Abstract
Several species of the Helicoverpa genus have been recognized as major agricultural pests from different regions of the world, among which Helicoverpa armigera species has been reported as the most destructive and cosmopolitan species in most regions of the world, including Iran. This pest is a polyphagous species and can cause damage to more than 120 plant species. Studying the internal microbiome of pests is very important in identifying species’ weaknesses and natural enemies and potential biological control agents. For genomic characterization of the microbial community associated with H armigera, the whole genome of insect larvae collected from vegetable fields in the northwest of Iran was sequenced using next-generation sequencing Illumina platform. Finally, about 2 GB of raw data were obtained. Using the MetaPhlAn2 pipeline, it was predicted that 2 endosymbiont bacterial species including Buchnera aphidicola and Serratia symbiotica were associated with H armigera. Alignment of reference strains sequences related to both endosymbiotic bacteria with raw data and subsequently, assembly analyses resulted in 2 genomes with 657 623 bp length with GC content of 27.4% for B aphidicola and 1 595 135 bp length with GC content of 42.90% for S symbiotica. This research is the first report on the association of B aphidicola and S symbiotica as endosymbiotic bacteria with H armigera worldwide.
Introduction
The cotton bollworm (Helicoverpa armigera) is an insect species belonging to the Noctuidae family in the Lepidoptera order, which its larva feed on a wide range of plants. It is one of the most devastating pests of cotton around the world. The cotton bollworm is a polyphagous species. The larvae can cause damage to more than 120 plant species, including cotton, alfalfa, tomato, potatoes, tobacco, maize, rice, sorghum, several peas, vegetables, and several ornamental plants. In Iran, the cotton bollworm is an economically destructive pest and can damage especially in cotton and tomato. 1
So far, at least 4 species of the genus Helicoverpa have been identified in Iran with overlapped geographical distributions in different regions of Iran, which sometimes makes it difficult to accurately identify the H armigera species, especially at the larval stage. 2 Molecular identification is a simple and very accurate way to distinguish species, but so far no one has attempted molecular identification of species of the genus Helicoverpa in Iran. On the contrary, like other animals, different types of microorganisms can be found in the body of insects, especially in their digestive system, which may have different relationships with the physiology of insects, such as symbiosis or pathogenicity. 3 The symbiosis interaction between insects and bacteria was comprehensively studied in recent years. 4 Bacterial endosymbionts are very common in insects. 5 The effects of endosymbiont bacteria on their insect host are very diverse such as protection against enemies, effects on the reproductive system, pheromone biosynthesis, and impact on the host’s genetic diversity. 6
Most endosymbiont bacteria are heritable and could transmit maternally to the next. 7 The endosymbiotic bacteria can be divided into 2 different groups including obligate or primary symbionts and facultative or secondary symbionts. 4 Primary symbionts have a nutritional function and occur in insects which feed on imbalanced diets such as plant saps or cellulose; therefore, they are mutualists. 8 However, facultative symbionts have several effects, ranging from mutualism to manipulation of reproduction. 9 Often several bacterial species occur simultaneously within the same host. The existence of multiple species is required for the survival of the insect while other associations are facultative. 4 So far, several gut bacteria have been isolated from H armigera, including Bacillus sp., 10 Corynebacterium sp., 11 Enterococcus sp., Lactococcus sp., Flavobacterium sp., Acinetobacter sp., and Stenotrophomonas sp. 12 using the cultural method and ribosomal RNA (rRNA) sequencing analyses. The role of many of these bacteria in the insect life cycle remains unknown; however, colonization of those dominant noninfectious microbes may restrict gut colonization by pathogens. 13 Some of these bacteria may act as endosymbionts; however, it has not yet been proven that these bacterial species act as a symbiont inside the insect’s body. 14
This study was conducted for molecular confirmation of the presence of H armigera in northwestern Iran and also for identification of the internal symbiotic bacteria of this insect in the region. Characterization of these symbiotic bacteria can be an introduction to understanding the reason for the wide host range of this insect and can be useful in the biocontrol approach. Identification of these bacteria helps scientists and plant pest management to better understand the life cycle of this pest and help to find new ways to control it.
Materials and Methods
Insect sampling and molecular identification
During the summer of 2021, several vegetable fields, including tomato, eggplant, and pepper, in East Azerbaijan province in the northwest of Iran were investigated to collect living and healthy H armigera larvae. About 52 instar larvae were separately collected from several vegetables. Due to the distribution of various species of the Helicoverpa genus (such as H peltigera, H zea, H viriplaca, and H armigera), accurate molecular identification of collected larvae was deemed necessary.
DNA was extracted from specimens using CTAB extraction buffer in methods described by Jangra and Ghosh with minor modifications. 15 The 10-mL extraction buffer consisted of 2.8 mL of 5M NaCl, 3.5 mL of 10% CTAB, 1 mL of 1M Tris-HCl, 400 μL of 0.5M EDTA (pH 8.0), 20 μL β-mercaptoethanol, and 2.6 mL sterile distilled water. About 2 g of surface-sterilized insect bodies was moved in a 2-mL tube and crushed using a mortar and pestle. Then, 1.5-mL extraction buffer was added to the microtubes. The tubes were incubated at 65°C for 1 hour and vortexed at every 10-minute interval. Then, an equal volume of isoamyl alcohol:chloroform (1:24) was added to microtubes and centrifuged at 10 000× for 15 minutes. The upper aqueous layer was transferred to a new micro-centrifuge tube and 0.8 volume of cold isopropanol was added and kept at −20°C for 10 minutes. The DNA was pelleted by centrifuging at 10 000× for 10 minutes. Then DNA pellet was washed with 70% ethanol. The pellets were dissolved in 30 μL sterile distilled water, kept overnight at 4°C, and stored at −20°C until use. DNA qualities and quantities were determined using spectrophotometry (UV-Vis 1280) by measuring the absorbance at 260 nm and ratios of 260 nm/230 nm and 260 nm/280 nm.
The extracted DNA was used for polymerase chain reaction (PCR) assays. The primer sets of 3373Ha_Hz_ITS1F (5′-gaggaagtaaaagtcgtaacaaggtttcc-3′) as forward primer and 3374Ha_ITS1-R (5′-cgttcgactctgtgtcctctagtgg-3′) as reverse primer for H armigera detection were used. 16 For PCR assays, the reactions consisted of 1 cycle at 95°C for 1 minute, followed by 40 cycles at 95°C for 15 seconds, 52°C for 10 seconds, and 72°C for 4 minutes, and a final extension step at 72°C for 5 minutes.
The PCR reaction mixture in 15 μL contained 5 μL of template DNA, 1 μL each of primer pairs (20 pmol), and 9 μL Ampliqon Taq DNA Polymerase Master Mix RED (Ampliqon, Odense, Denmark). After amplification, each PCR product was subject to electrophoresis in 1% agarose gel in 1× TAE buffer and stained with safe stain DNA Green Viewer. The PCR bands were visualized in UV gel documentation. Finally, the PCR product was purified and sequenced directly in Microsynth Company (Balgach, Switzerland). After sequencing, consensus sequences were generated from raw data files using MEGA-X software and a BLASTn search was performed for finding similar subjects in NCBI GenBank. The phylogenetic tree was drawn using the neighbor-joining algorithm with 1000 repetitions (bootstrap) The resulting sequences have been submitted to GenBank (NCBI) to get accession numbers.
Next-generation sequencing analyses
The DNA extraction from H armigera was performed in 3 replicates, and each replicate yielded about 200 µl of DNA with a concentration of 25 ng/µL. The DNA samples were purified and sent to Novogene, China, for library preparation and sequencing on the Illumina 1.9 Novaseq 6000 platform, generating paired-end reads. The quality of the Illumina raw sequencing data was checked using FastQC software version 0.73. To predict bacterial genomes and profile the composition of microbial communities, the MetaPhlAn2 pipeline version 2.6.6.17 was applied to the raw data.17, 18
To assemble bacterial genomes, the raw data were mapped to the reference genomes that had been previously deposited in GenBank, using Bowtie2 software version 2.5.0. The mapped reads were then assembled by metaSPAdes software version 3.15.4. To cluster the nucleotide sequences and improve the performance of sequence analyses, CD-HIT software version 4.8.1 was used to cluster the contigs. Finally, the assembled genomes were submitted to GenBank and assigned accession numbers, which are listed in Table 1. The genomes were annotated by 1 methods: (1) using the PGAP (Prokaryotic Genome Annotation Pipeline) in the NCBI database, which automatically trimmed and annotated the genomes before release, and (2) using the GhostKOALA online tool, which assigned KEGG Orthology (KO) terms to the predicted protein sequences. The protein sequences were predicted from the contigs using MetaGeneMark online program. The GhostKOALA annotation results for the assembled genomes are shown in Table 2, and an overview of the genomes annotation is presented in Figure 1.
Genomic characteristics of endosymbiotic bacteria genomes obtained from Helicoverpa armigera..

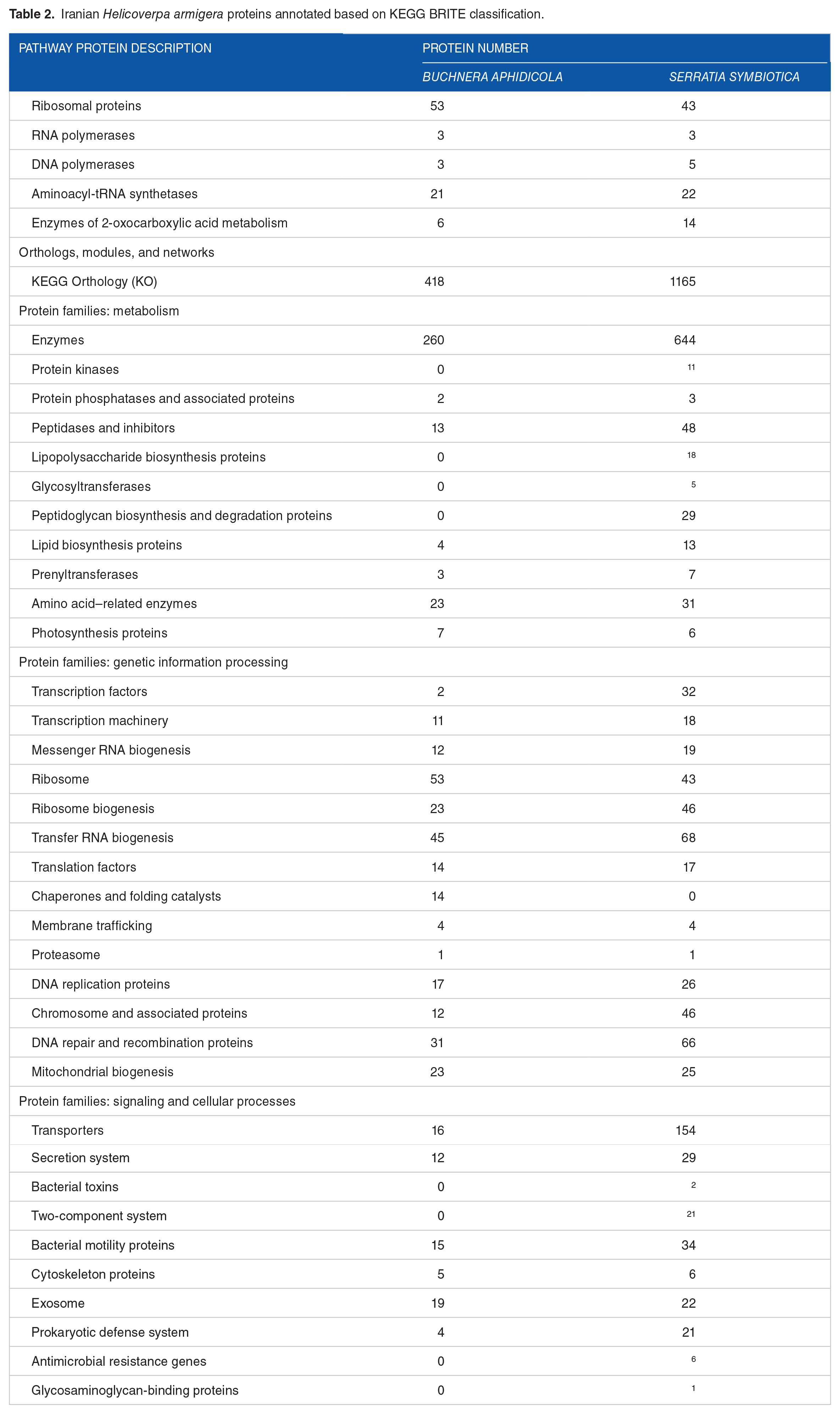
Iranian Helicoverpa armigera proteins annotated based on KEGG BRITE classification.
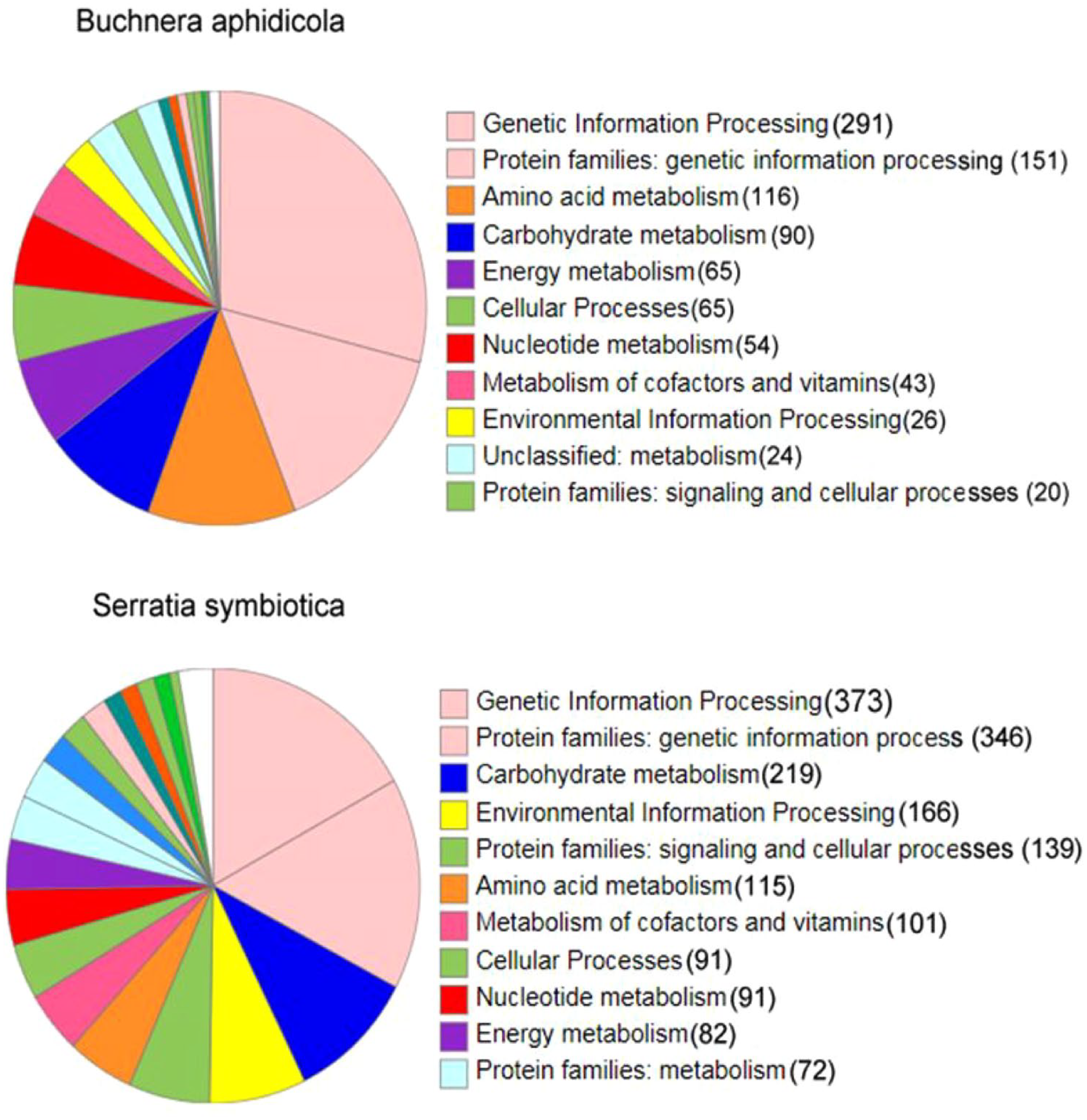
Functional category and pathways for annotated proteins of Buchnera aphidicola and Serratia symbiotica genomes.
Results
Insect collection and identification
Helicoverpa armigera is a polyphagous species, but in the northwest of Iran, its greatest damage is on vegetables, especially tomatoes, eggplants, and pepper, etc, so several vegetable fields in the Northwestern region of Iran were investigated for detecting H armigera damage. About 52 larvae were collected that could belong to H armigera. To ensure that the larvae belonged to the species H armigera, the extracted DNA was subjected to PCR using genus-specific primer pairs. The PCR results confirmed that the collected larvae belonged to H armigera. Finally, the Iranian H armigera rRNA gene sequence was submitted to GenBank under accession number OP806113.1.
Next-generation sequencing analyses
To characterize all bacteria accompanying H armigera causing damage to vegetables in East Azerbaijan, the DNA extracted from larvae was subjected to next-generation sequencing (NGS) using Illumina platform. Finally, raw data with 2 GB size and 21 178 248 read pairs were obtained from each replication. Each raw read length was 150 bp and the insert size was 350 bp. The GC content of the whole genome was about 30%.
The MetaPhlAn2 analyses results
In the first step, to estimate the bacterial population inside H armigera genome, the NGS raw data were subjected to MetaPhlAn2 analyses. The prediction results indicated that 2 endosymbiont bacterial species, B aphidicola and S symbiotica, existed in high quantity inside H armigera raw data, in which 3.8% of genome contents belonged to S symbiotica and about 4.2% of genome contents belonged to B aphidicola. Both identified bacteria are considered gram-negative and from Enterobacterales, gammaproteobacterial group.
Genomic characterization of endosymbiont bacteria associated with H armigera
The MetaPhlAn2 analysis results predicted 2 bacterial species in the raw sequencing data. To prepare the bacterial genomes, the raw data were mapped to a reference sequence of 653 353 bp, comprising the chromosome sequence (accession NZ_CP029205) and 2 plasmid sequences (accession NZ_CP029203 and NZ_CP029204) of B aphidicola isolate LNK. The assembly and clustering of the mapped reads produced 1224 contigs with a total length of 657 623 bp and a GC content of 27.4%. Similarly, the raw data were mapped to a reference sequence of 3 550 913 bp, consisting of the chromosome sequence (accession NZ_CP050855) and 2 plasmid sequences (accession NZ_CP050856 and NZ_CP050857) of S symbiotica strain CWBI-2.3. The assembly and clustering of the mapped sequences yielded 4308 contigs with a total length of 1 595 135 bp and a GC content of 42.90%.
Genome annotation
The NCBI PGAP annotation identified 2810 and 1050 proteins in the genomes of S symbiotica and B aphidicola, respectively. A further annotation analysis was conducted using the GhostKOALA online server to obtain more detailed information about the bacterial genomes. The GhostKOALA annotation results for the S symbiotica genome showed that 2196 proteins (about 78% of all proteins) were fully annotated and assigned KO numbers. In addition, 394 proteins received a second KO number and 220 proteins remained uncharacterized. The GhostKOALA annotation results for the B aphidicola genome revealed that 1003 proteins (about 95.5% of all proteins) were fully annotated and assigned KO numbers. Moreover, 37 proteins received a second KO number and 10 proteins remained uncharacterized. A summary of the genomes and annotation results is shown in Figure 1 and Table 2.
Each replicate generated about 2 GB of raw data. The quality of the raw data was evaluated using FastQC software version 0.73. To predict bacterial genomes and profile the composition of microbial communities, the MetaPhlAn2 pipeline version 2.6.6 was applied to the raw data. The prediction results indicated that 2 endosymbiotic bacterial species, B aphidicola and S symbiotica, were present in high abundance in the H armigera raw data.
Discussion
The cotton bollworm is a polyphagous pest which could damage up to 70% of several crops in Iran. The most important host of the pest is cotton which is cultivated in several regions in Iran. 19 Regarding wide host ranges of the pest on several crops and the very diverse weather conditions in different regions of Iran, it seems that the insect has found a high adaptation to the diverse climate of the country. One of the effective factors in creating adaptation to environmental conditions in insects is endosymbiotic bacteria. Endosymbiosis in insects is common. The effects of endosymbiotic bacteria in their insect host are including protection against natural enemies, providing essential nutrients, reproductive system process, etc. Endosymbiotic bacteria are usually transmitted vertically and maternally. Insect-bacterial symbiosis allows faster adaptation to change environments. 4
Like other insect species, H armigera shared a symbiotic relationship with microorganisms. However, it was previously reported that a densovirus was associated with insect populations which increase larval and pupal development rates, female lifespan and fecundity that increase larval resistance to baculovirus and biopesticides, Bacillus thuringiensis toxin. 3 Any other organism was not reported as a special symbiont for H armigera up to now.
In this research, the H armigera was collected from vegetable fields in the northwest of Iran. To characterize endosymbiont bacteria associated with insect, the DNA extracted from larvae were subjected to NGS analyses. The MetaPhlAn2 prediction indicated that 2 well-known endosymbiotic bacteria genomes, including B aphidicola and S symbiotica, existed inside the insect genome.
The B aphidicola has been reported as a free-living bacteria belonging to the Gammaproteobacteria class which was previously described as the obligate endosymbiont of aphids and has been isolated from the pea aphid (Acyrthosiphon pisum). 20 An adult aphid may carry more than 5 × 106 Buchnera cells. Aphids have developed a bilobed bacteriome containing 60 to 80 bacteriocyte cells; each bacteriocyte can contain multiple vesicles, symbiosomes derived from the cell membrane. 4 The Buchnera also can increase the transmission rate of plant pathogenic viruses by producing symbionin, a protein that binds to the viral coat and protects it inside the aphid. This protein makes the viral particles survive longer and be able to infect another plant when the aphid feeds. 21 The B aphidicola was reported as aphids’ primary endosymbiotic bacteria; there is not any report of its association with other insects up to now. This study is the first report on the association of B aphidicola with H armigera worldwide.
On the contrary, using NGS analyses another bacteria S symbiotica was found in Iranian H armigera bodies. The S symbiotica is a facultative anaerobic bacteria belonging to Gammaproteobacteria class 22 which was isolated from Aphis fabae for the first time 23 and lives as a symbiont in aphids’ bodies which coexist with B aphidicola. The S symbiotica helps insect cells to produce tryptophan 24 and improves host resistance to heat stress and wasp parasitism. 25 The S symbiotica is one of the main 3 facultative or secondary endosymbiotic bacteria which is found along with B aphidicola in aphids. 14 Although its existence is not very necessary for the preservation of the Buchnera-aphid association, S symbiotica is dependent on B aphidicola for amino acid provision when they infect insects. 24 The association of S symbiotica with H armigera is reported for the first time too.
In another genome annotation method, through KEGG Mapper reconstruction, for B aphidicola, 482 metabolism pathways including 74 amino acid metabolism, 61 carbohydrate metabolism, 53 energy metabolism, 38 metabolisms of cofactors and vitamins, 25 nucleotide metabolism, 5 lipid metabolism, 14 metabolisms of other amino acids, 2 glycan biosynthesis and metabolism, 3 metabolism of terpenoids and polyketides, 11 biosyntheses of other secondary metabolites pathways were recognized. Also, 3 RNA polymerases; 53 ribosomes; 22 aminoacyl-tRNA biosynthesis proteins; 14 folding, sorting, and degradation proteins; 35 replication and repair proteins; 4 ABC transporters; 5 phosphotransferase system (PTS) proteins; 7 bacterial secretion system proteins; 12 signal transduction proteins; 1 peroxisome; 8 cell growth and death proteins; 10 quorum sensing proteins; 8 biofilm formation proteins; 17 cell motility proteins; and 100 other proteins were recognized.
On the contrary, for S symbiotica, 482 metabolism pathways including 176 carbohydrate metabolism, 123 amino acid metabolism, 95 metabolisms of cofactors and vitamins, 94 energy metabolism, 64 ABC transporters, 13 PTS proteins, 60 nucleotide metabolism, 30 lipid metabolism, 27 metabolisms of other amino acids, 55 glycan biosynthesis and metabolism, 16 metabolism of terpenoids and polyketides, 28 biosyntheses of other secondary metabolites pathways were recognized. Also, 3 RNA polymerases; 43 ribosomes; 22 aminoacyl-tRNA biosynthesis proteins; 40 folding, sorting, and degradation proteins; 65 replication and repair proteins; 14 bacterial secretion system proteins; 65 signal transduction proteins; 1 lysosome; 2 peroxisomes; 14 cell growth and death proteins; 20 quorum sensing proteins; 38 biofilm formation proteins; 37 cell motility proteins; and 98 other proteins were recognized.
Conclusions
Both B aphidicola and S symbiotica were, respectively, characterized as primary and secondary endosymbiotic bacteria in aphids at present. In this study, we performed the NGS method to determine endosymbiotic bacterial genomes inside H armigera genome. The NGS method can be used as a very effective method for the detection of bacterial genomes with a very low population or fastidious and nonculturable bacteria inside the host genome. The results obtained in this research indicated that 2 endosymbiotic bacteria, B aphidicola and S symbiotica, which were previously detected only in aphid bodies were present inside lepidopteran insect bodies. The mechanism of action and multiplication of B aphidicola and S symbiotica inside the H armigera body is still unknown, but it is similar to the action of these bacteria inside the body of the aphids. This research is the first report from B aphidicola and S symbiotica association with H armigera worldwide. This research provides valuable data for elucidating the ecology and physiology of this major pest. The genome features of these symbiotic bacteria can facilitate genetic engineering applications and reveal insights into the intricate microbiome within insects.
Footnotes
Acknowledgements
We thank the Plant Bacteriology Lab of the University of Tabriz for providing laboratory equipment for this research.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
PS has done lab works; MJ has done conceptualization and supervision; SN was advisor for this work; RK has done writing, editing, validation and has acted as project administration. All authors have read and agreed to the published version of the manuscript.
