Abstract
Since the development of high-density microarray technology in the late 1990s, global host gene expression changes in response to various stimuli have been extensively studied. More than a dozen peer-reviewed publications have investigated the effect of Leishmania infection in various models since 2001. This review covers the transcriptional changes in macrophage models induced by various Leishmania species and summarizes the resulting impact these studies have on our understanding of the host response to leishmaniasis in vitro. Characterization of the similarities and differences between various model systems will not only further our understanding of Leishmania-induced changes to macrophage gene expression but also identify potential therapeutic targets in the future.
Introduction
Parasites of the genus Leishmania cause a variety of devastating and fatal diseases in humans and domestic animals worldwide. The spectrum of illnesses, collectively called leishmaniasis, ranges from cutaneous ulcerative lesions to fatal visceralizing infections and affects an estimated 12 million people worldwide.1,2 Middle Eastern conflicts and a combination of environmental changes have resulted in the rise of leishmaniasis epidemics worldwide.3–8 With the absence of effective vaccines, chemotherapy is the only avenue against leishmaniasis. Currently, this small arsenal of drugs is far from ideal due to a lack of selectivity and the emergence of drug resistance. Thus, there is an urgent need for a better understanding of host-parasite interactions to develop new therapeutic approaches.
Leishmania parasites are a heterogeneous family of 53 species, 20 of which are infectious to humans. 9 The type of disease that will manifest depends largely on the infecting Leishmania species, although the hosts’ genetics and immune responses can also play a role. 10 Cutaneous leishmaniasis is mainly caused by L. major and L. tropica in Europe, Africa, and Asia, and by L. mexicana, L. amazonensis, and L. braziliensis in the Central and South America.11,12 Parasites of the L. donovani complex cause visceral leishmaniasis, which affects inner organs like the liver and spleen with L. donovani being the causative agent in Asia and Africa, whereas L. chagasi (syn. L. infantum) is responsible for the disease in Latin America and the Mediterranean.12,13
Leishmania parasites exhibit a digenetic lifecycle, in which the extracellular promastigote form lives in the sand fly gut, whereas the intracellular amastigotes reside in the phagolysosome of infected mammalian macrophages. Although some Leishmania species may initially infect neutrophils and dendritic cells, macrophages are considered the main host cells.14,15 Parasites are taken up by phagocytosis, in which the phagosomes and lysosomes fuse to form phagolysosomes. This acidic intracellular compartment contains aggressive enzymes that usually eliminate foreign microorganisms. 15 Leishmania parasites are able to not only survive but replicate in this hostile environment by both adapting to the host environment and manipulating host responses, including metabolic, signaling, and immune pathways.14–19 Different Leishmania species bind to disparate receptors, such as complement, mannose, and fibronectin receptors. This variation in binding may influence the type of host response that is triggered. 14 In addition, some Leishmania species inhibit the maturation of the phagolysosome.14,20,21 This may, in part, be achieved by the parasites affecting macrophage lipid metabolism and lipid microdomains.14,17,22 Interestingly, some parasite species reside in communal vacuoles (L. mexicana and L. amazonensis), whereas others exist in single-parasite vacuoles that divide with the parasite (L. major and L. donovani).14,17,21
Leishmania parasites can trigger different host immune responses, which is ultimately responsible for disease severity. The T Helper cell 1 (TH1) cytokine response is associated with parasite clearance and cure in both murine models and clinical studies.23,24 In contrast, the T Helper cell 2 (TH2) cytokine response leads to increased parasite numbers and disease exacerbation.23,24 Parasite infections have also been shown to induce host arginase expression and activity.25,26 It is not completely understood how increased levels of arginase cause or contribute to disease exacerbation, but the decreased arginine availability may reduce the production of the anti-leishmanial agent nitric oxide and hinder local T-cell development.25-29 In addition, host arginase activity may produce more ornithine and/or polyamines for parasite salvage and proliferation.26,30 Another survival mechanism of Leishmania parasites appears to be the inhibition of host apoptosis.14,31
Several tools are available to study the global host response induced by parasite infections. One such tool is microarray technology, which can be used to measure global changes in the transcriptome. The purpose of this review is to summarize and discuss results that have been obtained with various Leishmania species in the 4 most commonly used host model systems: primary cells derived from C57BL/6 or BALB/c mice, the human THP-1 cell line, and human monocyte-derived macrophages (MDMs). Technical details of the reviewed studies, as well as their general conclusions, are presented in Table 1.
Model-based summary of host gene expression studies.
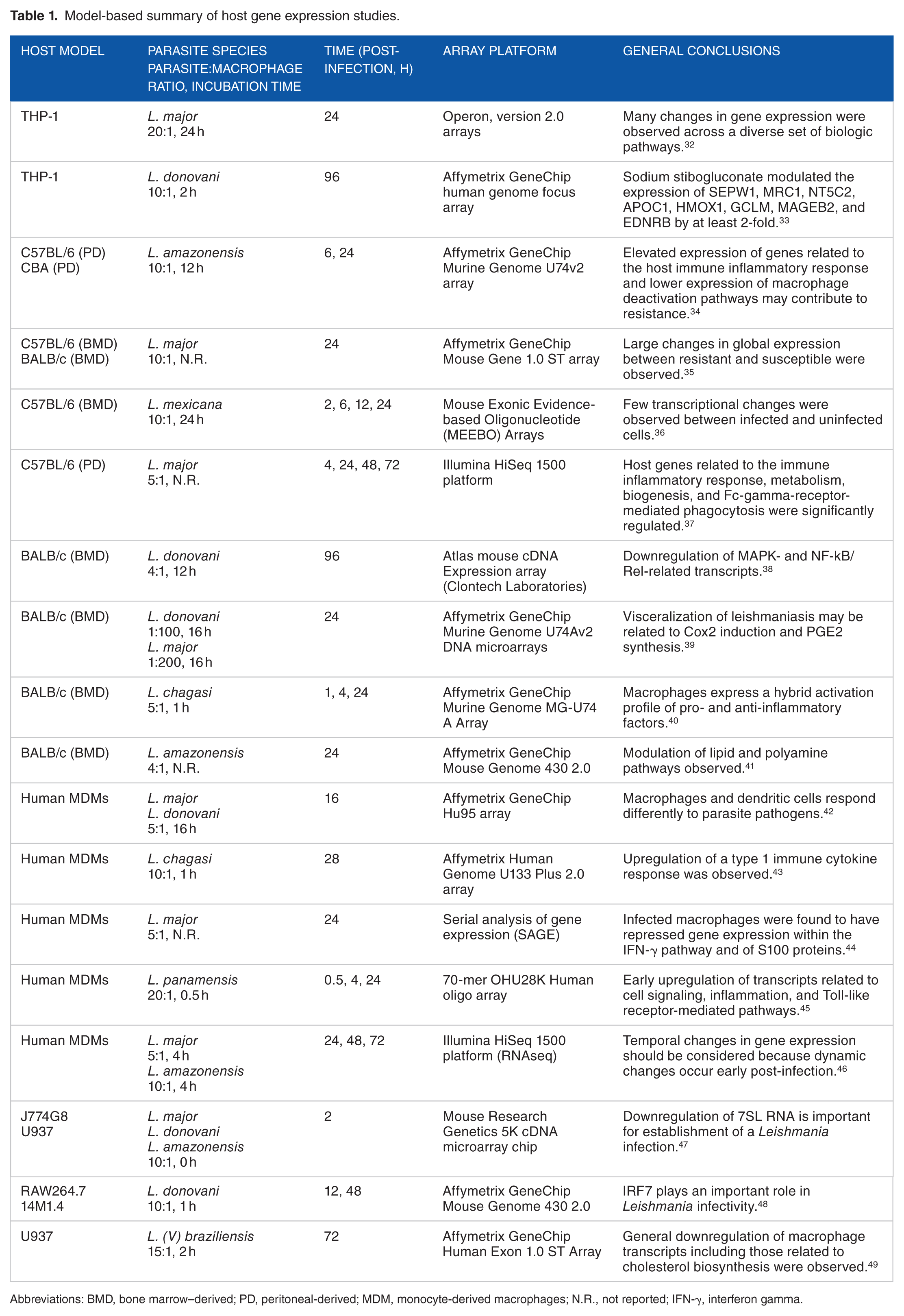
Abbreviations: BMD, bone marrow–derived; PD, peritoneal-derived; MDM, monocyte-derived macrophages; N.R., not reported; IFN-γ, interferon gamma.
C57BL/6 Macrophages
C57BL/6 mice are a widely used and well-studied inbred strain, from which peritoneal-derived (PD) or bone marrow–derived (BMD) macrophages are used for in vitro studies. In contrast to other mouse strains (eg, CBA and BALB/c), C57BL/6 mice display resistance to most Leishmania-based infections due to a TH1 response.34,35 Chemokine production following infection from multiple Leishmania species appears to be a common hallmark for macrophages from this murine model. In one study, the modulation of genes associated with the host inflammatory response that trigger cytokine and chemokine production was speculated to contribute to the ability of C57BL/6 PD macrophages to control L. amazonensis infection. 34 In a separate study investigating L. major infection, higher expression of certain chemokines as well as uniquely modified host signal transduction pathways such as mTOR, ErbB, and insulin signaling were observed. 35 Based on the fact that certain cytokines play crucial roles in macrophage activation, diverse inflammatory responses mediated by these factors have a high probability to contribute to the observed differences in leishmaniasis resistance. It is worth noting that the two aforementioned studies obtained their macrophages from different sources (PD in the L. amazonensis study and BMD in the L. major study). Although baseline gene expression is divergent depending on the macrophage source, evidence does suggest that both respond to cytokines similarly, which implies the potential overlap of gene expression between both sources could be valid. 36
More recently, RNAseq was employed to compare changes in the global transcriptome between C57BL/6 host macrophages and L major during the first 72 hours of infection. 37 Obtaining transcriptomic data simultaneously from the murine macrophage model and parasites was possible based on the fact that there is little sequence conservation between these two biologic systems. Changes in host expression were greatest both in number of transcripts and magnitude at the 4-hour timepoint. Pathway analysis revealed transcripts associated with immune responses were most highly induced, which included both anti- and pro-inflammatory cytokines. In fact, the interaction between cytokines and their receptors was the most significantly upregulated KEGG pathway, further cementing a role for these factors during L. major infection. 37
Changes in transcripts associated with other pathways have also been reported in C57BL/6 macrophages infected with various Leishmania species, such as electron transport/oxidative phosphorylation suppression, 36 MAPK signaling suppression, 35 and apoptosis-associated factor upregulation. 34 Although chemokine production appears to be a consistent response in this model, the identification of other common pathways from gene expression studies in C57BL/6 mice is somewhat limited. Perhaps, this is due to the fact that two of the four studies in C57BL/6 macrophages aimed to identify differentially expressed genes against various susceptible mouse strains (CBA and BALB/c).34,35 One might expect to identify a unique set of differentially expressed genes and pathways using macrophages from susceptible mice infected with Leishmania as the comparator rather than uninfected C57BL/6 macrophages. Additional studies are needed both to pinpoint the specific cytokines as well as chemokines critical to Leishmania infection and to clarify signaling pathways that may be consistently altered across different forms of leishmaniasis.
BALB/c Macrophages
BMD and PD macrophages from BALB/c mice are one of the most widely used experimental in vitro models to study leishmaniasis. 50 Briefly, L. infantum infection in this model induces an initial susceptibility, but at later stages, BALB/c mice are able to control parasite growth through acquired immune mechanisms. 51 To assess how L. donovani affects BMD macrophages from BALB/c mice, comparative microarray analysis was used to relate gene expression profiles of non-infected BALB/c macrophages with those that have been infected. 38 The initial study conducted by Buates and Matlashewski 38 determined L. donovani infection suppressed ~40% of detectable gene expression after four days. Of the 588 well-characterized genes that were assessed, there was a general downregulation in BALB/c gene expression. 38 A significant number of genes were downregulated by approximately two-fold, which suggests that although the magnitude of gene expression changes may not be large, the breadth of small changes to gene expression could have significant overall effects on host cell responses. The authors suggest that this overall suppression of gene expression could be associated with disruptions to both intracellular signaling cascades and general transcription factors. For example, the repression of many genes within a specific battery of pathways might allow the parasite to reproduce while keeping the macrophage viable. Although downregulation of gene expression was more pronounced during L. donovani infection, post-infection induction was also observed. For example, increased expression of macrophage inflammatory proteins (CCL3, CCL4, and CXCL2), known chemo-attractants for macrophages, was detected in L. donovani-infected macrophages compared with non-infected cells.
A separate study investigating both L. donovani- and L. major-induced gene expression after 24 hours (rather than four days) 38 also observed a general repression of gene expression in L. donovani-infected macrophages, but by relatively small amounts. 39 Experiments investigating L. major infection in BALB/c macrophages exhibited a similar gene expression profile to that of L. donovani with only 26 genes differentially expressed between the two Leishmania species using a two-fold threshold cut-off. 39 One of the differentially expressed transcripts was prostaglandin-endoperoxide synthase (Cox2), which was strongly induced by L. donovani, but downregulated by L. major. The authors postulate that the ability of L. donovani to induce Cox2, trigger PGE2 synthesis, and repress TH1 cytokines to favor a TH2 response might play an important role in its visceral nature. 39
A potential role for p53 signaling has been implicated by microarray studies using BMD macrophages from the BALB/c model infected with L. major. 35 Identification of the unique expression of transcripts associated with p53 signaling was seen in BALB/c-derived macrophages, but not C57BL/6-derived macrophages, suggesting that activation of this pathway could be important in susceptibility to L. major. In addition to the studies with L. donovani and L. major, work has also been performed in BALB/c-derived macrophages with L. chagasi. 40 Despite the fact that L. chagasi and L. donovani are both visceralizing species, the gene expression profile of L. chagasi infection was quite different from both L. donovani and L. major in the BALB/c model. Rather than global transcript repression, approximately 80% of genes were unchanged at all the timepoints assayed (1, 4, and 24 hours after infections). In the small percentage of genes that were altered, upregulation of anti-inflammatory response genes and a downregulation of pro-inflammatory genes were observed. 40 Specifically, the mRNA of CCL2, CCL3, and CCL4 decreased modestly, in contrast to the upregulation that was seen in L. donovani infection. 38 During L. chagasi infection, stromal CXCL12, CXCL2, and transforming growth factor-β receptor 2 subunit (TGFβR2) transcripts were all upregulated. In general, L. chagasi infections appear to have little global effect on gene expression in BALB/c macrophages; however, preferential expression of specific genes that enable an anti-inflammatory response could optimize the chance of parasite survival.
Transcriptomic profiling and biological network analysis have also been performed in BALB/c macrophages infected with L. amazonensis. 41 Amastigotes were able to control macrophage expression of early pro-inflammatory components with associated histocompatibility 60 (h60) downregulation and conversion of arginine metabolism to a parasite-supportive pathway. There was additional downregulation of the Toll-like receptor pathway associated with infection, likely preventing inflammatory signaling. This is of interest as Toll-like receptors play a critical role in sensing invasion of pathogens and activation of the innate immune system. 52 Moreover, promastigotes were shown to re-enter macrophages already hosting amastigotes, thus increasing parasitic load and severity of the infection. 41 The downregulation of genes encoding chemokine receptors may explain why L. amazonensis-infected macrophages have reduced emigration and remain cutaneous. Distinct upregulation of the macrophage fatty acid biosynthesis pathway was also observed in macrophages infected with L. amazonensis amastigotes compared with those not infected. The authors postulate that lipids might be impacting the recruitment and retention of key proteins to the area as well as acting as a source of nutrition for the parasites. 41
In summary, a general downregulation of gene expression was observed in BALB/c macrophages on infection with various Leishmania parasites, and many of the differentially expressed genes were involved with inflammatory processes. Some discrepancies do exist within these pathways. For example, upregulation of CCL3 and CCL4 was observed at later timepoints (24 and 96 hours) in both infected BALB/c and C57BL/6 macrophages compared with the uninfected controls,35,38 whereas modest downregulation of these factors was observed at earlier timepoints (4 hours) in a separate study using the BALB/c model. 40 Interestingly, these two studies in BALB/c mice with contradictory expression of CCL3 and CCL4 were in agreement regarding downregulation of CXCL2. Based on these results, more work is needed to elucidate the specific inflammatory factors that are regulated in the BALB/c model. It is worth mentioning that data from the initial study of L. donovani-infected BALB/c macrophages used a 96-hour timepoint and thus assessed gene expression during an established amastigote infection, 38 whereas subsequent studies assayed at ⩽24 hours and therefore might be capturing early-stage infectivity instead.
THP-1
The human mononuclear THP-1 cell line was originally derived from a patient with acute monocytic leukemia. It is used as a human macrophage model for leishmaniasis based on similarities to MDMs. Furthermore, use of THP-1-derived macrophages reduces genetic heterogeneity among samples by limiting the variability observed among MDMs derived from individual donors.
There have been two studies investigating the transcriptomic effects of Leishmania infection in THP-1 cells. The first analyzed global gene expression changes following either L. major infection alone or L. major infection followed by treatment with interferon gamma (IFN-γ). 32 L. major regulates the expression of various genes to facilitate infectivity and virulence while ensuring host cell survival. 53 L. major is capable of repressing IL-12 production, which in turn stimulates IFN-γ and TH1 cell differentiation, thus delaying the macrophage’s ability to mount an anti-Leishmania response. 54 When macrophages were treated with IFN-γ, L. major demonstrated increased survival, suggesting L. major may counteract the IFN-γ response in macrophages. 55 Furthermore, L. major infection was found to alter gene expression broadly, affecting genes involved in cell survival, metabolism, signal transduction, and transport. 32
In the second study, transcriptomic changes in THP-1 cells were investigated in response to either L. donovani exposure alone or L. donovani with the anti-Leishmania agent, sodium stibogluconate. 33 The putative mechanism of action of sodium stibogluconate lies in its ability to interfere with various metabolic and redox signaling pathways in the parasite, which are thought to contribute to its anti-parasitic effects. 33 Less robust gene expression changes were observed in this study compared with the previous report. 32 However, many of the genes modulated in response to L. donovani were also associated with the IFN-γ pathway, corroborating the involvement of IFN-γ and related factors during Leishmania infection. Interestingly, treatment of L. donovani-infected cells with sodium stibogluconate was found to attenuate parasite-induced changes in gene expression. 33
Human Monocyte-derived Macrophages
In an attempt to employ an alternative to the human THP-1 model system, human-based CD14+ peripheral blood-derived monocytes have been isolated and exposed to various pathogens. Human macrophages initiate an inflammatory response to infectious agents through utilization of interleukin 8 (IL-8), tumor necrosis factor alpha (TNF-α), and IL-1β. 42 Based on the examination of macrophage expression profiles post-infection with different Leishmania species, there is evidence that different species of Leishmania induce unique macrophage responses. 49 For example, decreased expression of IFN-γ-induced transcripts and increased gene expression associated with an inflammatory response by L. major compared with L. donovani have been observed. 42 Serial analysis of gene expression (SAGE) to quantitatively analyze the transcriptomes of L. major-infected MDMs suggested that intracellular multiplication of L. major parasites did not seem to dramatically shift the transcriptional programming of macrophages. 44 One transcriptional response that was apparent using SAGE analysis included differential regulation of cytokines, chemokines, and pro-inflammatory mediators. The authors postulate that this selective upregulation and downregulation of pro-inflammatory transcripts could allow L. major to better establish themselves within host macrophages by avoiding potential harm caused by an inflammatory response.
However, the estimation that different species of Leishmania induce unique macrophage responses could be overstated as other studies have found the transcriptional responses across different Leishmania species to be quite alike. For example, L. major and L. donovani infections have been shown to induce a similar transcriptomic response, 42 which was consistent with findings that gene expression profiles were similar between MDMs infected with either L. major or L. amazonensis. 46 In addition, upregulation of transcripts related to inflammatory cytokines and immunomodulation have been observed across multiple Leishmania species. 46 Taken together, these studies with L. major suggest the host macrophage response is relatively independent of the Leishmania species and that the response is more complicated than simply a unidirectional change in inflammatory pathway gene expression.
Moreover, when studying MDMs infected with L. chagasi, addition of autologous Leishmania-naïve T-cells resulted in a pattern of gene expression consistent with type 1 immune cytokine activation (IFN-γ, IL-6, IL-1α, and IL-1β). 43 The addition of T-cells (adaptive) to the infected macrophage (innate) culture was used to assess the unique immune response that occurs on first interaction. L. chagasi infection without T-cells led to the downregulation of transcripts associated with classical inflammation and cellular respiration, whereas the presence of T-cells led to an increase in differentially regulated genes. 43 One such example includes genes that promote acute inflammation, reflecting an environment favorable for TH1 immune cytokine responses. Based on these results, the authors concluded that L. chagasi infection in the presence of the adaptive immune system results in an upregulation of type 1 immune cytokine response and a decrease in genes associated with cellular regulation and classical inflammatory receptors. 43 These studies highlight the importance of the microenvironment for macrophage activation during infection. As referenced throughout this review, leishmaniasis has generally been associated with the type 1 immune cytokine response involving IFN-γ. In a study investigating post-infection with L. (Vianna) panamensis, early timepoints (⩽4 hours) were associated with the most robust changes in gene expression, which included upregulation of cell signaling pathways and inflammation compared with uninfected controls. 45 Consistent with previous findings utilizing other Leishmania species, an upregulation of genes encoding for Toll-like receptor pathway proteins was evident early on during infection as well. 45
Summary
This review covers the key findings from 4 most commonly used macrophage model systems for global transcriptomic studies investigating the host response to Leishmania spp. infection. However, it is worth noting that other human and murine model systems have provided valuable insight toward our understanding of host-parasite interactions at the transcriptomic level (eg, J774G8, U937, RAW264.7, 14M1.4) (Table 1).47-49,56 These studies have identified exciting biomarkers involved in the macrophage response to various Leishmania species, such as NFAT5-mediated TLR expression, 56 7S RNA,47,48 and IRF7. 48 Each of these three biomarkers appears to play a fundamental role in effective control of leishmaniasis by promoting the resistant phenotype. Furthermore, these factors provide detailed mechanistic information on important macrophage-Leishmania spp. interactions during an infection, thus serving as potential leads during drug development for combating not only leishmaniasis but also other pathogen-induced disease states as well. A few studies have investigated global gene expression in the hamster model, which, although less commonly used to study host-Leishmania interactions, more closely resembles infections in the human host. Here, it was found that gene expression varied substantially in vivo and in vitro, with a surprising surge of IFN-γ in hamster infections that did not reduce parasite load and disease progression.57,58 These and other studies demonstrate that significant differences exist between in vivo and in vitro models.
Most of the studies investigating the macrophage-Leishmania interactome from the transcriptomic level focus on the host response rather than parasite gene expression profiles. This is based on the fact that the functional significance of transcriptomic changes in Leishmania is questionable because functionality is regulated at the post-transcriptional level. 59 In other words, gene expression levels in Leishmania do not necessarily correlate with consequent protein abundance. An important factor that should be considered when measuring gene expression changes in macrophages is timepoint selection, as more dynamic changes were apparent at early timepoints.45,46 Therefore, it would appear that measuring early timepoints post infection is important as Leishmania seems to establish a niche within the macrophage with limited communication between host and parasite transcriptomes after the initial infection. 46
One of the major challenges in identifying common factors and pathways among the various gene expression studies of Leishmania-infected macrophages is the heterogeneity of the experimentation. The difficulty in identifying robust and consistent changes among these studies is often attributed to differences in the model system used, sources of macrophages, parasite species, levels of infectivity, transcriptome platforms, data analysis, and infectivity protocols (eg, parasite:macrophage ratios, incubation time of macrophages with parasites, and post-infection timepoints). For example, some macrophages are derived from human primary (MDMs) or in vitro models (THP-1), whereas others isolated from the different mouse models (BALB/c and C57BL/6) are harvested from either bone marrow or the peritoneum. Furthermore, parasite:macrophage ratios ranged from 4:1 to 200:1, and post-infection timepoints for transcriptome analyses ranged between 0.5 and 96 hours. The end result is a vague picture that underscores the challenges researchers face and caution that must be taken when interpreting diverse data sets in hopes of targeting specific genes or gene products for therapeutic purposes.
Despite these challenges, apparent hallmarks of Leishmania-induced expression profiles in macrophages point toward a “general signature of the Leishmania-macrophage infectome.” 46 For example, many of the studies investigating the effects of multiple species of Leishmania (L. major, L. donovani, and L. amazonensis) on an identical model system observed few differences within the host transcriptome.39,42,46 This consistency among three unrelated studies would seem to suggest that the host model system rather than Leishmania species plays a larger role in differentiating one gene expression study from another. This finding is especially surprising as the behavioral characteristics of the different strains as well as the clinical presentations differ, yet the differences in host response they elicit are minimal. Perhaps, one explanation for this observation is the fact that the Leishmania lifecycle is relatively identical across the genus and independent of species.
Regarding specific pathways and biomarkers that were consistently identified across multiple studies, factors related to macrophage activation were unsurprisingly affected. Particularly, cytokines associated with a pro-inflammatory response such as IFN-γ and those related to classic macrophage activation and the TH1 response were implicated. Different pathogens appear to use IFN-γ pathway inhibition as a mechanism to subvert the host response to invasion. 60 Leishmania appears to be no different in its ability to suppress this pathway (Figure 1) as this phenomenon was observed across multiple Leishmania species infecting multiple macrophage model systems. In the in vitro mouse models, some of the most robust changes were related to differential expression of transcripts encoding for cytokines related to inflammation. Macrophages from C57BL/6 mice differentially expressed numerous, yet specific pro- and anti-inflammatory chemokines with transcripts related to cytokine-cytokine receptor interactions consistently being highlighted.34,35,37 Similar mixed results were observed in the BALB/c model as well. This suggests that the changes induced by Leishmania in the macrophage represent a delicate balance to precisely control the host response for survival purposes during an infection. However, human models demonstrated a more consistent pattern of gene expression related to the TH1 response. In THP-1 cells, L. major was able to survive in an IFN-γ-enriched environment, and in human-derived MDMs, downregulation of subsets of IFN-γ-induced genes have been observed.42,44
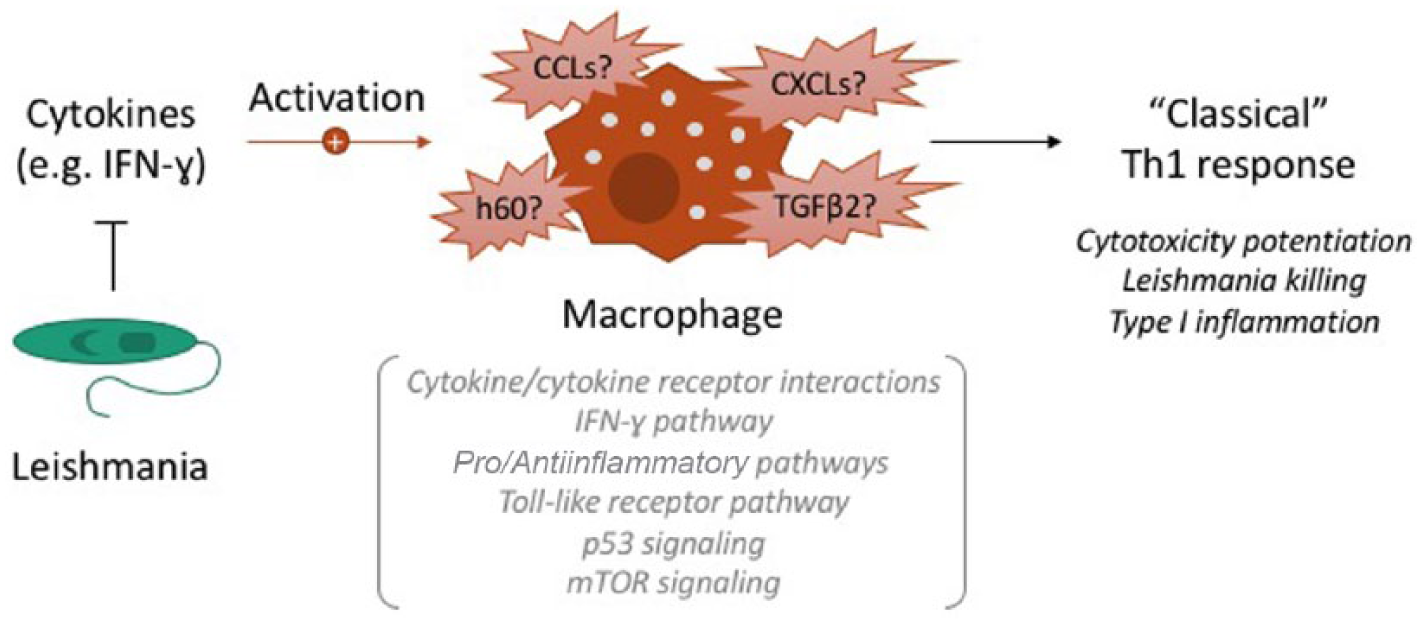
Basic model of transcripts and pathways reported to consistently change in their pattern of gene expression following exposure to Leishmania across transcriptomic studies. IFN-γ indicates interferon gamma; CCLs, C-C motif chemokine ligands; CXCLs, C-X-C motif chemokine ligands; h60, histocompatibility 60; TGFβR2, transforming growth factor-β receptor 2.
In conclusion, despite the challenges that different parasite strains, diverse host models, and varying experimental protocols present, common themes of how Leishmania influences host gene expression are emerging. Parasites affect the host response typically in the early phases of infection and influence mainly the cytokine and global inflammatory responses. Overall, the studies on transcriptional changes in macrophage models have provided only limited insight into the sophisticated mechanisms by which Leishmania parasites influence their hosts to facilitate a successful infection. The fact that no “silver bullet” was identified across all studies and all models is useful in highlighting an important point. When choosing an in vitro macrophage model system to study leishmaniasis, researchers are likely going to struggle to interpret their results across other studies due to differences in experimental design. However, this review does provide researchers with a comprehensive framework from which to make an informed decision on which model to choose based on their research question. The identification of novel transcriptomic targets, as well as the validation of previously implicated pathways involved in macrophage gene expression following Leishmania exposure, does contribute to our understanding of host-parasite interactions. This should, in the long term, lead to improved therapeutic strategies for the treatment of leishmaniasis.
Footnotes
Acknowledgements
The authors thank Jon Taylor, Research Assistant at the Pacific University School of Pharmacy research laboratory, for his insight and assistance in preparing this review article for publication.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Author Contributions
BDS and SCR wrote manuscript and performed literature review; MD compiled targeted literature reviews and contributed to first draft; SS, JT, CT, AS performed targeted literature reviews and helped with writing.
