Abstract
MicroRNAs play a significant role in the development of cancers, including lung cancer. A recent study revealed that smoking, a key risk factor for lung cancer, increased the levels of hsa-mir-301a in the tumor tissues of patients with lung squamous cell carcinoma (LUSC). The aim of the current study is to investigate the mechanism by which tobacco smoke increases hsa-mir-301a levels in LUSC tumor tissues using bioinformatics analysis. Bioinformatics tools and online databases, including The Cancer Genome Atlas (TCGA), LinkedOmics, and Encyclopedia of RNA Interactomes (ENCORI), were applied in this study. Our results showed a correlation between the upregulation of hsa-mir-301a in LUSC tissues and smoking exposure. However, no correlation was discovered between patients’ smoking status and the expression level of the hsa-mir-301a host gene, SKA2, prompting us to investigate possible changes in microRNA processing under tobacco smoke exposure. In silico results using online platforms suggest that post-transcriptional processes, which involve the RNA-binding proteins DGCR8 and FUS, contribute to the elevation of mature hsa-mir-301a levels in smoking patients with LUSC. Our findings suggest that RNA-binding proteins play a key role in controlling the processing of hsa-mir-301a, indicating a complex regulation of hsa-mir-301a in the LUSC tissues of smokers.

Introduction
Lung cancer is a common type of cancer worldwide and the leading cause of cancer death. 1 Histologically, it is classified as small-cell lung cancer or non-small-cell lung cancer. Non-small-cell lung cancer (NSCLC) makes up 85% of new cases. The most common forms of NSCLC are lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC). LUSC accounts for about 30% of new NSCLC cases. 2 LUSC is known for its aggressive nature, poor prognosis (with only a 10% 5-year survival rate), and limited availability of targeted drugs compared with LUAD. Lung cancer, including LUSC, has long-hidden clinical signs that usually manifest in the advanced stages of the disease, when treatment options are limited. Options for treatment typically include targeted therapy, chemotherapy, radiation therapy, and surgery; however, the possibility of a local surgical excision of the tumor largely depends on its local spread. 3 The current molecular testing landscape for advanced LUSC includes testing for EGFR, ALK, KRAS, ROS1, BRAF, NTRK1/2/3, MET, RET, and ERBB2 genetic alterations, although the overall frequency of targetable molecular alterations in LUSC ranges from 2% to 10%. 4 The number of approved targeted therapies for LUSC is significantly lower than for LUAD, and the success of targeted therapy may be significantly affected by the presence of previous treatment. Moreover, although the possible systemic treatment options for advanced stages of LUSC are still diverse, not all accepted treatment options have shown a significant difference in patient survival. 5 Thus, the specific targets responsible for the development and progression of LUSC are still not fully understood.
Epidemiological evidence indicates that most cases of lung cancer are straightforwardly connected to smoking. 6 Thus, less than 10% of lung cancer cases occur in individuals who have never smoked. Furthermore, smokers have a significantly higher risk, 10 to 25 times greater, of dying from lung cancer compared with non-smokers. Notably, smoking has the least effect on LUAD and the greatest effect on LUSC. 7 There is strong evidence that connects tobacco smoking to the development of lung cancer, specifically LUSC. However, the exact mechanisms by which tobacco smoke contributes to the development of LUSC are not yet fully understood.
In recent years, researchers have found that microRNAs play a significant role in the development of various cancers, including lung cancer.8,9 MicroRNAs are non-coding RNAs that are typically 20 to 24 nucleotides long. They have the ability to recognize and attach to specific mRNA transcripts based on their nucleotide sequence. Such interaction can result in the degradation of the mRNA or the inhibition of its translation. 10 Alterations in microRNAs’ downstream molecular targets then affect signaling pathway cascades. This, in turn, can have a profound impact on the overall network of gene expression and cellular processes. MicroRNAs are crucial for regulating cell growth, differentiation, development, and apoptosis. 11
It is well known that external factors can significantly alter the levels of microRNAs. Thus, tobacco smoking affects the levels of microRNAs in different human organs. 12 Our recent study has revealed a connection between smoking and higher levels of hsa-mir-301a in tumor tissues from LUSC patients. 13 Reports indicate that hsa-mir-301a is highly expressed in various tumor types and promotes cancer cell growth and invasion. 14 Moreover, hsa-mir-301a has been shown to be involved in the development of cisplatin resistance in patients with NSCLC. 15 Understanding the mechanism of hsa-mir-301a regulation may be crucial for diagnosing and selecting an effective therapy regimen for LUSC patients. The objective of this study was to examine how tobacco smoke affects the levels of hsa-mir-301a in LUSC tumor tissues. To accomplish this, we made use of several free online tools and databases.
Materials and Methods
UALCAN analysis
The Expression module of UALCAN 16 (The University of Alabama at Birmingham Cancer data analysis Portal; http://ualcan.path.uab.edu/analysis.html) was applied to analyze has-mir-301a and gene expression data in the TCGA (The Cancer Genome Atlas) dataset of LUSC tissues and normal lung tissue. UALCAN provides the ability to categorize patients into several subgroups using clinical patient data, including smoking status for patients with lung cancer. The Welch’s t-test was used to determine the significance of expression-level differences between normal lung tissue and primary LUSC tissues, and between different subgroups of LUSC based on smoking status.
LinkedOmics database analysis
The LinkFinder module of the online platform LinkedOmics 17 (http://www.linkedomics.org/) was used for the correlation analysis of microRNA and gene expression levels in the TCGA dataset of LUSC samples. The Spearman’s correlation coefficient was used to statistically analyze the results.
Prediction of RNA-binding proteins–miRNA interaction
The Encyclopedia of RNA Interactomes (ENCORI) online platform 18 (https://rnasysu.com/encori/) was used to predict the interactions between miRNA and RNA-binding proteins.
Results
Smoking is accompanied by an increase in hsa-mir-301a
We compared hsa-mir-301a levels in LUSC samples with values in normal samples using the UALCAN online tool, which included 503 primary LUSC cases. The results demonstrated that the LUSC tumor tissues had significantly higher levels of hsa-mir-301a when compared with the normal group. (Figure 1A). A subgroup analysis was conducted using UALCAN to examine the impact of smoking habits on LUSC patients. Patients who continued smoking during or after treatment were considered current smokers. Patients who had never smoked were categorized as non-smokers. Those who had stopped smoking before being diagnosed were classified as reformed smokers. There were two subgroups of reformed smokers: those who had stopped smoking more than 15 years before being diagnosed with LUSC and those who had stopped within 15 years of diagnosis. The research results showed that patients who smoked had significantly higher levels of hsa-mir-301a compared with non-smokers (Figure 1B). These findings suggested that cigarette smoke increases hsa-miR-301a levels in LUSC. We conducted further investigation to understand how tobacco smoke induces the levels of hsa-miR-301a in LUSC tissues.

The levels of hsa-mir-301 in LUSC (according to the UALCAN database) are shown in bars. (A) The bars compare the levels of hsa-mir-301 in primary tumors to normal tissues, and (B) in subgroups based on the patients’ smoking habits. The LUSC samples were divided into four subgroups: smokers, non-smokers, reformed smokers 1 (⩽ 15 years), and reformed smokers 2 (> 15 years).
A review of the UALCAN database’s data on hsa-mir-301a expression level and its impact on LUSC patient survival revealed no significant impact, suggesting that this microRNA could be more of a potential therapeutic target rather than a prognostic one (Supplementary figure 1) 16
Smoking does not affect the expression of SKA2, hsa-mir-301a host gene
Mature microRNA levels are controlled during the biogenesis cascade at both the transcriptional and post-transcriptional stages. 19 The genomic location of microRNAs plays a crucial role in the regulation of primary microRNAs during transcription. 20 The precursor of hsa-mir-301a is located in Intron 1 of the SKA2 (Spindle And Kinetochore Associated Complex Subunit 2) host gene. 21 This fact led us to analyze the levels of the SKA2 gene and investigate the correlation between SKA2 and hsa-mir-301a in LUSC tissues. The SKA2 gene expression levels in LUSC tissues were examined using the UALCAN tool. The findings showed that LUSC tissues had much greater amounts of SKA2 mRNA compared with normal tissues (Figure 2A). Next, we looked into the relationship between the levels of hsa-mir-301a and SKA2 in LUSC tissues using the LinkFinder module in the LinkedOmics database. The results showed a significant positive correlation between hsa-mir-301a and the SKA2 gene expression (Figure 2B). These results suggest that the regulation of the SKA2 gene promoter at the transcriptional level can affect the levels of hsa-mir-301a in LUSC tissues.

The levels of SKA2 gene expression in LUSC (according to the UALCAN database) are shown in bars. (A) The bars compare the levels of SKA2 gene in primary tumors to normal tissues, and (C) in subgroups based on the patients’ smoking habits. The LUSC samples were divided into four subgroups: smokers, non-smokers, reformed smokers 1 (⩽ 15 years), and reformed smokers 2 (> 15 years). (B) Spearman’s correlation analysis of the correlation between SKA2 and hsa-mir-301a. Data were obtained using LinkedOmics.
Next, we examined the levels of the SKA2 gene in several groups of LUSC patients, divided according to their smoking behaviors, to determine whether tobacco smoke can cause an increase in SKA2 expression. After splitting patient samples into subgroups depending on their smoking habits, the study of SKA2 gene levels showed that smoking had no significant effect on the rise in SKA2 mRNA levels in LUSC tissues (Figure 2C). The level of SKA2 mRNA was significantly higher in all subgroups of LUSC samples compared with normal tissue. However, there were no significant differences in SKA2 gene levels between subgroups of LUSC patients (Figure 2C). This suggests that smoking may cause an increase in hsa-mir-301a levels compared with non-smokers during the post-transcriptional stage of microRNA biogenesis, as there were no differences in SKA2 gene expression levels between subgroups of smokers and non-smokers with LUSC.
Smoking may contribute to hsa-mir-301a biogenesis
The process of microRNA biogenesis involves two RNases, named DROSHA and DICER. In the nucleus and cytoplasm, respectively, these RNases carry out a series of sequential processing steps. 19 There is growing evidence that RNA-binding proteins are involved in regulating post-transcriptional processing. 22 Specifically, these proteins can either positively or negatively affect the processing of pri-microRNA in the nucleus and pre-microRNA in the cytoplasm by binding to the microRNA precursor. We assumed that smoking may affect RNA-binding proteins, perhaps resulting in more hsa-mir-301a being processed.
Using the ENCORI platform, we have identified 39 proteins that can interact with SKA2 mRNA (Table 1). From this group, we specifically isolated proteins that can interact with Intron 1, which encodes the precursor of hsa-mir-301a. A total of 27 proteins have been identified to interact with Intron 1 of SKA2 mRNA (Table 1). Next, we used the UALCAN to compare the levels of genes encoding these 27 proteins (Table 1) between subgroups of current smokers and non-smokers. From our analysis, we found three genes (DGCR8, DICER1, and FUS) that were differentially expressed. These genes showed significantly higher expression levels in the subgroup of smokers compared with non-smokers, as shown in Figure 3.
RNA-binding proteins capable of binding to SKA2 mRNA.

Data were obtained from the ENCORI platform: The Encyclopedia of RNA Interactomes.
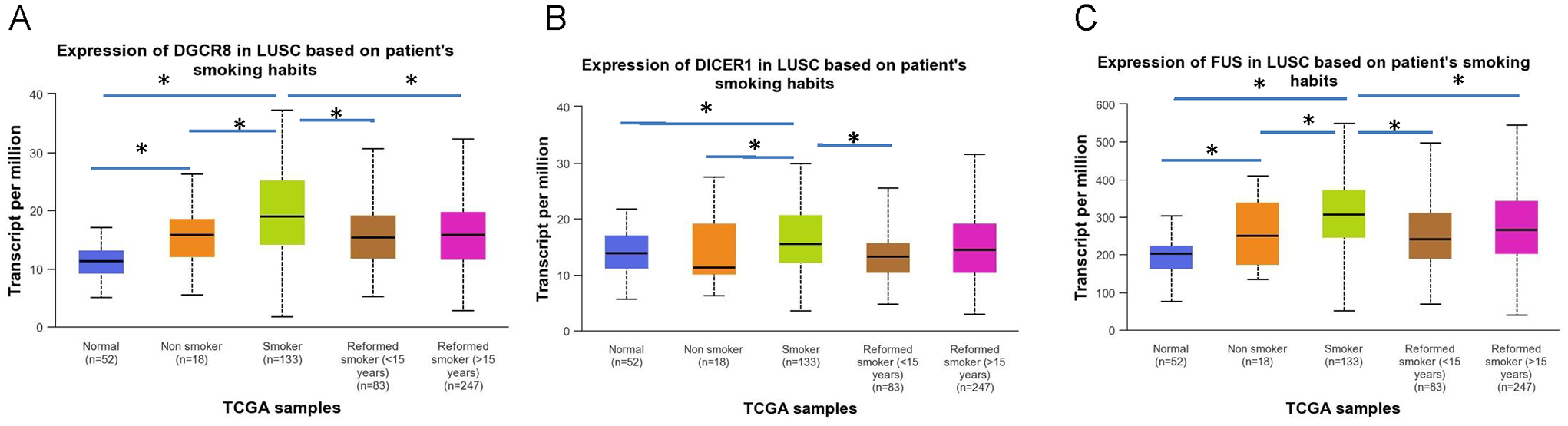
Gene expression levels of RNA-binding proteins. The bars compare the levels of (A) DGCR8, (B) DICER1, and (C) FUS genes in subgroups based on the patients’ smoking habits. The LUSC samples were divided into four subgroups: smokers, non-smokers, reformed smokers 1 (⩽ 15 years), and reformed smokers 2 (> 15 years).
The correlation between the expression levels of the genes DGCR8, DICER1, and FUS and the levels of the SKA2 gene and hsa-mir-301a was examined using the LinkFinder module of the LinkedOmics database. The findings showed that hsa-mir-301a significantly correlated with two of the three genes, DGCR8 and FUS (Figure 4A to C). The correlation analysis showed that there was no significant correlation between the expression levels of SKA2 and the levels of the DGCR8, DICER1, and FUS genes (Figure 4D to F).
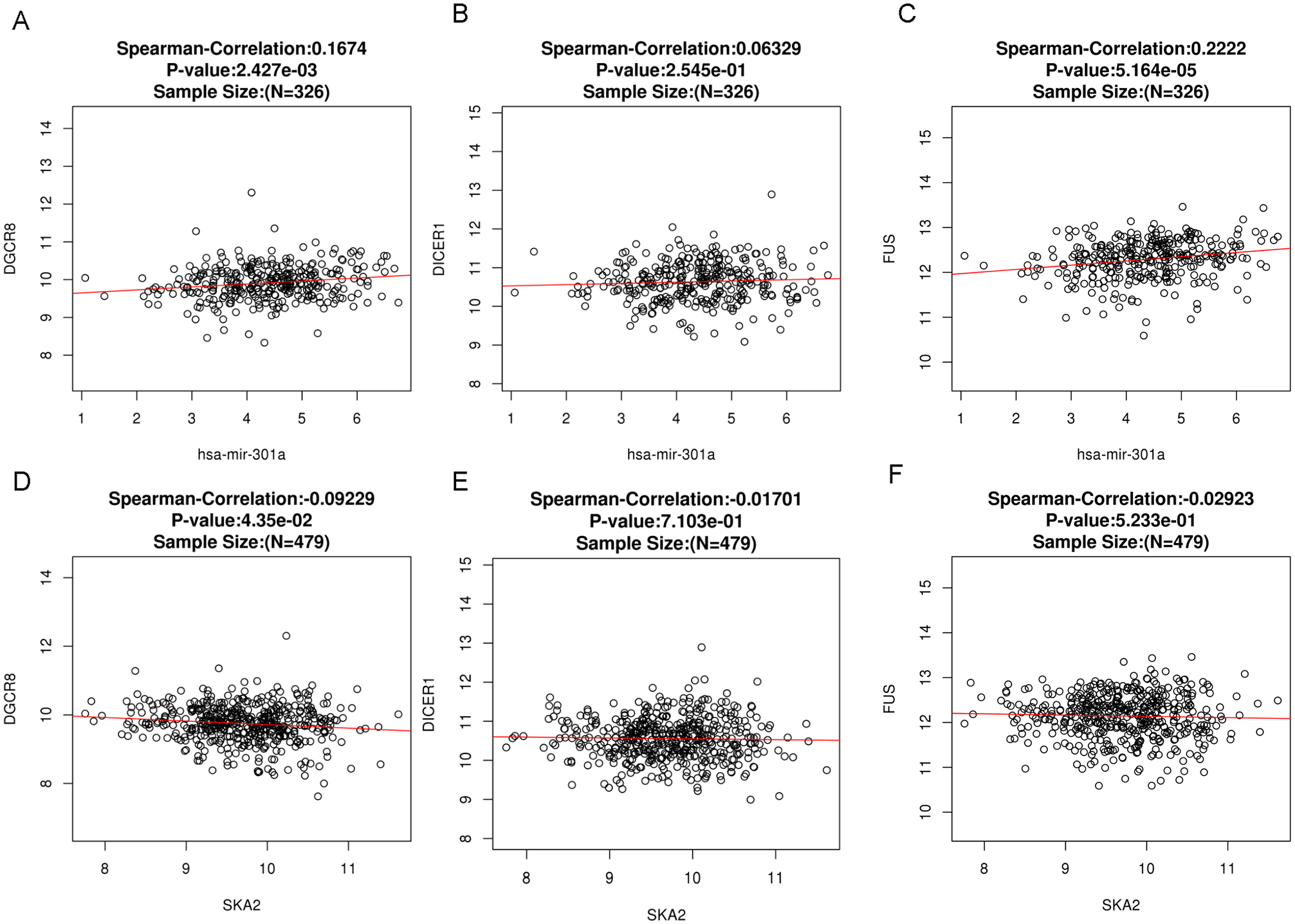
Spearman’s correlation analysis of the correlation between hsa-mir-301a (A to C) or SKA2 (D to F) and gene expression levels of RNA-binding proteins. Data were obtained using LinkedOmics.
A recent study revealed that smoking can reduce the expression of the tumor suppressor PTEN. 13 This reduction was associated with increased levels of hsa-mir-301a, which targets PTEN mRNA.23 -26 Therefore, we examined the correlation between PTEN levels and the hsa-mir-301a host gene, SKA2, and the genes that encode the RNA-binding proteins DGCR8 (DiGeorge syndrome critical region 8) and FUS (fused in sarcoma protein). The analysis revealed a significant negative correlation between the levels of hsa-mir-301a and its target gene, PTEN, in LUSC tumor tissues (Figure 5). However, no significant correlation was found between PTEN and SKA2. The results also showed significant negative correlations between PTEN and the DGCR8 and FUS genes in LUSC tumor tissues. These facts suggest that RNA-binding proteins encoded by the DGCR8 and FUS genes may be key contributors to the increased levels of hsa-mir-301a in tumor tissues of smoking LUSC patients by enhancing microRNA biogenesis.

Spearman’s correlation analysis of the correlation between PTEN (target of hsa-mir-301a) and (A) hsa-mir-301a, (B) SKA2, (C) DGCR8, and (D) FUS. Data were obtained using LinkedOmics.
Discussion
It has been widely recognized that smoking is strongly associated with the development of LUSC. 27 The treatment options for LUSC are currently limited. Therefore, it is crucial to identify the key molecules that are associated with smoking and contribute to the development of LUSC. Understanding the molecular mechanisms of LUSC is crucial for improving patient prognosis. Despite numerous studies on gene expression signatures associated with smoking, there is still a lack of comprehensive understanding in this area. A recent study revealed that smoking leads to a decrease in the tumor suppressor PTEN in LUSC tissues by increasing hsa-mir-301a. 13 The deregulation of hsa-mir-301a has been previously reported in several types of cancer. 14 Therefore, understanding the molecular mechanisms of this microRNA in cancer could help identify potential therapeutic targets for tumor treatment. In this study, we tried to propose a possible mechanism for smoking-induced increase in hsa-mir-301a levels. We conducted a bioinformatics analysis of TCGA data using free online tools for this purpose.
The location of microRNAs is a significant factor in determining the transcriptional regulation of primary microRNAs. Knowing the coexpression patterns between microRNAs and host genes in tumor tissues considerably improves our understanding of microRNA transcriptional and post-transcriptional control. The hsa-mir-301a precursor is found within the first intron of the SKA2 host gene. The SKA2 gene encodes the protein SKA2, which is necessary for the start of anaphase in mitosis and plays a role in maintaining the metaphase plate and silencing the spindle checkpoint. 28 Studies have demonstrated that SKA2 can control cell proliferation. 29 The expression of the SKA2 gene is increased in various cancer cell lines and clinical samples, including lung cancer. 30 Our findings revealed that SKA2 levels were significantly higher in LUSC tissues compared with normal tissues. In addition, we observed a significant positive correlation between SKA2 levels and hsa-mir-301a levels in tumor tissues. These results suggest that alterations in SKA2 expression may impact the expression of hsa-mir-301a in LUSC tumor tissues at the transcriptional level. However, when patient samples were divided into cohorts based on smoking status, the analysis of SKA2 gene expression levels revealed that smoking does not have a significant additive effect on the increase in SKA2 levels in LUSC tumor tissues.
In recent years, evidence has emerged that the level of mature microRNA is regulated not only by the transcription rate but also by the efficiency of processing of the microRNA precursor by DROSHA (in the nucleus) and DICER1 (in the cytoplasm). 22 Specific RNA-binding proteins can recognize sequences in microRNA precursors and modulate processing efficiency, depending on cellular context or external signals. 31 Our in silico data showed that 27 RNA-binding proteins can interact with the hsa-mir-301a precursor. Further analysis of the expression of genes encoding these RBPs identified three genes, DGCR8, DICER1, and FUS, the expression of which was significantly increased in a cohort of smoking patients. In addition, we observed a significant positive correlation between the DGCR8 and FUS genes and hsa-mir-301a levels in LUSC tissues, suggesting that proteins encoded by the DGCR8 and FUS genes may promote processing and consequently lead to increased hsa-mir-301a levels in a cohort of smokers. Previously, some RNA-binding proteins have already been proposed as potential therapeutic targets for the treatment of malignant tumors.32,33 Furthermore, they are considered to be involved in molecular pathways that link chronic inflammation in chronic obstructive pulmonary disease (COPD) with malignant cell transformation in the development of lung cancer. 34 However, further research is needed to identify the precise pathways or molecular targets that are most promising for potential clinical application.
The DGCR8 protein and RNase III (DROSHA) form the microprocessor complex, which is responsible for initiating microRNA maturation in the cell nucleus. 35 DGCR8 interacts with pri-microRNA, facilitating efficient and accurate processing. The FUS protein is involved in multiple cellular processes. 36 It has been discovered that FUS helps in the production of microRNA by assisting in the recruitment of Drosha to pri-microRNA. 37 It seems that smoking causes an increase in RNA-binding proteins, DGCR8 and FUS, in LUSC tissues. This increase stimulates the first step of hsa-mir-301a processing in the nucleus.
Conclusion
In summary, bioinformatics analysis of TCGA data using free online tools revealed that smoking is connected to the increased levels of hsa-mir-301a in LUSC tissues. Furthermore, higher levels of hsa-mir-301a in LUSC tissues are associated with increased expression of genes encoding RNA-binding proteins, DGCR8 and FUS, involved in hsa-mir-301a maturation. These facts indicate that post-transcriptional processes are likely the primary factor that regulates the level of hsa-mir-301a in the LUSC tissues of smoking patients. Undoubtedly, further experimental research is required to provide more evidence that post-transcriptional mechanisms play a major role in elevating hsa-mir-301a levels in LUSC tissues in a cohort of smokers.
Supplemental Material
sj-tiff-1-bbi-10.1177_11779322241302168 – Supplemental material for Mechanistic Insights of hsa-mir-301a Regulation by Tobacco Smoke in Lung Squamous Cell Carcinoma: Evidence From Bioinformatics Analysis
Supplemental material, sj-tiff-1-bbi-10.1177_11779322241302168 for Mechanistic Insights of hsa-mir-301a Regulation by Tobacco Smoke in Lung Squamous Cell Carcinoma: Evidence From Bioinformatics Analysis by Vladimir O Pustylnyak, Alina M Perevalova and Lyudmila F Gulyaeva in Bioinformatics and Biology Insights
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by RUSSIAN SCIENCE FOUNDATION (grant no. 22-15-00065).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Vladimir O. Pustylnyak: Conceptualization, Methodology, Investigation, Validation, Data curation, Writing – original draft, Writing – review and editing. Alina M. Perevalova: Methodology, Investigation, Validation, Data curation, Writing – original draft. Lyudmila F. Gulyaeva: Conceptualization, Methodology, Investigation, Validation, Data curation, Resources, Writing – original draft, Writing – review & editing, Supervision, Project administration, Funding acquisition.
Data Availability
The datasets used for bioinformatics analysis in this study are available from online platforms, such as UALCAN, LinkedOmics, and ENCORI.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
