Abstract
Background:
Explainability (xAI) is critical for fostering trust, ensuring safety, and supporting regulatory compliance in healthcare AI systems. Large Language Models (LLMs), with impressive capabilities, operate as “black boxes” with prohibitive computational demands and regulatory challenges. Small Language Models (SLMs) with open-source architectures present a pragmatic alternative, offering efficiency, potential interpretability, and alignment with data privacy frameworks. This study evaluates whether token-level attribution (TLA) methods can provide technical traceability in SLMs for clinical decision support.
Methods:
The Captum 0.7 attribution library was applied to a Qwen-2.5-1.5B model on 20 breast cancer cases from a publicly available dataset. Hardware requirements were profiled on consumer-grade GPU. Using perturbation-based integrated gradients, we analyzed how clinical input features statistically influenced token generation probabilities.
Results:
Attribution heatmaps successfully identified clinically relevant input features, with high-attribution tokens corresponding to expected clinical factors. The model occupied minimal storage, enabling local deployment without cloud infrastructure. This validates that SLMs can provide algorithmic traceability required for regulatory frameworks.
Conclusions:
This proof-of-concept demonstrates the technical feasibility of combining SLMs with perturbation-based xAI methods to achieve auditable clinical AI within practical hardware constraints. While TLA provides statistical associations, bridging toward causal clinical reasoning requires further research.
Keywords
Introduction
Explainable Artificial Intelligence (xAI) establishes trust, ensures safety, and upholds regulatory compliance within Artificial Intelligence (AI) systems in healthcare, with a particular focus on diabetes management. 1 Specifically, this prior research examined the essential technical and regulatory dimensions for developing explainable applications, including the application of SHAP-type xAI tool for the attribution of a level of contribution of a feature in the prediction of a specific diabetes dataset, such as the PIMA Indians Diabetes Dataset. 1
Historically, AI applications in medicine1,2 primarily leverage structured numerical data for disease analysis and prediction. 1 However, the growing integration of Large Language Models (LLMs) into clinical decision support, patient communication, and documentation introduces significant challenges.1,3 These models frequently operate as “black boxes,” 1 raising substantial concerns regarding transparency, bias, and safety, which are especially critical in high-stakes healthcare environments.1,3 -5 This disjunction highlights a pressing gap between technical advancements in xAI for language models 1 and the stringent requirements of clinical practice.
Central to addressing these concerns is the concept of trustworthy AI, 6 which mandates transparency supported by actionable insights and methodologies for implementing AI-enabled healthcare solutions. 1 This requires systems to be developed and utilized with adequate traceability and explainability,1,6,7 ensuring human awareness during AI interaction and comprehensive information for deployers regarding system capabilities and limitations.1,6 The advent of LLMs, specifically for processing complex clinical data, has indeed profoundly impacted healthcare. 3 Despite their advancements, often driven by the prevailing “Bigger is Better” paradigm, 4 this premise exhibits inherent limitations. 8 A more nuanced perspective on explainability is therefore necessary because while larger models possess impressive capabilities, their increased scale introduces substantial technical, ethical, and practical considerations. 8
Scaling LLMs incurs elevated computational costs demanding significant hardware resources and substantial energy consumption,4,8 which can impede their widespread deployment, particularly in resource-constrained healthcare settings.2,3,5 From a technical standpoint, training LLMs presents challenges such as the vanishing gradient problem, where the signal for updating model parameters diminishes as it propagates through deep networks, hindering effective learning. Research efforts are actively exploring more efficient architectures, such as DeepSeek R1, designed to process vast data volumes with reduced computational demands. 9 Furthermore, ongoing investigations into optimal scaling laws seek to elucidate the precise relationship between model size, training dataset size, and performance, with the objective of maximizing efficiency and minimizing resource waste. 4 Crucially, the inherent complexity of LLMs often results in a significant lack of transparency5,6,8 regarding their decision-making processes. Ethical concerns, 7 including bias in training data and the potential for misuse, demand careful consideration, especially given the sensitive nature of clinical data3,5 and the imperative for fair and equitable healthcare outcomes.6,8 If an AI system provides recommendations without a clear rationale, 6 its integration into clinical practice, particularly for high-stakes decisions like diagnoses or treatment plans,1,3,5,6 is unlikely to gain acceptance. While explainability in traditional machine learning often centers on feature importance, 1 particularly in numerical datasets, identifying the specific words, phrases, or even subword tokens contributing to a “feature” in natural language text remains a formidable challenge.6,7
In light of these challenges, SLMs are emerging as a compelling alternative, offering a pragmatic balance among efficiency, interpretability, and practicality.2,3,5 It is imperative to clarify that while the inherently “small” size of SLMs enhances their efficiency and reduces computational requirements,2,8 their open-source nature is arguably more pivotal for fostering the transparency and explainability of their decision-making processes, qualities that are indispensable in the clinical setting.1,3,5 The selection of appropriate xAI methods and tools for these models is non-trivial.3,4,7
The pursuit of explainability in AI has gained renewed urgency in clinical settings, where professional responsibility, patient safety, and regulatory compliance converge.1,3,5,6 As AI systems are increasingly considered for healthcare applications, the question extends beyond model performance to whether their decisions can be understood, trusted, and appropriately overseen.1,6 This need for “explainable AI” draws from social and cognitive sciences, which conceptualize explanation as a contrastive, context-dependent, and social process.6,7 As argued by Miller, 7 individuals seek to understand not just why an outcome occurred, but why it occurred instead of an expected alternative. This aligns with calls for rigorous research into how AI models can justify their outputs using clinically meaningful features rather than opaque computational artifacts.1,8 It is also crucial to distinguish between interpretability, the technical description of a model’s function, and explainability, which is the ability to communicate the reasoning behind a specific decision to a human user.7,10
Despite impressive performance across various benchmarks,3,5,9 the inherent opacity of LLMs presents significant epistemic challenges for clinical integration. Models such as GPT-43,5,8,9 or PaLM-2 are trained on extensive text corpora to capture statistical patterns, but they do not inherently encode causal relationships or verifiable logical structures. 7 This lack of grounding means their outputs, while contextually fluent, do not reveal a human-accessible decision path, making it impossible to determine why specific information was prioritized or excluded. 6 This “black-box” nature is particularly problematic due to their documented tendency toward “hallucination”—generating plausible but factually incorrect content. 11 Furthermore, existing explanation methods, such as those based on gradients or attention, often fail to produce attributions that align with expert domain knowledge. 6
From a regulatory perspective, this opacity creates challenges for auditability and validation.1,3,5 The European AI Act, 12 for example, emphasizes requirements for traceability and human oversight in high-risk applications, including clinical decision support.6,8 The massive parameter spaces and emergent behaviors of LLMs complicate both ex ante validation and ex post auditing processes.3,4,8
Beyond epistemic and regulatory concerns, LLMs also face substantial practical deployment challenges in healthcare settings. 3 While commercial LLMs demonstrate impressive linguistic fluency,3,4 they often lack the structured reasoning capabilities required for clinical tasks involving multistep inference or causal explanation. 3 Their black-box nature,5,8 combined with the risk of hallucinations, limits their epistemic reliability. 3 These constraints, along with hardware requirements that exceed typical hospital infrastructure, make large models impractical for real-time, privacy-sensitive clinical deployments.3,5 This further motivates the exploration of lightweight, locally-deployable SLMs 2 as a more sustainable 8 and transparent alternative.3,5
A text-based explainability tool featuring a broad set of functions for xAI in deep learning has been adopted.5 -7 Captum 0.7 was chosen for this study as it is open-source, requires low computational power, and is part of a well-established ecosystem. This aligns with the focus on Textual Explainability, or Semantic xAI (SxAI), which refers to a model’s capacity to provide rationales that are not only technically accurate but also readily understandable to non-AI experts, particularly healthcare professionals requiring alignment with clinical reasoning.1,3,5,6 The present study builds upon and integrates the methodologies outlined in Geukes Foppen et al, 1 bridging the gap toward Large Concept Models (LCM; Figure 1). This shift toward concept-based explanations aims for greater alignment with human understanding,1,5 -7 as good explanations are characterized by clarity, precision, understandability, sufficiency of detail, completeness, usefulness, accuracy, and trustworthiness.3,6
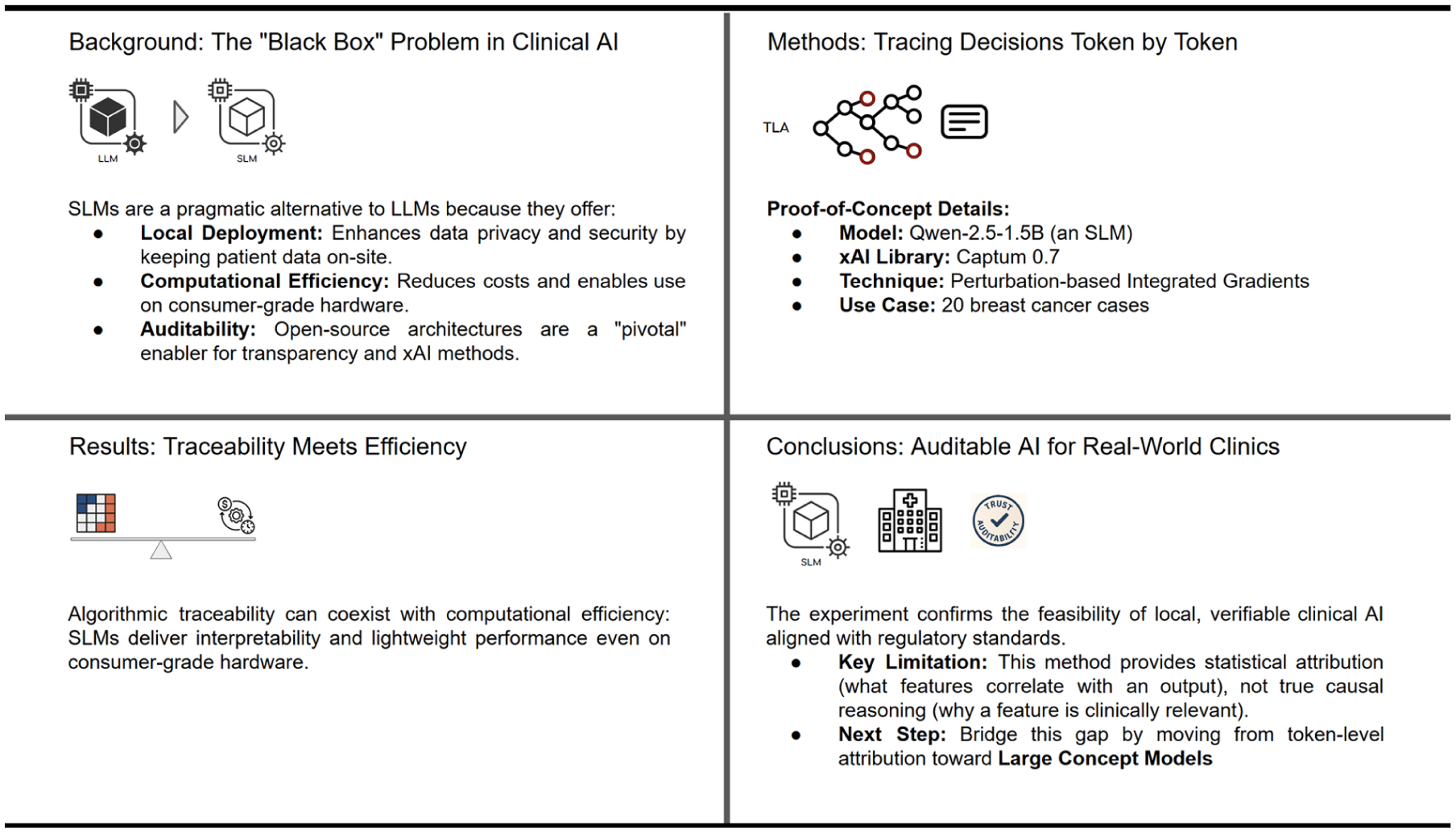
The path to verifiable clinical AI: An open-SLM proof of concept.
Clinical Landscape of Language Models: From Efficiency to Explainability
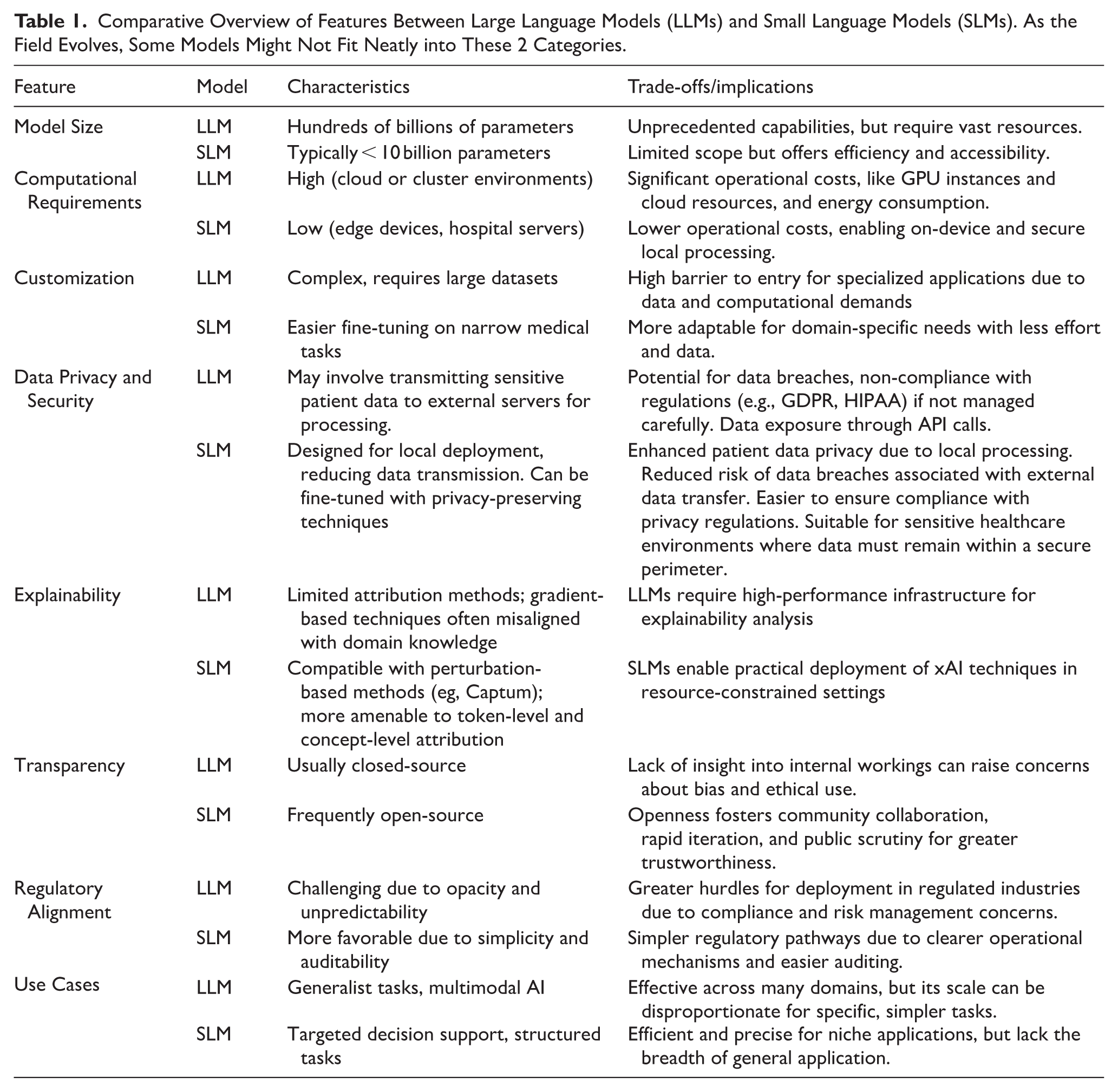
SLMs, typically defined by parameter counts below 7 to 10 billion 3 and open-access architectures,3,5 represent a potentially more tractable approach for clinical AI applications. 3 Recent architectures, including Phi-2, Mistral, and similar models, demonstrate that reduced parameter counts can maintain competitive performance on specific tasks 5 while offering computational efficiency advantages.3,4 The reduced computational footprint of SLMs may facilitate compliance with regulatory requirements for transparency and auditability.4,5 Their computational efficiency enables local deployment and fine-tuning, 2 potentially reducing the risk surface compared to large-scale, centralized models.3,5 Additionally, SLMs may be more amenable to existing explainability techniques such as perturbation analysis and attribution methods. 5 Moreover, SLMs align well with sustainability goals. 8 Given the rising environmental and financial costs of training and hosting LLMs, 8 the use of lightweight models is both an ecological and ethical imperative in healthcare. 13 To facilitate a clear understanding of the architectural and operational distinctions driving the focus on SLMs, Table 1 provides a comparative overview of key characteristics between LLMs and SLMs relevant to healthcare applications.
Comparative Overview of Features Between Large Language Models (LLMs) and Small Language Models (SLMs). As the Field Evolves, Some Models Might Not Fit Neatly into These 2 Categories.
Reduced complexity in SLMs doesn’t inherently lead to interpretability or increased interpretability. Like LLMs, most SLMs utilize transformer-based architectures, meaning their internal representations remain opaque despite their smaller scale.5,6,8 Consequently, explainability must be a deliberate engineering effort, rather than an assumed benefit of their size.
Recent advances have demonstrated that open-access SLMs can rival or even surpass large proprietary models in clinical benchmarks. The clinical applicability of SLMs has been demonstrated with notable results:
Meerkat, a family of medical SLMs (from 7 to 70 billion parameters), 3 addresses concerns about the privacy and security of proprietary models, allowing for on-premises implementation on relatively common GPUs (such as a single NVIDIA GeForce RTX 3090 with 24 GB for the 7B and 8B models). Meerkat-7B, for the first time surpassed the United States Medical Licensing Examination passing threshold (60%) for a model of its size (achieving 77.1% accuracy), and Meerkat-70B outperformed GPT-4 by an average of 1.3% across 6 medical benchmarks. In complex clinical cases, Meerkat-70B correctly diagnosed 21 cases, exceeding the human average (13.8) and nearly matching GPT-4 (21.8). Its responses to daily clinical questions are more detailed and complete than those of other smaller models. Its excellent performance stems from the use of high-quality Chain-of-Thought data, synthesized from 18 medical textbooks via GPT-4.
Diabetica-7B, a specialized model for diabetes management, 14 outperformed all other open-source models of similar size and proved comparable to proprietary models (GPT-4, Claude-3.5) in diabetes-specific tasks (multiple-choice questions, “fill-in-the-blank,” open dialog). Its clinical evaluations showed superior performance to human experts in medical consultation, education, and clinical record summarization, with significant time savings. Diabetica employs a meticulous data processing pipeline and a self-distillation method to align the model with human preferences and mitigate “catastrophic forgetting.”
The BC-SLM (Breast Cancer Small Language Model), adapted to specific guidelines and based on Retrieval-Augmented Generation, demonstrated high concordance (86%) with the recommendations of a multidisciplinary tumor board, providing transparent decisions and referencing sources. 5
SLMs also show great potential in integrating multimodal data (radiological images, clinical notes, laboratory data) for holistic disease risk prediction, outperforming single-source models and offering personalized decision support. 5
These rapid tech advantages highlight the synergistic value of models that are both small and open: efficient, auditable, 1 and fit for integration into resource-constrained healthcare environments, 2 as highlighted in Table 1.
An Approach to Explainable AI in Text Analysis
Widespread clinical adoption of language models remains hindered by regulatory demands for algorithmic traceability (eg, EU AI Act Article 13, FDA SaMD guidelines),3,5 which fundamentally conflicts with the opacity of current LLMs. This study tests whether SLMs with open weights, code for the model architecture,3,5 and modern xAI techniques can provide technical traceability within real-world clinical constraints (hardware, latency), 2 distinct from aiming for textual explainability.
Pragmatic clinical AI adoption necessitates balancing 3 competing demands: performance (model accuracy), auditability (traceability of decisions), and feasibility (resource constraints). 2 Here, the focus is on SLMs as the most suitable GenAI models for meeting these demands, particularly given their lower computational constraints.2,3,8 These constraints make SLMs advantageous for both text generation and xAI algorithms, especially in environments where LLM deployment is often impractical due to resource limitations.2,8
Proof-of-Concept
Starting from the hypothesis that the combined application of SLM and xAI techniques based on tokenization could offer a significant advantage in the interpretability of decisions in the medical field, we developed a Proof-of-Concept (PoC). This PoC integrates SLM and xAI techniques.
This PoC describes structured steps to interpretability (Figure 2). As a starting point, patient-specific clinical data, from a publicly available dataset of 20 breast cancer patients and the prompt engineering framework, 5 is organized into structured prompts. This sample specifically enables a pragmatic evaluation of the explainability mechanisms within this PoC.
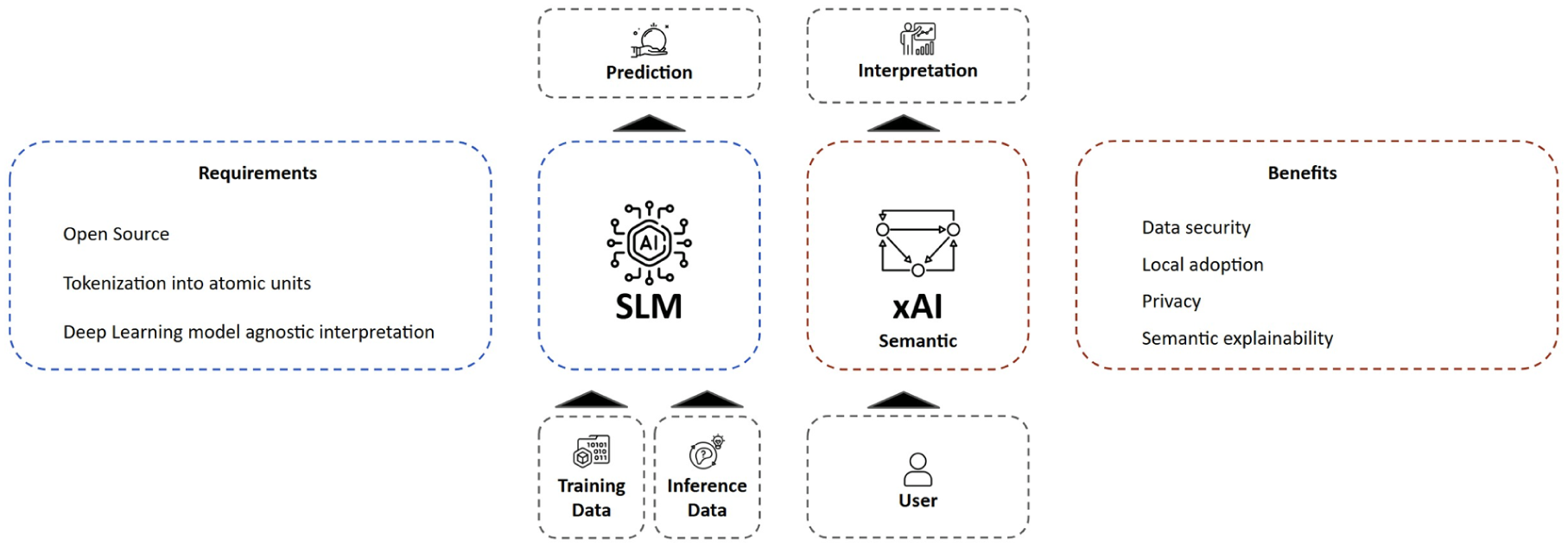
Flow diagram showing the overall system architecture, highlighting the interaction between SLM components and xAI analysis.
To bridge the gap between technical feasibility and clinical utility, this investigation employs a structured, two-phase methodology designed to systematically evaluate whether token-level attribution (TLA) in SLMs can meet the dual requirements of technical traceability and practical deployability in oncology use cases. This design, while acknowledging the system’s non-interpretative nature, comprises:
• First methodology phase: Technical Validation: ○ Model: Qwen-2.5-1.5B (8-bit quantized) ○ Methods: ■ Controlled token perturbation (20% masking rate) ■ Captum’s LLM Attribution with integrated gradients ○ Enhancement: Dataset stratified into 15 training cases (represented as prompts) ○ Metrics: ○ Hardware profiling (VRAM usage, latency)
• Second methodology phase: Clinical Relevance Assessment ○ Dataset: 20 breast cancer cases
5
○ Focus Features: ■ Receptor status (ER/PR/HER2) ■ TNM staging ■ Oncological family history ○ Analysis: ■ Heatmap attribution versus clinician-annotated gold labels ■ Sensitivity analysis for critical clinical variables
To ensure a focused and manageable scope for this initial evaluation, we concentrated on specific aspects, meaning some important considerations were beyond the purview of this study. These include:
Human-centered evaluation: No clinician usability studies or cognitive load assessments (eg, NASA-TLX) were conducted to evaluate real-world user experience1,3;
Causal inference validation: The Captum attribution methods provide correlational insights rather than establishing causal relationships between input features and model decisions;
Cross-model performance comparison: The focus is exclusively on SLM characteristics without benchmarking against larger foundation models or alternative architectures.
Textual Attribution Analysis for Healthcare SLMs: A Technical Backbone for Enhanced Explainability
SHAP is a well-recognized xAI tool and uses a mathematical model in the interpretation processes of machine learning models. Where SHAP is designed for many machine learning tasks and family of models, Captum is specific for deep learning models and fits well with language models internal workings. Captum version 0.7 is an open-source library within the PyTorch ecosystem, designed to work seamlessly with PyTorch models. It provides tools for analyzing how models process inputs and generate outputs. The native compatibility of Captum in the PyTorch ecosystem makes this library a ready-to-use solution for analyzing individual tokens and understanding model decision pathways.
To understand this study, and in general, how Language Models work, it is important to understand what tokens are. Tokens are the fundamental input units to language models; they can be words, character sequences, multiple words, or punctuation marks. Tokenization into atomic units (these tokens) enhances efficiency, accuracy and allows the model to manage rare words (if a model has never seen a word, it can be broken down into known parts), morphological variations, and keep the internal vocabulary size manageable. Also, breaking down words into components allows models to learn root and affixes, enhancing their ability to generalize across similar words. Each token is mapped to an integer so that a machine can process it. Language models then use conditional probabilities over tokens to generate new text.
Our approach leverages what we term semantic explainability, using Captum to analyze how meaningful text units influence model outputs. This method identifies statistical associations between input tokens and predictions. Captum segments text into manageable tokens and enables analysis of input variations to assess their impact, offering statistical, not semantic, insight into model behavior.
Trust and ethical considerations are at the core of successful AI integration in life sciences. By offering explainable mechanisms, xAI fosters confidence in AI systems, ensuring that healthcare providers and patients can understand the rationale behind clinical suggestions. Moreover, xAI helps mitigate concerns related to algorithmic bias, enabling the identification and correction of disparities in model training and deployment. This is especially crucial in ensuring equitable healthcare delivery and compliance with regulatory standards.
Experimental Setup
Initially, a Qwen 2.5 7B, and Qwen 2.5 3B 15 model was employed before transitioning to a Qwen 2.5 1.5B 8-bit quantized model, designed to operate on resource-constrained environments. This version was used in a zero-shot learning setup, no prior fine-tuning was applied, and no examples were provided in prompts, deliberately replicating the limitations of commercial LLMs when dealing with specialized medical cases. All these models are used with Hugging Face, the most widespread library for open source GenAI and NLP models (and more). Using perturbation techniques through Captum’s attribution methods, we analyzed the statistical importance of text components in influencing model outputs. It’s important to keep in mind that perturbation-based attribution reveals feature importance, but not the clinical rationale behind predictions.
Technical Implementation Details
Rather than perturbing individual character sequences (which would create unnatural text because they are sub-words), Captum allows definition of interpretable features at word and sentence levels, computing attribution for these complete components.
To perturb, in the context of attribution methods, means using a certain feature or removing it. In the context of the text, a possible way to “remove” a feature is to use a “baseline” reference value. The default value Captum uses for the baseline is the empty string, but this is not optimal as it would produce unnatural texts and thus output. We adopt a different strategy, our baseline are those patient values that effectively contributed to generate the target response text. Changing text features shows which tokens will be less likely to be produced with the new input prompt.
The result of the analysis with Captum mapped relationships between input text features and the generated tokens. By systematically varying text features, critical decision factors were identified and compared their importance across diverse clinical scenarios. This approach provided insights into the model’s reasoning processes and enabled us to refine its interpretability.
It is important to understand that for these tests extremely SLMs are used for computer efficiency, to demonstrate that SLM can be used with constrained hardware resources, and to simulate the aforementioned limitations relating to LLMs, for matched comparison. Therefore, this PoC is aimed at demonstrating the impact of this approach on decision-making with Language models, rather than evaluating the accuracy.
Attribution Analysis Results
The perturbation techniques effectively revealed the relative importance of different clinical parameters in the model’s decision-making process, providing insights into how the SLM processes and weights various medical factors.
The analysis (Figure 3) mapped statistical relationships between input clinical features (horizontal) and generated tokens (vertical). By systematically varying text features, we identified which inputs most strongly influenced token generation probabilities.
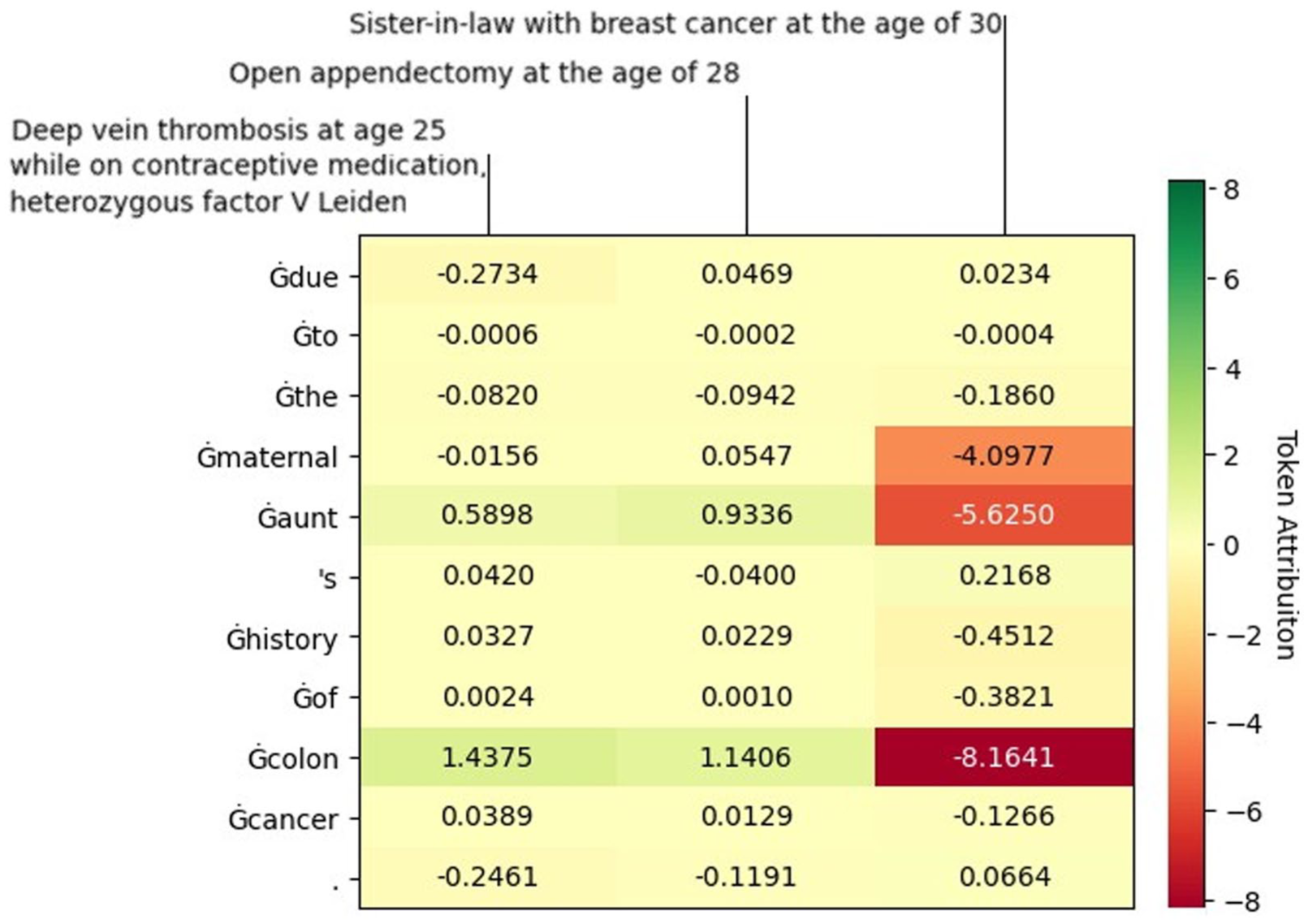
Token-level attribution heatmap for clinical input features. Input features are displayed on the horizontal axis (PI = Previous Illness, PST = Previous Surgical Treatment, OFH = Oncological Family History); generated tokens appear on the vertical axis. Color scale indicates attribution values: blue represents negative attribution (reduced token probability), red represents positive attribution (increased token probability).
To validate that the model learns genuine feature-outcome associations rather than spurious correlations, we conducted a counterfactual analysis. Counterfactual analysis validated that the model learns genuine feature-outcome associations rather than spurious correlations.
The predominantly negative attribution values demonstrate that the model correctly recognizes feature-target mismatches; features from Patient A appropriately receive negative attribution when predicting outcomes for Patient B. This sensitivity to patient-specific context confirms the model’s reliance on learned clinical patterns rather than dataset artifacts or positional biases. Conversely, if mismatched inputs had shown neutral or positive attributions, this would indicate problematic behavior where the model ignores input features.
The PoC validated the initial hypothesis that TLA methods, implemented through Captum, can be technically applied to language models. This demonstrates the feasibility of combining SLMs with statistical xAI techniques to audit model behavior, though not to fully explain clinical reasoning.
Discussion
This study has demonstrated that SLMs, particularly those with open-source architectures, present a pragmatic pathway toward achieving auditable and trustworthy AI solutions within resource-constrained healthcare environments (Figure 2). By addressing the limitations of LLMs, including their inherent opacity, 11 prohibitive computational demands, 8 and associated challenges in clinical integration, this work highlights the synergistic value of “small” and “open” models for high-stakes medical applications. The PoC, utilizing semantic explainability techniques with an SLM in an oncology use case, underscores the technical feasibility of providing vital insights into AI decision-making.2,3
Pragmatic Course for Healthcare AI Including LLMs and SLMs
The evolution of language models in healthcare is undergoing a significant transformation. While LLMs continue to demonstrate impressive capabilities, 14 SLMs emerge as a complementary solution that effectively addresses the specific needs of the healthcare sector, particularly in terms of security, privacy, and regulatory compliance. 1 (S)LMs can be analyzed with the xAI lens when they are open-source, thus allowing for the weights and architecture to be inspected. 5 By focusing on what these models can achieve within limited computational environments, the concept of “small” is redirected in a way that emphasizes their transformative potential across a wide range of applications, and thus extends the application value continuum of language models. This shift toward SLMs offers a pragmatic balance among efficiency, interpretability, and practicality, crucial for widespread adoption (Table 1).
Technical Feasibility of SLM-Enabled Semantic xAI
While previous work 1 established the conceptual and regulatory importance of xAI for ensuring trust and safety with structured numerical data, this follow-up study critically shifts the focus to (unstructured) text data and the unique “black box” challenges introduced by language models. 7 The present technical PoC, moves beyond the “why” of xAI to the “how” by proposing an actionable framework for auditing SLMs in a clinical context. Specifically, using a Qwen-2.5-1.5B model on a breast cancer dataset, this PoC confirms the technical feasibility of applying token-level attribution analysis via Captum. This implementation of text feature analysis and semantic explainability enables detailed tracking of how SLMs process and weight clinical information, potentially preventing model drift through continuous monitoring of decision models. By focusing on a small, open-source model, the study also addresses the practical constraints of deploying AI in resource-limited hospital environments. 3 While this PoC confirms the technical feasibility of applying TLA analysis to SLMs in healthcare contexts, significant methodological and interpretability challenges remain to be studied. 6
Defining and Achieving Explainability Versus Auditability
The integration of AI into clinical medicine demands rigorous scrutiny of model decision-making processes regarding both interpretability and explainability. 11 While LLMs remain fundamentally opaque, 14 SLMs achieving auditability through technical traceability is a critical and achievable goal. 5 This study specifically employed Captum, a PyTorch-based library, to generate attribution heatmaps using methods such as Integrated Gradients (Figure 3). This PoC confirms the feasibility of tracking input-output dependencies in SLMs at the token level, identifying statistical associations between clinical inputs and model outputs. However, it should be emphasized that these visualizations do not offer semantic explanations in the ontological or causal sense, but rather serve as statistical heuristics. 1 While current xAI methods often fail to align with human semantic intuitions or clinical reasoning, 7 this pragmatic, infrastructure-aligned first step provides a controlled, auditable approximation of transparency. Further research explores Concept-based Explainability, which aims to align internal representations with human-interpretable concepts, but such methods remain underdeveloped for SLMs in the clinical domain.
xAI Drives Regulatory Compliance and Trust
The EU AI Act proposes a risk-based approach to regulating AI systems. 16 Importantly, the Act also addresses general-purpose AI models like LLMs. Recognizing their unique characteristics, the Act stipulates separate rules regarding transparency requirements for these models. 1 This emphasis on transparency is crucial for building trust and ensuring accountability in the use of LLMs in healthcare and other sensitive domains. 6
Regulatory compliance and fostering trust are paramount for the successful integration of AI into healthcare. Regulatory bodies like the FDA 17 and EMA 18 have recently issued guidelines addressing the use of AI in drug development and the medicinal product lifecycle, reflecting the growing recognition of AI’s potential impact and the need for a framework to ensure its safe and effective implementation. “Human in the loop” is a recommended practice by both the FDA and EMA, supporting regulatory compliance by involving humans in interacting with, testing, auditing, and validating AI models, with the level of involvement increasing with the model’s risk.
This complex regulatory landscape underscores the need for an adaptable approach in integrating AI principles, 19 emphasizing explainability and transparency tailored to diverse AI applications. 1 While a growing trend toward alignment exists, distinctions remain, as seen in the comparative approaches to transparency between Health Canada (as seen in Health Canada’s “Pre-Market Guidance for Machine Learning-Enabled Medical Devices”) 20 and FDA frameworks (as seen in FDA’s “Draft Guidance on AI-Enabled Device Software Functions”) 21 for AI-powered medical devices. Health Canada mandates detailed explanations of ML model workings, whereas the FDA recommends transparency without the same level of detail. This heterogeneity, coupled with the institution-specific AI processes in healthcare, poses significant challenges for transferring expertise and creating a unified methodology. 5
Regulatory considerations are crucial for ensuring the safety and effectiveness of drugs and clinical decision-making. The inherent opacity of LLMs is a key reason why the FDA has not yet authorized any LLMs for clinical use, 22 as their broad capabilities and unpredictable outputs require careful scrutiny. In healthcare, language models must demonstrate robustness to errors and resistance to cyber-attacks to prevent compromise of clinical decisions. 1
Multidisciplinary tumor boards exemplify the critical need for transparent and auditable AI by integrating vast amounts of data and automating analysis, enhancing the accuracy, timeliness, and personalization of clinical decisions. 23 These boards are a crucial component of comprehensive cancer care, ensuring that patients receive evidence-based, personalized treatment recommendations based on the latest clinical guidelines and evidence, which promotes high-quality care. 24 Similarly, Data Monitoring Committees and other oversight groups can significantly benefit from xAI-centered SLMs in their oversight functions, 25 as these models can analyze complex data, identify patterns, and provide insights into their decision-making processes, enhancing trust and ensuring scientific soundness. Open-source SLMs, with their potential for greater interpretability, could facilitate regulatory approval processes, particularly where decision-making transparency is crucial. 5
Bridging the Gap Toward Large Concept Models
Translating these technical insights into relevant applications in healthcare requires future research. The fundamental challenge of connecting statistical token attribution to truly interpretable explanations remains largely unresolved in current xAI approaches for healthcare applications. 1
While TLA provides statistical associations, establishing causal relationships requires fundamentally different methodological approaches. Current attribution methods identify which input features correlate with model outputs, but cannot determine whether these features causally drive decisions or merely co-occur with causal factors. Bridging this gap may require hybrid architectures combining statistical models with causal inference frameworks. 26 Applying causal mediation analysis in clinical SLMs is an exciting endeavor that remains largely unexplored.
By grouping related features into higher-level clinical concepts, the clarity and applicability of AI-driven recommendations can be improved. This direction aligns with emerging approaches like LCMs introduced by Meta, which focus on predicting and manipulating concepts rather than individual tokens. A “concept” is defined as a language and modality-agnostic unit corresponding to a sentence or higher-order idea.
Unlike traditional LLMs that predict next tokens, LCMs predict and manipulate semantic concepts—units more aligned with clinical reasoning (eg, “receptor status” rather than “ER+,” “PR+,” “HER2-” as separate tokens). Adapting LCM architectures to healthcare-specific ontologies (eg, SNOMED-CT, ICD-11) could enable SLMs to generate explanations grounded in clinical conceptual frameworks. Applying an LCM-inspired approach to healthcare AI could help bridge the gap between word-based reasoning and holistic clinical decision-making, where complex patient assessments require an integrated, concept-driven interpretability framework.
Concept-level explainability methods complement LCM architectures by providing attribution at the level of semantic units rather than isolated tokens. Recent research introduces ConceptX, a model-agnostic framework that identifies semantically rich concepts (drawn from resources like ConceptNet 27 within the prompt and assigns them importance based on the semantic similarity of generated outputs.
Technological challenges also require the evolution toward increasingly accurate and efficient domain-specific models, optimized for specific medical areas, on which to perform explainability through more intuitive user interfaces for healthcare professionals. 3 Furthermore, it is expected that regulatory frameworks will continue to evolve to specifically address xAI in healthcare, encompassing requirements for accessible model weights, training and test datasets, and training methodologies throughout the entire lifecycle for full transparency. 1 Integrating clinical expertise will be crucial to contextualize technical findings and ensure practical applicability. 11
SLMs represent a technological alternative and enable a new paradigm in the application of AI in healthcare, where the balance between performance, interpretability, and safety is crucial (Figure 1). This responsible adoption of AI in clinical practice will be significantly accelerated by their continued development, particularly when driven by advancements in robust textual explainability frameworks designed to provide increasingly intuitive and clinically relevant insights.
Footnotes
Ethical Considerations
N/A.
Author Contributions
Remco Jan Geukes Foppen, Ph.D., Alessio Zoccoli, Vincenzo Gioia, M.S. All authors are equal contributors.
Remco Jan Geukes Foppen: Writing—review & editing, Writing—original draft, Methodology, Investigation, Conceptualization.
Alessio Zoccoli: Writing—review & editing, Methodology, Investigation, Formal analysis.
Vincenzo Gioia: Writing—review & editing, Writing—original draft, Methodology, Investigation, Formal analysis, Conceptualization.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
