Abstract

A prominent resistance mechanism in Gram-negative bacteria is drug efflux. These mechanisms expel solutes from the cell. Antimicrobials, metabolites, and quorum-sensing signal molecules are expelled through efflux pumps helping bacteria to remove these toxic products from their interior environments. 1 The flux of chemicals mediated by efflux pumps is inhibited by drugs known as efflux-pump inhibitors (EPIs). Antibiotic efflux inhibitors are in the nascent stage of drug development, and these efflux pumps can be used as significant therapeutic targets. 2 In underdeveloped nations, the indiscriminate use of antibiotics could lead to the emergence of more highly resistant bacteria strains linked to periodontal microbial resistance. 3 Allosteric sites are the active site of an enzyme where a molecule can bind and change the shape of the active site. Antibiotic resistance usually arises when the target protein’s active site loses its affinity for the antibiotic; enzymes chemically alter it, or when efflux mechanisms are employed. 4 Understanding allosteric mechanisms are essential, specifically finding Allosteric sites, which is a pre-requisite for drug discovery and design. Using pocket characteristics and protein dynamics, many computational approaches predict Allosteric sites. 5
Porphyromonas gingivalis produces a multidrug-resistant efflux protein called a bifunctional NAD(P)Hhydrate repair enzyme. This enzyme catalyzes the epimerization of the S- and R-forms of NAD(P)HX, and the dehydration of the S-form of NAD(P)HX at the cost of ADP, turned into AMP. This efflux protein structure was not solved with X-ray crystallography or NMR and was identified using the alpha fold. Besides, its active, hydrophobic, and Allosteric sites still need to be solved. 6 Hence, this study aimed to identify potential Allosteric sites for drug designing for efflux inhibitors.
Using PASSer (https://passer.smu.edu/) is an ensemble learning tool designed to predict Allosteric sites accurately (Figure 1). 7 Further, Dogsitescorer (https://proteins.plus) was used for detecting Binding pockets (Figure 2). DoGSiteScorer is a grid-based method that uses a Difference of Gaussian filter to detect potential Binding pockets—solely based on the 3D structure of the protein—and splits them into subpockets. Uniprot id—A0A0K2J2N6_PORGN, Porphyromonas gingivalis multi-efflux protein was selected, downloaded, and submitted to the passer tool to identify druggable sites.
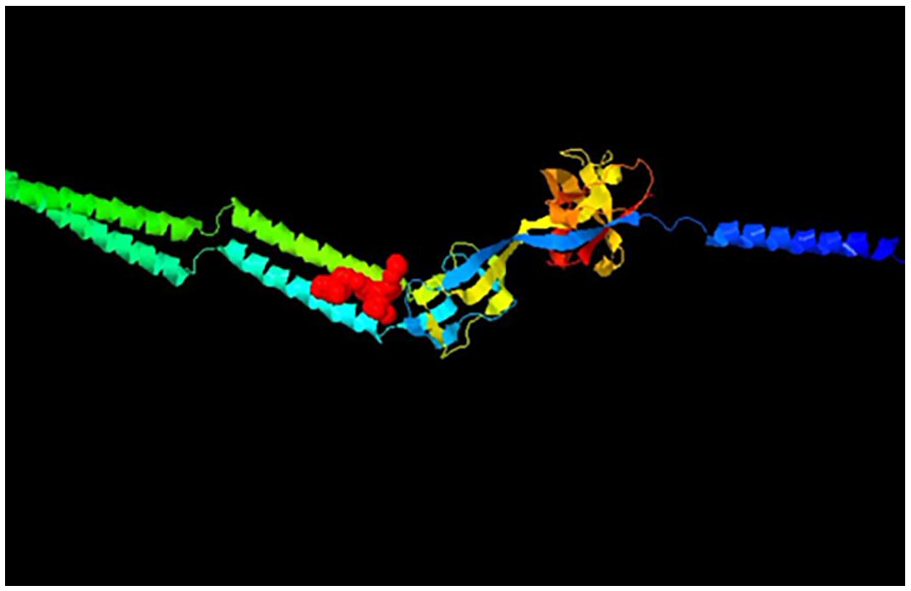
Allosteric Site of Porphyromonas gingivalis (PASSer) (The red color pocket indicates a higher probability with a higher likelihood of the Binding pocket being an allosteric site. Other color pockets have less probability of druggability).

Binding pockets (DoGSiteScorer) (The yellow and gray areas indicate Binding pockets).
Ensemble learning method predicted druggable Binding pockets with Druggability Score: 0.022 and Hydrophobicity score: 32.000. AlloPocket 1 shows a Score of 0.229, Druggability Score: of 0.022, Number of Alpha Spheres: 30, Volume: 391.230, Mean local hydrophobic density: 12.222, Hydrophobicity score: 32.000. The alpha model confidence score was high (pLDDT > 90). DoGSiteScorer is a grid-based approach that detects possible Binding pockets based on the 3D structure of the protein and divides them into sub-pockets using a Difference of Gaussian filter. Binding pockets showed with volume surface area of 2547.99 A and a surface area of 4052.69 A (Table 1).
Characteristics of identified Binding sites.

In conclusion, the current study identifies five potential druggable sites for small molecules to target Resistant efflux pump Porphyromonas gingivalis protein.
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
