Abstract
The advent of biobanks with vast quantities of medical imaging and paired genetic measurements creates huge opportunities for a new generation of genotype–phenotype association studies. However, disentangling biological signals from the many sources of bias and artifacts remains difficult. Using diverse medical images and time-series (ie, magnetic resonance imagings [MRIs], electrocardiograms [ECGs], and dual-energy X-ray absorptiometries [DXAs]), we show how registration, both spatial and temporal, guided by domain knowledge or learned de novo, helps uncover biological information. A multimodal autoencoder comparison framework quantifies and characterizes how registration affects the representations that unsupervised and self-supervised encoders learn. In this study we (1) train autoencoders before and after registration with nine diverse types of medical image, (2) demonstrate how neural network-based methods (VoxelMorph, DeepCycle, and DropFuse) can effectively learn registrations allowing for more flexible and efficient processing than is possible with hand-crafted registration techniques, and (3) conduct exhaustive phenotypic screening, comprised of millions of statistical tests, to quantify how registration affects the generalizability of learned representations. Genome- and phenome-wide association studies (GWAS and PheWAS) uncover significantly more associations with registered modality representations than with equivalently trained and sized representations learned from native coordinate spaces. Specifically, registered PheWAS yielded 61 more disease associations for ECGs, 53 more disease associations for cardiac MRIs, and 10 more disease associations for brain MRIs. Registration also yields significant increases in the coefficient of determination when regressing continuous phenotypes (eg, 0.36 ± 0.01 with ECGs and 0.11 ± 0.02 for DXA scans). Our findings reveal the crucial role registration plays in enhancing the characterization of physiological states across a broad range of medical imaging data types. Importantly, this finding extends to more flexible types of registration, such as the cross-modal and the circular mapping methods presented here.
Introduction
The concept of registration—mathematical maps between coordinate systems—has ancient roots, dating back at least as far as ancient Egypt. 1 There, land was surveyed with regularly knotted ropes that were gathered together—physically implementing isomorphic scaling. 2 In the medical imaging domain, registration has long been used to align different individuals to a common prototypical example of the data (ie, an atlas), or an idealized parameterization of an anatomical feature. In general, registration helps models target biological variance of interest. Variation from non-biological sources is common in medical images: for example, magnetic resonance imagings (MRIs) encode many technical and environmental artifacts, such as operator dependence or patient position in a scanner. In the supervised learning setting, there is a large literature on how to adjust for such nuisance variation. These approaches include: leveraging data augmentations, 3 adjustment based on an assumed causal model, 4 adjustment using auxiliary labels, 5 and regularizing prediction functions across different domains. 6 The unsupervised learning setting, meanwhile, presents a greater challenge as there is no outcome variable to help focus the model on relevant signals.
To ameliorate this, we demonstrate how registration affects the representations that deep neural networks learn, vastly increasing the strength of genetic, diagnostic, and phenotypic associations. This is shown across a broad range of registration methods from hand-crafted templates to nonlinear deformation fields, and a correspondingly broad range of medical data from the electro-temporal waveforms of electrocardiograms (ECGs), to two-dimensional (2D) images such as dual-energy X-ray absorptiometries (DXAs) and brain MRI slices, as well as three-dimensional (3D) spatio-temporal cardiac MRI movies.
The recent availability of large-scale multimodal measurements in biobanks7,8 provides an opportunity to systematically study how registration affects representations of physiology. Because biobanks often assess individuals with many different modalities they allow us to compare and contrast different image types. 9 Leveraging multimodal data from the UK Biobank, this article makes the following contributions (1) quantify the benefit of registration across a broad range of medical image modalities, (2) show that several neural network-based methods (VoxelMorph, 10 DeepCycle, 11 and DropFuse) 12 can effectively learn registrations allowing for more flexible and efficient processing than is possible with hand-crafted or atlas-based registrations, and (3) demonstrate the breadth of downstream phenotypic analyses enriched by registration, scaling to thousands of statistical tests that comprise a phenome-wide association study (PheWAS) and the millions that constitute a genome-wide association study (GWAS).
Anatomical registrations
Many anatomical features have been used for registration. For example, 3D anatomical atlases of the brain are used to register brain MRIs. 13 Likewise, the cardiac cycle of a single heartbeat can serve as a template for both ECGs and cardiac MRIs. 14 For example, the full 10 s of the resting ECG can be registered by template matching the QRS complex, followed by alignment, scaling, and median computation.15,16 The DXA scans can be registered with rigid homeomorphic mappings to an exemplar individual. 17 Figure 1 shows visual examples of anatomical registration by template matching and atlas alignment in five different modalities. The top row shows each modality from three different individuals overlaid before registration, while the bottom row shows the modalities overlaid after registration.

Examples of three individuals overlaid as red, green, and blue color channels before and after registration. The top row shows the original modality and the bottom row shows the registered version. From left to right, the modalities are the resting ECG, DXA 5 (hip), DXA 2 (lumbar spine), T1 brain MRI, and DXA 11 (whole-body skeletal).
Learned registrations
Classical registration methods solve a new optimization problem for every pair, and they are therefore computationally expensive.18-21 Recent deep learning methods propose to learn the alignment between the medical image and reference instead10,22-24; supervised methods train the network and compare the output with pre-computed alignments, while unsupervised methods train their networks by learning a transformation from the image to reference, applying it and then comparing how well the aligned image matches the reference. 25 VoxelMorph, for instance, uses convolutions and spatial transformations together with a UNet architecture to learn a deformation field for each image/reference pair. It learns amortized registration by jointly maximizing agreement between the aligned image and the reference, with a transformation smoothness regularizer. 10
While deformation fields can preserve or even increase parameterization of a modality, many registration techniques reduce dimensionality. Taking reduction to its logical extreme, DeepCycle uses a single-parameter autoencoder. This one-dimensional latent space is encoded with the inductive bias of periodicity, registering single-cell RNA expression data to the mitotic cell cycle. 11 Multimodal fusion methods, such as DropFuse, use contrastive cross-modal learning to register different modalities into a 256-dimension latent space, also greatly reducing data size. 12 Similar strategies have been pursued in a line of recent works where contrastive encoders learn joint representations of multimodal data such as natural images and their captions,26,27 or paired clinical measurements.28,29 Figure 2 provides a graphical summary of the learned registration techniques.

Three deep-learning methods used to register medical images. DeepCycle registers MRI frames to the cardiac cycle by encoding each from with a single parameter, θ, VoxelMorph learns spatial warps via pairwise amortized registration while retaining overall image dimensions and DropFuse uses dropout and cross-modal fusion to register multiple modalities together into a 256-dimensional latent space.
Genetic analysis of medical images
The advent of large genotyped Biobanks enables whole-genome association testing with traits derived from medical images such as MRIs, retinal images, and ECGs.30-32 Active research has extended these genomic analyses from traits to spaces, performing the association tests in unsupervised ways. For example, GWAS of each dimension in a variational autoencoder, or the principal components of that encoding or even with ECG voltages from medians over every millisecond of input.33-36 Building on this work, we demonstrate how registration uniformly results in richer phenotypic and genotypic associations over a diverse set of methods and modalities.
Methods
We train modality-specific DenseNet-style 37 convolutional encoders and decoders to reconstruct both registered and unregistered medical images from latent space bottlenecks. Table 1 details the registration techniques considered and the modalities involved. Many implementations of registration are considered, including registration learned from scratch, optimized from a predefined type of mathematical transformation (eg, homeomorphic or warp fields), or algorithmically hard-coded. The learned registrations map between individuals via deformation fields (VoxelMorph), with the inductive bias of periodicity (DeepCycle) or contrastively across modality (DropFuse). These previously described models are briefly summarized below. Code for these experiments is available in the Broad Institute’s ML4H github repository: https://github.com/broadinstitute/ml4h/tree/master/model_zoo/registration_reveals_genetics.
Modalities and registrations.

DXA, dual-energy X-ray absorptiometry; ECG, electrocardiogram; MRI, magnetic resonance imaging.
Each modality, its original shape, shape after registration, the resulting reduction in dimensionality, the method used to register the modality, and the way the parameters of the registering transformation are derived (ie, hard-coded with domain knowledge, optimized parameters of a known transformation or learned a new transformation via neural net approximation).
DeepCycle
DeepCycle learns to encode data using a single-parameter latent space registered to the unit circle, demonstrating the extreme reductions possible with registration. A convolutional encoder learns a single parameter bottleneck,
VoxelMorph
VoxelMorph is trained to minimize both smoothness and similarity losses. The smoothness loss encourages anatomical plausibility while the similarity loss ensures the fidelity of the learned registration. We trained VoxelMorph with four-chamber long axis cardiac MRI (cMRI) cine series and smoothness loss weight of 0.5. For purposes of comparison, the individual with median body mass index (BMI) was selected as an exemplar and all cMRI movies were VoxelMorphed to them.
DropFuse
DropFuse is cross-modal autoencoder trained to minimize a reconstruction loss combined with a contrastive loss, which ensures that paired ECG and MRI samples are mapped to nearby points in the latent space, while discordant modality pairs are pushed away. The embeddings from each modality are fused with random dropout at each latent space coordinate. The model is trained with ECG, MRI, and DXA series pairs. The encoders for each modality are serialized separately so that inference requires only one modality to be available.
Results
To quantify the overall biological signal captured by a representation, we aggregated a broad range of phenotypes of clear biological import, including age, sex, BMI, heart rate, disease diagnoses, and principal components of genetic ancestry, see Supplementary Figures 1 to 3 for the complete lists of phenotypes used with each modality. Importantly, the phenotypes considered include both general biological features such as age and sex, along with more modality-specific phenotypes, such as the QT interval of the ECG. We then build linear probes to detect how much each representation has learned about each phenotype. It has been shown that linear separability increases monotonically as we probe deeper into the model 40 ; thus by analyzing the deepest bottleneck layer, we get the best estimate for how much each phenotype is recoverable from each latent space. Both continuous and categorical phenotypes are considered. The linear probes are fivefold cross-validated to bootstrap confidence intervals on the linear and logistic regression models trained on representations of each modality before and after registration. These results are summarized in Table 2.
Registered modalities capture more phenotypic and genetic associations.
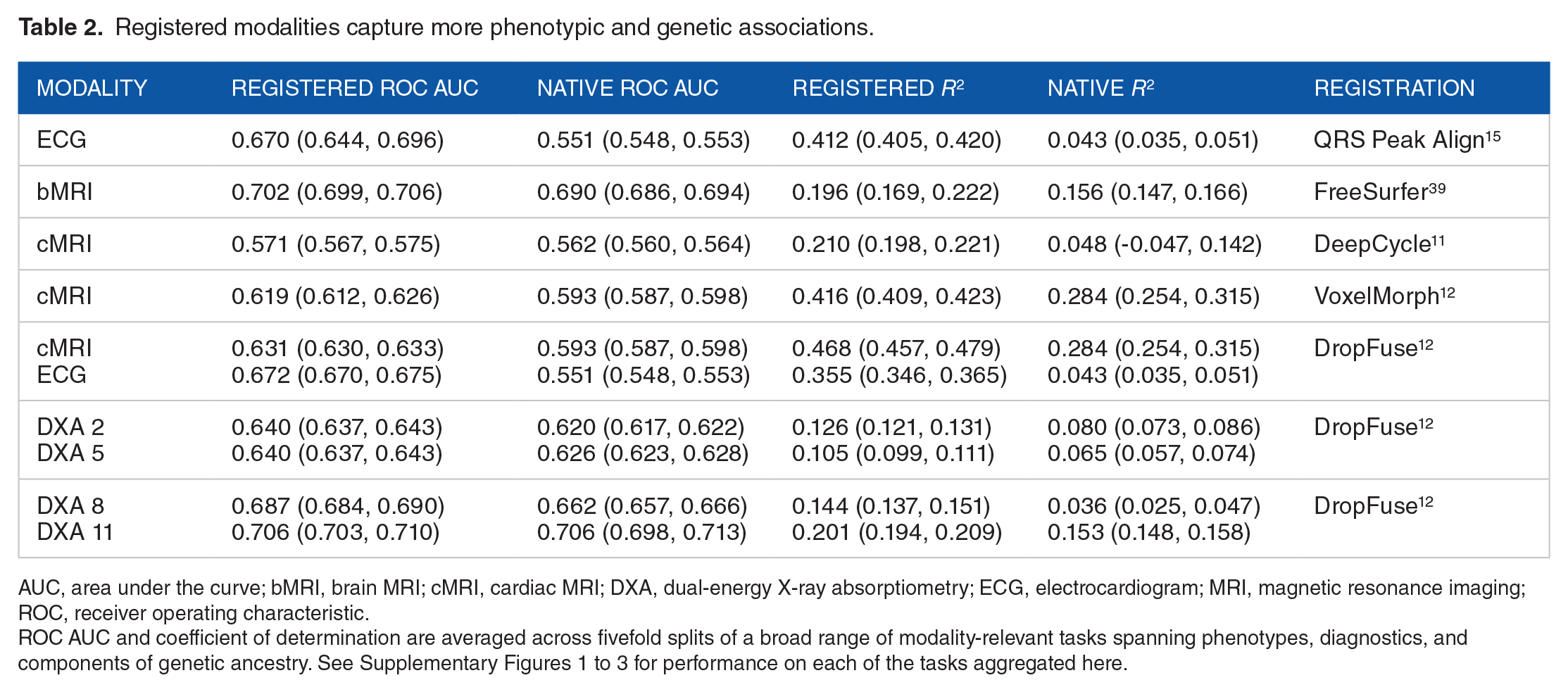
AUC, area under the curve; bMRI, brain MRI; cMRI, cardiac MRI; DXA, dual-energy X-ray absorptiometry; ECG, electrocardiogram; MRI, magnetic resonance imaging; ROC, receiver operating characteristic.
ROC AUC and coefficient of determination are averaged across fivefold splits of a broad range of modality-relevant tasks spanning phenotypes, diagnostics, and components of genetic ancestry. See Supplementary Figures 1 to 3 for performance on each of the tasks aggregated here.
Improvements in the area under the ROC curve and the coefficient of determination,
Notably, even lossy registrations can yield more biological signals, for example from the 5000 time points of the 10 s resting ECG to the 600 time points in the registered median waveform. Although much lower in overall dimensionality, registration greatly reduces intra-individual variations such as phase and baseline drift. This frees up the latent space to “spend” its expressive capacity representing more population variation, which powers downstream analyses.
Registered modalities reveal more genetic variation
The prediction of principal components of ancestry described above indicated that registration often resulted in stronger genomic association. To precisely pinpoint the genetic loci involved, we perform association tests between the latent spaces and millions of single-nucleotide polymorphisms (SNPs) throughout the entire genome. This “unsupervised” GWAS works directly on the autoencoder representations. Specifically, for each SNP, we check if the latent space centroids of the three diploid genotypes (homozygous variant, heterozygous, and homozygous reference) are separable, as quantified by multivariate analysis of variance (MANOVA). Prior to performing MANOVA across the sets, we account for confounders from population stratification and batch effects, by removing these sensitive features from the latent space with iterated nullspace projection. 41
The GWAS results are shown with Manhattan plots from registered and unregistered modalities superimposed in Figure 3. In general, registered modalities have both more and stronger associations. One notable exception for the brain MRIs is the gene WNT16, which is a site previously associated with bone density, height and body plan, not specific to the brain. In contrast, the top peak in the registered GWAS, labeled c15orf54 (chromosome 15 open reading frame 54), has previously been associated with brain region volumes and cortical structure, and this site is not significant in the native coordinate GWAS.42,43 Similarly, the loci in genes FOXD2, GMNC, DAAM1, and PTCH1 all have previously reported associations with brain-specific traits including white matter microstructure, cortical surface areas, and Alzheimer’s disease biomarkers,44-46 see Supplementary Table 1 for all T1 brain MRI lead SNPs, LocusZoom for the full GWAS summary statistics: https://my.locuszoom.org/gwas/761153/.

Manhattan plots of the T1 brain MRI (top) and the resting 12-lead ECG (bottom) with unregistered representations (lead SNPs shown in red) and with registered representations (lead SNPs shown in purple). For P-values and exact loci, see Supplementary Tables 1 to 3.
The GWAS of the median-waveform registered 12-lead resting ECG revealed 86 genome-wide significant loci in contrast to 0 genome-wide loci for the full 10 s unregistered ECG. The 86 loci have many previous associations with cardiovascular and specifically electrocardiographic traits. Note the large number of ion channel genes identified, specifically the sodium channels SCN5A and SCN10A and the potassium channels KCNQ4, KCND3, KCNH2, and KCNQ1. These genes play a critical role in cardiac conduction and have been associated with many cardiovascular disorders including atrial fibrillation, Brugada syndrome, long QT syndrome, and other cardiac conduction diseases.47-49 Besides the ion channels, many of the other loci identified with nearest genes including TTN, TBX3, PITX2 have extensive previous associations with cardiovascular traits and disorders.33,50-54 Supplementary Table 2 contains the complete list of resting ECG lead SNPs, and full GWAS results are available at: https://my.locuszoom.org/gwas/520108/.
“Miami” plots, which superimpose two Manhattan plots after multiplying the y-axis of the unregistered modality by –1 for DXA series 12 scans, are shown in Supplementary Figure 4. 55 We identified 21 genome-wide significant loci, including the genes CPED1, AKAP11, and GDF5, which have previous associations with bone mineral density, body height, and osteoarthritis. 56 All registered DXA 12 lead SNPs are listed in Supplementary Table 3. In contrast, the GWAS of the DXA 12 latent space learned before registration identifies just two loci. The full GWAS results for the DXA 12 modality are available at: https://my.locuszoom.org/gwas/204146/.
Clustering SNPs in latent space elucidates genetic architectures
The SNP representations can be clustered to identify genetic structure in latent spaces. In particular, we perform hierarchical clustering based on the direction from the mean embedding of the homozygous reference group to the mean embedding of the heterozygous and homozygous variant groups for each lead SNP identified in a GWAS. Figure 4 contrasts latent space GWAS on three different brain MRI representations learned from the cerebellum, using only white matter, only gray matter, and the whole cerebellum. The consistency of the SNP clustering between training runs and sub-regions of the cerebellum demonstrates the reproducibility of both the SNP findings and their high-dimensional latent space representations. Specifically, the blue arrows in Figure 4 highlight an SNP in the gene SLC35B3, which is significant in all three brain representations and consistently in an outgroup—a distal branch of the hierarchical clustering. This gene, of the Solute Carrier Family 35, member B3 has previous associations with motion sickness, 57 working memory, 58 and autosomal dominant ophthalmic outbursts. 59 In contrast to the consistent singularity of SLC35B3, the clustering also finds a consistent grouping of genes highlighted with blue squares in Figure 4. In that square, across all three cerebellar representations, we find the genes LHX1, MSX1, RELN, EPHB1, and SASH1. Compellingly, these loci all have previous associations with brain morphology 60 and cognitive traits. 61 The full GWAS results for the cerebellum are available on LocusZoom: https://my.locuszoom.org/gwas/181958/.

At top left is clustering of SNPs from autoencoders trained only on the white matter of the cerebellum, in the middle from latent spaces using only the gray matter of the cerebellum, and at top right from models trained on the whole cerebellum. Bottom panel shows Manhattan plots of the three cerebellar latent spaces with distinct but overlapping architecture.
Hierarchical clustering in cross-modal spaces can identify modality-specific and modality-shared genetic clusters. For instance, working in the cross-modal ECG space, we find clusters corresponding to SNPs affecting the QT interval (SNPs associated with NOS1AP and KCNQ1) and SNPs related to the P-wave (SNPs associated with SCN10A and ALPK3), highlighted with green boxes in Figure 5 left. We also identify a large cluster corresponding to SNPs affecting multiple cardiac traits such as those associated with BAG3, SLC35F1, or GOSR2. This larger group is shared between the MRI and ECG spaces, as highlighted by the two large blue squares in Figure 5. The full GWAS results for the ECG and cardiac MRI are available at: https://my.locuszoom.org/gwas/908783/.

Hierarchical clustering of lead SNPs from the cross-modal latent space of the ECG (top left) and cardiac MRI (top right). The blue square highlights a similar clade in both models with SNPs from CASQ2, BAG3, GOSR2, NKX2-5, while the green squares highlight two clades in the ECG latent space not seen with the MRI. The bottom panel shows a Miami plot of ECG (red) and cardiac MRI (purple).
Registered modalities reveal more diagnostic variation
Phecodes are a taxonomy of billing codes aggregated into diagnostic labels to be more reflective of true disease phenotypes. 62 After determining phecode status from billing codes for the UK Biobank population, we conducted PheWAS before and after registration. In 50% of the subjects, we derived latent space vectors between the centroids of individuals with and without each diagnosis. 63 The remaining 50% of the individuals were embedded into the latent space and projected onto this vector. The resulting phecode vector component was tested for association with the phecode diagnosis from the EHR using a logit model corrected for age, sex, and race. After Bonferroni corrections for multiple testing, we show associations in QQ plots colored by phecode category with significant phecode associations labeled. Figure 6 shows a comparison between the PheWAS of the cross-modally registered cardiac MRI latent space and the cardiac MRI latent space trained in native coordinates. Registered latent spaces consistently identified more significant associations, specifically for cardiac MRIs 104 compared to 51 and for ECGs 63 compared to 2 (see Supplementary Figure 5), and for brain MRIs 38 compared to 28 (see Supplementary Figure 6).

Phenome-wide association study QQ plots showing −log10(P-value) for the cardiac MRI registered by DropFuse (104 associations) and unregistered (51 associations). Phecodes are grouped and colored by diagnostic category.
DeepCycle Groks the cardiac cycle
The most lossy registration method considered (arguably the lossiest registration possible) is with the DeepCycle convolutional encoder, which uses only a single parameter to encode the entire frame from a cardiac MRI movie. In comparison with a convolutional autoencoder without the inductive bias of periodicity, the DeepCycle MRI representations capture more biological signal, as quantified in row 3 of Table 2. Visual inspection of the DeepCycle decoder’s reconstructions reveal that the parameter,
Discussion
This work was motivated by the observation that autoencoder latent spaces trained from medical images in their native coordinate systems used much of their expressive power encoding aspects of the images of limited biological significance. The top principal components in these spaced encoded information like limb orientation in DXAs or the baseline wander of ECGs. We show that registration allows models to learn representations, which use more expressive power encoding biological information such as physiological state and genetic background. This finding is consistent across diverse registration methods, imaging modalities, and organ systems.
Notably, even lossy registrations result in more biological signals. Although much lower in overall dimensionality, registration greatly reduces intra-individual variations. This frees up the latent space to “spend” its expressive capacity representing the population variation that powers the downstream biological association studies. Still, the linear probes that we used to predict phenotypes only provide a floor on how predictable a phenotype is; nonlinear reconstruction methods might be able to recover some phenotypes that linear models cannot. More expressive explainer models, such as sparse autoencoders 65 might more cleanly predict the phenotypes present in a representation. Future methods can explicitly disentangle the learned phenotypes, for instance, by factorizing modality-specific vs modality-shared signals into separate subspaces.
Limitations
There are situations where registration can introduce bias and distort associations. Anatomical atlases constructed in one population may not be appropriate for other populations with different demographics, ancestry, or disease states. An advantage of the registration learning methods (VoxelMorph, DeepCycle, and DropFuse) over template-matching methods is that they do not require prototypical individual(s) to be selected as reference. Still, learning methods are limited by the variation present in their training data. The UK Biobank, studied here, is not representative of the world at large. Larger, more representative biobanks need to be built and existing models need to be inspected and/or corrected for potentially harmful bias. In our genetic analysis, we removed known confounds with iterative nullspace projection after model training, but this can also attenuate biological associations.
Conclusions
The best registration method for a given analysis will depend on many factors including computational resources, availability, quality and applicability of anatomical atlases, as well as the level of interpretability desired. Some registration techniques are complementary. For example, two modalities registered in space by VoxelMorph can later be cross-modally fused with DropFuse. Building up unified, biologically informative latent spaces by combining many modalities and types of registration is an exciting avenue of future research.
Supplemental Material
sj-pdf-1-bbi-10.1177_11779322241282489 – Supplemental material for Genetic Architectures of Medical Images Revealed by Registration of Multiple Modalities
Supplemental material, sj-pdf-1-bbi-10.1177_11779322241282489 for Genetic Architectures of Medical Images Revealed by Registration of Multiple Modalities by Sam Freesun Friedman, Gemma Elyse Moran, Marianne Rakic and Anthony Phillipakis in Bioinformatics and Biology Insights
Footnotes
Acknowledgements
The authors thank the UK Biobank participants for their time and generosity in providing the data for this study. Data were accessed under the Broad Institute’s UK Biobank application number 7089. The authors would like to thank Eric Lander for helpful discussions on the DeepCycle model, which motivated this work.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: SFF received funding from IBM research, Bayer Healthcare, and a Broad Ignite Award. GEM received support from the Eric and Wendy Schmidt Center at the Broad. AP is currently an employee of Google Ventures.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
SFF gathered the data, trained the models, created figures, tabulated the results and drafted the manuscript. SFF and AP concieved of the study. GEM identified related work and edited the manuscript. MR wrote the VoxelMorph section and guided the VoxelMorph implementation.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
