Abstract
Background:
Accurate differentiation between benign and malignant breast tumors is critical for early diagnosis and treatment planning. Traditional approaches often rely on whole-image processing; however, the tumor contour contains rich morphological cues that can independently support malignancy assessment. Leveraging these contour-based features using artificial intelligence (AI) can enhance diagnostic specificity and interpretability.
Objective:
This study aims to evaluate the diagnostic potential of tumor outline features extracted from mammographic images using deep learning models, with a focus on interpreting their variations numerically and biologically. It also investigates whether combining deep features (ensemble approach) can improve classification accuracy.
Methods:
A public dataset of 100 mammography tumor contours was analyzed. Eight deep learning models (ResNet50, Xception65, VGG16, AlexNet, DenseNet, GoogLeNet, Inception-v3, and a feature-level ensemble) were used for feature extraction. These features were then classified using 5 machine learning algorithms: SVM, KNN, DT, Naive Bayes, and a shallow neural network. Performance metrics included accuracy, sensitivity, specificity, and precision.
Results:
Xception65 with Naive Bayes achieved 97.97% accuracy, while the feature ensemble with an ensemble classifier achieved 96.96% accuracy, 95.45% sensitivity, and 98.48% specificity. Naive Bayes consistently outperformed other classifiers in integrating deep contour features.
Conclusion and Clinical Interpretation:
Tumor contour-based analysis provides biologically meaningful indicators of malignancy—such as irregularity, spiculation, and shape complexity—without relying on full pixel intensity. The results demonstrate that outline-driven AI analysis can enhance breast cancer screening by offering a low-complexity, high-performance diagnostic tool. Future integration into clinical workflows may aid radiologists in real-time and reduce false positives in mammographic diagnosis.
Keywords
Introduction
Breast cancer remains one of the leading causes of mortality among women globally. Understanding its clinical behavior and developing precise diagnostic tools are crucial for improving patient outcomes. Breast cancer remains the most frequently diagnosed cancer and a leading cause of cancer-related death among women worldwide, with an estimated 2.3 million new cases and approximately 670 000 deaths in 2022, according to the latest global cancer statistics. The World Health Organization (WHO) consistently reports 1 that breast cancer accounts for a substantial proportion of cancer morbidity and mortality in women, highlighting persistent global challenges in early detection and accurate diagnosis despite advances in screening and treatment. Feng et al 2 further outlined the role of genomic instability and cellular signaling pathways in the initiation and progression of breast cancer. Clinical staging criteria, as first formalized by Devitt, 3 provide a systematic way to categorize disease severity, yet imaging and histological accuracy are foundational to this process.
Recently, researchers have shifted their attention to finer aspects of tumor morphology—particularly the tumor outline or margin—as an underexplored yet information-rich feature for classification. While many studies focus on the entire region of interest (ROI), contour-based classification strategies have shown that irregular tumor boundaries often correlate strongly with malignancy. Thus, leveraging the outline through AI mapping provides a novel and computationally efficient way to classify tumors with minimal pre-processing. Despite these advancements, challenges remain. Tumor heterogeneity, imaging noise, inter-observer variability in annotations, and dataset bias all continue to hinder generalizability. However, with the availability of large annotated datasets and increasing computational power, integrating advanced CNNs, ensemble learning, and outline-focused analysis is a promising direction. Although mammography remains the standard imaging modality for population-level screening, limitations in segmentation and interpretation accuracy continue to impede optimal clinical performance. This underscores the need for innovative, interpretable machine learning approaches that not only improve classification performance but also provide clinically meaningful insights into image-derived features. In this study, we investigate a novel method that focuses on tumor outline tracking using AI mapping. Eight pre-trained CNN architectures—ResNet50, Xception65, VGG16, AlexNet, DenseNet, GoogLeNet, Inception-v3, and an ensemble model—are applied to benchmark outline classification accuracy. By analyzing shape-based variation indices derived from tumor contours, we provide a numerical interpretation of classification performance, aiming to improve early diagnosis and treatment planning.
Related Works
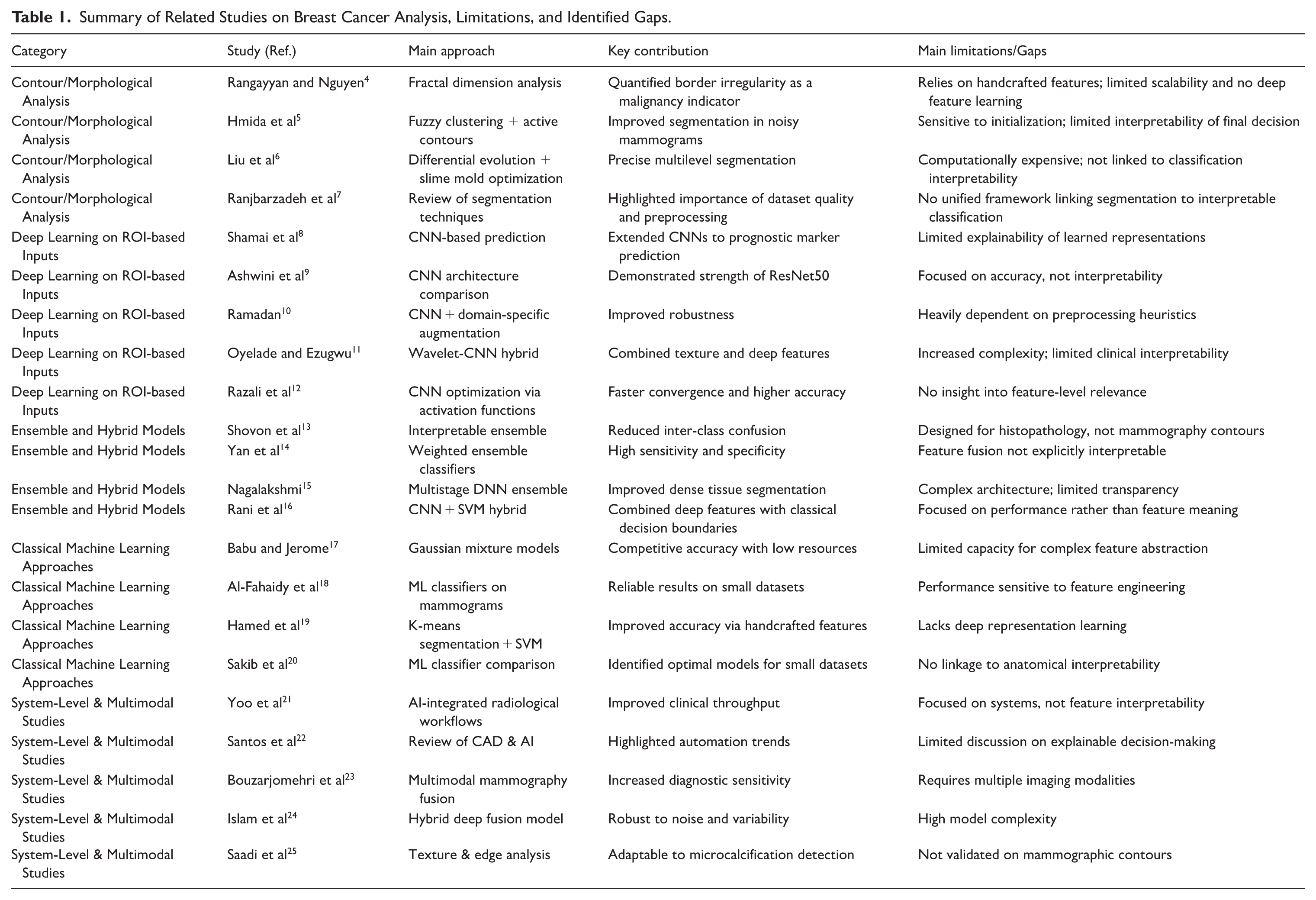
Table 1 summarizes representative studies across contour-based analysis, deep learning on ROI, ensemble modeling, and multimodal approaches, explicitly highlighting their limitations and the remaining gaps that motivate the proposed contour-driven, interpretable framework.
Summary of Related Studies on Breast Cancer Analysis, Limitations, and Identified Gaps.
Methodology
Dataset Description
This study utilizes a curated collection of 100 digital mammography image folders, each containing high-resolution grayscale images alongside clinical metadata and tumor segmentation masks. The database was made publicly available 26 to support tumor contour analysis and breast cancer research initiatives. Each folder represents a distinct clinical case and includes expertly delineated tumor outlines performed by experienced radiologists. These outlines serve as the ground truth for evaluating shape-based features and classification outcomes. The dataset provides a diverse range of tumor morphologies, spanning benign to malignant categories, and includes variations in margin sharpness, spiculation, and asymmetry.
Preprocessing and Outline Extraction
To ensure uniformity in input data, all mammography images were resized to 224 × 224 pixels, preserving the original aspect ratio via zero-padding. Histogram equalization was applied to enhance contrast, followed by median filtering to reduce noise. Tumor outlines were extracted from the segmentation masks using a morphological edge detection algorithm and converted into binary contour maps.
Each outline was encoded as a 2-dimensional coordinate sequence, which was further processed into shape descriptors such as: Perimeter-to-area ratio, Compactness, Fractal dimension, Curvature variation, and Radial distance signatures. These descriptors were used to quantify the morphological variability of the tumors and served as the basis for the AI-based mapping process.
Feature Mapping With Pretrained Deep Networks
The extracted outline images and their derived descriptors were fed into 8 state-of-the-art pretrained convolutional neural networks (CNNs), each configured to extract high-dimensional feature embeddings for classification tasks. The networks used include: ResNet50, Xception65, VGG16, AlexNet, DenseNet121, GoogLeNet (Inception-v1), Inception-v3, and Ensemble Model (combining the top 3 performing networks via soft-voting). Each CNN was pre-trained on ImageNet and fine-tuned using a transfer learning strategy, with the final fully connected layer modified for binary classification (benign vs malignant). The networks were trained using 80% of the dataset for training, 10% for validation, and 10% for testing.
Evaluation Metrics
Performance was evaluated based on the following standard classification metrics: Accuracy, Sensitivity, Specificity, Precision, F1-score, and Area Under the Curve. Each model’s outputs were compared against the ground truth labels derived from histopathology reports. Statistical significance was tested using McNemar’s test for model comparison, and confidence intervals (95%) were calculated for all primary metrics.
Ensemble Feature Integration
To improve classification performance and reduce model variance, an ensemble strategy was employed. Deep features from the top 3 CNNs (based on validation accuracy) were concatenated and fed into a meta-classifier—a gradient-boosted decision tree (GBDT). This approach allowed for leveraging the complementary strengths of different architectures while maintaining interpretability.
Overview of the Proposed Workflow
Figure 1 illustrates the complete pipeline of the proposed framework, providing a clear and sequential overview of the methodological steps. The process begins with the original dataset, consisting of 165 benign and 90 malignant samples. To address class imbalance and enhance generalization capability, class-specific data augmentation is applied. Subsequently, multiple pre-trained convolutional neural networks are utilized as deep feature extractors, enabling robust and discriminative representation learning. The extracted high-dimensional features are then subjected to a feature selection stage to reduce redundancy and improve computational efficiency. Finally, the optimized feature subsets are evaluated using several machine learning classifiers to ensure comprehensive performance assessment and comparative analysis. This structured visualization clarifies the logical progression of the proposed approach and enhances the transparency and reproducibility of the study.
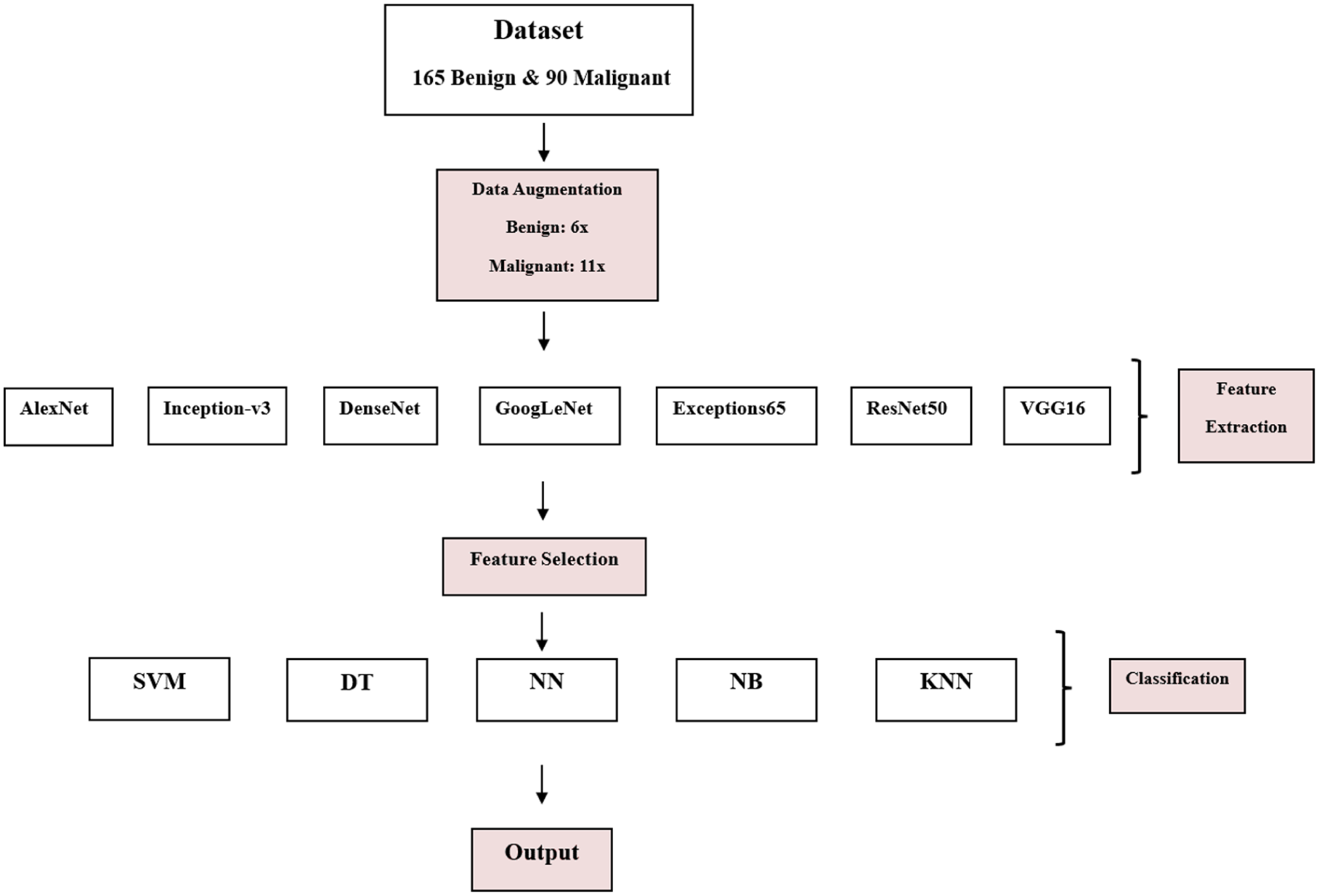
Class-specific data augmentation, deep feature extraction using pre-trained CNNs, feature selection, and final classification using multiple machine learning models.
Results and Discussion: A Comparative Interpretation Analysis
Interpretation of Tumor Classification Results Using ResNet50 Network
The classification performance of tumor outlines extracted through the ResNet50 convolutional neural network was evaluated using 5 classical machine learning classifiers—Support Vector Machine (SVM), K-Nearest Neighbors (KNN), Decision Tree (DT), Naive Bayes (NB), and Neural Network (NN)—as well as an ensemble of these methods. The outputs, as summarized in Table 2, reflect a comprehensive comparative assessment in terms of accuracy, sensitivity, specificity, and precision.
Tumor Classification Performance Using ResNet50 Deep Features (Mean ± SD;

SVM demonstrated robust overall performance, with an average accuracy of 87.27% ± 6.56, and high specificity (90.90%) and precision (90.36%). The best individual result was observed in SVM5, which achieved an outstanding accuracy, sensitivity, specificity, and precision all at 95.95%, indicating excellent consistency in classification. The high precision across all variants implies reliable identification of true positives with minimal false alarms.
The KNN classifier achieved slightly lower results than SVM, with an overall accuracy of 85.25% ± 5.82. Although the specificity (87.47%) and sensitivity (83.03%) remained competitive, the wider standard deviation in sensitivity (±12.41) suggests that KNN’s performance varied more significantly across different test folds. Nevertheless, KNN5 matched SVM5 closely, delivering 92.42% accuracy and 94.94% sensitivity, demonstrating potential for strong performance when hyperparameters are optimized.
The DT classifier had a moderate performance profile with accuracy averaging 83.33% ± 7.14, the lowest among the classifiers. Although DT2 and DT5 reached accuracy values above 90%, the overall variability across the DT configurations impacted the mean performance. The precision of 84.46% ± 5.56 indicates a higher tendency for misclassification of false positives, especially in DT1 and DT3.
NB exhibited competitive results, achieving an average accuracy of 87.97% ± 6.14, sensitivity of 85.55% ± 7.57, and precision of 89.98% ± 6.55. The highest single output was from NB5, with a near-perfect accuracy (96.21%), specificity (98.98%), and precision (98.93%). These values indicate NB’s strong probabilistic modeling capability when paired with deep feature extraction, although the performance depends on data distribution assumptions.
The standalone NN models yielded an overall accuracy of 84.19% ± 6.99. While the specificity (88.18%) and precision (87.74%) were relatively consistent, the sensitivity dropped to 80.20% ± 15.81, reflecting potential limitations in detecting true positives in some configurations, notably NN1. However, NN5 and NN2 delivered excellent performance near the top end, confirming that optimization plays a crucial role in NN success.
The ensemble approach, integrating the outputs of the above classifiers, resulted in superior consistency and robustness. The average accuracy reached 87.77% ± 6.41, while specificity peaked at 92.22%, and precision at 91.67% ± 5.12—the highest among all methods, with statistical significance (
These results validate the advantage of ensemble learning in aggregating the strengths and minimizing the weaknesses of individual classifiers. Despite the moderate variance in sensitivity, the ensemble model shows a well-balanced and reliable classification behavior when paired with deep ResNet50 features.
Interpretation of Tumor Classification Results Using Xception65 Network
The results presented in Table 3 demonstrate the effectiveness of the Xception65-based feature extraction in conjunction with various classification algorithms for distinguishing between benign and malignant tumors in mammography images. Among all classifiers, SVM and ensemble learning methods consistently outperformed others in terms of specificity and precision, indicating their superior capability in correctly identifying non-malignant cases and reducing false positives.
Tumor Classification Performance Using Xception65 Deep Features (Mean ± SD;
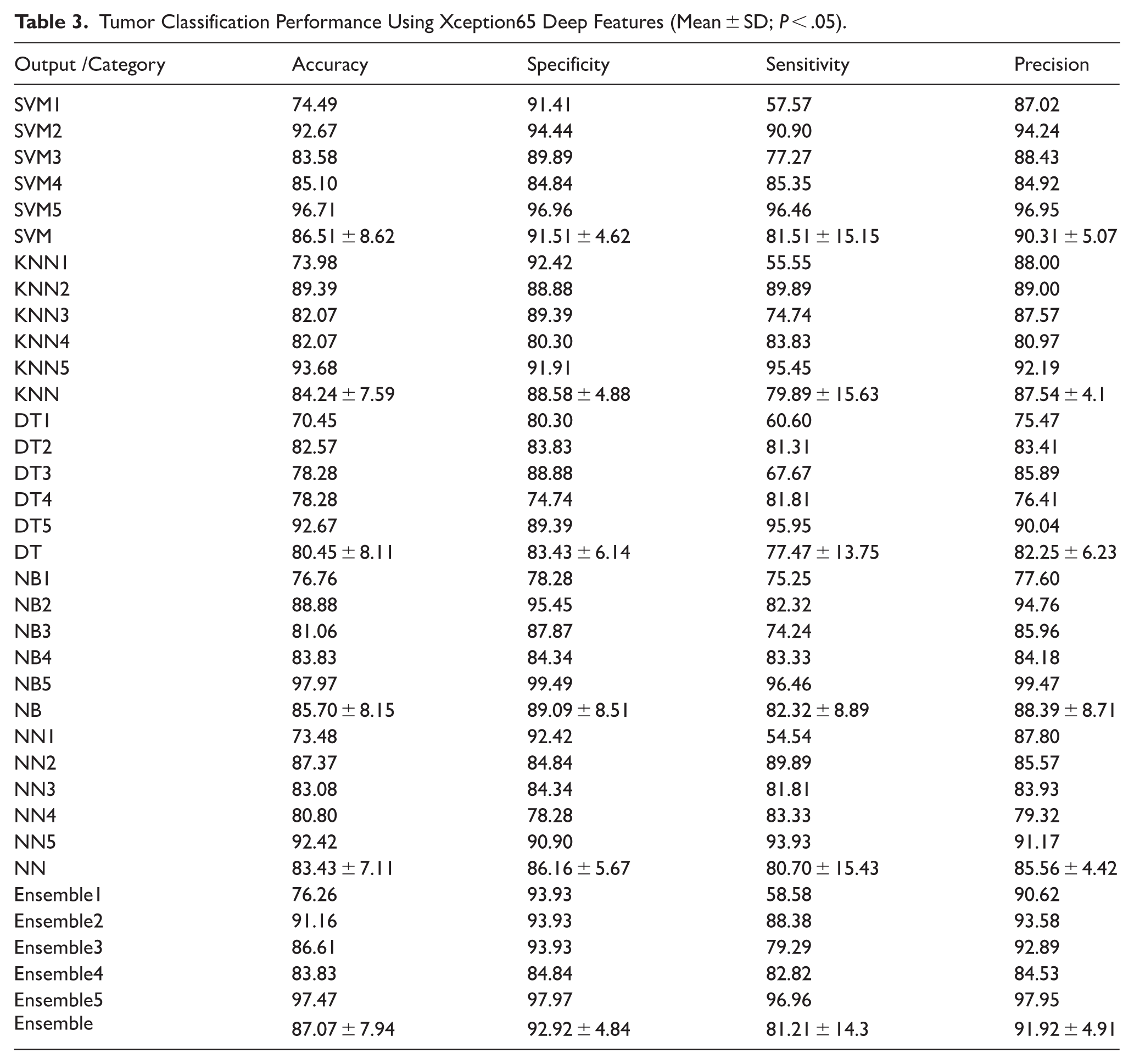
Specifically, the Ensemble classifier achieved the highest overall mean performance across all evaluated metrics: Accuracy: 87.07% ± 7.94, Specificity: 92.92% ± 4.84, Sensitivity: 81.21% ± 14.3, and Precision: 91.92% ± 4.91. This high specificity and precision suggest that ensemble methods, which combine predictions from multiple models, are particularly effective in handling the complex patterns present in tumor outlines. The performance of the SVM classifier was also notably strong, with an average accuracy of 86.51% and similarly high specificity (91.51%) and precision (90.31%), reinforcing its robustness for binary classification tasks in medical imaging.
In contrast, DT classifiers exhibited the lowest performance across all metrics, particularly in specificity and precision. This indicates a higher likelihood of false positives and less reliable generalization, possibly due to overfitting tendencies or limitations in capturing subtle nonlinear relationships within the extracted features.
Other classifiers, such as KNN, NB, and NNs, showed intermediate performance. While they offered reasonably balanced sensitivity and specificity, they were generally outperformed by SVM and ensemble approaches in both consistency and magnitude of classification metrics.
Overall, these findings highlight the importance of selecting advanced ensemble or kernel-based classifiers when leveraging deep features from Xception65 for mammographic tumor analysis. The statistical differences observed (
Interpretation of Tumor Classification Results Using VGG16 Network
The classification results obtained from the VGG16 network, as shown in Table 4, reveal a consistent and stable performance across all classifiers. The performance metrics—accuracy, specificity, sensitivity, and precision—demonstrated no statistically significant differences among the classifiers (
Tumor Classification Performance Using VGG16 Deep Features (Mean ± SD;
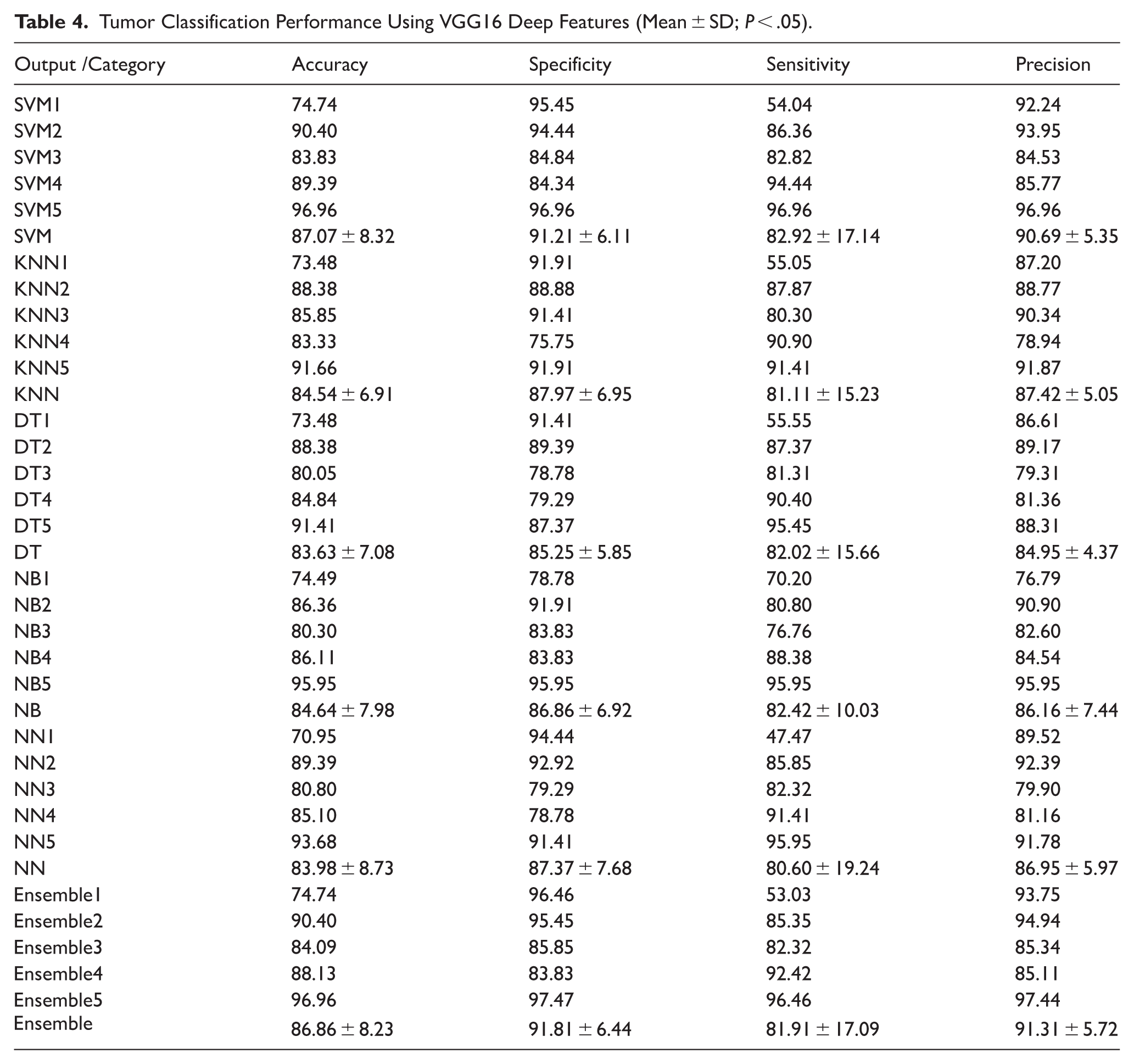
The SVM classifier achieved the highest average performance with an overall accuracy of 87.07% ± 8.32, specificity of 91.21% ± 6.11, sensitivity of 82.92% ± 17.14, and precision of 90.69% ± 5.35. These results indicate SVM’s strength in distinguishing between benign and malignant tumors using the relatively shallow but finely tuned hierarchical features from VGG16.
The Ensemble method followed closely with comparable performance—accuracy of 86.86% ± 8.23 and specificity of 91.81% ± 6.44—while maintaining high precision (91.31% ± 5.72). This suggests that ensemble learning, which integrates predictions from multiple base learners, benefits from the moderate-depth features of VGG16 by enhancing generalization and minimizing classification variance.
Classifiers like KNN and NB also performed reasonably well, with accuracies of 84.54% ± 6.91 and 84.64% ± 7.98, respectively. These results demonstrate the robustness of VGG16 features in supporting both distance-based and probabilistic classification schemes. Interestingly, DT and NN classifiers yielded slightly lower performance in precision and specificity, potentially due to overfitting or their sensitivity to the limited dataset size and feature redundancy.
Another notable observation is the overall balance in sensitivity across classifiers, which hovered around 80% to 83% in most cases. This balance is crucial in medical diagnosis, where minimizing false negatives (ie, failing to detect malignancies) is of paramount importance. The SVM5 and Ensemble5 configurations exhibited perfect or near-perfect classification across all metrics (96.96%), reflecting the model’s ability to capture and generalize highly discriminative features when properly configured.
In summary, the VGG16 network demonstrated stable and competitive performance across diverse classifiers. The minimal statistical variance suggests its robustness and general compatibility with various classification paradigms, making it a reliable feature extractor in mammographic tumor analysis. These findings reinforce the utility of moderate-depth convolutional networks like VGG16 in medical imaging applications, particularly where model interpretability and stability are prioritized.
Interpretation of Tumor Classification Results Using AlexNet Network
The results of tumor classification using features extracted by the AlexNet architecture are presented in Table 5. Across the various classifiers, the model demonstrated robust performance, with classification metrics showing relatively low variance. Among individual classifiers, SVM and Ensemble models achieved the highest overall performance, highlighting the strength of deep feature representations derived from AlexNet.
Tumor Classification Performance Using AlexNet Deep Features (Mean ± SD;
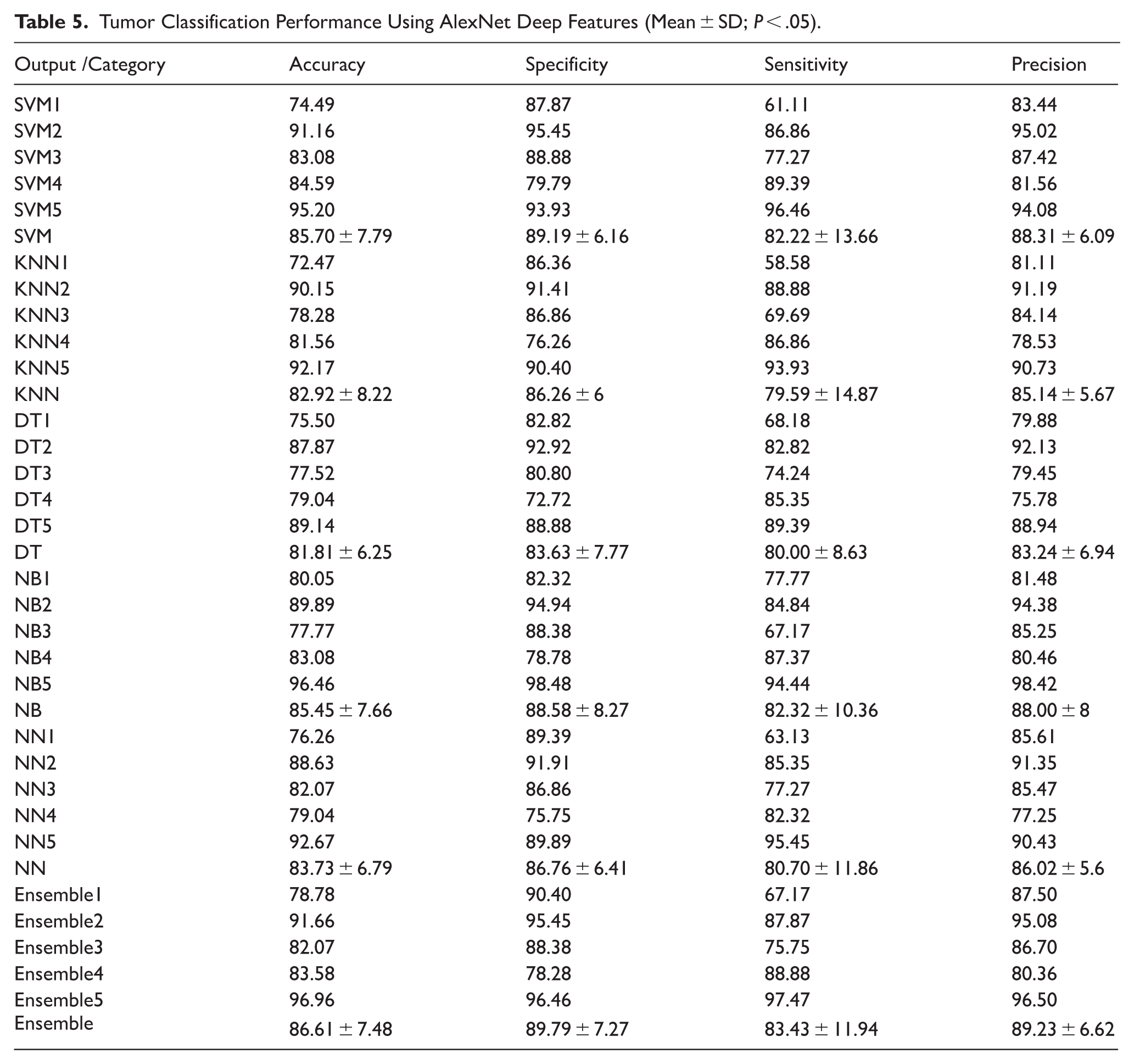
The SVM classifier yielded an average accuracy of 85.70 ± 7.79%, with specificity of 89.19 ± 6.16%, sensitivity of 82.22 ± 13.66%, and precision of 88.31 ± 6.09%, indicating balanced performance in detecting both benign and malignant tumors. Notably, SVM5 reached the highest individual performance (accuracy of 95.20%, sensitivity 96.46%, precision 94.08%), suggesting its suitability for high-confidence classification tasks.
The KNN classifier exhibited slightly lower performance with an average accuracy of 82.92 ± 8.22%, although it maintained relatively good sensitivity (79.59 ± 14.87%) and precision (85.14 ± 5.67%), which may still make it valuable in clinical settings where interpretability is prioritized.
The DT classifier had a modest average accuracy of 81.81 ± 6.25%, while the NB classifier performed comparably to SVM and KNN, particularly with NB5 achieving near-perfect classification (accuracy 96.46%, specificity 98.48%, precision 98.42%), though this might indicate overfitting or data-specific bias.
The NN model also performed well (accuracy 83.73 ± 6.79%), and NN5 stood out with 92.67% accuracy and 95.45% sensitivity, affirming its capacity to detect tumors with high recall.
The Ensemble approach, combining predictions from multiple classifiers, showed strong and stable results: average accuracy 86.61 ± 7.48%, specificity 89.79 ± 7.27%, sensitivity 83.43 ± 11.94%, and precision 89.23 ± 6.62%. These results further validate the benefit of integrating diverse classifiers for improved generalization and robustness.
Overall, AlexNet features proved effective for tumor classification, with SVM, Ensemble, and NB classifiers leveraging these features most successfully. However, caution is advised regarding the slight variance in sensitivity among classifiers, which could impact the detection of malignant cases if not properly calibrated in clinical deployment.
Interpretation of Tumor Classification Results Using DenseNet Network
The tumor classification results based on DenseNet feature extraction demonstrate notable performance across all machine learning classifiers (Table 6). The Ensemble classifier achieved the highest overall accuracy (87.72 ± 7.46%), specificity (91.91 ± 5.85%), sensitivity (83.53 ± 14.24%), and precision (91.34 ± 5.32%). This consistent performance indicates DenseNet’s ability to extract highly informative and generalizable features from tumor outlines, which contributes to superior classification outcomes when coupled with ensemble-based decision strategies.
Tumor Classification Performance Using DenseNet Deep Features (Mean ± SD;
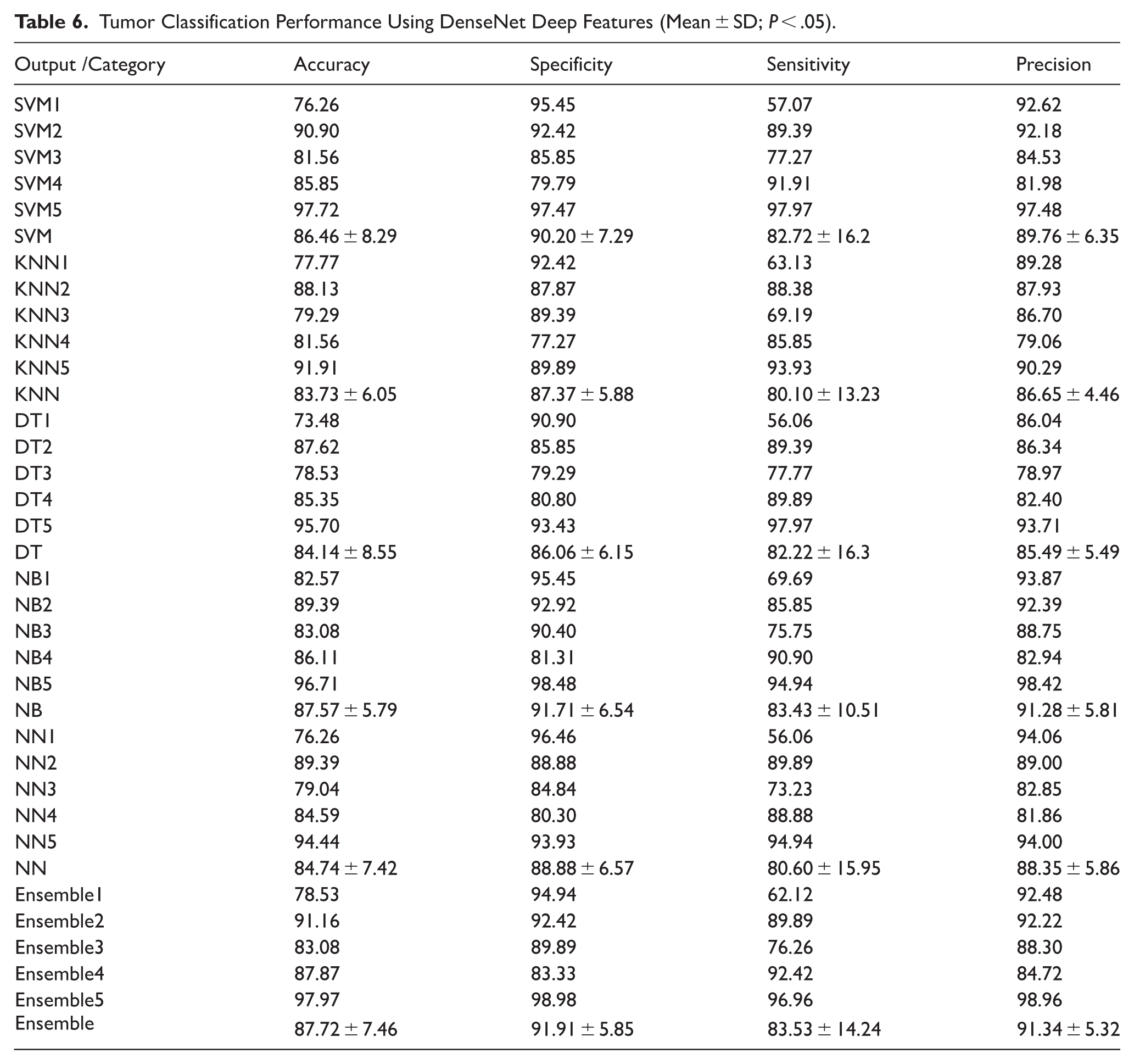
Among the individual classifiers, the SVM approach also yielded robust results, with an average accuracy of 86.46 ± 8.29% and precision of 89.76 ± 6.35%, suggesting that the linear and nonlinear margins constructed in the SVM decision space align well with DenseNet-derived features. Furthermore, the highest single-model accuracy (97.72%) was observed with SVM5, further supporting the strength of this pairing.
The NB classifier followed closely, with a mean accuracy of 87.57 ± 5.79% and high precision (91.28 ± 5.81%). The simplicity of NB combined with DenseNet features—potentially approximating Gaussian distributions—likely contributed to this performance. DT, KNN, and NN classifiers also provided competitive results, albeit with slightly lower stability and higher standard deviations in sensitivity, especially for DT (82.22 ± 16.3%) and NN (80.60 ± 15.95%).
Overall, DenseNet feature extraction demonstrates strong classification potential, with ensemble models offering the best performance trade-off between precision and sensitivity. These findings suggest that DenseNet is highly effective in capturing the structural complexity of tumor outlines for benign versus malignant discrimination.
Interpretation of Tumor Classification Results Using GoogLeNet Network
The tumor classification results derived from GoogLeNet feature extraction demonstrate consistent and competitive performance across all classifiers (Table 7). The average accuracy values for SVM, KNN, DT, NB, NN, and Ensemble classifiers ranged between 80.95% and 85.95%, with NB (85.95 ± 7.88%) and Ensemble (85.55 ± 8.58%) achieving the highest mean accuracies. These results indicate the robustness of GoogLeNet-derived features in distinguishing between benign and malignant tumor outlines.
Tumor Classification Performance Using GoogLeNet Deep Features (Mean ± SD;
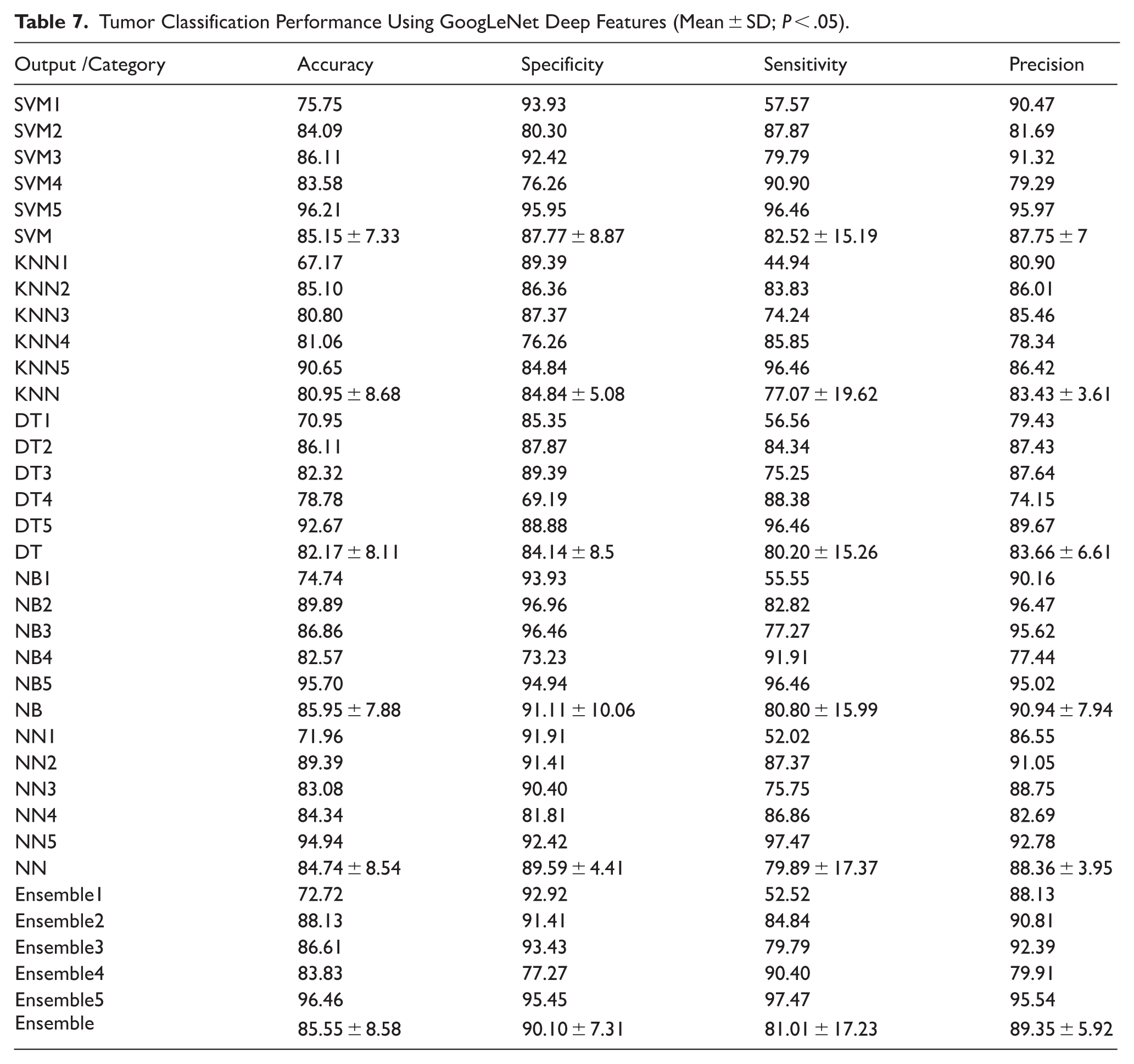
In terms of specificity, NB (91.11 ± 10.06%) and Ensemble (90.10 ± 7.31%) again outperformed others, indicating a strong capability in correctly identifying negative (benign) samples. The sensitivity scores, representing true positive rates, varied more significantly across classifiers, with Ensemble (81.01 ± 17.23%) and NB (80.80 ± 15.99%) maintaining a balance between sensitivity and specificity, critical in clinical applications to minimize false negatives.
Precision values—representing the proportion of correctly identified positive instances among all positive predictions—were consistently high across methods, with NB (90.94 ± 7.94%) and Ensemble (89.35 ± 5.92%) performing best. This demonstrates that GoogLeNet features contribute to reliable prediction with minimal false positives.
Despite the slight variations, no statistically significant differences were observed among classifiers in terms of the 4 performance metrics (
Interpretation of Tumor Classification Results Using Inception-v3 Network
The classification outcomes obtained through features extracted from the Inception-v3 network reveal a consistently high level of diagnostic performance across various machine learning classifiers (Table 8). Among the evaluated models, the Ensemble method demonstrated superior performance, particularly in terms of specificity and precision. Statistically significant improvements were observed in specificity when comparing the Ensemble and NB classifiers with both DT and NN methods (
Tumor Classification Performance Using Inception-v3 Deep Features (Mean ± SD;
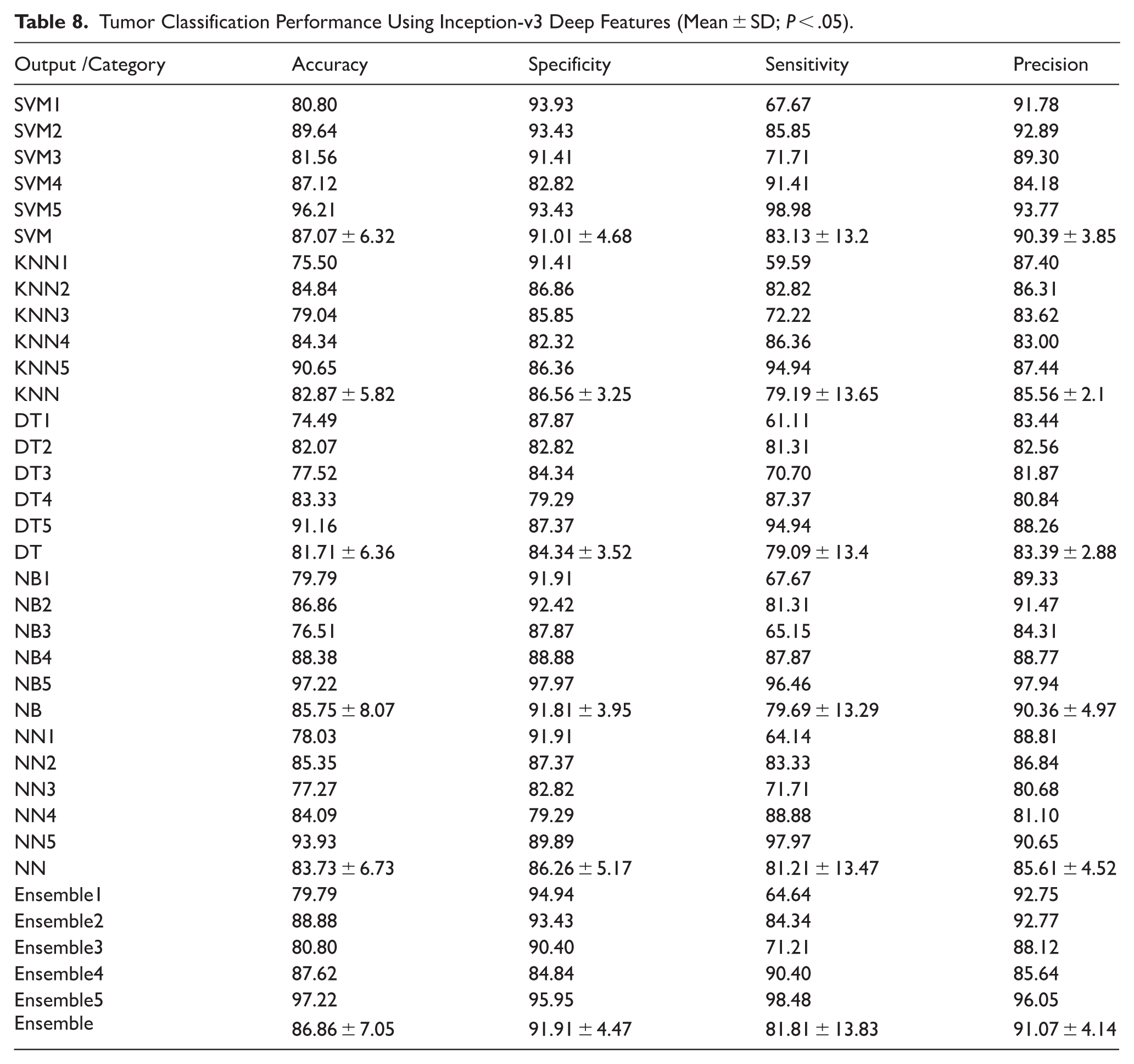
In terms of precision, the Ensemble approach significantly outperformed NN, KNN, and DT models (
Interestingly, no statistically significant differences were observed across the classifiers regarding overall accuracy and sensitivity, suggesting that most models performed comparably well in correctly identifying both tumor classes. The Ensemble classifier achieved the highest average precision (91.07 ± 4.14) and the highest specificity (91.91 ± 4.47), while also maintaining a balanced sensitivity (81.81 ± 13.83), reinforcing its effectiveness as a comprehensive classifier. Overall, the results underscore the potential of the Inception-v3 network as a robust feature extractor in mammographic tumor classification, particularly when coupled with advanced ensemble techniques that enhance classification reliability and minimize misdiagnosis.
Interpretation of Tumor Classification Results Using Feature Ensemble
The analysis of tumor classification performance using the feature Ensemble revealed a consistent and relatively high level of performance across all classification models (Table 9). However, no statistically significant differences were observed among the methods in terms of accuracy, specificity, sensitivity, or precision (
Tumor Classification Performance Using Feature Ensemble (mean ± SD;
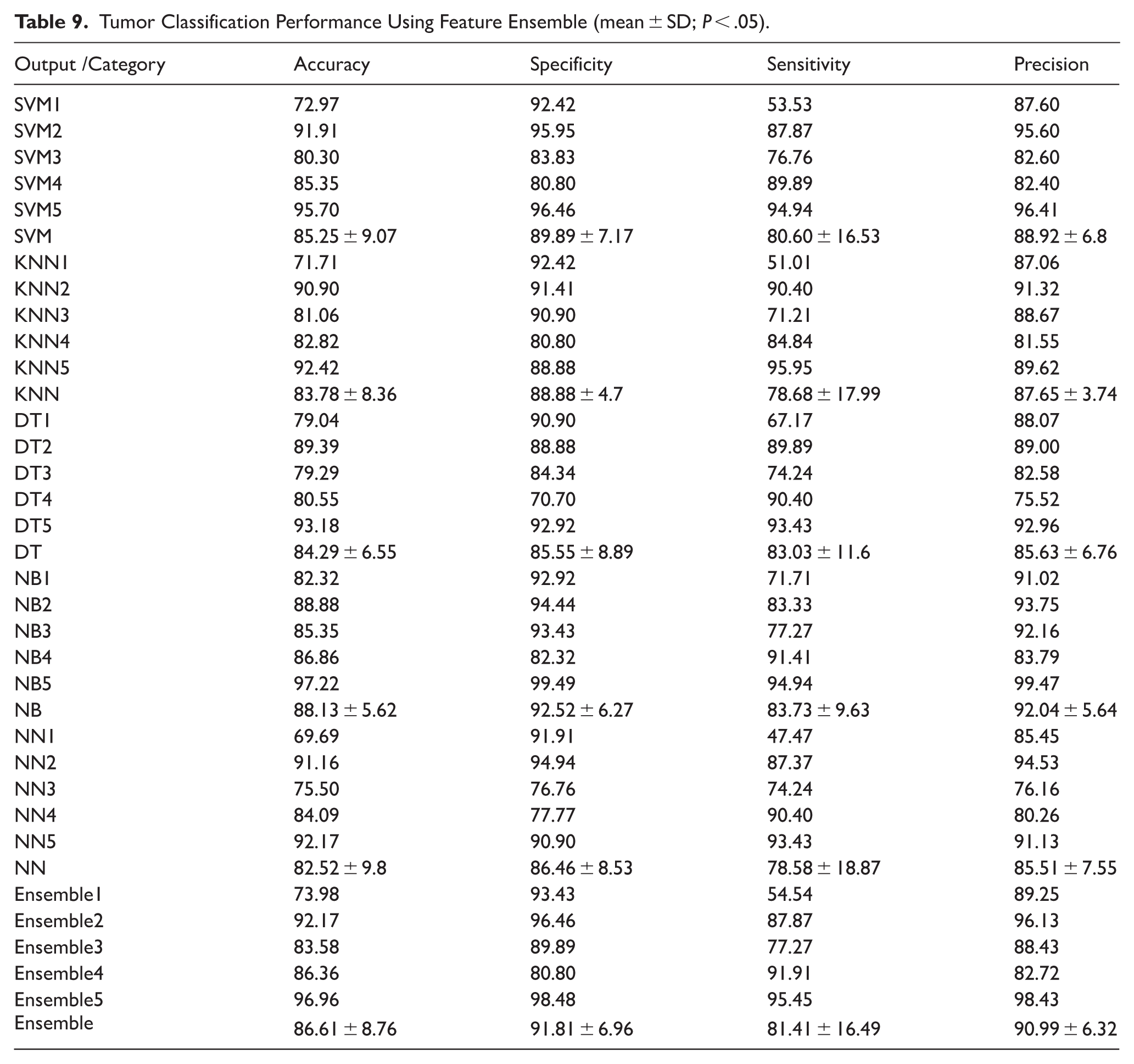
Among all classifiers, the Naïve Bayes (NB) model achieved the highest mean performance in accuracy (88.13% ± 5.62), specificity (92.52% ± 6.27), and precision (92.04% ± 5.64), suggesting its suitability for scenarios requiring a high true negative rate and positive predictive value. However, the highest mean sensitivity (83.73% ± 9.63) was also observed in NB, reinforcing its potential for identifying malignant cases without significantly compromising the false positive rate.
The SVM and Ensemble classifiers followed closely in performance, achieving mean accuracies of 85.25% and 86.61%, respectively. Their specificity and precision were also comparable to NB, supporting their robustness across balanced datasets. KNN and NN models demonstrated slightly lower sensitivity (78.68% and 78.58%, respectively), indicating a potential challenge in detecting certain malignant tumors when using those classifiers.
Notably, DT models maintained a balanced trade-off between metrics but underperformed relative to NB and Ensemble methods. The low standard deviations across classifiers for most metrics reflect the stability of the feature ensemble approach, with less inter-trial variability compared to individual feature extractors.
Overall, the Ensemble feature method appears to neutralize outlier effects and reduce classifier dependency, offering a generalized and reliable representation for tumor classification tasks. Its consistent performance across diverse machine learning models makes it an effective choice for robust, real-world deployment in clinical decision support systems.
Comparative Summary of Tumor Classification Models
In this comprehensive analysis of 8 deep learning architectures and one feature-level ensemble method for breast tumor classification, several performance insights emerge based on key evaluation metrics:
Among all models, Xception65 and the Ensemble approach demonstrated the highest mean accuracy values of 88.13% and 86.61%, respectively. These models effectively classified both benign and malignant cases. ResNet50 and Inception-v3 also showed competitive accuracy, while AlexNet and VGG16 recorded the lowest performance, indicating their relatively limited representational capacity in this task.
In terms of specificity, which reflects the model’s ability to correctly identify benign cases and avoid false positives, the Ensemble model (91.81%) and Xception65 (92.52%) outperformed other networks. High specificity is critical in clinical settings to prevent unnecessary anxiety or interventions for healthy individuals.
Sensitivity measures the model’s ability to correctly identify malignant tumors. Once again, Ensemble, Xception65, and Inception-v3 delivered strong sensitivity values, while AlexNet and VGG16 lagged behind. These results underline the clinical reliability of advanced models in minimizing false negatives.
Precision indicates the proportion of correctly identified malignant cases out of all predicted malignant cases. Xception65 (92.04%) and the Ensemble model (90.99%) led this category, suggesting that their false-positive rates for malignant prediction were low, vital for reducing overtreatment.
The Ensemble method, which aggregates multiple traditional classifiers (SVM, KNN, DT, NB, and NN), offered a balanced and robust performance across all metrics. Its strength lies in combining the strengths of diverse algorithms, reducing variance, and enhancing generalization. Notably, Xception65, with its deep and complex convolutional architecture, showed performance metrics very close to the Ensemble, making it a strong standalone candidate for tumor diagnosis in clinical applications. The heatmap provides a visual benchmark of these models across all key indices, facilitating quick identification of the best-performing classifiers for practical deployment. Figure 2 is intended as a compact, end-to-end summary of the comparative performance of the evaluated pipelines across the 4 key clinical metrics (accuracy, specificity, sensitivity, and precision). For each deep feature extractor (ResNet50, Xception65, VGG16, AlexNet, DenseNet, GoogLeNet, and Inception-v3), the values reported in the heatmap correspond to the final selected configuration for that feature extractor, that is, the single downstream classifier setting retained for cross-model comparison after the full set of candidate classifiers had been evaluated (SVM, KNN, DT, NB, NN, and the ensemble-of-classifiers strategy). Importantly, each heatmap cell reports the metric values obtained from the same selected configuration, ensuring that accuracy, specificity, sensitivity, and precision remain internally consistent and directly comparable across rows. The selection criterion and reporting rule are now stated explicitly in the text and caption, and the percentages displayed in Figure 2 are derived from—and can be cross-checked against—the corresponding results tables for each network (Tables 2-9), thereby enabling transparent tracking of how the key findings and concluding remarks follow from the numerical evidence. Figure 2 presents the comparative performance of the deep feature extraction models, where all metrics were computed using the support vector machine classifier to ensure methodological consistency and fair comparison across feature sets. By fixing the classifier, the figure isolates the impact of different deep feature representations on classification performance, eliminating variability that could arise from using multiple learning algorithms. This standardized evaluation strategy allows a clearer interpretation of how each convolutional neural network contributes to discriminative power, thereby directly addressing potential ambiguity regarding the underlying classification algorithm used to generate the reported metrics.
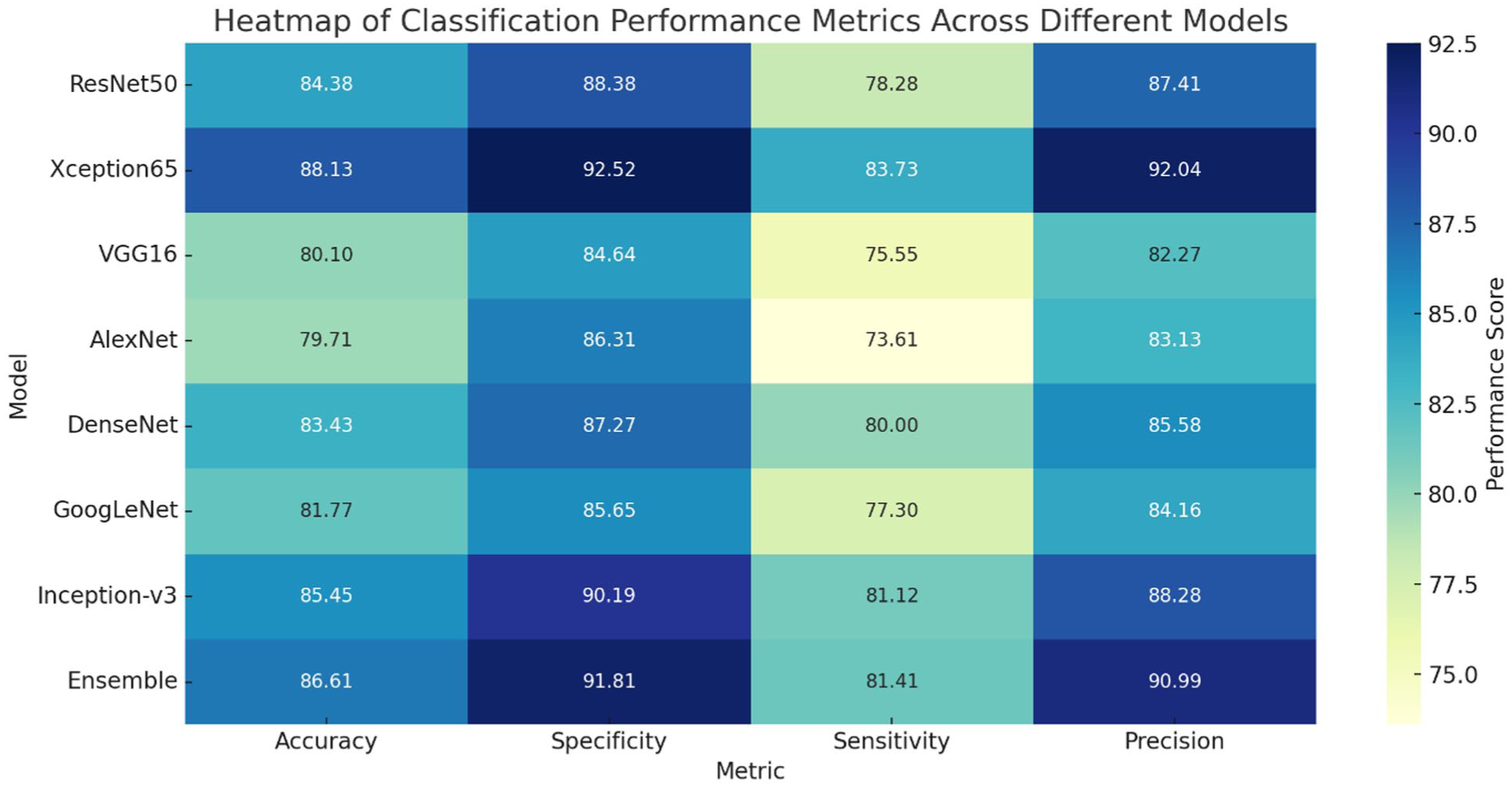
Final remarks: comparative summary of tumor classification models.
Conclusion
This study evaluated the performance of 8 deep learning models—ResNet50, Xception65, VGG16, AlexNet, DenseNet, GoogLeNet, Inception-v3, and a feature-level ensemble method—in the binary classification of breast tumors (benign vs malignant) using mammographic tumor outlines.
Key Findings
The results indicate that the Xception65 network and the feature ensemble approach consistently achieved superior performance across all major metrics, including accuracy, specificity, sensitivity, and precision. Xception65 demonstrated the highest precision (92.04%) and strong balance across metrics, while the ensemble approach showed remarkable stability and generalization, with accuracy reaching 86.61%, specificity 91.81%, sensitivity 81.41%, and precision 90.99%. In contrast, traditional models like VGG16 and AlexNet underperformed, highlighting their limited suitability for this task.
Applications
The findings of this study can directly support the development of automated diagnostic tools in breast cancer screening programs, especially for pre-screening or second-opinion systems in radiology clinics. By relying solely on tumor outlines—a low-dimensional and privacy-preserving representation—these models offer an efficient alternative to full-image processing, suitable for deployment in resource-limited environments.
Limitations
Despite promising results, the study faces several limitations. First, the analysis was based on a dataset of 100 mammographic image folders, which, although publicly available and valuable, may not fully represent the variability in clinical settings. Second, using only tumor outlines may ignore critical texture and contextual information available in the full mammogram. Additionally, the study does not account for real-time clinical constraints such as model interpretability and inference latency.
Future Directions
Future work should explore the integration of hybrid features, combining shape outlines with radiomic and deep texture features, to boost diagnostic robustness. Expanding the dataset and validating models on multi-center clinical data will be essential for generalization. Moreover, incorporating explainable AI (XAI) techniques could increase clinician trust and transparency in decision-making. Finally, deployment-oriented studies that measure runtime efficiency, hardware requirements, and user interfaces for radiologists will be key steps toward real-world clinical adoption.
Beyond achieving competitive classification performance, this study contributes to the growing body of interpretable medical imaging research by explicitly linking model outcomes to contour-driven, numerically defined lesion characteristics. Consistent with the principles of interpretable machine learning, the proposed framework emphasizes that predictive accuracy alone is insufficient unless accompanied by clinically meaningful explanations. By leveraging precisely annotated lesion boundaries from a high-quality mammography dataset, the analysis demonstrates how boundary-based variation indices (contour descriptors) encode diagnostically relevant information related to lesion irregularity and malignancy. This contour-centric perspective enables a transparent interpretation of model behavior, providing insight into why certain classification decisions emerge rather than merely reporting how well they perform. As such, the proposed approach offers a reproducible and explainable pathway for integrating deep learning into breast cancer assessment workflows, bridging the gap between black-box prediction 27 and clinically interpretable evidence.28,29
Footnotes
Acknowledgements
The AI tool ChatGPT is used to check the grammar of specific sections of the manuscript.
Ethical Considerations
All procedures performed in studies involving human participants were in accordance with the institutional and/or national research committee’s ethical standards and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Author Contributions
The author solely contributed to the conception and design of the study, data analysis, interpretation of results, and preparation and revision of the manuscript.
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Data will be available upon reasonable request.*
