Abstract
BRIP1 (Breast Cancer 1 Interacting Helicase 1) is a tumor suppressor gene that has vital function in preserving the genetic stability by repairing DNA damage though have significant associations with the onset of breast cancer (BC) if mutated or overexpressed. In this study, the prognostic value of BRIP1 gene was evaluated and validated through bioinformatics approaches utilizing transcriptomic (mRNA expression) data from several BC databases. To determine the prognostic value, the expression level of mRNA transcript was analyzed in context of comparison between breast tumor and normal tissues regarding clinical features, breast tumor subtypes, promoter methylation status, correlation level, mutation frequency, and survival of BC patients. BRIP1 expression was found to be significantly overexpressed in various BC molecular subtypes (e.g. PAM50, Sorlie’s) and clinical status (estrogen and progesterone receptor) than associated normal tissues which correlated with prognosis. Also, in promoter methylation level, its expression was observed as upregulated-hypomethylated regarding various clinicopathological features. Multiple data mining exhibited positive correlation between BRIP1 and INTS2 (Integrator Complex Subunit 2) expressions in BC. Further, mutation analysis revealed that BRIP1 gene was altered by acquiring both somatic and germline mutations. In addition, a total of 42 mutations; 24 missense, 8 fusion, 7 truncating, and 3 inframe mutations in BC patients was detected in BRIP1 protein. Moreover, higher BRIP1 expression was found to be correlated with poor disease-specific, disease metastasis-free, relapse-free, and overall survivals of BC patients. Since, overexpression of BRIP1 was identified to be associated with different clinical features, breast tumor subtypes, promoter methylation status, and survival of BC patients that may provide a risk of ensuing malignant transformation. Thus, lower expression of BRIP1 might hinder BC prognosis. We consider that this analysis will present a proof for BRIP1 gene to be a noteworthy molecular biomarker for BC prognosis.
Introduction
Cancer is anticipated to remain the foremost reason for death over the next 20 years.1,2 Among all the types, breast cancer (BC) is the most predominant cancer and the second chief reason of death in women3-6 though prognosis and proper treatment can significantly better the conditions by attaining enough time to prepare for combating the situation. 7 In 2019, the anticipated number of BC cases was 271,270 (male 2670 and female 268,600) and the predicted deaths were 42,260 (male 500 and female 41,760) in United States.8,9 Although better screening, early diagnosis and treatment for BC are accessible, the prognosis is not much fruitful10 -12 because BC can have an assorted appearance with enormous multifariousness and pervasiveness tendencies regarding morphological natures, clinical results, and subtypes. 13
Conventional prognostic and analytical characteristics like the condition of lymph node, dimension of the tumor, histological grade, and categories of the hormonal receptors (estrogen and progesterone receptors) are not adequate for prognosis and early-stage diagnosis of BC. 14 Therefore, identification of potential molecular biomarkers to predict the prognosis of BC patients is an urgent necessity. In the past years, there have been significant improvements in molecular genetics, transcriptomics and linkage study of the BC.15 -17 Several analyses have explained that BC risk factors are linked to the interaction between definite genes.5,6,10 Understanding such genes included tumorogenesis and their pathways are essential in development of preventive and therapeutic targets to battle BC. Yet, the vast numbers of BC cases are not linked to a higher penetrance mutated gene, but to genes of lower penetrance such as BRIP1 which are often mutated in the general population.18,19
The BRIP1 gene is morphologically expressed in both malignant and healthy cells. BRIP1 interacts with numerous proteins associated with regulation of DNA damage responses and checkpoint signaling, which are vital for retaining chromosomal and genomic constancy. 20 BRIP1 encodes a protein belonging to the RecQ DEAH helicase family 21 which helps to repair damaged DNA by interacting with the BRCA1 (Breast Cancer 1). 19 That’s how BRIP1 plays an incredibly vital function in preserving constancy of the genetic information of the cell and acts as a tumor suppressor. 19 The conserved motif, Q (DEAH box DNA helicase type), also essential for organizing ATP binding, BRIP1 catalytic function and DNA repair purposes 22 but could have associations with the onset of BC if mutated. Based on a range of mutations (truncated, germ-line and missense) recognized in BC in diverse populations studied, BRIP1 emerges to have a chief function in the onset of BC.
The very first proof of BRIP1 as a BC susceptibility gene was found by analyzing mutational screening of the whole BRIP1gene sequence in a British group of familial BC cases and controls. 21 Also, a current analysis has recommended that mutations in DNA repair related genes like BRIP1are associated with elevated BC risk. 5 Mutation creates short non-functional BRIP1 protein which not succeeds to interact with BRCA1 protein to repair DNA damage, and stimulate cellular growth and proliferation, activating the onset of BC. The preliminary evidence for the clinical significance of BRIP1 was the recognition of germ-line mutations within BRIP1in patients connected with early BC, thus signifying a link between BRIP1mutations and low penetrance BC. 21 In addition, the overexpression of BRIP1 protein was found to be associated with an augmented cell proliferation rate, 23 which may also be linked to earlier tumor recurrence.24,25 Moreover, a significantly poorer overall survival (OS) was observed while BRIP1 was overexpressed (P <.05). 26 The function of BRIPI gene in the pathogenesis of BC is still contentious, though slight impacts cannot be ignored.
In this current research, the prognostic and predictive value of BRIP1 expression will be evaluated in context of transcriptomics data (mRNA expression) compared in breast tumor and normal tissues regarding clinical-pathological parameters, tumor subtypes, and promoter methylation level, co-expression, mutation and survival status by extracting and evaluating all presently obtainable data and subsequently validating their predictive potentiality in large BC-allied cohort in silico.
Materials and Methods
Comparative analysis of BRIP1 expression in cancer and normal tissues
Gene expression pattern of BRIP1 was analyzed in several cancerous and normal human body tissues by employing different bioinformatics tools used for cancer database analysis. Expression of BRIP1 transcript in different cancerous and normal tissues was analyzed comparatively. Oncomine
27
(https://www.oncomine.org), GEPIA2
28
(http://gepia2.cancer-pku.cn) and GENT2
29
(http://gent2.appex.kr) databases were utilized to observe the expression of BRIP1 gene. In Oncomine database, 526 datasets from 70,905 samples were selected to analyze transcriptomic divergence of BRIP1 gene in cancer and normal tissues. The threshold selected for cancer microarray database analysis of Oncomine was; P-value
BRIP1 expression analysis in BC and normal breast tissues
The transcription analysis of BRIP1 gene was analyzed comparatively among BC and normal tissues using ULCAN 30 (http://ualcan.path.uab.edu/), GEPIA2 and HPA 31 (https://www.proteinatlas.org/) server. Cancer transcript data is statistically analyzed in ULCAN server which provides different cancer transcriptome data such as TCGA datasets. The GEPIA2 server contains 1,085 tumors and 291 normal tissue data related to BC whereas the UALCAN web server provides access to data from 1097 cancers and 114 normal tissues for analysis. Expression pattern of BRIP1 gene was analyzed in these servers from TCGA datasets and cox p-value of less than 0.05 (P < .05) was considered significant. Also, immunohistochemistry of BRIP1 protein expressed in BC and normal tissues was observed according to HPA005474 dataset by employing HPA web server.
BRIP1 expression analysis through profiling molecular subtypes and clinicopathological parameters
Transcriptome data of BRIP1 gene based on molecular subtypes of BC patients in TCGA data was analyzed by employing BC gene-expression miner v4.5 (bcGenExMiner v4.5) (http://bcgenex.ico.unicancer.fr) which is a web-based independent server that provides access to published genomic data. Three types of statistical analysis can be done using bcGenExMiner v4.5; expression, prognosis, and correlation analysis. TCGA data of bc-GenExMiner v4.5 was utilized to determine BRIP1 targeted expression through gene expression profiling subtypes; PAM50 (Parker’s molecular 50), HU’s, Sorlie’s, SCMGENE (subtype clustering model GENE), SCMOD2 (subtype clustering model2), SCMOD1 (subtype clustering model1), RSSPC (robust single sample predictors classification), RSCMC (robust subtype clustering model classification), and RIMSPC (robust intrinsic molecular subtype predictors classification) subtypes.
Analysis of relationship between BRIP1 transcript and different clinicopathological features such as receptor status ER (estrogen receptor) and PR (progesterone receptor) in IHC (immunohistochemistry) was analyzed in TCGA data (
Study of association between BRIP1 and promoter methylation
1247 TCGA breast cancer tests dataset were employed for analysis of promoter methylation status of BRIP1 mRNA using UCSC Xena server 32 (https://xenabrowser.net/) and also based on their clinicopathological characteristics BRIP1 promoter methylation was analyzed in TCGA datasets with the UALCAN server. The analysis was carried out comparing normal and cancer patient’s dataset.
Identification of correlated genes associated with BRIP1
Genes correlated with BRIP1 transcript in BC was identified and confirmed by employing different web-based server; ULCAN, UCSC Xena, GEPIA2, and bc-GenExMiner v4.5 server. To identify the genes highly correlated with BRIP1 gene TCGA dataset of ULCAN server was utilized for data mining and Pearson’s correlation analysis was carried out. Genes that are highly associated with BRIP1 were further assessed for identifying the relationship level by generating a heat map comparing BRIP1 and expression of highly associated genes using UCSC Xena server. Moreover, Pearson’s correlation plot was evaluated for assessing the correlation level using UCSC Xena, GEPIA2, and bc-GenExMiner v4.5 server. The p-value when it is less than 0.05 was considered statistically significant.
Identification of mutation and copy number alteration in BRIP1
Mutation, somatic copy no. alteration in BRIP1 gene of human BC was analyzed by utilizing different datasets from BC studies. Three recent studies named “Juvenile Papillomatosis and Breast Cancer (2020),” ‘the Metastatic Breast Cancer Project (2020),’ and “Breast Cancer (2020)” containing a total of 383 samples were chosen, retrieved and evaluated for observing mutation and copy no. alterations of BRIP1 gene in human BC. Another study of mutations and copy no. alterations in BRIP1 gene in BC were also performed specifically on four TCGA database studies which were Cell 2015, Firehose legacy, Nature 2012, and PanCancer Atlas. These four TCGA database contains 3834 samples for breast cancer genomics analysis. GISTIC algorithm (Genomic Identification of Significant Targets in Cancer) was utilized for all these analyses of mutation and copy no. alterations of BRIP1 gene in BC by using cBioPortal server 33 (https://www.cbioportal.org/) which provides database for analyzing multiple types of human cancer genomics.
Profiling survival of BC patients in relation to BRIP1
Finally, to determine the prognostic value of BRIP1, impact of expression level on breast cancer patient’s survival was analyzed and survival plot on different variables was generated using PrognoScan server 34 (http://dna00.bio.kyutech.ac.jp/PrognoScan/) and KM-Plotter (https://kmplot.com/analysis/). PrognoScan server is an independent web server for analyzing prognostic value of genes expressed in different types of human cancer while KM-Plotter can determine the impact of 54,675 genes on survival rate in 21 types of cancer. Distant metastasis-free survival plot was analyzed using KM-Plotter and Disease-specific survival, Overall survival, and Relapse free survival plot was generated by employing PrognoScan server where the hazard ratio (HR) with 95% confidence intervals (CI) was considered as significant. In all analyses, p-value less than 0.05 were regarded as statistically significant.
Results
Comparative analysis of BRIP1 expression in several cancers and healthy cohorts
To analysis the expression of BRIP1 gene in different types of cancer and corresponding normal patients, databases of three web-based platforms Oncomine, GENT2, and GEPIA2 were utilized. Among 349 unique analysis, only 17 analysis was significant in the Oncomine database, where downregulation and upregulation of BRIP1 gene was observed comparatively between different cancer and corresponding healthy tissues. Data showed that upregulation of BRIP1 gene was occurred in case of BC and also in colorectal and brain cancer tissues compared to respective healthy patients (Figure 1A). Upregulation of BRIP1 gene was observed in various breast carcinomas including male breast carcinoma, invasive ductal breast carcinoma, invasive breast carcinoma, invasive lobular breast carcinoma, ductal breast carcinoma, invasive breast carcinoma stroma (Table 1). mRNA transcript of BRIP1 gene was also analyzed utilizing the Affymetrix HG-U133pLUS2 platform from the GENT2 database showed that BRIP1 gene was unregulated in BC patients including adrenal gland, bladder, brain, cervix, colon, gallbladder, kidney, ovary, pharynx, skin, tongue and vaginal cancer patients in comparison of healthy people (Figure 1B). Further analysis of the data was performed in 16 different cancer types than normal tissues which showed that BRIP1 gene was over-expressed in every type of cancer tissue compared to normal tissue using GEPIA2 server (Figure 1C).

BRIP1 expression in various cancer and normal tissues analyzed by Oncomine, GENT2, ULCAN, GEPIA2 and HPA databases. (A) BRIP1 up-regulation (red) or down-regulation (blue) in different cancerous and corresponding normal tissue datasets generated by Oncomine database. (B) BRIP1 mRNA expression presented in a box-plot form representing in cancerous and normal tissues from GENT2 database where boxes = median, dots = outliers, red-boxes = tumor tissues, and blue-boxes = normal tissues. (C) mRNA transcript (TPM) profile of BRIP1 gene in case of 16 different cancers and corresponding normal tissues of human bodies obtained from GEPIA2 database. (D and E) Comparative BRIP1 transcriptome analysis expressed in BC and normal datasets as box plot form using ULCAN and GEPIA2 server respectively. (F and G) Immunohistochemistry image of BRIP1 protein expressed in normal and cancerous breast tissues respectively obtained from HPA database. BC indicates breast cancer; BRIP1, Breast Cancer 1 Interacting Helicase 1; GENT2, gene expression across normal and tumor tissue. Here, HPA database: Human Protein Atlas database, TPM: Transcript Per Million, mRNA: messengerRNA, UALCAN is a comprehensive, user-friendly, and interactive web resource for analyzing cancer OMICS data, GEPIA2 web server: Gene Expression Profiling Interactive Analysis web server.
Transcription level of BRIP1 gene in different BC tissues regarding significant changes due to cancer in Oncomine datasets.
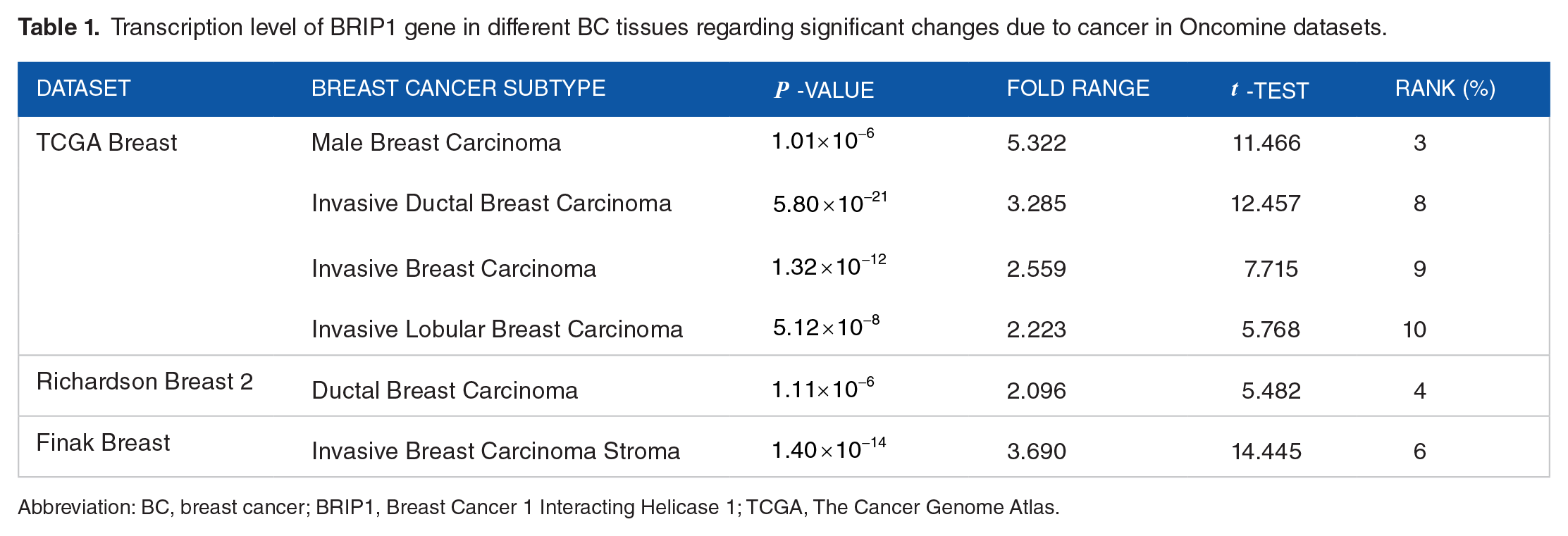
Abbreviation: BC, breast cancer; BRIP1, Breast Cancer 1 Interacting Helicase 1; TCGA, The Cancer Genome Atlas.
Comparative analysis of BRIP1 gene was also analyzed specifically in BC and normal people’s TCGA data using ULCAN and GEPIA2 web-based server exhibited that BRIP1 gene was upregulated in BC patients compared to healthy people (Figure 1D and E). Immunohistochemistry images of BC and normal people were also retrieved from HPA005474 dataset in HPA server and analyzed comparatively. Among 23 BC patients patient ID 2392 showed high staining signal and strong intensity while normal breast myoepithelial and adipocyte cell showed medium staining signal representing down-regulation of BRIP1 gene (Figure 1F and G).
BRIP1 expression analysis through profiling molecular subtypes and clinicopathological parameters
In this analysis, BRIP1 expression was shown to be upregulated in BCs compared to normal breast. In case of PAM50 subtype, Sorlie’s subtype and RSSPC subtype there was no BRIP1 expression was observed in normal breast tissues. All subtypes; PAM50, HU’s, Sorlie’s, SCMGENE, SCMOD2, SCMOD1, RSSPC, RSCMC, and RIMSPC subtypes showed that BRIP1 is expressed at a high level in highly aggressive BCs like basal-like, HER2 enriched, and the luminal B subtype in comparison to luminal A subtype Figure 2A to I.

Molecular expression evaluation of BRIP1 gene through profiling of intrinsic molecular subtypes in TCGA data using bc-GenExMiner v4.5 web server. Intrinsic molecular subtyping of BC patients according to (A) PAM50 subtype (Parker’s molecular 50), (B) HU’s subtype, (C) Sorlie’s subtype, (D) SCMGENE subtype (subtype clustering model GENE), (E) SCMOD2 subtype (subtype clustering model2), (F) SCMOD1 subtype (subtype clustering model1), (G) RSSPC subtype (robust single sample predictors classification), (H) RSCMC subtype (robust subtype clustering model classification), and (I) RIMSPC subtype (robust intrinsic molecular subtype predictors classification) subtypes. BRIP1 expression of BC using bc-GenExMiner v4.5 web server was analyzed where different clinicopathological parameters considered were; Estrogen receptor (ER) status (J), and Progesterone receptor (PR) status (K). Significant difference between groups was measured employing Welch’s t test to generate p-values following Dunnett-Tukey-Kramer test by utilizing bc-GenExMiner v4.5 server. BC indicates breast cancer; BRIP1, Breast Cancer 1 Interacting Helicase 1; IHC, immunohistochemistry; ER, estrogen receptor status; PR, progesterone receptor status; N, nodal status; TCGA, The Cancer Genome Atlas.
BRIP1 mRNA transcript in TCGA dataset of BC was showed negative correlation with both cases of ER and PR status. BRIP1 expression was found to be significantly higher in case of ER- and PR- status compared to ER + and PR + status (Figure 2J and K, Table 2). Welch’s t test was employed to generate p values and the Dunnett–Tukey–Kramer test was performed by employing bc-GenExMiner v4.5 server to plot the significant difference in each test.
Relationship analysis between BRIP1 transcription and clinicopathological parameters of BC using bc-GenExMiner v4.5 server.

Abbreviations: BC, breast cancer; BRIP1, Breast Cancer 1 Interacting Helicase 1; ER, estrogen receptor status; PR, progesterone receptor status. Here, mRNA: messengerRNA
Analysis of promoter methylation status of BRIP1
Promoter methylation is a substantial factor in case of genetic and epigenetic expression. The methylation pattern is related to specific physiological conditions like hypo-methylation is reported to be positively related to cancer conditions. Here, by employing TCGA dataset of UCSC Xena server BRIP1 gene expression and promoter methylation level were evaluated by creating a heat map in case of PAM50 BC subtypes. The comparison demonstrated that BRIP1 expression was negatively associated with promoter methylation in case of PAM50 BC subtype of 1247 TCGA breast cancer tests dataset (blank frame) (Figure 3A). ULCAN based data analysis revealed that promoter methylation was significantly low in case of all clinical subtypes of BC; sample type (Figure 3B), individual cancer stages (Figure 3C), patients race (Figure 3D), gender (Figure 3E), age (Figure 3F), major subclasses (Figure 3G), menopause status (Figure 3H), and TP53 mutation status (Figure 3I) compared to respective normal patients (Supplementary Table 1). Therefore, the analysis exhibited that BRIP1 transcript was upregulated and that can be occurred due to lower level of promoter methylation in BC patients.

Promoter methylation level of BRIP1 in BRCA of TCGA datasets. Heat map of BRIP1 expression and promoter methylation comparison in PAM50 BC subtypes of TCGA datasets using UCSC Xena server where red color specifies high expression while blue color specifies low expression (A). Box plot analysis of promoter methylation level of BRIP1 in TCGA breast cancer (BRCA) datasets using ULCAN server based on sample type (B), individual cancer stages (C), patients race (D), gender (E), age (F), major subclasses (G), menopause status (H), and TP53 mutation status (I). Data were analyzed comparatively regarding significant fluctuations in normal and other variables. The beta value shows the level of DNA methylation ranging from 0 (unmethylated) to 1 (fully methylated). Different beta value cut-offs were considered to point out hyper-methylation [beta value 0.7-0.5] or hypo-methylation [beta-value 0.3-0.25]. BC indicates breast cancer; BRCA, Breast Cancer 1; BRIP1, Breast Cancer 1 Interacting Helicase 1; DNA, deoxyribonucleic acid; HER2, human epidermal growth factor receptor 2; PAM50, Parker’s molecular 50; TCGA, The Cancer Genome Atlas; TNBC; Triple negative breast cancer. UALCAN is a comprehensive, user-friendly, and interactive web resource for analyzing cancer OMICS data, TP53: Tumor Protein P53, UCSC Xena server: University of California Santa Cruz Xena server.
Correlation level of genes associated with BRIP1
At first, to identify the correlated gene with BRIP1, data mining was performed using TCGA (BRCA) dataset which revealed that INTS2 gene was highly correlated with BRIP1 gene utilizing ULCAN server (Figure 4A). Then Pearson’s correlation analysis was performed comparing the expression of BRIP1 and INTS2 gene where the coefficient value was 0.6 (Figure 4B). A heat map of Pearson’s (r = 0.64) and Spearman’s (r = 0.62) correlation analysis was created utilizing the UCSC Xena server for identifying the intensity of the correlation level of INTS2 and BRIP1 gene (Figure 4C and D). Pearson’s correlation analysis was employed again to confirm the highly positive association of INTS2 gene with BRIP1 gene by employing GEPIA2, (R = 0.64) and bc-GenExMiner v4.5 (R = 0.41) server respectively. These analyses revealed that INTS2 might be correlated with BRIP1 mediated signaling pathways which can cause progression of BC (Figure 4E and F). p value of less than 0.05 was considered as statistically significant.
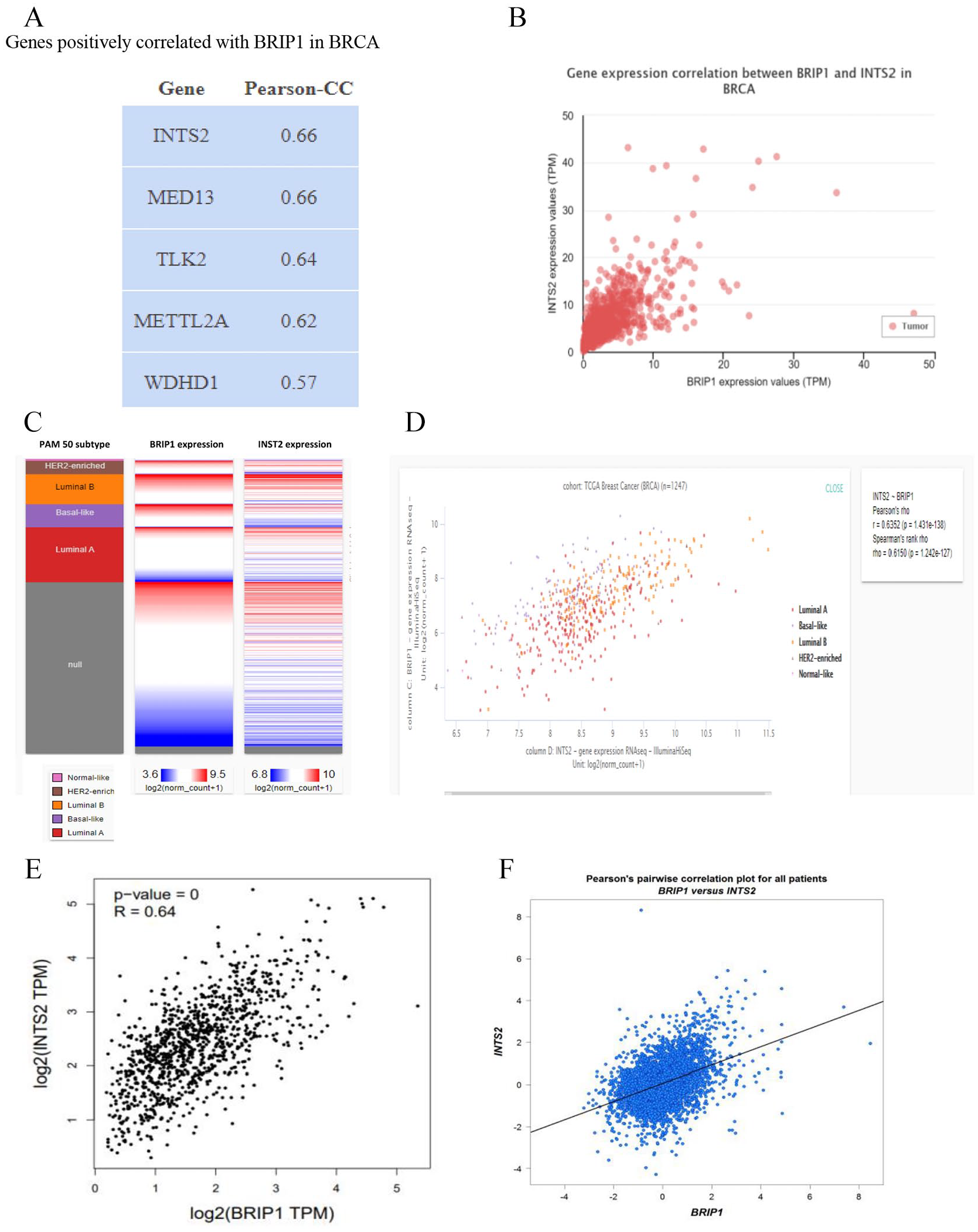
Identification of correlation level of genes associated with BRIP1 expression in BC using different web server. Data mining and Pearson’s correlation analysis were employed to identify the genes which showed high correlation with BRIP1 gene in TCGA dataset of ULCAN server respectively (A and B). Genes which is highly associated with BRIP1 expression was further assessed by generating a heat map and Pearson’s correlation with the UCSC Xena server respectively (C and D). Pearson’s correlation plot was generated for confirming the correlation level using GEPIA2 and bc-GenExMiner v4.5 server respectively (E and F). BC indicates breast cancer; BRIP1, Breast Cancer 1 Interacting Helicase 1; TCGA, The Cancer Genome Atlas. TPM: Transcript Per Million, UALCAN is a comprehensive, user-friendly, and interactive web resource for analyzing cancer OMICS data, GEPIA2 web server: Gene Expresssion Profiling Interactive Analysis web server, UCSC Xena server: University of California Santa Cruz Xena server.
Mutation and copy number alteration of BRIP1
Identification of mutation and copy no. alteration in BRIP1 gene was performed twice by utilizing four TCGA databases (Cell 2015, Firehose legacy, Nature 2012, and PanCancer Atlas; n = 3834) and then three different recent breast cancer genome project databases which are: Juvenile Papillomatosis and Breast Cancer (MSK, 2020), The Metastatic Breast Cancer Project (Provisional, February 2020), and Breast Cancer (MSK, Nature Cancer 2020; where n = 383) by employing web based independent server cBioPortal. In case of four studies of TCGA database, alteration frequency calculated was 10.74% in 1099 cases which was found to be highest for Breast (TCGA) study among four studies (Figure 5A). Whereas, in recent three studies of BC, copy number alteration frequency measured was 13.08% in 237 cases as highest frequency among the database of three studies in form of bar diagram (Figure 5B). Lollipop plot was generated for both TCGA and recent database analysis of mutation status in specific region of BRIP1 gene in human BC by cBioPortal. In analysis of utilizing TCGA database, BRIP1 gene was altered in 364 (10%) of queried patients (3827) and in 364 (9%) of queried samples (3834) in 4 studies having both somatic and germline mutation frequency 0.9% and 0.1% respectively. Also, a total of 42 mutations (24 Missense, 8 Fusion, 7 Truncating and 3 Inframe mutations) including 19 duplicate mutations in patients with multiple samples was detected in DEAD_2 and Helicase_C_2 domain of BRIP1 protein within 1-1249aa residues (Figure 5C). In case of breast cancer database analysis of three recent studies, a total of 3 mutations (2 Missense and 7 Truncating) were observed in DEAD_2 and Helicase_C_2 domain of BRIP1 gene within 1-1249aa residues. In addition to that, BRIP1 protein was altered in 30 (13%) of queried patients (343) and 33 (9%) of queried samples (383) in 3 studies having somatic mutation frequency 0.8% (Figure 5D). Moreover, the types of mutation in BRIP1 transcript vs analyzed studies of origin were plotted in both analyses (Figure 5E and F). Hereafter, all these findings exhibited that types, levels and frequency of mutations and copy no. alterations in BRIP1 protein may be correlated to BC progression.
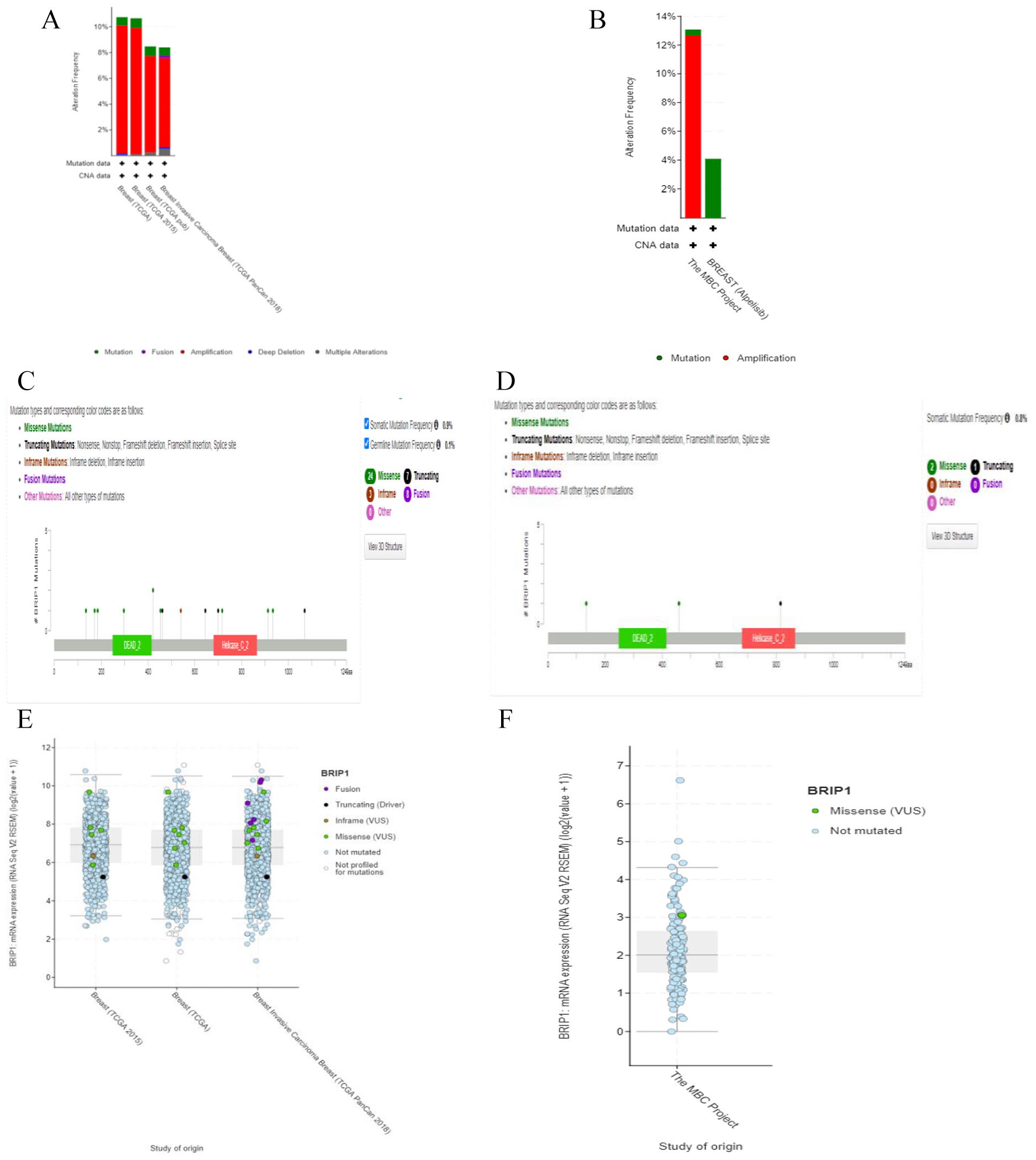
Mutation and copy no. alteration identification in BRIP1 gene of human BC tissue. (A and B) Bar diagrams representing the mutation types of BRIP1 gene in terms of alteration frequency in TCGA and different cancer project databases respectively. (C and D) Types of mutations in different BRIP1 region exhibited by lollipop diagram in TCGA and different cancer project databases respectively. (E and F) BRIP1 transcript expression showing the alteration level comparatively in TCGA and different cancer project databases respectively. All these analyses of mutation and copy no. alterations in BRIP1 gene of BC were analyzed by using cBioPortal server. BC indicates breast cancer; BRIP1, Breast Cancer 1 Interacting Helicase 1; CAN, copy number alteration; TCGA, The cancer genome atlas.
Relationship of BRIP1 expression and survival of BC patients
BRIP1 expression and survival status of BC patients were analyzed employing PrognoScan and KM-Plotter web server (significant Cox p value < 0.05). Results showed that in all cases of suvival analysis, overall survival, relapse free survival, distant metastasis-free survival, and disease-specific survival, with the high expression status of BRIP1 the survival rate of several BC cancer patients was significantly (p value < 0.05) decreased (Supplementary Table 2, Figure 6A to F). In addition, hazard ratio (HR) was observed to more than 1 (HR > 1) in all types of survival status, which means that survival probability was more in those patients who showed a lower level of expression of BRIP1 and at low risk. But BC patients who showed higher BRIP1 expression are at high risk and showed a significant sharp decrease in survival time compared to the low expression of BRIP1 BC patients. The survival analysis was performed in different BC data cohort in different types of survival plot with different web-server to properly predict and validate the survival status of BC patients.

Analyzing the relationship between BRIP1 expression and BC patient’s survival status in terms of different variables. (A) Distant metastasis-free survival, (B) Disease-specific survival, (C) Overall survival (D) Relapse-free survival (Probe ID: 221703_at), and (E) Relapse free survival (Probe ID: 18395). Distant metastasis-free survival plot was analyzed using KM-Plotter where the hazard ratio (HR) with 95% confidence intervals (CI) was considered as significant (A) and other survival plots were generated by employing PrognoScan server (B to E). In all analyses, p value less than 0.05 was regarded as statistically significant. BC indicates breast cancer; BRIP1, Breast Cancer 1 Interacting Helicase 1.
Discussion
The function of BRIP1 gene in the onset, development and prognosis of BC was verified by analyzing BRIP1gene expression and survival level regarding various parameters between cancer and normal tissues. In this present study, BRIP1 expression was found to be overexpressed in numerous cancer types including breast, brain, cervix, colon, kidney, ovary, pharynx, skin, and vaginal cancer patients in comparison to healthy people. Over expression of BRIP1 protein has association with earlier tumor recurrence which may be due to elevated cell proliferation rate.24,25,35 In addition, anomalies in BRIP1expression were recognized to have a significant connection with the progression of breast cancer (BC), ovarian cancer, and type J Fanconi anemia. 21 In contrast, other studies showed that downregulation of BRIP1 gene was found in different cancers; specifically in ovarian and cervical cancer.36,37 It was also predicted that BRIP1 gene may be susceptible to breast, prostate and ovarian cancer.21,36,38 Other studies found that the presence of the SNPs in BRIP1 gene has association with the commencement of prostate cancer.39,40
In this present study, BRIP1 gene was observed as highly expressed in BC patients compared to healthy people which has similarities with the findings of other study. 41 BRIP1 gene was showed an average of five fold upregulation in BC and was identified as a strong candidate gene responsible for BC progression. 26 Another study also suggests the BRIP1 gene as an oncogene that has a relation in malignancy of BC tissue when overexpressed.41,42 Moreover, Immunohistochemistry images of BC tissue showed a high staining signal and strong intensity representing BRIP1 upregulation while normal breast cells showed medium staining signal representing down-regulation of BRIP1 gene. In accordance with the study, an immunohistochemistry analysis found that BRIP1 expression was increased following the elevation of BC grade.26,43
In our study, we also utilized gene expression profiling subtypes containing several molecular intrinsic subtypes for analyzing BRIP1 expression. BRIP1 overexpression was analyzed in every intrinsic molecular subtype compared to normal sample along with the overexpression level in highly aggressive BCs such as basal-like, HER2 enriched, and the luminal B subtype than luminal A subtype. In accordance with our result, BRIP1 overexpression was reported to associated with clinical features and lower survival of luminal BC subtypes. 26 Luminal A subtype had maximum upregulation of BRIP1 (6.5-fold) along with Luminal B (5.2-fold) and both the Her2 + and Triple-negative subtype had an analogous level of BRIP1 expression (3.8-fold) than normal samples using RT-qPCR analysis. Whereas, BRIP1 expression was upregulated in the Luminal A subtype (11.43-fold), followed by Luminal B (4.54-fold), Triple-negative (4.86-fold) and Her2 + (5.6-fold) subtypes using western blot analysis. 26
The growth of BC is frequently coordinated by regularly analyzing the concentrations of ER and PR in the tumor which acts as predictors of first-rate prognosis. BRIP1 expression was analyzed in case of ER and PR status of BC patients in this study to confirm the prognostic value of BRIP1 gene. The ER and PR are dimeric, gene-regulatory endocrine steroids considered as chief significance for tumors originating in the reproductively competent periods. The ER pathway has crucial task in the pathophysiology of BC. 44 Also, ER-positive BCs are connected with slow tumor growth, worse histology ranking, DNA diploidy, and hence an enhanced overall prognosis. 45 Another study reported that the ER is expressed in most cases of BC. 46 Around 75% of all BCs express ER and/or PR, where about 20% of BC exhibits an overexpression of HER2. In addition, in case of prognosis, ER-positive BC is considered as more approving compared to ER-negative BC. 5 In this study, BRIP1 expression was observed as higher in case of ER- and PR- status than ER + and PR + status which presents a potential BC prognosis value of BRIP1 gene depending on ER and PR status.
Level of promoter methylation could have association with alteration in gene expression that related to onset of cancer. Epigenetic alteration like DNA methylation in genes plays significant role in tumorigenesis, cancer progression and metastasis.47,48 DNA methylation takes place with the accumulation of a methyl group from S-adenosylmethionine (SAM) into cytosine residues of the DNA template. 49 Silencing of BC related genes by hypermethylation was observed as related to BC carcinogenesis and progression. 5 Also, in a study of BC prognostic value analysis of differentially expressed-aberrantly methylated hub genes, 677 upregulated-hypomethylated and 361 downregulated-hypermethylated genes were observed. 50 In this present study, BRIP1 expression was observed as upregulated-hypomethylated regarding different clinical subtypes of BC. Thus, hypomethylation of promoter region related to overexpression of BRIP1 gene expresses its prognostic value. In line with our interpretation, a study suggested that both DNA hypermethylation and hypomethylation can be potentially engaged with varied progressions of BC expansion and prognosis. 51
BRIP1 gene mainly acts as DNA damage repairing gene,52,53 interacts with BRCA1 and together they reconnect broken DNA strands.19,21 BRIP1 acts as tumor suppressor and that’s how they avert cells from acquiring mutations.20,54 If BRIP1 protein is incomplete or lost it doesn’t intermingle with the BRCA1 protein and not succeeds to fix the damaged DNA. 55 Mutation in one copy of BRIP1 gene can produce unusually tiny and nonfunctional BRIP1 protein. 56 A recent study showed association of rare missense alleles in BRIP1 with BC risk 57 and, also, two SNPs in BRIP1 gene showed significant link with BC. 22 In this study, mutation in BRIP1 protein was observed and that provide prognostic value to this particular gene. In accordance with this result, several researches showed that mutation,5,21,54,58,59 missense mutation57,58 or slight variation in nucleotide of BRIP1 are associated with BC development or increases the risk of BC highly. Also, BRIP1 truncating and functionally deleterious missense mutations were found to be augmented the risk of BC by comparing more than 48,000 BC and 43,000 healthy cases.20,54
In this present study, a total of 42 mutations (24 Missense, 8 Fusion, 7 Truncating and 3 Inframe mutations) were detected in BRIP1 protein. In accordance with our study, 7 germline mutations; three missense, two silent, two intronic mutations in BRIP1 were recognized in a study. 60 Moreover, two independent germline (missense) mutations in of BRIP1 were observed which were also found to be linked to the early commencement of familial BC. 21 Furthermore, BRIP1 truncating mutations were observed where the pervasiveness of BRIP1 truncating mutations was around 0.7% and the relative risk for onset of BC was 2.0 in patients with familial BC.20,54,61 In current study, in case of recent three BC studies, copy number alteration frequency of BRIP1 gene was found 13.07%. Therefore, mutation and copy number alteration observed in BRIP1 gene in our study could have provide potential basis for diagnosis and prognosis of BC.
Finally, to estimate the prognostic value of BRIP1, overexpression level was analyzed in association with survival rate of BC patients. Results showed that overall survival, disease-specific survival, disease metastasis-free survival and relapse-free survival were reported to be low when BRIP1 was overexpressed in BC patients and vice versa. In accordance with this result, a significantly poorer overall survival (OS) was observed while BRIP1 was overexpressed (P < 0.05). 26 In addition, survival analysis was also executed in luminal A and luminal B molecular subtypes and the OS of patients was significantly worse showing BRIP1 overexpression than patients with low BRIP1 expression. 26 In this study, overexpression of BRIP1 gene has showed direct positive association with poor survival status of BC patients. This result significantly validates the prognostic value of BRIP1 gene.
The present study was performed utilizing the experimental data accessible in public databases. Further investigations including large scale and comprehensive analyses will verify our results and will provide justification for the clinical utility of BRIP1 in BC prognosis assessment and designing therapeutical approaches against BC.
Conclusion
To analyze the candidacy of BRIP1as a prospective BC prognostic marker, we studied the BRIP1mRNA expression in diverse clinical features, BC subtypes, promoter methylation status, correlation level, mutation frequency, and survival of BC patients. In summary, we presented the verification result that there is significant association between BRIP1 mRNA overexpression levels with several BC clinical features and subtypes (e.g. ER and PR status), promoter methylation status, and survival of BC patients. In addition, significant mutation in BRIP1 protein related to BC prognosis was also observed. Thus, lower expression of BRIP1 might hinder BC prognosis. These findings suggest that BRIP1 gene may not be only a favorable predictive molecular biomarker of development and prognosis of BC but also could perform as a latent therapeutic target.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322211055892 – Supplemental material for Prognostic Value Estimation of BRIP1 in Breast Cancer by Exploiting Transcriptomics Data Through Bioinformatics Approaches
Supplemental material, sj-docx-1-bbi-10.1177_11779322211055892 for Prognostic Value Estimation of BRIP1 in Breast Cancer by Exploiting Transcriptomics Data Through Bioinformatics Approaches by Umama Khan and Md. Salauddin Khan in Bioinformatics and Biology Insights
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
