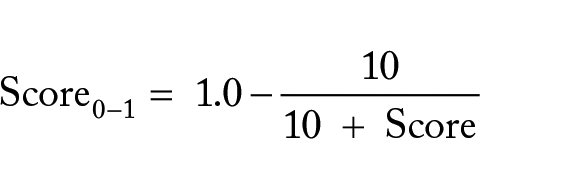Abstract
Ovarian clear cell carcinoma (OCCC), one of the histopathological types of ovarian cancer, has a poor prognosis when it recurs; however, it is difficult to precisely predict the risk of recurrence. Here, we analyzed pathological images of OCCC to elucidate the relationship between pathological findings and recurrence, and using machine learning, we established a classifier to predict the recurrence and several other prognosis indicators of this disease. In total, 110 patients with OCCC treated with primary surgery at a single institution were enrolled in this study. We used the deep-learning neural networks to process the whole slide images of OCCC obtained by digitally scanning the original hematoxylin and eosin-stained glass slides. The images were preprocessed and used as input to the machine learning pipeline. We fine-tuned its parameters to predict the recurrence, progression-free survival, and the overall survival days of all patients. We predicted the recurrence of OCCC with an overall accuracy of 93%, area under the receiver operating characteristic curve of 0.98, and sensitivity/specificity above 0.92 using Resnet 34. Furthermore, we predicted progression-free survival/overall survival of the patients with ~90% accuracy. In conclusion, our study demonstrates the feasibility of using a machine learning system to predict different features of OCCC samples using histopathological images as input. This novel application provides accurate prognosis information and aids in the development of personalized treatment strategies.
Keywords
Introduction
Ovarian cancer is the eighth most common cancer and the eighth most common cause of death from cancer in women worldwide. 1 It is estimated that there were over 313,000 cases of ovarian cancer and almost 207,000 deaths worldwide in 2020. 1 Epithelial ovarian cancer (EOC), the most common type of ovarian cancer, is classified according to its histopathological profile into 5 major distinct subtypes: high-grade serous, low-grade serous, mucinous, endometrioid, and clear cell carcinoma. According to the World Health Organization (WHO) classification of tumors of the ovary in 2014, different histotypes exhibit different morphology, etiology, and biological behavior. 2 Along this line, EOC is surgically and pathologically categorized by the International Federation of Gynecology and Obstetrics (FIGO) system, and the current standard of care consists of either primary debulking surgery, or interval debulking surgery following neoadjuvant chemotherapy. 3 Both histopathology and FIGO staging are considered the gold standard for classification of EOC subgroups and are relevant prognostic factors for stratification. 4 Many studies have explored the molecular mechanisms underlying this malignancy,5,6 however, accurate prediction of treatment outcome and anticipated personalized prognosis is still lacking.7,8
Among the histotypes of EOC, ovarian clear cell carcinoma (OCCC) exhibits unique clinicopathological and molecular features, and therefore is recognized as a distinct entity, presenting unique challenges for treatment. 9 Among its features, OCCC tends to be diagnosed at an earlier stage than ovarian serous carcinoma, with 57% to 81% and 19% to 22%, respectively, presenting at stages I or II.10-12 Patients with OCCC usually present with a large, unilateral pelvic mass, and occasionally have thromboembolic vascular complications or hypercalcemia.13,14 The prevalence of OCCC varies geographically; while accounting for only 1% to 12% of EOC cases in Western countries, OCCC is relatively frequent among Asian women, accounting for 25% to 30% and 10.3% to 11.6% of EOCs in Japan and Korea, respectively.15-17 Owing to its early diagnostic, the 5- year survival rate is relatively favorable in early stage (~90%),18-21 on the other hand, advanced stage cases result in prognoses worse than those for high-grade serous carcinoma, a major histotype in ovarian cancer.11,22 However, some patients show poor prognosis even after detection at early stages of OCCC, highlighting the need of novel and accurate prognosis prediction tools.
Common statistical methods familiar to clinicians are ill-suited for handling complex information; until recently, this has been a major limitation that prevented the extraction of meaningful patterns from large data sets with multiple input variables. Machine learning, a branch of artificial intelligence (AI) technology that allows computers to identify patterns from complex and large sets of past examples, is expected to achieve better performance. In the field of cancer research, a large body of evidence demonstrates that application of deep learning algorithms has the potential to improve diagnostic accuracy.23,24 Particularly, in ovarian cancer research, a study employing machine learning provided critical diagnostic and prognostic prediction for patients with EOC even before initial intervention. 25 Therefore, we hypothesized that AI technology can contribute to establish a more sophisticated prognosis prediction model for OCCC.
In this study, we developed an OCCC-specific prognosis prediction classifier based on machine learning algorithms using pathological tumor images of patients with OCCC. We named this system O3C Glass-Class (“Glass” comes from the fact that the input for the system are pathology images from glass slides). We used this system to predict the recurrence of OCCC and obtained an overall accuracy of 93%, area under the ROC curve (AUC) of 0.98 and sensitivity/specificity above 0.92. Moreover, we found that only a few images are necessary for accurate predictions, an aspect that is helpful for treatment centers without access to abundant resources. Importantly, we designed our machine learning framework as an open-source application, with an internal architecture that can be easily extended or modified for other purposes. In summary, this study demonstrates the feasibility of using a machine learning system to predict the prognosis of OCCC, and serves as a platform for cancer-related analyses involving whole-slide histopathological images.
Materials and Methods
Patients and clinical samples
In this study, we enrolled patients with OCCC who were treated by primary surgery with/without chemotherapy between 2000 and 2012 at the Department of Obstetrics and Gynecology, The Jikei University School of Medicine, Tokyo, Japan. We excluded patients with insufficient clinical data and those without pathological image for analysis. All available hematoxylin and eosin-stained slides of the OCCC were reviewed by a gynecological pathologist to confirm the histological tumor type based on the WHO classification of 2014, 2 and tumors were staged in accordance with the FIGO system (2014). We also retrospectively reviewed medical records in 2019 and investigated clinicopathological parameters: age at diagnosis, ascites volume, ascites or peritoneal washings cytology, surgical method, chemotherapy administration, patient progression-free survival (PFS), and patient overall survival (OS). Patients who have achieved remission following initial therapy and who subsequently experience a return of cancer cells after treatment were defined as recurrence. Progression-free survival was defined as the duration from the date of surgery to the last follow-up without progression or the date when diagnosis of recurrence was established. Overall survival was defined as the interval between the date of surgery and the date of death if the patient died of disease or the last follow-up date if the patient was still alive with or without disease. Optimal surgery for OCCC included total abdominal hysterectomy, bilateral salpingo-oophorectomy, subtotal omentectomy, and pelvic and para-aortic lymphadenectomy. Suboptimal surgery provided less than optimal staging except as a fertility-sparing surgery. The Ethics Review Committee of The Jikei University School of Medicine and National Center for Child Health and Development approved this study protocol (31-253 [9752], 2020-157).
Pathological image acquisition
Whole slide imaging for OCCC cases were obtained by digitally scanning the original hematoxylin and eosin (HE)-stained glass slides at 20× magnification using an Aperio AT2 scanner and a ScanScope digital pathology system. The images were saved using the SVS format (Leica Biosystems; Buffalo Grove, IL, USA).
Image processing
To process the images used as input for the machine learning classifier, we scaled each image to reduce their size 32 times. We applied filters to remove annotation marks (e.g. green, blue, or red pens), removed small dirt and cropped each image into 255 × 255 pixels tiles. Next, we ranked the images using two criteria, first, according to their tissue percentage composition, measured by the difference between colored pixels and the background color; second, by a score adapted from https://github.com/CODAIT/deep-histopath (visited in January 2021).
where t is the tissue percentage in the tile, c is the color factor, s is the saturation value factor, and q is the quantity factor (i.e. the more tissue a tile has, higher will be the score). Next, we normalize the score in the [0, 1] interval by calculating
Machine learning
We divided our image tiles into training, validation, and test sets (70%, 15%, 15%, respectively), making sure that the tiles from each patient were used only for training or testing, to eliminate the risk of training and predicting tiles from the same patient. To balance the data sets used for training, we randomly selected the same number of patients for both classes (e.g. 15 patients with or without relapse cases), and repeated this procedure at least 10 times, with different images used for training, validation, and testing. We used the pre-trained deep learning neural network Resnet 18, 34, 50, and 101, 26 then trained for 30 epochs, aiming to maximize the F-Score measure, using the Adam optimizer, with maximum learning rate of 0.01, clip gradient of 0.1, and weight decay of 10–4. Finally, the predictions of the different tiles from the same patient were combined using the median value of the probability prediction of each tile. We used Python version 3.6 and the Pytorch framework version 1.1. 27
Statistical analysis
Wilcoxon rank sum test was used to compare differences of continuous variables in two groups. Characteristics between two groups were compared using the Pearson’s chi-square test with Yates’ continuity correction for categorical variables. A two-sided P value of less than .05 was considered to be statistically significant for all analyses. The R statistical package version 3.6.2 (https://www.r-project.org/) was used for the statistical analyses.
Results
Clinicopathological characteristics in OCCC
One-hundred thirteen patients with OCCC were enrolled in this study, then 2 patients with insufficient clinical data and 1 patient without available pathological image were excluded, we finally investigated the data from 110 OCCC patients for analysis. The clinicopathological characteristics of the OCCC patients are summarized in Table 1. Patient age was 51.6 ± 10.5 years (mean ± standard deviation) and 75.5% of them were at early stages of the disease (stages I and II). Of 110 OCCC patients, 76 patients were optimally staged and underwent systematic lymphadenectomy, whereas 28 and 6 patients underwent suboptimal and fertility-sparing surgery, respectively. Ninety-seven patients received more than 3 cycles of platinum-based adjuvant chemotherapy, and the follow-up period of our cohort was 2794.1 ± 1777.4 days (mean ± standard deviation).
Clinicopathological parameters of the patients with ovarian clear cell carcinoma.
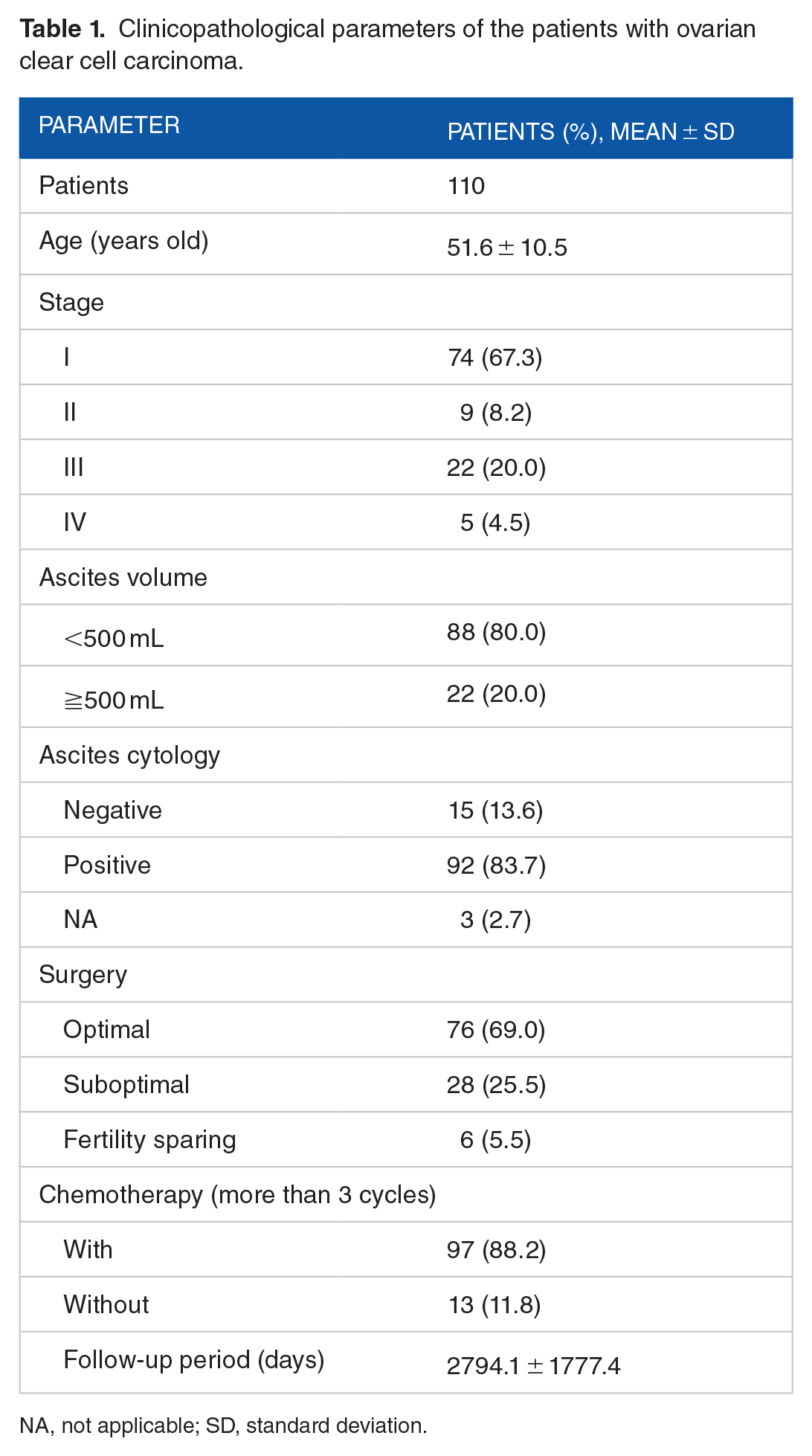
NA, not applicable; SD, standard deviation.
Machine learning classifier predicts multiple cancer features
For patients, caretakers, and their families, it is essential to understand what are the chances of relapse, and the overall prognosis and survival at initial diagnosis. In addition to the decisions about treatment options and further clinical steps, this information also has a critical influence on the patients’ quality-of-life; however, for OCCC and other forms of cancer, strategies to accurately predict the relapse of these malignancies are not well-established. To address this issue, we developed a customized AI pipeline that receives as input HE-stained images from OCCC samples and outputs several features related to disease outcome (Figure 1A). We applied several filters to the input images, cropped them into smaller tiles, and ranked the tiles according to two criteria: (1) a score quantifying the image composition and diversity and (2) according to the tissue percentage in each tile (Figure 1B and C).

The O3 C Glass-Class AI pipeline: (A) from each patient, we obtained a tumor sample with hematoxylin and eosin staining. Next, we obtained a digital image and applied a series of filters to improve the overall quality of the sample and to remove unwanted characteristics (e.g. manual annotations and noise). We cropped each whole-slide image into smaller tiles (n = 10, 50, 100, 500, 1000), and ranked them either by tissue composition or by a score, quantifying the overall quality and quantity of information displayed by each tile. Finally, we subject the tiles to the deep-learning neural network ResNet 34), 26 using images from 70% of patients for training, 15% for validation (i.e. parameter optimization), and 15% for testing the accuracy of the model. We repeated this procedure at least 10 times to assess the robustness of the model. (B and C) Example tiles ranked by (B) tissue composition or by (C) score, and associated either with recurrence or no recurrence of the disease.
We divided the dataset into two groups in the view point of whether recurrence was observed or not, and trained a deep learning framework using as input images from 110 patients (1 image per patient, cropped into smaller tiles) (Table 2). We used a strict training approach, dividing the data into training, validation and test sets, and optimizing the algorithm parameters only on the validation set; we repeated this routine 10 times to ensure independence of the combination used for training and testing, the results were always consistent.
Clinicopathological parameters of patients with ovarian clear cell carcinoma with or without recurrence.

NA, not applicable; SD, standard deviation.
Using this setup, we predicted the recurrence of OCCC and obtained an overall accuracy of 93%, area under the receiver operating characteristic (ROC) curve (AUC) of 0.98 and sensitivity/specificity above 0.92 using Resnet 34. The AUC indicates the trade-off between detecting true-positives while rejecting and false-positives. The sensitivity measures the proportion of relapse cases that were correctly identified, and the specificity measures the proportion of nonrelapse cases correctly identified (Figure 2A to C). Overall, these results indicate that the HE-stained histological images contain information that can be used to assess the odds of disease relapse, and the similarity of the accuracy obtained even using a different number of tiles and ranking criteria attest to the consistency of the O3C Glass-Class framework. In this process, we compared the performance of other classifier including Resnet 18, 50, and 101, then found that Resnet 34 showed the best performance compared with other three classifiers (Supplemental Figure 1). Considering this result, we decided to use Resnet 34 for the further analyses.

Results of the classification: (A) prediction of recurrence or no recurrence of the disease. The comparison of the accuracy obtained by the deep-learning framework when using a different number of tiles obtained from patient tumor hematoxylin and eosin staining and ranked either by tissue percentage composition or by a quality score (“Methods”). (B) The receiver operating curves, depicting the overall classifier performance. Values close to (0,1) are ideal, because it indicates perfect classification—namely, no mistakes for both classes. (C) The sensitivity/specificity curves, showing the ability of the deep-learning model to correctly classify instances from both classes without making mistakes (values close to 1,1 are ideal). (D to F) Using the same indicators to predict the progression-free survival (PFS days), and (G to I) the overall survival of the patient (OS days).
Next, we wondered if our machine learning system could be used to predict the PFS of the cohort participants. For this purpose, we divided the data set into 2 groups by median of PFS, namely, short PFS (under 2788 days) and long PFS (longer than 2788 days; Table 3). This arrangement of exactly half of our data assigned to each group helped us avoid a notorious problem in machine learning (i.e. class imbalance), that often bias classifiers and lead to inflated but incorrect results. Once more, we compared different setups, including the number and the criteria for ranking the tiles (Figure 2D to F) and could predict whether patients are more likely to have a short- or long-PFS outcome with ~93% accuracy (AUC: 0.98, sensitivity/specificity 0.93). Interestingly, we observed that while only 50 tiles were enough to yield accurate predictions of the disease recurrence, the best results for predicting the PFS were obtained with 1000 tiles, indicating that more information is necessary for the machine learning framework to correctly determine the long or short progression of OCCC.
Clinicopathological parameters of patients with ovarian clear cell carcinoma with short- or long PFS days.

NA, not applicable; PFS, progression-free survival; SD, standard deviation.
Finally, we challenged the machine learning algorithm to predict the OS of the cohort participants. Similar to the processing we performed for the PFS days, we also divided the input data by median of OS as short OS (less than 3012 days) or long OS (more than 3012 days) (Table 4). We found that either ranking the image tiles according to tissue percentage (50 tiles), or by composition score (1000 tiles), yielded good accuracy values—in both cases, accuracy of ~90%, and AUC of ~0.95 (Figure 2G to I). This finding was particularly interesting because it suggests that there are multiple signs in the HE-stained images that indicate the severity of the disease.
Clinicopathological parameters of patients with ovarian clear cell carcinoma with short- or long-OS days.

NA, not applicable; OS, overall survival; SD, standard deviation.
Taken together, these results indicate the feasibility of using a machine learning system to predict different features of OCCC samples, using histopathological images as input. While these disease characteristics are often difficult for specialists to detect at initial assessment and are basically only determined in a retrospective manner, the machine learning algorithms enables doctors to prospectively identify patterns that accurately determine the progression and outcome of this malignancy.
Discussion
In this study, we established a machine learning prognosis prediction tool for a specific subtype of ovarian cancer (OCCC) that is very common in Asian countries. The input for this system was the patient’s tumor sample stained with hematoxylin and eosin. Our system (named O3C Glass-Class) was able to process whole-slide images and accurately predict the recurrence, PFS, and OS days of more than 100 patients.
To date, in addition to several other types of cancer,7,25,28 different approaches for the diagnostic and prognosis of patients with ovarian tumors were devised. For instance, a combination of serum tumor markers and age, with or without ultrasound findings was developed to predict ovarian cancer in patients with adnexal masses, 29 and another study reported that preoperative serum C-reactive protein (CRP) levels offer additional evidence to CA125 in the differential diagnosis of ovarian tumor. 30 Together, these studies highlight that the approaches using AI are important tools in the realm of precision medicine. However, the development of machine learning algorithms that provide accurate prognostic for OCCC was difficult due to its low occurrence worldwide. Owing to a high occurrence of OCCC in Japan, we have an access to a relatively large number of samples, leveraged on this resource to strengthen the training of the O3C Glass-Class framework and developed a framework that is specific and fine-tuned for the prognosis of this malignancy.
It is well accepted that residual disease following primary debulking surgery strongly correlates with patient survival and that complete gross cytoreduction to no residual disease status appears to be associated with the best overall outcomes in ovarian cancer. 31 Although primary surgery followed by platinum-based chemotherapy improves prognosis in early-stage OCCC, the risk of relapse remains substantially. Even in stage I patients, there are microdisseminations those are not detected by general imaging studies or pathological examinations. 19 Hence, it is clear that the prediction of recurrence independently of clinical stage has a profound impact on the treatment decision-making.
In this sense, O3C Glass-Class enables gynecologists to obtain high-quality information about the patient’s ill-status and support doctors and patients in devising a personalized treatment plan. At present, we found only a handful of studies specific for ovarian cancer, aiming to establish predictive models using machine learning. In a study for risk prediction model for ovarian cancer, the algorithm that used XGBoost analyzing gene signature showed a sensitivity of 74% to 100%. 32 In another study for diagnosis of ovarian cancer, the algorithm that used XGBoost analyzing results of preoperative examinations showed an AUC of 0.80. 33 Furthermore, high accuracy and AUC (92.4% and 0.968, respectively) for segregating EOC from benign ovarian tumors were obtained by Random Forest. 25 Strict comparison is impossible because our study focused on prognostic prediction of OCCC using pathology images; however, we believe that the accuracy and AUC obtained in our study is comparable or better. Recently, studies in other type of cancer have emerged to predict genetic abnormalities from pathological image, 34 and we are planning to develop the study with the same concept in OCCC.
Furthermore, information other than histopathological images was not necessary for our prediction model, an aspect that facilitates an efficient collaboration between pathologists and clinicians to devise treatment strategies. Although further testing is required before this framework is adopted in clinical settings, we anticipate that the O3C Glass-Class will allow the identification of OCCC patients who need more aggressive treatment due to the possibility of recurrence.
Evidently, every new technology has limitations. As it happens with other deep-learning frameworks, the O3C Glass-Class still lacks a proper explanation of the reasons underlying classifications; in the machine learning field, this is an ongoing research topic. In addition, we did not incorporate other clinical features known to be predictors of favorable or unfavorable prognosis of EOC (e.g. monocyte to lymphocyte ratio, 35 and plasma levels of D-dimer, fibrinogen and platelets). 36 Together with genomic information and blood plasma biomarkers, this will likely enhance the predictive power of O3C Glass-Class even further. Development of therapeutic strategy for OCCC in clinical practice and precise pathological diagnosis of OCCC would be acknowledged as a limitation for application of O3C Glass-Class. Continuous accumulation of real-world data with accurate diagnosis is critical for prediction of the prognosis of OCCC. Furthermore, in this study, we recruited only Japanese patients. Future studies involving a larger population will uncover the role played by ethnic differences in the prognosis of OCCC and will enable the O3C Glass-Class to take these differences into account.
Nevertheless, this study demonstrates that AI-based algorithms are powerful tools to provide critical information for prognostic assessment of patients with OCCC. Its accuracy above 90% corroborate that deep learning is a technology that works even for rare types of ovarian cancer. In summary, with its open and extensible architecture, O3C Glass-Class provides precise predictions and facilitate the development of a personalized treatment strategy in OCCC.
Conclusion
Our study demonstrates the feasibility of using a machine learning system to predict different features of OCCC samples using histopathological images as input. This novel application provides accurate prognosis information and aids in the development of personalized treatment in OCCC.
Supplemental Material
sj-tif-1-bbi-10.1177_11779322221134312 – Supplemental material for O3C Glass-Class: A Machine-Learning Framework for Prognostic Prediction of Ovarian Clear-Cell Carcinoma
Supplemental material, sj-tif-1-bbi-10.1177_11779322221134312 for O3C Glass-Class: A Machine-Learning Framework for Prognostic Prediction of Ovarian Clear-Cell Carcinoma by Ryo Yokomizo, Tiago JS Lopes, Nagisa Takashima, Sou Hirose, Ayako Kawabata, Masataka Takenaka, Yasushi Iida, Nozomu Yanaihara, Kei Yura, Haruhiko Sago, Aikou Okamoto and Akihiro Umezawa in Bioinformatics and Biology Insights
Footnotes
Acknowledgements
The authors thank E. Suzuki and K. Saito for secretarial work.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by KAKENHI; by the Grant of National Center for Child Health and Development.
Declaration of conflicting interests:
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: AU is a co-researcher with MTI Ltd., Terumo Corp., BONAC Corp., Kaneka Corp., CellSeed Inc., ROHTO Pharmaceutical Ltd., SEKISUI MEDICAL Ltd., Metcela Inc., PhoenixBio Ltd., Dai Nippon Printing Ltd. AU is a stockholder of TMU Science Ltd., Morikuni Ltd., and Japan Tissue Engineering Ltd. The other authors declare that there is no conflict of interest regarding the work described herein.
Author Contributions
Conceptualization: RY, TJSL, and AU, Data curation: RY, NT, TJSL, and AK, Formal analysis: RY, NT, and TJSL, Funding acquisition: AO and AU, Investigation: RY, NT, TJSL, AK, MT, and YI NY, Methodology: RY, NT, and TJSL, Project administration: AO, HS, and AU, Software: NT and TJSL, Supervision: KY, HS, AO, and AU, Writing–original draft: RY, Writing–review & editing: TJSL, SH, HS, AO, and AU.
Supplemental Material
Supplemental material for this article is available online.