health, Ontario Institute of Regenerative Medicine, Stem Cell Network, SickKids Foundation, Peterson Foundation, MS Society of Canada and the MS Scientific Research Foundation. She has received funding for investigator-initiated research from Biogen and has served on scientific advisory boards for Biogen, Alexion and Hoffman-LaRoche. B.B. serves as a consultant to Novartis, UCB and Roche. She provides non-remunerated advice on clinical trial design to Novartis, Biogen and Teva Neuroscience and is funded by the NMSS, NIH and the Canadian MS Society. E.W. is funded by the NMSS, the NIH, PCORI and the Race to Erase MS. E.W. has received consulting honoraria from Jazz Pharma, Emerald and DBV. She volunteers on a clinical trial committee for Novartis. H.T. is the Canada Research Chair for Neuroepidemiology and Multiple Sclerosis. Current research support was received from the National Multiple Sclerosis Society, the Canadian Institutes of Health Research, the Multiple Sclerosis Society of Canada and the Multiple Sclerosis Scientific Research Foundation. In addition, in the past 5 years, she has received research support from the UK MS Trust; travel expenses to present at CME conferences from the Consortium of MS Centres (2018), the National MS Society (2016, 2018), ECTRIMS/ACTRIMS (2015, 2016, 2017, 2018, 2019, 2020) and American Academy of Neurology (2015, 2016, 2019). Speaker honoraria are either declined or donated to an MS charity or to an unrestricted grant for use by H.T.’s research group.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This study was supported by the Multiple Sclerosis Scientific and Research Foundation (#EGID: 2636; PI: Tremlett), a grant from the Foundation of the Consortium of Multiple Sclerosis Centers’ MS Workforce of the Future program, and an award from the University of British Columbia Faculty of Medicine Summer Student Research Program. The funding source was not involved in the study design, the collection, analysis and interpretation of the data, or in the decision to submit this article for publication.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
