Abstract
Background:
Acute myeloid leukemia (AML) has a heterogeneous molecular profile, clinical presentations, and response to treatments and outcomes. DNA methylation is conducted by DNA methyltransferases including DNMT3B. Poly ADP-ribose polymerase 1 belongs to a family of enzymes that mediate important cellular processes including DNA repair, transcription, and cell death/cell proliferation, and it is involved in the development, spread, treatment, and prognosis of some cancers. The objective of this study is to assess the impact of PARP1 and DNMT3B genes expression on laboratory characteristics, response to treatment and survival in Egyptian cytogenetically normal AML patients.
Methods:
This study included 67 Egyptian CN-AML patients in addition to 8 healthy bone marrow donors. Measurement of DNMT3B and PARP1 gene expression was done on bone marrow samples via real-time semiquantitative polymerase chain reaction.
Result:
Expression of both DNMT3B and PARP1 genes was significantly upregulated in AML (P = .001, P = .036, respectively). Upregulated DNMT3B was associated with higher total leukocyte count (TLC), PB, and BM blast cell%. Also, upregulated PARP1 correlated with higher TLC, PB, and BM blast cell%. High expression of both DNMT3B and PARP1 correlated with greater frequencies of FLT3-ITD. High DNMT3B expression, and combined upregulation of both PARP1 and DNMT3B genes associated significantly with ELN stratification. But no correlation was found with response (CR), overall survival (OS), disease-free survival (DFS), or event-free survival (EFS).
Conclusion:
Our findings highlight the importance of considering DNMT3B and PARP1 expression levels as potential prognostic biomarkers for progression and aggressiveness of CN-AML patients in AML. Assessing their expression levels could be an indicator to guide treatment decisions and potentially improve patient outcomes.
Background
Acute myeloid leukemia (AML) is an abnormal clonal proliferation of myeloid progenitors in the bone marrow and/or peripheral blood and other tissue. 1 Acute myeloid leukemia derives from a dominant mutation, and then develops cooperative transformative mutations and other mechanisms such as epigenetics modifications leading to neoplastic transformation and clinical/biological heterogeneity. 2 The unfavorable prognosis of AML is primarily due to the heterogeneity of leukemic cells and their ability to develop resistance to chemotherapy. Therefore, it is crucial to explore novel therapeutic approaches that target specific genes involved in apoptosis and tumor progression. 3
Epigenetic modifications are inheritable changes in gene expression that occur without altering the DNA sequence. These modifications include DNA methylation, histone deacetylation, and miRNA regulation. 4 DNA methylation is carried out by DNMT enzymes, which include DNMT1, DNMT3A, and DNMT3B. These enzymes regulate epigenetic changes by methylating cytosine, predominantly at CpG dinucleotides, resulting in the formation of 5-methylcytosine. Abnormalities in DNA methylation are closely linked to tumor progression and patient prognosis, particularly in hematological malignancies such as AML. 5
Poly (ADP-ribose) polymerases (PARPs) are essential enzymes that regulate various cellular processes, including the DNA damage response, chromatin remodeling, and transcriptional regulation, thereby preserving genetic stability. Approximately 90% of poly-ADP-ribosylation (PARylation) within a cell is catalyzed by PARP1. PARP1 facilitates PARylation by covalently attaching ADP-ribose units to specific amino acid residues on target proteins. While PARP1 was originally recognized for its role in detecting and repairing single-strand DNA breaks, its PARylation activity also promotes the recruitment of other DNA repair proteins to sites of damage. 6
DNMT3B and PARP1 have been implicated in cancer progression and their expression levels have been observed to be altered in various types of cancer.4,7,8 In AML, few studies explored the expression of DNMT3B and its prognostic significance, whereas, limited research investigated the role of PARP1 and postulated that it may function as a standalone prognostic determinant. 9
Notably, results from published clinical trials revealed that PARP inhibitors (PARPi) in hematopoietic cancers are promising therapies, however, patient-specific factors should be considered, and further studies on the regulation and expression of PARPs are necessary to provide more accurate data on their ideal therapeutic use. 4 Therefore, we conducted a prospective analytic study with the aim of this study sought to measure DNMT3B and PARP1 genes expression in de novo adult AML patients, and to assess their utility as prognostic biomarkers by correlating the results with established prognostic parameters and with patients’ clinical outcome.
Patients and Methods
The study population comprised 67 newly diagnosed (de novo) adult CN-AML patient who presented to the outpatient clinic of the National Cancer Institute (NCI), Cairo University, from April 2019 to May 2020. The range of patients’ ages was (19-73) years. In addition, 8 age and sex matched healthy subjects were chosen from the BM donors at the bone marrow transplantation (BMT) unit and included as a control.
Exclusion criteria included Pediatric AML, previous treatment, AML M3 subtype and association with other neoplasms. All cases were diagnosed based on standard diagnostic methods of examination and cytochemical staining of the BM, immunophenotyping and cytogenetic analyses according to WHO classification, 2017. Patients were classified according to ELN risk stratification of AML 10 into favorable risk (FR), and intermediate risk (IR).
The standard care of treatment in our institute for AML fit patients younger than 56 years (apart from APL) remains intensive chemotherapy with 7 + 3 protocol (Ara-C/Idarubicin or Doxorubicin) aiming for remission induction, while unfit or elderly patients who are ineligible for intensive chemotherapy protocol, receive less aggressive treatment on palliative intent including HMAs or low dose Ara-C. Indications of HSCT in AML patients in our center, patients with unfavorable cytogenetics (eg, FLT3) in the first CR & Refractory/Relapsed Patients after achieving complete remission.
Response to induction therapy was assessed both clinically and by BM examination at day 28. Complete remission was defined according to the standard criteria, 10 then the patients were divided into responsive (CR), and nonresponsive (No CR). Patients were followed up for at least 1 year after treatment.
RNA extraction, and cDNA synthesis and qPCR reaction
A sample of 1 to 2 mL of BM was collected on EDTA vacutainers under complete aseptic precautions before the start of treatment. Total cellular RNA from Human BM was extracted by using QIAamp RNA blood Mini Kit for total RNA purification (QIAGEN®Austin, Texas, USA) following the manufacturer’s instructions. RNA concentration and samples’ purity were detected by using the Nanodrop spectrophotometer. One microgram of RNA was used to perform Reverse transcription via Applied Biosystems™ High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific, USA), then cDNA was stored at −20°C until required.
For amplification of the cDNA to detect DNMT3B, PARP1 genes, the TaqMan® Universal PCR Master Mix at 2× concentration (Catalog no.: 4440040, Thermo Fisher Scientific, Applied Biosystems, USA) was used in addition to readymade assay, TaqMan primer probes for test genes DNMT3B (Hs00171876_m1), PARP1(Hs00242302_m1) and β-Actin gene as a reference gene. Real time PCR amplification was performed with programming the computerized thermocyclers (ABI step one-Applied Biosystems) as follows: Initial denaturation at 95°C for 10 minutes, followed by 40 cycles of denaturation at 95°C for 15 seconds, and (Combined Annealing/Extension) at 65°C for 1 minute.
Data from the amplification plot was obtained and analyzed. Using the 2 -ΔΔCt method, the data are presented as the fold change in DNMT3B, PARP1 genes. Acute myeloid leukemia patients were categorized into high and low expressers according to the median mRNA expression of PARP1 9 and DNMT3B. 5
Statistical analysis
Data management and analysis were conducted using SPSS version 24. Numerical data were summarized as means with standard deviations or medians with ranges, depending on the distribution. Categorical data were presented as frequencies and percentages. Normality of numerical data was assessed using the Kolmogorov-Smirnov and Shapiro-Wilk tests. Group comparisons for categorical variables were made using the Chi-square or Fisher’s exact tests, as appropriate. The Kaplan-Meier method was employed to estimate EFS, DFS, and OS, while predictor and prognostic variables were analyzed in relation to survival using the log-rank test. All tests were 2-sided, with P-values < 0.05 considered statistically significant.
Results
Patients’ characteristics
The current study comprised 67 patients, out of them 43(64%) were male patients, and 24(36%) were females. The range of patients’ ages was (19-73) years with mean ± SD of 44 ± 14.15 years. Patients’ characteristic and initial laboratory findings were shown in Supplemental Table 1.
Expression levels of DNMT3B and PARP1 in BM of AML patients
PARP1 and DNMT3B mRNA transcript expressions were tested in all cases and control groups. DNMT3B expression was successfully detected in 64 cases while PARP1 detection was successful in 65 cases. DNMT3B expressions showed significant upregulation in AML cases [median: 11.78, range: 0.05-97.93] compared with the control group [median: 1.32, range: 0.09-9.36, P = .001]. Similarly, PARP1 expressions were significantly upregulated in the patients [median: 2.26, range: 0.05-7.33] compared with the control group [median: 0.65, range: 0.25-2.56, P = .036] (Table 1).
Expression levels of DNMT3B and PARP1 in BM of AML patients.

Association between the expression DNMT3B and PARP1 and patients’ characteristics
As regards DNMT3B, 32 (50%) of cases were above 11.77 (high expression group) and 32 (50%) were below or equal to 11.77 (low expression group), while for PARP1, 30 cases (46.2%) were above 2.26 (high expression group) and 35 (53.8%) were below or equal to 2.26 (low expression group). As regards the combined expression of PARP1 and DNMT3B, both were downregulated in 20 (32.3%) cases, both were upregulated in 18 (29%) cases, upregulated PARP1 and downregulated DNMT3B in 10 (16.1%) cases and 14 (22.6%) cases was upregulated DNMT3B and down regulated PARP1 in relation to control group.
Correlation between the expression DNMT3B and patients’ characteristics
Patients with high DNMT3B expression had higher TLC levels (P = .031), more blasts in the PB and BM (P = .004, P = .005, respectively) and more frequent FLT3-ITD (41.4% vs 11.5%, P = .013) than those with low expression. Also, high DNMT3B expression associated significantly with intermediate risk (IR) stratification according to ELN classification (93.1% vs 69.2% in the low expressers; P = .035), but DNMT3B expression did not significantly correlate with any other clinicolaboratory variable (Table 2).
Correlation between DNMT3B gene expression and patients’ characteristics.

Abbreviations: FR, favorable risk; IR, intermediate risk.
Correlation between PARP1 gene expression and patients’ characteristics
Patients overexpressing PARP1 displayed increased TLC levels (P = .024), more PB and BM blasts (P = .012, P = .007, respectively) and more frequent FLT3-ITD (50% vs 10.3%, P = .001) compared with those with under-expression. However, PARP1 levels showed no significant association with any of the remaining parameters (Table 3).
Correlation between PARP1 gene expression and patients’ characteristics.
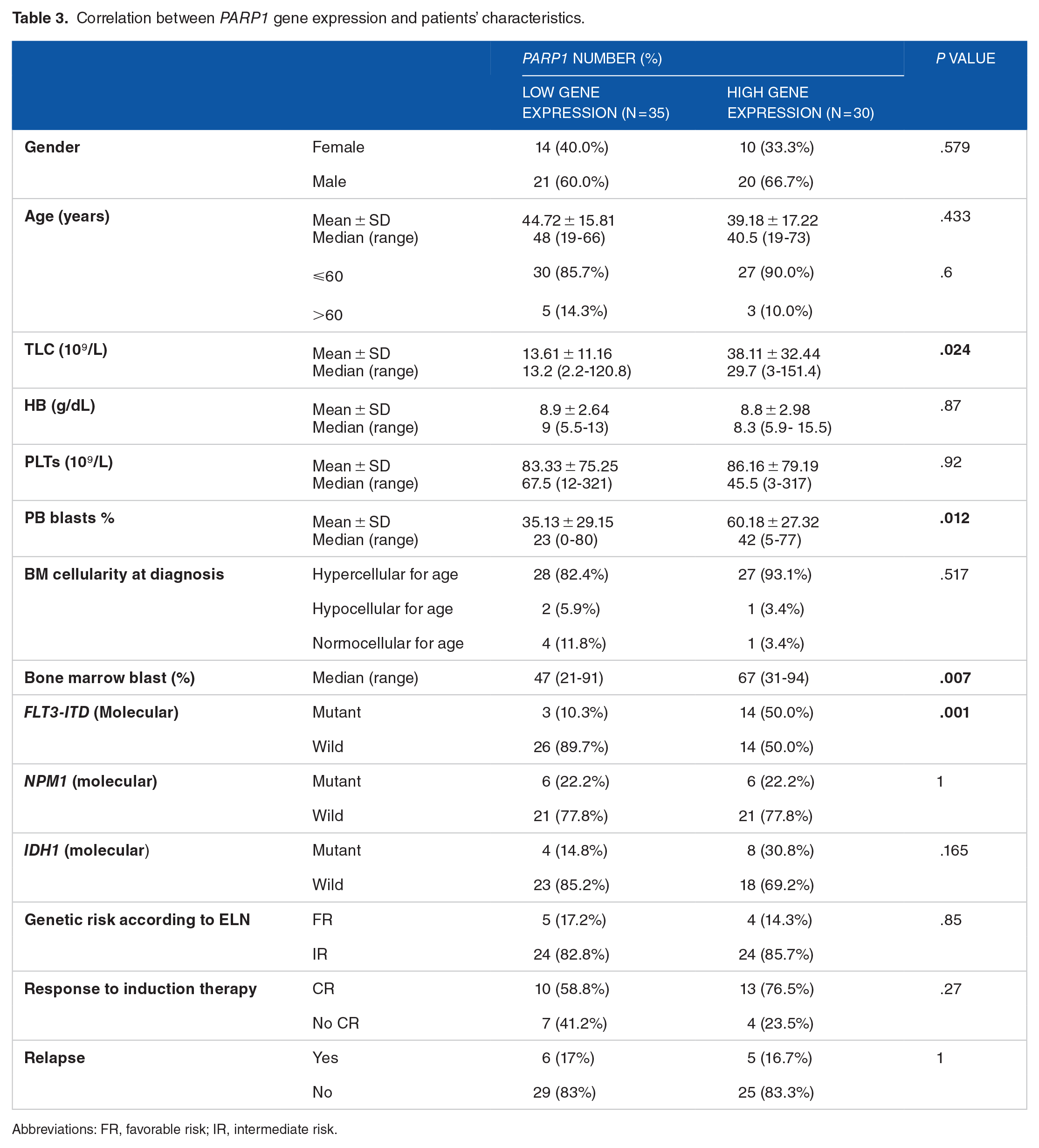
Abbreviations: FR, favorable risk; IR, intermediate risk.
Correlation of combined expression of both gene and patients’ characteristics
We found that cases with downregulation of both PARP1 and DNMT3B genes associated with lower TLC (P < .001), decreased PB blast count (0.01) and lower BM blast cell count (P = .037), compared with cases with upregulation of both genes and cases with upregulation of one gene and downregulation of the other. On the other hand, upregulation with of both PARP1, DNMT3B genes correlated with more prevalent FLT3-ITD (0.006) and higher frequency of ELN intermediate risk (P = .023) compared with downregulation of both genes and to upregulation of one gene and downregulation of the other, but no statistically significant association was found with the other clinicolaboratory variables (Table 4).
Correlation of combined expression of both PARP1 and DNMT3B and patients’ characteristics.

P value <.05 is considered statistically significant.
Cells that are sharing 2 letters are not statistically significant.
The relevant median value of the variable is used as a cutoff.
Impact of PARP1 and DNMT3B expression on CR
No statistically significant difference was found between high and low expression levels of either PARP1, DNMT3B genes, or the combined expression of both genes and CR (P = .27, P = .81, P = .76, respectively). Also, no significant association was found between the levels of expression of PARP1, DNMT3B, or the combined expression of both genes and relapse (P = 1, P = .96, P = 1, respectively).
Impact of DNMT3B and PARP1 expression on patients’ survival
The patients’ median OS was 1.18 months (range, 0.03-38.8 months). No significant difference could be detected between low or high expressers of either DNMT3B, or PARP1 in overall survival (OS), disease-free survival (DFS) or event-free survival (EFS). As regards DNMT3B, [median OS of the low expressers: 0.16 months vs 1.1 months for the high expressers (P = .281)], [the median DFS of the low expressers: 6.1 months vs 3.3 months for the high expressers (P = .53)], and [median EFS of the low expressers: 3 months vs 1 month for the high expressers (P = .21)] (Supplemental Figure 1).
Regarding PARP1, [median OS of the low expressers: 0.86 months vs 1.35 months for high PARP1 expressers (P = .42)], [median DFS of the low expressers: 6.8 months vs 6.1 months for the high expressers (P = .51)], and [the median DFS of the low expressers: 1.3 months vs 3 months for the high expressers (P = .36)] (Supplemental Figure 2).
In univariate analysis, better DFS associated significantly with lower blast percentage in the BM [median DFS: 13.1 months vs 4.4 months for the patients with higher blast percentage (P =
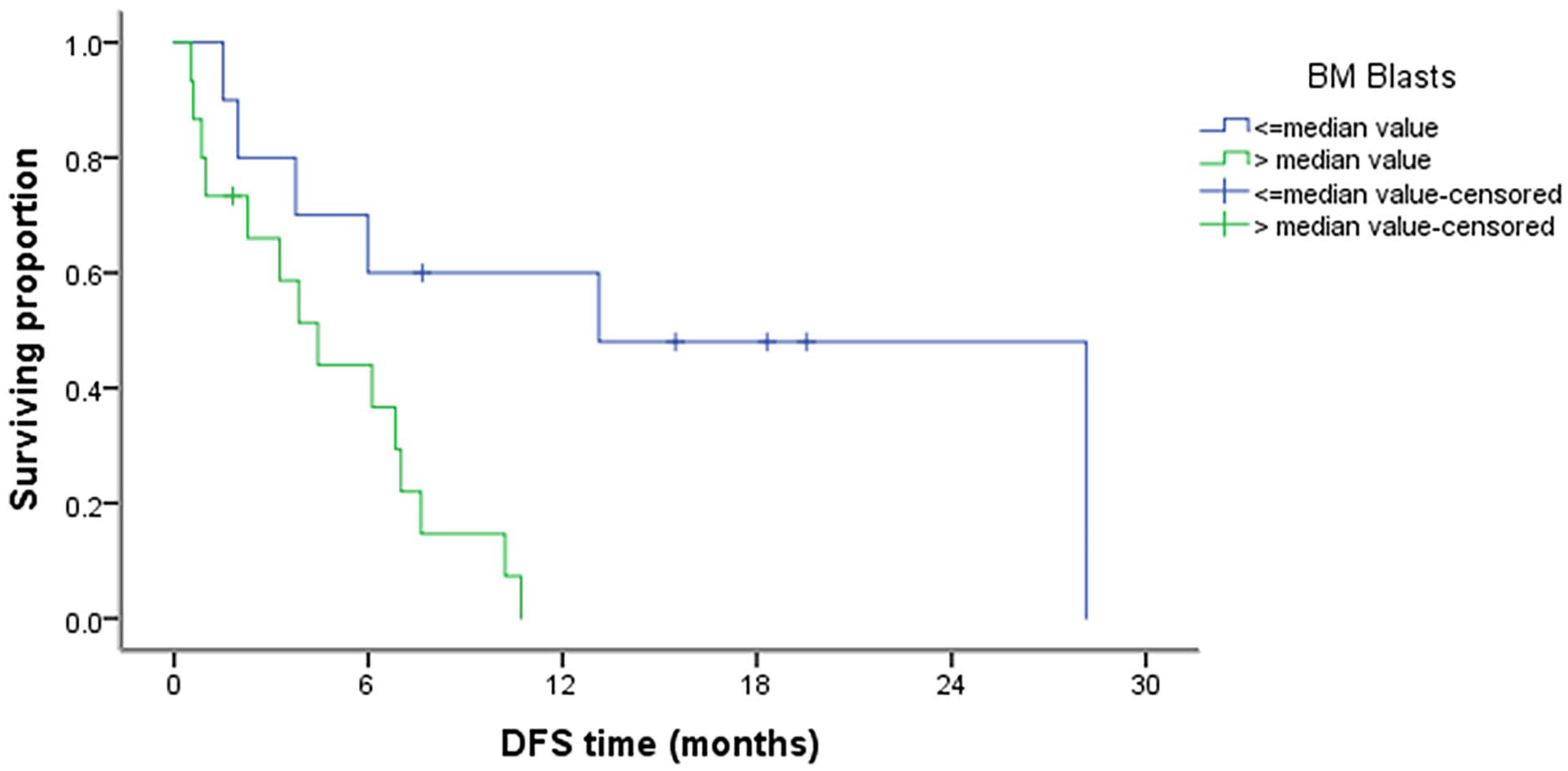
DFS in relation to blast % in the BM (P = .009).

EFS in relation to CD64 (P = .032).
Discussion
Despite recent advancements in refinement of predictive and prognostic markers, the mortality rate of AML has remained largely unchanged over the past 2 decades with an approximate median overall survival of 8.5 months, representing the fifth worst cancer type in terms of 5-year OS. This underscores the urgent need to identify novel biomarkers and molecular mechanisms driving leukemogenesis and AML progression to develop innovative therapeutic strategies that can improve patient outcomes. 11
It has been discovered that PARP1 is overexpressed in several human malignancies, including nasopharyngeal carcinoma, gastric cancer, hepatocellular carcinoma, breast cancer, and prostate cancer. In addition, the overall prognosis is inversely linked with its overexpression. It is believed that PARP1 uses a variety of molecular routes to carry out its carcinogenic function.12-15 While DNMT3A mutations are common in AML occurring in approximately 1/5 of the de novo patients and causing changes in DNA methylation, the incidence of DNMT3B mutation in AML is rare and the influence of DNMT3B expression on AML treatment and outcome is poorly defined and controversial. 5
The current study showed significant upregulation of DNMT3B expression in AML cases compared with the control group (P = .001), a significant correlation between DNMT3B overexpression and higher total leukocyte count (TLC) (P = .031), as well as higher percentages of bone marrow and PB blasts (P = .004, P = .005, respectively). Our observation aligns with previous research suggesting a potential involvement of DNMT3B in AML pathogenesis and a potential correlation between elevated DNMT3B expression and higher proliferation rate or disease progression and aggressiveness.16,17
Overexpression of DNMT3B in AML was also reported by Mizuno et al, who, in addition, showed that AML cells with methylated p15INAK4B had a tendency to express greater levels of DNMT3B. They concluded that, increased expression of DNMT3B might assist leukemogenesis via the induction of abnormal regional hyper methylation. 18 Likewise, a study on pediatric AML by Lamba et al, 19 reported that increased DNMT3B expression was linked to worse prognosis and higher genome-wide methylation burden (GWMB) that might be implicated in the development and hostility of the disease.
In contrast to our results, Zhang et al, 5 found that the expression of DNMT3B was reduced in AML patients and DNMT3B down regulation associated with higher TLC and lesser PB blasts. Others suggested that DNMT3B had a basic role in the pathogenesis of AML yet, may not behave as an oncogene based on observations from earlier studies that showed a tumor suppressive function for DNMT3B in inv(16)(p13;q22) AML and MLL-AF9 AML.20,21 On the contrary, DNMT3B was oncogenic in AML cell lines and mouse models; therefore, Wong et al, 8 proposed that the role played by DNMT3B in AML appeared to be subtype specific.
Also in this study, higher DNMT3B expression was significantly linked to the presence of FLT3-ITD (P = .013). However, no significant association between DNMT3B expression and NPM1 or IDH1 could be identified. It should be noted that cases harboring FLT3-ITD are stratified as intermediate risk (IR) patients in the recent ELN risk classification regardless of the allelic ratio or the presence of concurrent NPM1 mutation. 10 On stratifying our population according to the recent ELN classification, we noticed that DNMT3B overexpression significantly correlated with (IR) (P = .035), hence, we confirm DNMT3B expression levels may be a useful prognostic biomarker in AML with FLT3-ITD, and the incorporation of its quantification might be helpful in the genetic profile used for the categorization of patients into specific therapeutic strategies.
Our findings agree with Hayette et al 16 and Niederwieser et al, 17 who reported DNMT3B expression associated with FLT3ITD mutations through the association with HOXA9. The association of DNMT3B expression with the HOXA family of genes was also reported by Lamba et al, 19 in pediatric AML suggesting that epigenetic modifications of the DNMT3B locus could take a part in transcriptional deregulation that possibly helps the evolution of the disease. In contrast Zhang et al, 5 reported that AML with DNMT3B under- expression displayed decreased frequencies of IDH1 but DNMT3B levels did not significantly correlate with NPM1 or FLT-ITD.
As regards the prognostic value, we did not observe any correlation between DNMT3B expression and disease-free survival, event-free survival, or overall survival in our studied AML patients which is in agreement with Zhang et al, 5 who stated that DNMT3B gene expression may not be a significant prognostic biomarker and its potential role in AML remained poorly defined. On the contrary, overexpression of DNMT3B was reported as a significant independent poor prognostic indicator.16,17 The divergent outcomes observed between the results could plausibly be attributed to different population’ demographic features (young or older AML patients), dissimilarities in AML characteristics, different sample sizes, and methodologies employed. Furthermore, discrepant outcomes may have originated from the specific cutoff values used to define high or low genes expression. We suggest larger multicenter studies should be conducted for better evaluation and standardization.
The present study demonstrated significant upregulation in PARP1 expression levels in AML patients compared with the controls (P = .036), significant associations between increased PARP1 expression and higher TLC (P = .024), higher counts of blasts in the peripheral blood and bone marrow (P = .012, P = .007, respectively). Our results are concordant with previous studies 9,15,22,23 that showed overexpression of PARP1 in AML, particularly in the poor prognosis genetic sub-group and inhibition of PARP1 suppressed the proliferation of AML cells, brought about apoptosis in vitro, and enhanced the prognosis in mice. These results suggest a potential involvement of PARP1 in AML development.
Furthermore, our results revealed that elevated PARP1 expression significantly correlated with FLT3-ITD (P = .001), but, did not significantly associate with either NPM1 or IDH1. These findings are congruent with Li et al 9 and suggest that PARP1 expressions may hold potential prognostic value in AML. Interestingly, some research studies have provided valuable insights into the molecular mechanisms and possible associations of PARP1 in AML. Where, some studies have shown an increase in genomic instability in FLT3-ITD mutated AML through PARP1 associated with MYC expression. 4
In addition, a favorable correlation between PARP1 expression and myeloproliferative leukemia virus oncogene (MPL) was previously demonstrated and overexpression of both PARP1 and MPL associated with poor outcome suggesting that activation of MPL by PARP1 is another mechanism by which PARP1 affects AML prognosis. 15 Furthermore, Pashaiefar et al 23 found overexpression of PARP1 in AML patients with chromosomal translocations when compared with those without translocations, and in patients with 2 or more translocations when compared with those with an isolated translocation, suggesting that PARP1 is most likely associated with severity of genetic risk.
Moreover, analysis of concomitant DNMT3B and PARP1 expression revealed that cases with the downregulation of both genes displayed significantly reduced TLC, and blast count in peripheral blood and in BM, in comparison to those with upregulated DNMT3B and PARP1 genes. Conversely, the elevated levels of DNMT3B and PARP1 genes were correlated with a higher TLC (P < .001), and elevated blast count in the bone marrow (P = .037), and peripheral blood (P = .01).
To the best of our knowledge, our results are the first one to demonstrate a significant association between the combined upregulation of DNMT3B and PARP1 and the presence of FLT3-ITD mutations in AML patients. Notably, a higher proportion of cases exhibiting upregulated expression of both genes were found to harbor FLT3-ITD mutations (58.8%), as opposed to those with downregulated expression (6.3%) (P = .006). These observations imply a plausible association between the combined expression of DNMT3B and PARP1 and the occurrence of FLT3-ITD mutations in AML patients. In addition, upregulation of both PARP1 and DNMT3B genes associated with higher frequency of ELN intermediate risk (IR) (P = .023). However, a significant correlation between the combined expression of DNMT3B and PARP1 and the presence of NPM1 or IDH1 mutations could not be revealed.
On the same line, a previous study found the use of combination of drugs that traps PARP1 and inhibits DNMT promoted apoptosis of the lymphomatous and leukemic cells via various mechanisms including inhibition of cell productivity and expansion, and increase of reactive O2 molecules. 24
Limitations of this study
The main limitation of this study was small sample size (67) and the unavailability of involving patients from centers other than NCI, and we focused on gene expression levels without delving into the mutational status of DNMT3B and PARP1 in AML due to limited budget. Multi-center study including large number of patients is necessary to approve our results.
Conclusions
Our results confirm results from previous studies and provide valuable insights into the significance of DNMT3B and PARP1 gene expression in AML, suggesting a potential role in disease progression and aggressiveness and supports their interplay in AML. Our study’s findings confirm the potential of DNMT3B and PARP1 as predictive markers for identifying AML patients with FLT3-ITD and highlight their involvement in AML pathogenesis.
Our findings might have significant implications in identifying high-risk AML patients who may benefit from targeted treatments or more intensive therapies. By assessing the expression levels of DNMT3B and PARP1, clinicians can obtain additional prognostic indicators to guide treatment decisions and potentially improve patient outcomes.
Supplemental Material
sj-docx-1-onc-10.1177_11795549241295649 – Supplemental material for Roles of DNMT3B and PARP1 Genes Expression in Cytogenetically Normal Acute Myeloid Leukemia
Supplemental material, sj-docx-1-onc-10.1177_11795549241295649 for Roles of DNMT3B and PARP1 Genes Expression in Cytogenetically Normal Acute Myeloid Leukemia by Hager A Mahmoud, Shahira KA Botros, Abdelhamid Mohamed Fouad, Mahmoud M Kamel and Rania S Abdel Aziz in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
The authors give special thanks to our patients and control who paved the way for us to carry out and finish our project.
Author contributions
HAM and RSAA designed the study and wrote the original drafts and coordinated specimen collection and transport. SKAB and RSAA managed kits purchasing tasks and supervised laboratory analysis workflow. HAM, MMK, and RSAA performed the laboratory analysis. AMF ensured appropriate patient selection and supervised data collection. All authors have read and approved the final manuscript.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
Approval (no.2306-403-026) of the current study was obtained from review board (IRB) of NCI, Cairo University. All the measures were performed in accordance with Helsinki guidelines.
Availability of data and materials
All data and materials are available and can be submitted when needed. Corresponding author is the responsible person who should be contacted if someone wants to request the data from this study.
Informed consent
The study objectives were explained then and written informed consents were got from all contributors (patient and control) before bone marrow sampling. Confidentiality of patient data was guaranteed.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
