Abstract
CenA is an endoglucanase secreted by the Gram-positive cellulolytic bacterium, Cellulomonas fimi, to the environment as a glycosylated protein. The role of glycosylation in CenA is unclear. However, it seems not crucial for functional activity and secretion since the unglycosylated counterpart, recombinant CenA (rCenA), is both bioactive and secretable in Escherichia coli. Using a systematic screening approach, we have demonstrated that rCenA is subjected to spontaneous cleavages (SC) in both the cytoplasm and culture medium of E. coli, under the influence of different environmental factors. The cleavages were found to occur in both the cellulose-binding (CellBD) and catalytic domains, with a notably higher occurring rate detected in the former than the latter. In CellBD, the cleavages were shown to occur close to potential N-linked glycosylation sites, suggesting that these sites might serve as ‘attributive tags’ for differentiating rCenA from endogenous proteins and the points of initiation of SC. It is hypothesized that glycosylation plays a crucial role in protecting CenA from SC when interacting with cellulose in the environment. Subsequent to hydrolysis, SC would ensure the dissociation of CenA from the enzyme-substrate complex. Thus, our findings may help elucidate the mechanisms of protein turnover and enzymatic cellulolysis.
Keywords
Introduction
Since the discovery of cellulolytic microbes during World War II, 1 hundreds of examples of all 3 types of cellulases: including endoglucanases (Eng) (EC 3.2.1.4), exoglucanases (Exg) (EC 3.2.1.91) and β-glucosidases or cellobiases (EC 3.2.1.21), which act cooperatively in the hydrolysis of cellulose substrates, have been characterized.2,3 One of the best-studied cellulase producers is the Gram-positive soil bacterium, Cellulomonas fimi. Since the report of the first cellulase gene cloned from C. fimi, 4 a good collection of gene determinants encoding both Eng and Exg have been cloned from this bacterium. 5 Among the characterized cellulases of C. fimi, the first Eng, designated CenA, 6 has been expressed using various host systems.6-8 The availability of recombinant CenA (rCenA), for example, from E. coli, does not only facilitate its application to the studies of enzymatic hydrolysis, but also comparative performance between itself and its native counterpart isolated from C. fimi. Moreover, employing bioinformatics tools for structural and functional characterization of proteins,9,10 rCenA and other recombinant cellulases have been analyzed to be composed of structural/functional motifs including (i) a substrate/cellulose binding domain, (ii) a catalytic domain and (iii) a linker sequence that is rich in hydroxyl amino acids.9,11,12
Proteins expressed in E. coli are devoid of post-translational modifications, and therefore, rCenA is non-glycosylated, despite the presence of glycosylation in native CenA.6,12,13 Consistent with previous observations for other non-glycosylated recombinant cellulases, rCenA expressed in E. coli and other host systems is enzymatically active and secretable.8,14 Therefore, glycosylation appears to play a role in maintaining the structural integrity of cellulases instead.13,15,16 However, there has been little concrete evidence to support this idea.
Convenient and specific assessment assays have been developed for both qualitative and quantitative assays of cellulase activities. For example, qualification and quantification of Eng activity is readily achieved using the carboxymethylcellulose (CMC) plate assay and the dinitrosalcylic acid detection method, 6 respectively, both of which employ CMC as the substrate. Despite the convenience of these protocols and the fact that they may be exploited to determine the potency of an enzyme, the outcomes are non-dynamic and represent only collective results. Thus, some subtle activities, for example, structural changes that occur continuously in proteins, may easily be overlooked. In this communication, employing a systematic approach with immunological, substrate binding and structural analysis, we report that rCenA undergoes spontaneous cleavages (SC), which proceed continuously, with efficiencies dependent upon local environmental conditions. Using the cellulose binding domain (CellBD) of rCenA as a model, it was demonstrated that CellBD was subjected to attack by SC. Since these cleavages were identified to be located in close proximity to N-linked glycosylation sites, it was postulated that the Asn residue of the Asn-Gly dipeptide on these sites underwent deamidation,17-19 thus leading to chain breakage. Our findings may help elucidate the mechanisms underlying the turnover of heterologous proteins in E. coli and the dissociation of CenA from the enzyme-substrate complex.
Materials and Methods
Bacterial strain and chemicals
E. coli JM101 employed as the host for DNA manipulations and recombinant protein expression was previously described. 20 DNA oligos were purchased from Invitrogen (Carlsbad, CA, USA). The Phusion PCR kit and restriction enzymes were purchased from NEB (Ipswich, MA, USA). Chemicals were purchased from Sigma-Aldrich Corporation (St. Louis, MO, USA) unless otherwise specified. Antibodies against CenA were raised in New Zealand white rabbits.
Construction of pFC for CenA expression in E. coli
To facilitate CenA expression in E. coli, construct pFC (Figure 1a) was engineered through genetic modification of an in-house Bacillus subtilis expression plasmid, pM2VegCenA 8 employing 2 rounds of PCR. The nucleotide A at the −11 position of the veg promoter in pM2VegCenA was first changed to a C, forming the vegC promoter. Subsequently, the spa leader sequence was precisely fused at the 3ʹ terminus of the vegC promoter-lac operator region of pM2VegCenA to form intermediate product I, with the help of pM2VegCenA as the template and a pair of PCR primers: −11C (5ʹ-CCGAATTCTA,ATTTAAATTT,TATTTGACAA,AAATGGGCTC,GTGTTGTCCA,A-3ʹ) and Spa-b (5ʹ-AGCATTTGCA,GCAGGTGTTA,CGCCACCAGA,TATAAGTAAT,GTACC-3ʹ). On the other hand, employing plasmid pEE23 7 as the template and another pair of primers: spa-cenA (5ʹ-TAACACCTGC,TGCAAATGCT,GCTCCCGGCT,GCCGCGTCGA,CTACGCCGT C-3ʹ) and cenA-BamHI (5ʹ-GCTGGATCCG,CTGGCCTGCG,GTGTAGGTCC,AGTCGAGCTT,CCACG-3ʹ), PCR was undertaken to obtain intermediate product II. Products I and II were joined by overlap extension PCR (OE-PCR). The extended fragment was then restricted with Eco RI and Bam HI, followed by its insertion into vector pM2VegCenA treated with the same 2 enzymes to attain construct pFC (Figure 1a).
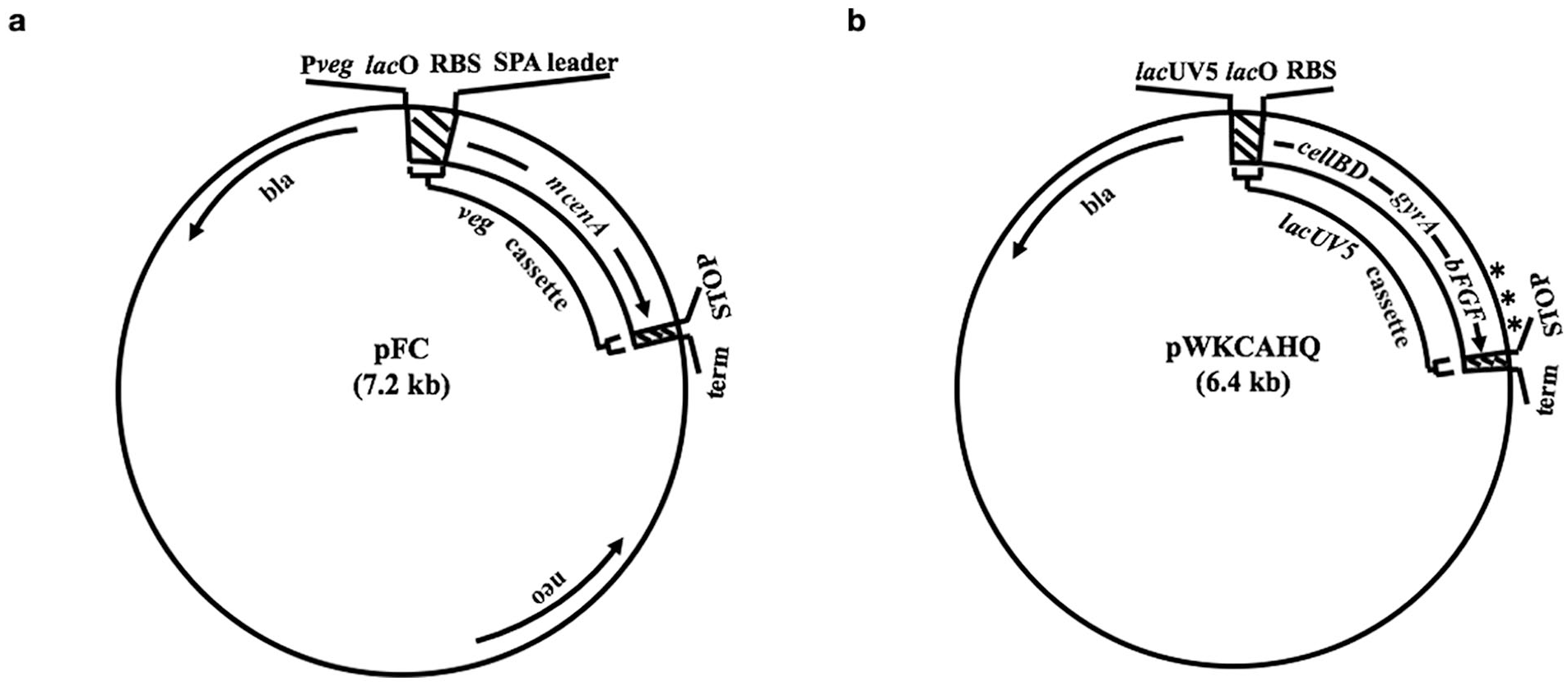
Schematic representation of DNA constructs: (a) pFC and (b) pWKCAHQ. Plasmids pFC and pWKCAHQ were employed to express recombinant CenA and CellBD-GyrA-bFGF precursor, respectively, in E. coli. Components present in (a) pFC include: mCenA = coding sequence for mature CenA (starting from Ala1; see Figure 4); bla = bla gene conferring resistance to ampicillin; genetic elements of the Veg cassette [hatched bars including vegC promoter (Pveg); lac operator (lacO); consensus ribosome binding site (RBS); coding sequence for Staphylococcal protein A leader peptide (SPA leader); stop codons in all 3 reading frames (STOP); transcription terminator (term)]. Components in (b) pWKCAHQ include: cellBD-gyrA-bFGF = coding sequence for the CellBD-GyrA-bFGF precursor; genetic elements in the lacUV5 cassette [hatched bars including lacUV5 promoter; lacO; RBS; STOP and term; *denoting epitopes on bFGF reacting with anti-bFGF antibodies]. The arrows indicate the directions of gene expression.
Cultivation of E. coli (pFC) transformants
E. coli (pFC) transformants were cultivated in 100 ml of either 2× YT, LB or MMBL culture medium prepared as described previously. 21 All the cultures were cultivated at different temperatures supplemented with 70 μg mL−1 of ampicillin and shaken at 250 rpm until an OD550 value reached 8.0. After induction with a final concentration of 0.1 mM IPTG for 10 hours, the cultures were fractionated into culture supernatant (SN) and lysate (Ly) samples. The latter was prepared using chemical treatments as described previously. 21
Binding of SN and Ly proteins to Avicel
SN and Ly proteins were mixed with pre-washed, swollen microcrystalline Avicel powder (Cas No.9004-34-6, Sigma-Aldrich) in a Falcon tube in a 2:1 volume ratio. A final concentration of 1 mM PMSF was then added to the tube, followed by mixing the content on a rocker shaker at 4°C for 1 hour. The unbound material was reserved for Western blot analysis. The Avicel matrix was then rinsed with 5 column volumes of washing buffer (20 mM Tris-HCl, 0.5 M NaCl, pH 8.0), and the bound content was eluted using 1 column volume of 8 M urea.
Western blot and zymographic analysis
SN and Ly proteins were analyzed by Tricine-SDS-PAGE and Western blotting as described previously 22 with the following modifications. The proteins were resolved on a 10% (w/v) gel. To study whether the protein bands were endoglucanase-positive, a zymographic technique 23 was conducted as follows with minor modifications. The agar replica was cast between 2 glass plates with 2% (w/v) agar and 1% (w/v) CMC dissolved in 50 mM disodium hydrogen phosphate, pH 7.0. The separated protein bands were then transferred to the CMC agar by capillary transfer as described. 24 CMC hydrolysis was allowed to proceed at 37°C for 2 hours. Visualization of CMCase activity by Congo red staining was performed as described previously. 23
Engineering of constructs pWKCAHQ, pWKC1A and pWKHQ
The construction of the plasmids pWKCAHQ (Figure 1b), pWKC1A and pWKHQ were described in our previous study. 25 In short, a couple of nucleotide mutations were introduced to the 2 fusion junctions in pWKCAHQ, with the first one located at the junction between the coding sequence for the cellulose binding domain (CellBD) and the gyrA gene, and the other one at the junction between the gyrA and bfgf gene sequences. Correspondingly, the 2 nucleotide mutations resulted in a pair of amino acid substitutions: Cys1Ala at the junction between CellBD and the GyrA intein (GyrA) and His197Gln at the junction between GyrA and bFGF. The 2 substitutions were shown to be effective in preventing auto-catalytic or spontaneous cleavage (SC) from occurring at the 2 mentioned junctions in the precursor fusion protein: CellBD-GyrA-bFGF (CGFP). Expression of pWKC1A, a variant of pWKCAHQ, would produce a CGFP derivative containing a single amino acid substitution, Cys1Ala, at the junction between CellBD and GyrA. On the other hand, expression of another variant of pWKCAHQ, pWKHQ, would yield a new CGFP derivative containing another amino acid substitution, His197Gln, at the junction between GyrA and bFGF (Figure 2b).
Analysis of CGFP and its derivatives
The conditions for the cultivation of E. coli transformants containing plasmids pWKCAHQ (Figure 1b), pWKC1A and pWKHQ were described previously. 25 The techniques of mechanical cell disruption of the cells, Western blotting and Avicel binding were described. 25 The 39 kDa deletant expressed by pWKCAHQ was retrieved from a Coomassie Brilliant Blue stained gel 21 and subjected to liquid chromatography-tandem mass spectrometry analysis (Instrumental Analysis Center of Shenzhen University, Lihu Campus).
Results
The complex composition of recombinant CenA products expressed in E. coli
Expression of pFC (Figure 1a) in E. coli was found to result in a SPA-mature CenA (mCenA) fusion precursor in the cytoplasm. However, analysis of lysate samples prepared from the culture by Western blotting revealed a complex pattern comprising multiple bands, including that of the precursor and many smaller bands (Figure 3ai). The intensity of the bands was amplifiable by modulating the growth conditions. For example, when the culture was grown at a higher temperature in the presence of succinic acid, which was previously shown to cause protein instability,26,27 interestingly, the band pattern of the proteins was revealed to be markedly enhanced (Figure 3ai).
In testing the ability of the products to bind to microcrystalline cellulose, Avicel, quite unexpectedly, despite being appreciably larger, the 36 kDa deletant was shown to bind less efficiently than a smaller 20 kDa counterpart (Figure 3ai). The results suggested that the former contained either a smaller cellulose-binding domain (CellBD) (Figure 2a) or a less critical portion for binding than did the latter.
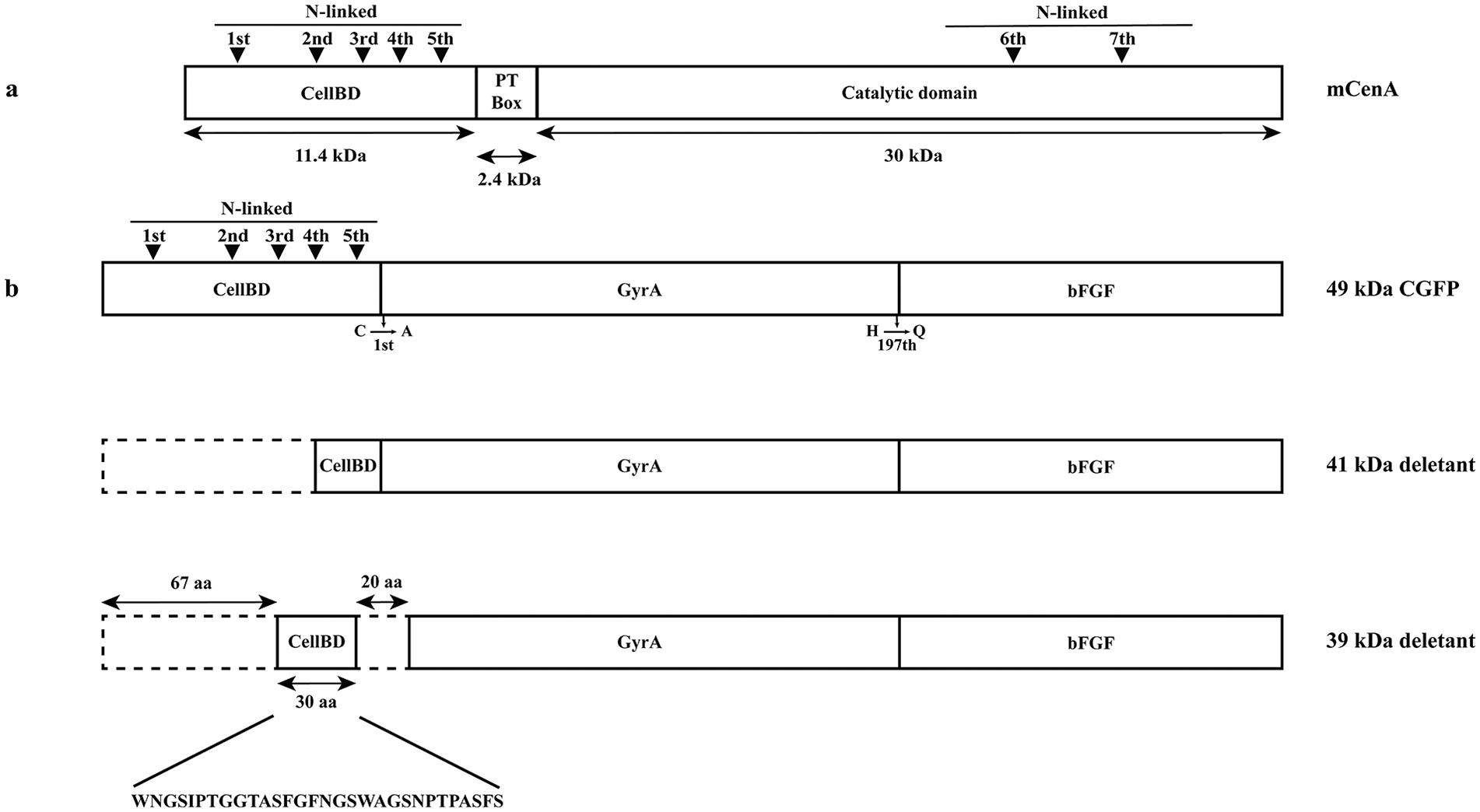
Schematic representation of mature CenA and various fusion proteins resulting from the expression of plasmid pWKCAHQ. The bar diagrams show the components of different recombinant proteins. (a) Mature CenA (mCenA) containing an N-terminal cellulose binding domain (CellBD) and a C-terminal catalytic domain connected by a Pro-Thr repeat (PT box). The ▼ represent the potential N-linked glycosylation sites identified within CellBD and the catalytic domain (see Figure 4). (b) The 3 fusion proteins resulting from the expression of plasmid pWKCAHQ (Figure 1b): 49 kDa CellBD-GyrA-bFGF precursor (CGFP); 41 kDa deletant; 39 kDa deletant. In engineering CGFP, a CellBD deletant lacking the last 5 amino acid residues (VPTTS; Figure 4) was fused with GyrA. The 41 kDa deletant was predicted to result from spontaneous cleavage (SC) occurring proximal to the fourth N-linked glycosylation site in CellBD. The 39 kDa deletant was revealed by sequencing to be generated by SC occurring close to the third and fifth N-linked glycosylation sites in CellBD. After SC, a 30 residue remnant CellBD segment (sequence shown below; see Figure 4) was spliced with a GyrA deletant (missing the N-terminal 11 residues) to form the 39 kDa deletant. The Cys to Ala and His to Gln substitutions engineered at the CellBD-GyrA and GyrA-bFGF junctions, respectively, were shown to offer protection to the junctions against SC. 25 Dotted boxes show the deleted sequences.
To examine whether the resolved bands were active on CMC substrate, a qualitative zymographic assay involving the use of a CMC overlay 23 was undertaken. The results revealed that the precursor and the 36 kDa derivative were highly active (Figure 2aii), whereas the smaller variants were inactive.
The results suggested the interpretation that the 36 kDa derivative likely possessed a complete catalytic domain (CD) of mCenA (Figure 2a) due to the previous observation that a small deletion of the C-terminus of CD resulted in the abolition of positive detection of Eng activity. 6 Thus, the Avicel binding and zymographic results together supported the conclusion that the 36 kDa derivative possessed a full-length 30 kDa CD, the 2.4 kDa Pro-Thr box with a 3.6 kDa remnant CellBD fused at its N-terminus (Figure 2a). On the other hand, despite being enzymatically inactive, the ability of the 20 kDa variant to bind to Avicel supported the conclusion that a major or an entire portion of it was derived from CellBD (Figure 2a).
Analysis of extracellular mCenA products
In E. coli (pFC), the SPA signal peptide worked well to mediate the excretion of mCenA to the culture supernatant (SN), in which background proteins and proteolysis were expected to be highly reduced. Despite the much ‘cleaner’ SN background where proteolysis was highly lessened, interestingly, the band patterns of mCenA proteins prepared from SN (Figure 3bi) and the cytoplasm (Figure 3ai) were shown to be highly similar.
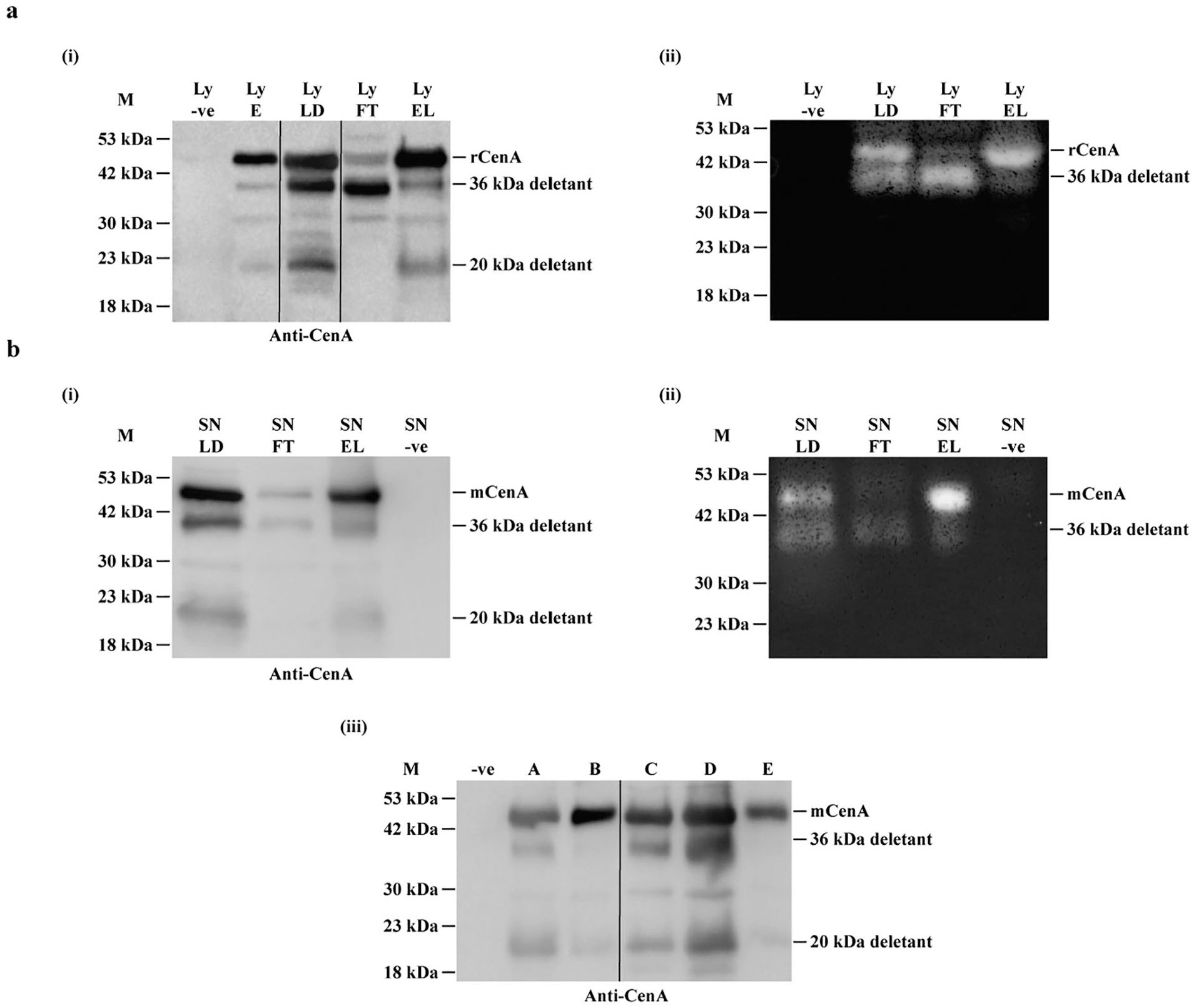
Western blotting and zymographic analysis of CenA expressed in E. coli (pFC) transformants. (a) Analysis of cell lysate samples: (i) Western blot results. Lanes: Ly –ve: negative control, prepared from a clone lacking CenA expression; Ly E: transformants grown in MMBL at 34°C; Ly LD: transformants grown in MMBL supplemented with 0.1 M succinic acid at 37°C; Ly FT: unbound proteins collected after Ly LD sample was used to bind to Avicel; Lane Ly EL: bound proteins of Ly LD sample eluted from the Avicel column. Each lane was loaded with a volume equivalent to 5 μl of cell culture. Ly LD, a re-arranged lane, is denoted by using 2 lines to separate it. (ii) Zymographic analysis of the samples, except Ly E, of panel (i). Each lane was loaded with a volume equivalent to 20 μl of culture for the assay. The presence of a halo indicated that the corresponding protein band on the gel of panel (i) was CMCase-positive, thus enzymatically active. (b) Analysis of culture supernatant samples, of which the cells were grown in MMBL supplemented with 0.1 M succinic acid at 37°C. (i) Western blot results. Lanes: SN LD: analysis conducted prior to Avicel binding; SN FT: unbound proteins collected after SN LD sample was used to bind to Avicel; SN EL: bound proteins of SN LD sample eluted from the Avicel column; SN –ve: negative control. Each lane was loaded with 20 μl of cell culture. (ii) Zymographic analysis of the samples of panel (i). Each lane was loaded with 40 μl of cell culture. (iii) Western blot results of culture supernatant samples of which the cells were cultivated under different growth conditions. Lanes: –ve: negative control; A: 2 × YT, 34°C; B: LB broth, 34°C; C: MMBL + 0.1 M succinic acid (pH 7.0), 34°C; D: MMBL + 0.1 M succinic acid (pH 7.0), 37°C; E: MMBL, 34°C. Each lane was loaded with 20 μl of the cell culture. Re-arranged lanes, A and B, are separated from other lanes. M stands for protein markers.
In addition, the SN and cytoplasmic proteins were shown to share the same propensity of affinity to Avicel. All these results supported the conclusion that mCenA was susceptible to the same mode of non-enzymatic attack in both the cytoplasm and SN, leading to auto-catalytic or spontaneous cleavages (SC) of mCenA. Different intensities of the protein bands reflected that the peptide bonds in mCenA were susceptible to unequal rates/frequencies of cleavage, which were affected by changes in the environmental factors.
The occurrence of SC in CellBD
The ability of the 36 kDa and 20 kDa deletants to bind to Avicel (Figure 3) and the fact that the N-terminal portion of the 36 kDa deletant stretched out to CellBD supported the interpretation that CellBD formed a considerable part of them (Figure 2a). Thus, we employed CellBD, which is around one-quarter the size of mCenA (Figure 2a), as a model to help define the locations of SC. To investigate whether SC would take place at the junction fused between intein GyrA (GyrA) and basic fibroblast growth factor (bFGF), construct pWKCAHQ (Figure 1b) was previously engineered. 25 However, in engineering pWKCAHQ, a coding sequence for a CellBD deletant lacking the last 5 amino acids (Figure 4) was employed to fuse with the gyrA gene (Figures 1b and 2b). The 49 kDa CellBD-GyrA-bFGF precursor (CGFP) encoded by pWKCAHQ was shown to bind to Avicel efficiently (Figure 5b), and bFGF could then be released from the bound CGFP using various conditions. 25

Amino acid sequence of mature CenA (mCenA). The 7 potential N-linked glycosylation sites of mCenA (UniProtKB/Swiss-Prot accession number: P07984), Asn-X-Ser/Thr, 5 being identified in CellBD and 2 in the catalytic domain, are boxed in red. The PT-box (112-134 residues) is underlined, whereas CellBD (1-111 residues) and the catalytic domain (135-418 residues) are located ahead of and after the PT-box, respectively. The 30 residue remnant CellBD segment of the 39 kDa deletant denoted in Figure 3b is highlighted in grey.
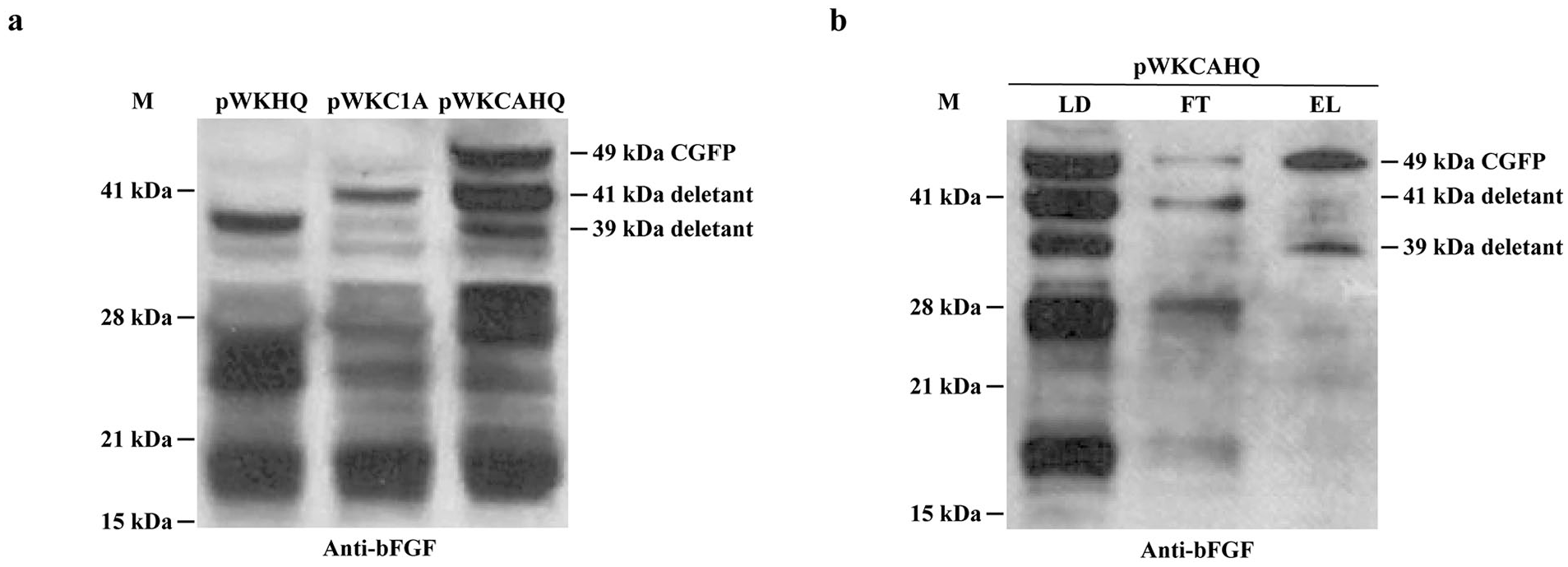
Western blot analysis of CGFP and its variants using bFGF as a probe. (a) Reactivities of cell lysate samples prepared from cell cultures containing the 3 expression constructs: pWKHQ, pWKC1A and pWKCAHQ with anti-bFGF antibodies. Each lane was loaded with a volume equivalent to 5 μl of cell culture sample. (b) Avicel binding results of cell lysate prepared from transformants containing plasmid pWKCAHQ. Lanes: LD: total proteins; FT: unbound proteins collected after LD sample was used to bind to Avicel; EL: bound proteins of LD sample eluted from the Avicel column. The 3 products: 49 kDa precursor (CGFP), 41 kDa deletant and 39 kDa deletant detected in the various preparations are indicated. M stands for protein markers.
On the other hand, if deletions occurred in CellBD of CGFP, they would be detectable by Western blot analysis employing bFGF as the probe (Figure 1b). Close examination of the 2 deletants of CGFP derived from the expression of pWKCAHQ (Figure 2b) and a 41 kDa product from a variant of pWKCAHQ, pWKC1A (with an A nucleotide mutated to C at the gyrA-bfgf junction), 25 the locations where SC took place were revealed. Firstly, expression of pWKCAHQ resulted in not only CGFP, but also a 41 kDa deletant and a 39 kDa deletant (Figure 5a). Interestingly, the 41 kDa deletant was found to bind to Avicel much less efficiently than the 39 kDa counterpart (Figure 5b). Sequencing results of the 39 kDa deletant revealed that both CellBD and GyrA were subjected to deletions, followed by the splicing of a 30 residue remnant CellBD (extending from Trp68 to Ser97) (Figure 4) with a GyrA deletant missing the N-terminal 11 residues (Figure 2b). On the other hand, despite a bigger size, the poor binding performance of the 41 kDa deletant to Avicel suggested that it contained a full-length GyrA-bFGF fusion, but only a 25 to 30 residue CellBD deletant that did not bind well to Avicel.
Secondly, the 41 kDa product (Figure 5a) detected from the expression of the variant construct pWKC1A 25 resulted likely from SC, which appeared to originate from the same point of attack as that for the development of the 41 kDa deletant encoded by pWKCAHQ described above (Figure 5a). Thirdly, the 39 kDa product (Figure 5a) encoded by another variant of pWKCAHQ, pWKHQ (with a GC doublet mutated to TG at the cellBD-gyrA junction) 25 resulted apparently from a normal event of auto-catalytic cleavage occurring at the junction between a recombinant protein, CellBD and an intein, GyrA, 25 thereby leading to the expected separation between CellBD and GyrA (Figure 2b).
Target sites for SC: N-linked glycosylation sites
Close examination of the mCenA sequence revealed the presence of 7 potential N-linked glycosylation sites: Asn-Xaa-Ser/Thr (Figure 4), where Xaa could be any amino acid except Pro. 28 Interestingly, 5 of these sites are located in the tiny CellBD (only one-quarter the size of mCenA), whereas the remaining 2 are found in the terminal portion of CD (Figure 4). More intriguingly, among the 7 sites, 5 of them possess Gly as the middle residue (Figure 4). Our findings described above support the hypothesis that the N-linked glycosylation sites, in particular the Asn-Gly-Ser/Thr tripeptide, function both as an ‘attributive tag’ and an initiation point for SC to occur (see Discussion). The analysis of the 36 kDa derivative resulting from the expression of pFC (Figure 3), and the 39 and 41 kDa deletants resulting from the expression of pWKCAHQ (Figure 5) revealed that they all contained a CellBD deletant where the point of cleavage was shown to occur in close proximity to an Asn-Gly-Ser/Thr sequence (Figures 2b and 4). As a matter of fact, Asn residues in a polypeptide have been shown to be prone to attack by deamidation, of which the onset is highly influenced by both the protein structure and environmental factors.17-19
Important role played by N-linked glycosylation in mCenA
Although native CenA is glycosylated,6,12,13 mCenA is not. It is envisaged that the N-linked glycosylation sites on mCenA, being a heterologous product in E. coli, serve as the points of differentiation so that foreign and endogenous proteins are ‘distinguishable’ under the influence of different environmental conditions and protein structural factors. SC is postulated to result from initially deamidation of Asn of the Asn-Gly dipeptide located in an N-linked glycosylation site (see Discussion). This intrinsic mechanism offers an efficient and cost-effective approach for the cells to eliminate dispensable heterologous proteins.
Despite its unimportance in mCenA, N-linked glycosylation presumably plays a crucial role in stabilizing and protecting a native CenA molecule, which is secreted to the unfriendly natural environment to meet up with and bind onto a cellulose substrate (Figure 6). Once the mission of hydrolysis is fulfilled by CD, the CenA molecule is required to be dissociated from the enzyme-substrate complex so that another cellulase molecule can take its turn. The most cost-efficient manner to fulfill the mission of dissociation is through SC. Although this process will take place in both CellBD and CD, nature’s design is so perfectly done that CellBD, being crucial for CenA to adsorb onto cellulose, must also be responsible for its detachment from the substrate. To ensure the dissociation, it is clear now why CellBD possesses a majority of 5, of the 7 N-linked glycosylation sites present in CenA (Figure 4).
Discussion
Western blot analysis revealed that rCenA products retrieved from the cytoplasm and growth medium of E. coli (pFC) transformants shared a similar band pattern (Figure 3). The results argued for the speculation that the smaller derivatives resulted from spontaneous cleavages (SC) rather than proteolytic processing of rCenA, based on the fact that proteolysis in the culture medium is expected to be uncommon. The cleavages of rCenA were sensitive to changes in environmental conditions, for example, temperature, pH and chemical composition. Succinic acid, which has a high solvent dielectric constant of 80, 29 is postulated to be able to increase the rate of deamidation of an Asn residue in a polypeptide, thus promoting the deprotonation of the peptide bond nitrogen anion, followed by the formation of initially a tetrahedral intermediate and finally a succinimide product.19,30,31 Since deamination is capable of causing protein unfolding and spontaneous degradation of proteins,17,18,30-32 it was speculated that succinic acid exerted a positive impact on enhancing the prevalence of SC, thus resulting in the formation of more intense bands instead of pristinely cleaved bands of rCenA (Figure 3).
Avicel binding and zymographic screening provided a facile analysis to help track down the locations where SC occurred in mature rCenA (mCenA). The findings that (i) the 36 kDa derivative of mCenA possessed both the cellulose binding domain (CellBD) and the catalytic domain (CD) (Figures 2 and 3), and (ii) the presence of CellBD in the 20 kDa variant since it bound well to Avicel (Figure 3), supported the conclusion that CellBD is an inevitable target of SC and would be suitably employed as a model for further study.
A fusion formed among CellBD, the GyrA intein (GyrA) and bFGF (Figure 2b) provided a feasible means for defining the locations of SC in CellBD, with bFGF serving as the probe. It was interesting to note that even single-residue mutations engineered at the CellBD-GyrA and GyrA-bFGF junctions (Figure 2b) could strongly affect the efficiency of SC occurring at the 2 junctions. 25 When the amino acid substitutions, Cys1Ala and His197Gln, were introduced to the 2 junctions, CellBD-GyrA and GyrA-bFGF, respectively, of the CellBD-GyrA-bFGF precursor (CGFP), 25 the mutations were able to offer protection to the 2 junctions from getting attacked by SC during expression. As a result, the 49 kDa full-length precursor, CGFP, was identified (Figure 5a). However, quite unexpectedly, 2 deletants where CellBD was shown to be attacked by SC were co-retrieved with CGFP (Figure 5a). The larger 41 kDa derivative was predicted to result from SC occurring close to the fourth N-linked glycosylation site (Figures 2b and 4). The 28 to 30 residue remnant CellBD polypeptide remaining attached to GyrA appeared insufficient to enable this derivative to bind well to Avicel (Figure 5b). However, the smaller 39 kDa deletant, which was confirmed to possess a CellBD segment comprising 30 residues spanning from Trp68 to Ser97 (Figures 2b and 4), despite being smaller, was shown to bind much better than the 41 kDa derivative to Avicel (Figure 5b). Interestingly, Trp68 and Ser97 were also identified to be close to 2 N-linked glycosylation sites, which were the third and fifth sites instead (Figure 4). In accord with these findings, the 41 kDa deletant encoded by the variant, pWKC1A (Figure 5a), was generated presumably in the same manner as the 41 kDa derivative of CGFP described above.
Recombinant CenA has been characterized since the mid-1980s. 6 However, attempts to crystallize its structure for in-depth analysis have been unsuccessful. 33 It had been commented that CellBD was ‘extremely labile’, 12 and the instability of CellBD reported above could be a significant reason accounting for the difficulty in crystallizing rCenA. Despite strenuous attempts, including the employment of specific proteases11,12 and site-specific mutagenesis 34 to study CellBD, the mystery of its high lability to degradation has been long kept. Nevertheless, it was reported that a CellBD deletant lacking the N-terminal 64 residues was still capable of binding to cellulose. 35 After integrating our research results into these previous findings, a much clearer picture of CellBD emerges and supports the following interpretations. First, CellBD is highly susceptible to breakage by SC, and the target sites are presumably the N-link glycosylation sites (Figures 2 and 4). Second, not all target sites will experience the same attack rate, which depends on structural conformation of the site concerned and the environmental conditions. Third, the C-terminal portion of CellBD, in particular the last 3 (third–fifth) N-linked glycosylation sites (Figure 4), as substantiated by the finding of the importance of the 30 residue remnant CellBD in binding to Avicel (Figures 2b, 4 and 5b), is concluded to play a crucial role in cellulose binding.
Asn residues in a polypeptide are susceptible to deamidation, which may lead to succinimide formation and finally breakage of the peptide chain.17,18,30-32 Just a slight change in the environmental pH is sufficient to trigger the onset of deamidation.18,19,31 An Asn residue has been shown to undergo fast rates of deamination, particularly when it is followed by a Gly residue, which presumably exerts only a tiny steric effect.17-19 Interestingly, among the 5 N-linked glycosylation sites distributed along CellBD, except the first site, the remaining 4 of them (80%) consist of the Asn-Gly dipeptide (Figure 4). More intriguingly, our findings showed that SC occurred near the last 3 N-linked glycosylation sites (Figures 2 and 4), which are located in a region crucial for cellulose binding.
It has been postulated that deamidation serves as a signal for the turnover of cytochrome c in eukaryotic cells.18,36 Our findings lend support to this conjecture. The finding of Asn69 at the N-terminus of the 30 residue remnant CellBD of the 39 kDa deletant (Figures 2b and 4) suggested that the Asn69 residue and the entire N-linked glycosylation site, Asn69-Gly70-Ser71, might serve as an ‘attributive tag’. A slight change in the environmental conditions, for example, pH, could trigger the onset of Asn deamidation. The N-linked glycosylation sites comprising Gly as the middle residue were envisioned to be readily picked out for deamidation. Since the initial formation of a tetrahedral intermediate from Asn was predicted to be fast and reversible, 19 the attack of the Asn residue might affect the structural stability of nearby residues through modifications of the hydrogen bonding network. 30
Consequently, the peptide bond of residues adjacent to the attacked Asn was destabilized and subjected to SC, which would prevent deamidation from proceeding further with the rate-limiting step of formation of the succinimide product.17,18,30-32 The interpretation might explain why the cleavages observed in CellBD occurred at peptide bonds proximal to the attacked Asn residues instead of at the bond formed between the Asn and Gly residues concerned (Figure 4). However, the exact details of the chemical reactions mentioned remain to be determined.
The low occurrence of the tripeptide, Asn-Gly-Ser/Thr, in 3 commonly encountered E. coli proteins: zero counts for both β-galactosidase 37 (1023 residues) and alkaline phosphatase 38 (471 residues), and only once for β-lactamase 39 (377 residues), reflects that the presence of the tripeptide in E. coli proteins is rare. However, Asn-Gly dipeptides are frequently found in the 3 polypeptides. Thus, the mechanism of deamidation proposed above, which employs the Asn-Gly-Ser/Thr tripeptide as recognition tab and proceeds under the influences of structural conformation and environmental conditions, explains how E. coli may take advantage of a rapid, cost-effective and non-enzymatic approach to discern and eliminate dispensable heterologous proteins.
In addition, the findings described above may provide new insights into the roles of N-linked glycosylation in facilitating native CenA to interact with cellulose during hydrolysis. First, unglycosylated mCenA is susceptible to attack by SC, which was proposed to be initiated from deamidated Asn residues of the N-linked glycosylation sites: Asn-Gly-Ser and Asn-Gly-Thr. Although the Asn-Gly dipeptide appeared to serve well as the target for Asn deamidation, an intact tripeptide, Asn-Gly-Ser or Asn-Gly-Thr, was presumably required to specify the details needed for glycosylation. Second, the glycans attached to these tripeptides appeared to play a crucial role in protecting the recognition sites from being attacked by SC. Third, it was previously reported that a C-terminal fragment containing the C-terminal 47 residues (Ser65 to Ser111) of CellBD (Figure 4) was capable of binding to cellulose. 35 In support of this finding, more precisely, our sequencing results showed that a 30 residue remnant CellBD spanning from Trp68 to Ser97 (Figures 2b and 4) worked well to enable the 39 kDa deletant to bind to Avicel (Figure 5b). Thus, the 2 tripeptides: Asn69-Gly-Ser (the third site) and Asn83-Gly-Ser (the fourth site), which form an integral part of the 30 residue remnant (Figure 4), appear to play an indispensable role for CellBD to bind to cellulose. Fourth, the last 2 potential N-linked glycosylation sites: Asn317-Thr-Ser (the sixth site) and Asn352-Gly-Ser (the seventh site) identified in the catalytic domain (CD) (Figure 4), which might also be involved in substrate binding (as well as catalysis), were also expected to be protected during hydrolysis.
In light of the above considerations, it is concluded that the functionally important modules in native CenA, which are exposed to the hostile environment during cellulolysis, are required to be well protected through glycosylation. Presumably, the N-linked glycans offer protection to CellBD and CD against degradative activities13,15,16,40 and help the domains acquire the required folding or conformation15,16,40 so that CenA can interact appropriately with the substrate. The glycans must unveil the functional domains before the interaction can occur. The binding of CenA to the substrate is expected to be transient and the complex must be dissociated rapidly and cost-effectively, thus non-proteolytically, so that another cellulase molecule can speedily gain access to interact with the substrate. According to this mechanism, the entire CenA molecule will be desorbed from the substrate in a trice. Therefore, only a non-enzymatic approach, SC, can fulfill such a mission (Figure 6). Although many models have been postulated to account for the interaction between an Eng enzyme and a cellulose substrate,41-43 our proposed mechanism represents the first model to explain how a cellulase-substrate complex may be swiftly dissociated to enable the hydrolysis to continue sustainably and cost-effectively.
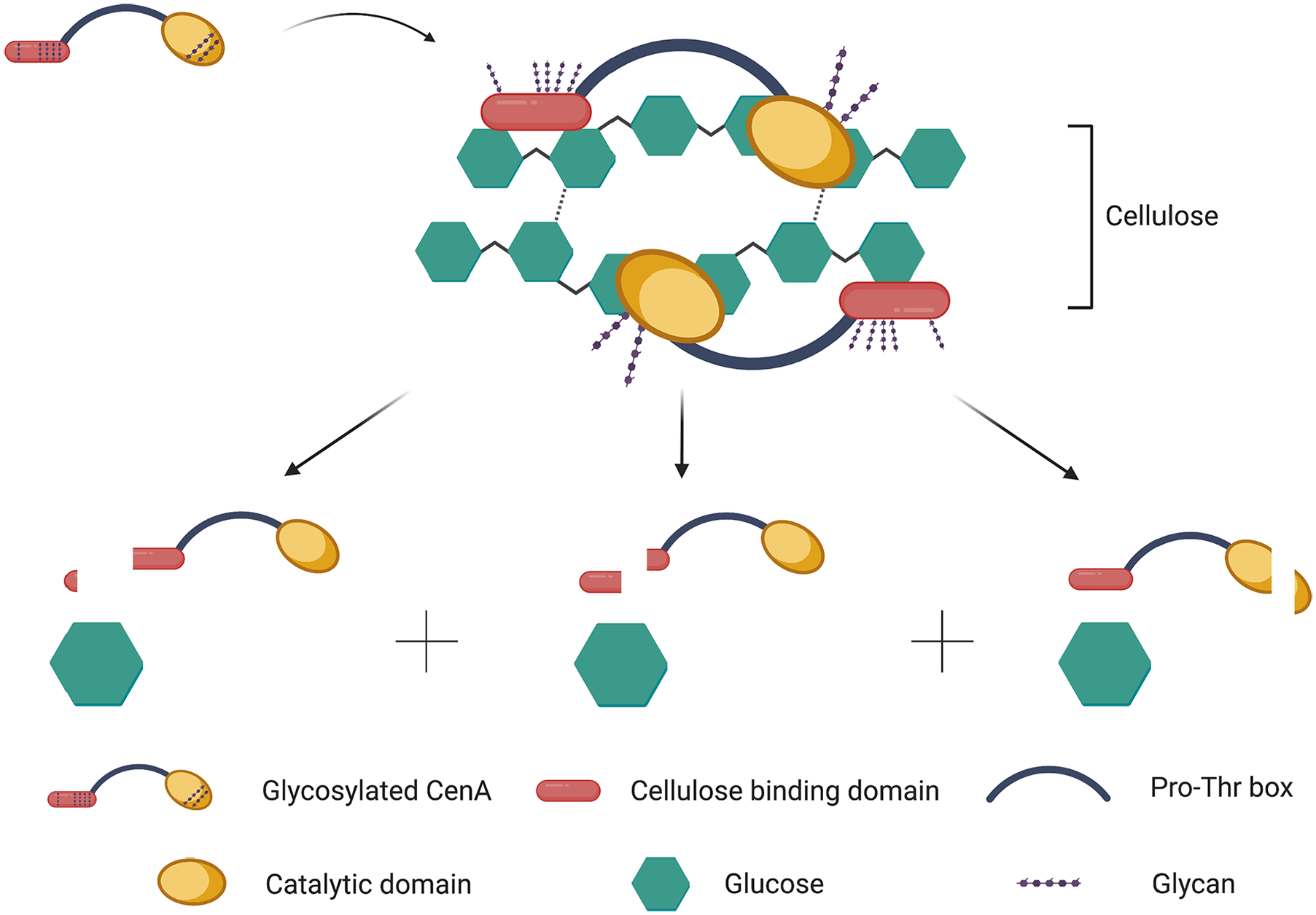
A model explaining dissociation of CenA from the enzyme-substrate complex after hydrolysis.
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by RGC project: GRF16101515.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
