Abstract
Background:
Clinical biomarkers, allow better classification of patients according to their disease risk, prognosis, and/or response to treatment. Although affordable omics-based approaches have paved the way for quicker identification of putative biomarkers, validation of biomarkers is necessary for translation of discoveries into clinical application.
Objective:
Accordingly, in this study, we emphasize the potential of in silico approaches and have proposed and applied 3 novel sequential in silico pre-clinical validation steps to better identify the biomarkers that are truly desirable for clinical investment.
Design:
As protein biomarkers are becoming increasingly important in the clinic alongside other molecular biomarkers and lung cancer is the most common cause of cancer-related deaths, we used protein biomarkers for lung cancer as an illustrative example to apply our in silico pre-clinical validation approach.
Methods:
We collected the reported protein biomarkers for 3 cases (lung adenocarcinoma-LUAD, squamous cell carcinoma-LUSC, and unspecified lung cancer) and evaluated whether the protein biomarkers have cancer altering properties (i.e., act as tumor suppressors or oncoproteins and represent cancer hallmarks), are expressed in body fluids, and can be targeted by FDA-approved drugs.
Results:
We collected 3008 protein biomarkers for lung cancer, 1189 for LUAD, and 182 for LUSC. Of these protein biomarkers for lung cancer, LUAD, and LUSC, only 28, 25, and 6 protein biomarkers passed the 3 in silico pre-clinical validation steps examined, and of these, only 5 and 2 biomarkers were specific for lung cancer and LUAD, respectively.
Conclusion:
In this study, we applied our in silico pre-clinical validation approach the protein biomarkers for lung cancer cases. However, this approach can be applied and adapted to all cancer biomarkers. We believe that this approach will greatly facilitate the transition of cancer biomarkers into the clinical phase and offers great potential for future biomarker research.
Plain Language Summary
Biomarkers, which are routinely used in clinics, allow better classification of patients according to their disease risk, prognosis, and/or response to treatment. Although affordable omics-based approaches have paved the way for quicker identification of putative biomarkers, validation of biomarkers is necessary for translation of discoveries into clinical application. This research article highlights the challenges of translating cancer biomarkers into clinical practice and summarizes feasible step toward “in silico pre-clinical validation” using the example of lung cancer types. Accordingly, protein biomarkers proposed for lung cancer are being investigated using the “in silico pre-clinical validation” approach to determine whether they have cancer altering properties (i.e., oncoprotein, tumor suppressor, and cancer hallmark), are expressed in body fluids (i.e., plasma/serum, saliva, urine, and bronchoalveolar lavage) and can be targeted with FDA-approved drugs. We believe that the step of in silico pre-clinical validation is the future of biomarker research for all professionals involved in clinical, biological, epidemiological, biostatistical and health research, and that it will greatly facilitate the transition of biomarkers to the clinical phase.
Keywords
Introduction
Cancer is an uncontrolled cell division process that leads to cell transformation due to the occurrence of various genetic changes. 1 It is one of the most common causes of death and the most frequently diagnosed disease in almost every country in the world. According to Global Cancer Statistics, 20 million new cancer cases and 9.7 million cancer deaths were observed in 2022. Among the different types of cancer, lung cancer is the most common cause of cancer death according to Global Cancer Statistics. 2 Recently, the American Cancer Society estimated that lung cancer will be responsible for 234 580 new cancer cases and 125 070 cancer deaths in 2024. 3
The diagnosis of early-stage lung cancer is around 15% and the cancer is usually only diagnosed in the late stages. In addition, the prognosis in the late stage is poor and is less than 5%. 4 Smoking is one of the main causes of lung cancer. In addition, genetic factors and exposure to environmental pollutants increase the risk of developing lung cancer. 5 Histopathologically, lung cancer is divided into 2 main types, small cell lung cancer and non-small cell lung cancer. 6 The prevalence of non-small cell lung cancer is much higher than that of small cell lung cancer. For example, small cell lung cancer accounts for around 13% to 15% of all lung cancer cases. In addition, non-small cell lung cancers are essentially categorized into different subtypes. The most common forms are lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC). 7
Statistics on the incidence and high mortality rate of cancers such as lung cancer show that new clinical strategies are needed, including the development of newer biomarkers or the discovery of novel biomarkers in diagnostics or drug development. For a trait to be called a biomarker, it must be measurable with valid tests, instruments, or assays, indicate differences between different phenotypes and be used specifically for medical practice (i.e., for diagnosis, prognosis, or response to a treatment). 8 But are these properties sufficient to indicate that these biomarkers have clinical utility and are ready to be analyzed in various clinical qualification studies? In other words, once biomarkers have been discovered, should the analytical and clinical testing of biomarkers begin immediately? In our opinion, they should not. Many conditions can be investigated or qualified before promising biomarkers can be transferred to clinical validation, which we have termed “in silico pre-clinical validation” in this study. With the advent of next-generation technologies, omics approaches (genomics, transcriptomics, proteomics, metabolomics, lipidomics, and microbiomics), bioinformatics, and big data, biomarker discovery is now moving quite rapidly toward clinical translation, and we believe that any in silico pre-clinical validation initiative can help solve the current challenges in validating biomarkers9,10 and pave the way for their use in the clinic.
Although each method has advantages and disadvantages depending on the application, the advantages of in silico approaches can be listed as follows: (i) it is a high-throughput method that allows researchers to screen large amounts of data efficiently, quickly, and simultaneously, (ii) since it is a high-throughput method, the time and resources required are much less than compared to other approaches,(iii) combining in silico studies with other approaches is much easier than with other approaches.11,12 Since promising biomarkers that are discovered should have specific characteristics and stringent properties, these properties can be efficiently analyzed with the in silico step (in silico pre-clinical validation step) and then the effects on human health outcomes should be approved experimentally (i.e., with clinical trials) for routine use in clinics.
Generally, the priority in the field of biomarkers is to validate the discovered biomarkers in an appropriate clinical context and to integrate them into the clinic. Considering the advantages of in silico methods and the requirements in the field of biomarkers, our main objective in this study is to apply and illustrate 3 different sequential in silico pre-clinical validation steps in lung cancer cases that we have proposed to help the possibility of translating cancer biomarkers into the clinic. In brief, we collected the reported protein biomarkers for 3 lung cancer types and performed 3 in silico pre-clinical validation steps: (i) evaluating whether the biomarkers function as tumor suppressors or oncoproteins and act as cancer hallmarks (i.e., cancer altering properties), (ii) evaluating whether they are expressed in body fluids, and (iii) evaluating whether they can be targeted with FDA-approved drugs (Figure 1). Overall, with this study we have proposed 3 novel sequential in silico pre-clinical validation steps that can be easily adapted for all cancer biomarkers to improve the quality of biomarkers and give them the confidence to qualify for clinical trials.

The overall approach implemented for the lung cancer types in the study.
Materials and Methods
Collection of lung cancer protein biomarkers
We have collected the reported protein biomarkers for lung cancer, LUAD, and LUSC from the BIONDA database (BIOmarker and biomarker caNdidates DAtabase) (as of May 2023). This database provides researchers with molecular biomarkers such as gene, protein, and miRNA biomarkers as well as biomarker candidates, which are published in scientific article abstracts and updated monthly using text mining. 13 We focused on protein biomarkers in this study and collected only the protein biomarkers. When we compare the different types of molecular biomarkers, we can say that mRNAs are less stable than DNAs due to the differences in molecular structure, which is important for the reproducibility of biomarker measurements. 14 However, proteins, which are among the most important functional molecules, can be highly dysregulated under different conditions. Therefore, the differential expression of proteins in cells under different conditions is important for identifying and understanding biological information. For these reasons, we believe that protein biomarkers can take a step forward compared to other types of molecular biomarkers in the clinic.
In addition, the reported protein biomarkers for lung cancer collected include all forms of lung cancer without specifying subtypes and may therefore include cases of LUAD and LUSC. Since it was hypothesized that the biomarkers for lung cancer currently used or recommended for use in the clinic should be different depending on the histological subtype of lung cancer,15 -18 we included the unspecified lung cancer cases, LUAD, and LUSC cases in the study to investigate the similarities and differences of the reported biomarkers and the extent of their overlap.
Moreover, in order to understand the molecular functions of the collected protein biomarkers for 3 types, the PANTHER (protein analysis through evolutionary relationships) classification database 19 was used.
Evaluation of protein biomarkers cancer altering properties
Cancer is a multi-stage development characterized by genetic changes that eventually lead to abnormal cell growth. Oncogenes and tumor suppressors are the 2 different classes of genes that can promote the development of malignant cells through different mechanisms such as gain of function and loss of function. 20 While oncogenes are genes whose activation contributes to the development of cancer, tumor suppressor genes are genes whose suppression contributes to the development of cancer. 21 Since the presence or absence as well as the expression level and pattern of oncogenes and/or tumor suppressors may correlate with malignant transformation (i.e., disease development and progression) or treatment, their contribution to clinical outcome is evident. Therefore, to assess the cancer altering properties of the biomarkers, we first used the Catalog of Somatic Mutations in Cancer (COSMIC) database (v.95) 22 and obtained a list of proteins that have tumor suppressor or/and oncogene functions.
In 2000, researchers wondered that while there are many types of cancer, how many or whether the same regulatory mechanisms or signals affect normal cells and turn them into cancer. After pondering these questions, the researchers proposed the concept of “cancer hallmarks,” which states that these hallmarks play a crucial role in cancer development and are probably present in all types of cancer. The 10 cancer hallmarks approved today include (i) proliferative signaling, (ii) suppression of growth, (iii) escaping immune response to cancer, (iv) cell replicative immortality, (v) tumor promoting inflammation, (vi) invasion and metastasis, (vii) angiogenesis, (viii) genome instability and mutations, (ix) escaping programed cell death, and (x) change of cellular energetics. 23
Mostly, the hallmarks of cancer genes are considered as driver genes of tumorigenesis. These genes can orchestrate for the most basic phenotypic features of tumor initiation and progression. Focusing on genes that play an important role in carcinogenesis and considering confirmed associations with cancer, in short, using genes that have been presented as cancer hallmarks as measurable indicators, such as biomarkers, can be very beneficial for the diagnosis and treatment of cancer. Therefore, it can be assumed that biomarkers that exhibit cancer hallmark features allow the identification of the cancer, the characterization of the cancer subtype or the detection of its biological characteristics in a more effective way. Accordingly, it can be assumed that a cancer biomarker with cancer hallmark properties will increase the quality of biomarkers and take them a step further in terms of their reliability in clinical studies. In this study, ten cancer hallmark genes were retrieved from a publicly available biological repository, the Cancer Hallmark Genes (CHG) database. 24 GeneCards: The Human Gene Database 25 was used to determine the proteins encoded by the cancer hallmark genes.
Evaluation of protein biomarkers presence in biological body fluids
Biological body fluids are the fluids that the human body either excretes or secretes. Biological fluids such as blood, plasma, urine, and saliva have high potential for non-invasive biomarker measurements compared to tissue, organ samples or cell extracts.26,27 It can be assumed that the collection of body fluids is non-injurious, simple, fast and inexpensive and that the examination of biomarkers from biological fluids in clinics is an important feature and advantage that should be considered. To discover the invasiveness of the culminated protein biomarkers with cancer altering features, the expression of proteins in plasma/serum, saliva, urine and bronchoalveolar lavage (BAL) was investigated using HBFP: a new repository for human body fluid proteome source. 28
Evaluation of protein biomarkers druggabilities
Once biomarkers have been discovered, the next step would be to find out if these biomarkers are druggable, that is, if the biomarker can be targeted with a drug, especially an FDA-approved drug. In this way, oncologists can offer their patients an appropriate choice of drugs and/or drug combinations, and patients can be assured that the drugs they use are carefully controlled and approved for use.29,30 Therefore, to investigate whether protein biomarkers can be used as drug targets, we used the Pharos database 31 to identify the presence of approved drug targets of biomarker proteins.
Results
Extraction of protein biomarkers
A total of 3407 different protein biomarkers for lung cancer (3008 protein biomarkers), LUAD (1189 protein biomarkers), and LUSC (182 protein biomarkers) were extracted from the database. To reveal the potential of the biomarkers for stratifying lung cancer patients into different types, we also comparatively analyzed the identified biomarkers. Of the 3407 protein biomarkers, 105 were identified for all analyzed types. In addition, 364 and 16 protein biomarkers specific to LUAD and LUSC, respectively, were identified. Also 701 common protein biomarkers for lung cancer and LUAD, 42 common protein biomarkers for lung cancer and LUSC and 19 common protein biomarkers for LUAD and LUSC were identified (Figure 2A).
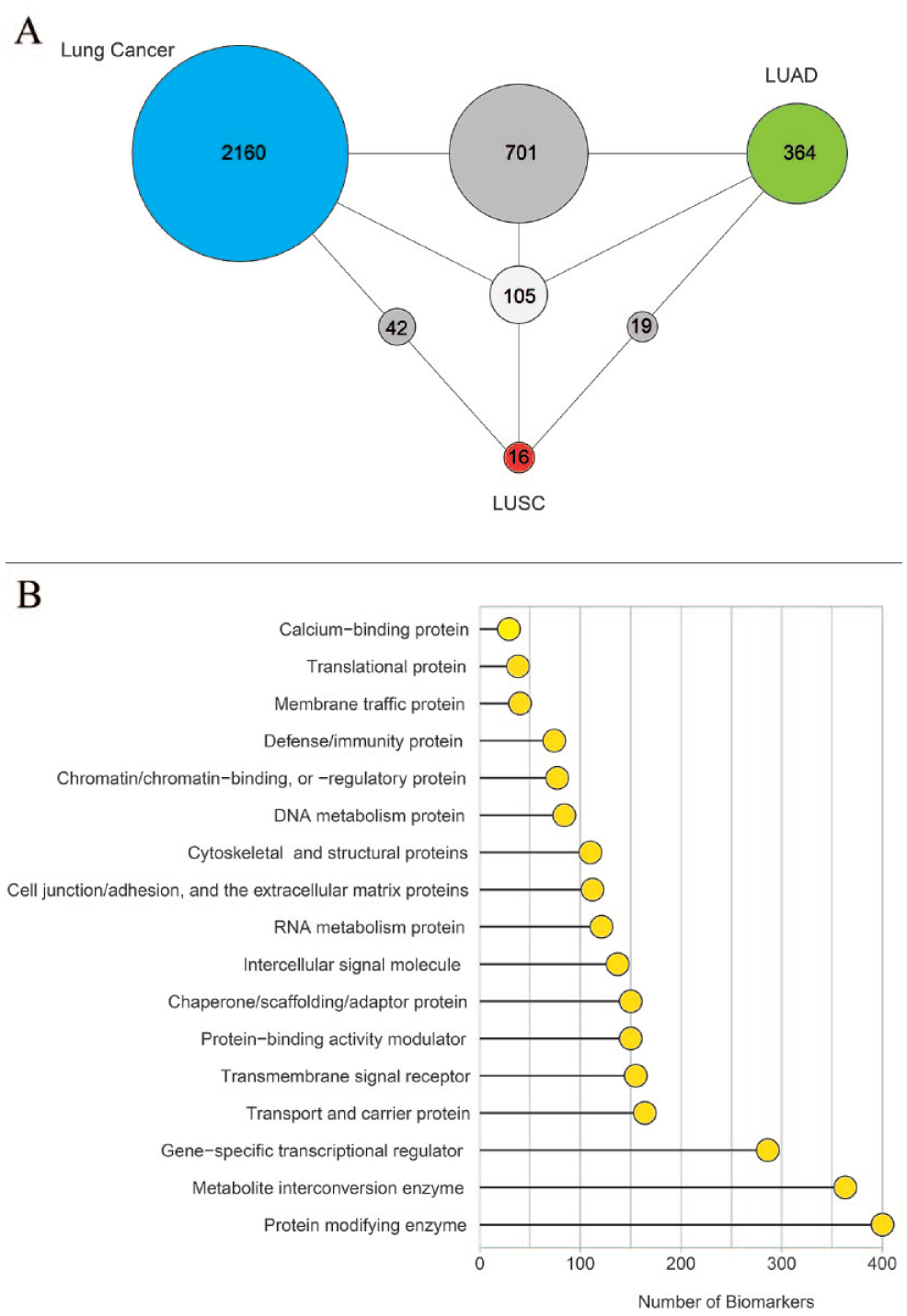
The number and molecular functions of the obtained protein biomarkers for lung cancer: (A) the Venn diagram representing the number of protein biomarkers collected for lung cancer, LUAD, and LUSC from the database and (B) the diagram representing the molecular functions of all protein biomarkers obtained.
It was found that the identified biomarkers have different molecular functions. However, the focus is on the functions of protein modification (400 protein biomarkers), metabolite conversion (363 protein biomarkers) and gene-specific transcriptional regulation (286 protein biomarkers) (Figure 2B).
In silico pre-clinical validation step 1: Lung cancer biomarkers cancer altering properties
It is clear that oncogenes and tumor suppressors play an important role in cancer development through loss of function or gain of function. If we compare a protein with oncogenic and tumor suppressor function with a protein without these functions, we can assume that the protein with oncogenic or tumor suppressor function is a more suitable candidate for a biomarker and can be used more efficiently as a cancer biomarker in clinics. Moreover, since the genes involved in these ten cancer hallmarks are drivers of tumor development, it is important to study these genes in different cancer types to understand the process of tumor development and use them as biomarkers. 32 Accordingly, a total of 557 different proteins that have an oncoprotein function, a tumor suppressor function or both functions were extracted from the COSMIC database 22 and a total of 374 different hallmark proteins were extracted from the CHG database 24 and these lists were integrated with our protein biomarker lists respectively.
In lung cancer, only 8.57% of the 3008 protein biomarkers, in LUAD only 11.85% of the 1189 protein biomarkers and in LUSC only 15.38% of the 182 protein biomarkers have at least oncoprotein or tumor suppressor functions. Of these protein biomarkers, 23 are common in all the 3 cancer types. In addition, 98 protein biomarkers common to lung cancer and LUAD and 5 protein biomarkers common to lung cancer and LUSC were identified that have at least oncoprotein or tumor suppressor functions (Figure 3A). Among these functions, the protein biomarkers with an oncoprotein function (46.90%, 49.65%, and 46.43% of protein biomarkers of lung cancer, LUAD, and LUSC, respectively), then the protein biomarkers with a tumor suppressor function (36.82%, 35.46%, and 35.71% of protein biomarkers of lung cancer, LUAD and LUSC, respectively) and finally protein biomarkers with both functions (16.28%, 14.89%, and 17.86% of protein biomarkers of lung cancer, LUAD, and LUSC, respectively) are the most represented in each of the 3 types (Figure 3B). It is assumed that a tumor suppressor gene is mutated, inactivated, or downregulated, while oncogenes are activated or upregulated in cancer cells. Since tumor suppressors are reduced in cancer cells under normal conditions, it is more difficult to detect tumor suppressors as biomarkers in cancer patients compared to oncoproteins. 33 Although many proteins are eliminated at this stage, it is therefore desirable that biomarkers with oncoprotein functions become evident.

In silico pre-clinical validation Step 1: The investigation of the cancer altering properties of biomarkers for lung cancer: (A) the comparative plots of protein biomarkers for 3 lung cancer types (lung cancer, LUAD, and LUSC) showing at least one oncogenic or tumor suppressive feature, (B) the distribution of oncogenic and/or tumor suppressive protein biomarkers, (C) the comparative diagrams of protein biomarkers for 3 types showing both oncogenic or/and tumor suppressive and cancer hallmark features, and (D) the percentage distribution of protein biomarkers, which cancer hallmark feature they have among the 10 different cancer hallmarks.
When these protein biomarkers were examined to determine whether they function as cancer hallmark proteins, 64.34%, 65.24%, and 78.57% of the protein biomarkers in lung cancer, LUAD, and LUSC, respectively, were found to function as hallmark proteins in addition to their oncoprotein and/or tumor suppressor function. Of these protein biomarkers, 18 are found in all 3 types. In addition, 64 protein biomarkers common to lung cancer and LUAD and 4 protein biomarkers common to lung cancer and LUSC were identified that have at least an oncoprotein or tumor suppressor function and were reported as cancer hallmarks (Figure 3C). Of the ten cancer hallmarks for all 3 cancer types, we can report that protein biomarkers play a prominent role in the following 3 hallmarks: invasion and metastasis (66.87%, 63.04%, and 72.73% of protein biomarkers of lung cancer, LUAD, and LUSC, respectively), escape from programed cell death (63.86%, 69.57%, and 63.64% of protein biomarkers of lung cancer, LUAD and LUSC, respectively), and proliferative signaling (57.23%, 60.87%, and 40.91% of protein biomarkers of lung cancer, LUAD, and LUSC, respectively) (Figure 3D). It is evident that one of the main targets of cancer cells in the early stages of malignancy development is profound proliferation. Moreover, targets that play an important role in promoting proliferation are still one of the best targets in cancer therapy. 34 Since we are studying a cancer, we believe that the biomarkers that have been or are being used in daily life for this disease should also have oncogenic features to accurately represent the disease.
We collected a total of 3407 different protein biomarkers for 3 cases, of which 176 (5.16%) have at least an oncogenic or tumor suppressive function and are presented as cancer hallmarks, that is, have cancer altering properties (S. Table 1).
In silico pre-clinical validation step 2: Detection of lung cancer biomarkers in body fluids
We can assume that one of the biggest challenges in transferring biomarkers to the clinic is clinical sampling. Non-invasive or minimally invasive collection and measurement of biomarkers is important for rapid, economical, and repeatable analyses. 35 Overall, the protein biomarkers identified for 3 types that have cancer altering properties (176 protein biomarkers) can also be obtained from at least 1 of the 4 biological fluids analyzed, with a value of about 72% (127 protein biomarkers) (S. Table 1). Accordingly, 58 of the protein biomarkers specific to lung cancer, 5 of the protein biomarkers specific to LUAD, 48 of the protein biomarkers common to lung cancer and LUAD, 3 of the protein biomarkers common to lung cancer and LUSC, and 13 of the protein biomarkers common to 3 cancer types were expressed in at least 1 of the body fluids investigated (Figure 4A). To determine more precisely which body fluid(s) (i.e., plasma/serum, saliva, urine and BAL) provide the best results, we comparatively analyzed the expression of the 127 protein biomarkers. This showed that for biological sampling, fluids such as urine (expressed 96 of the 127 protein biomarkers) and plasma/serum (expressed 92 of the 127 protein biomarkers) alone are more suitable for analyzing/measuring protein biomarkers. Of the protein biomarkers examined, 25.81% of the proteins in lung cancer, 28.36% of the proteins in LUAD and 50% of the proteins in LUSC were found to be expressed in both plasma and urine (Figure 4B). In contrast, 25 of the 127 protein biomarkers and 34 of the 127 protein biomarkers were expressed only in saliva and BAL, respectively.
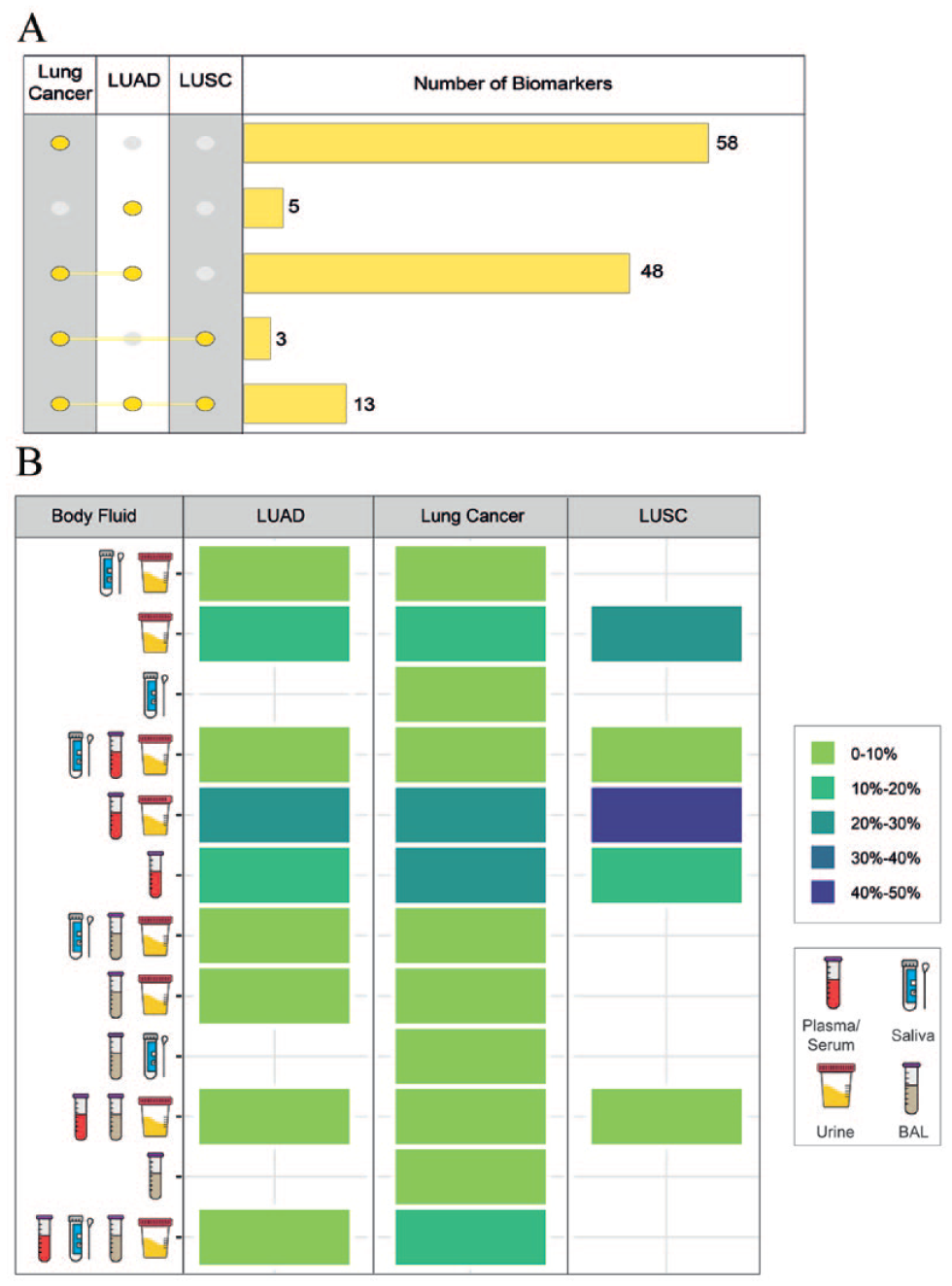
In silico pre-clinical validation Step 2: The analysis of lung cancer biomarkers in body fluids (A). The comparative visualization of protein biomarkers that exhibit oncogenic properties and are present in at least one of the four body fluids. (B) The distribution of protein biomarkers according to their occurrence in the body fluids.
In silico pre-clinical validation step 3: Druggability of lung cancer biomarkers
The fact that a biomarker has an approved target drug makes that biomarker a frontrunner, and this is a feature that is desired by oncologists. Accordingly, a total of 30 of the 127 protein biomarkers (23.62%) that have cancer altering properties and are expressed in at least 1 of the biological fluids analyzed also have FDA-approved drug target proteins. Of the 30 protein biomarkers, 2 biomarkers (FES and XPO1) were found to be specific for LUAD and 5 biomarkers (POLE, ATP1A1, PDGFRA, KDR, and ESR1) were found to be specific for lung cancer. In addition, 17 biomarkers were found to be present in both lung cancer and LUAD, and 6 biomarkers (EZH2, PIK3CA, MET, FGFR1, FGFR2, and EGFR) were found in all 3 cases studied (Figure 5). There are at most 27 FDA-approved drugs for the protein biomarker ESR1, followed by 26 FDA-approved drugs for EGFR and 17 FDA-approved drugs for KDR.

In silico pre-clinical validation Step 3: the investigation of druggability of lung cancer biomarkers. The bubble diagram shows the number of protein biomarkers targeted by FDA-approved drugs.
Discussion
New technologies are constantly being developed to identify molecular markers for complex diseases. Researchers have used various systems biology approaches, bioinformatics, and machine learning pipelines36 -40 and experimental techniques41,42 to discover biomarkers for all types of cancer. Although these new technologies have enabled researchers to find biomarker candidates using simpler approaches, these biomarkers should be pre-validated and specialized for use in clinical practice and have specific properties to be identified as actionable biomarkers.
Strengthening cancer prevention and prognosis measures and creating a sustainable infrastructure for cancer treatment are crucial to the global fight against cancer. This health challenge can be reduced with the help of the potential biomarkers. As biomarkers can guide the optimization of decisions in clinical practice, these can meet the needs of the healthcare system. 43 However, the question is whether the researchers who discover new biomarkers can also apply them in clinical practice? Contrary to scientific belief, the translation of biomarkers into the clinic is low. For example, the PD-L1 biomarker was approved by the FDA in 2018, the EGFR protein biomarker was approved by the FDA in 2012, the ALK biomarker was approved by the FDA in 2011, and the CEA biomarker was approved by the FDA in 1985 and is currently used in clinical practice for lung cancer. 44
To translate the potential biomarkers into clinical benefit, extensive standard requirements must be met, which can be efficiently assessed by in silico pre-clinical validation approaches, of which we have proposed 3 in silico pre-clinical steps in the study. We believe that when we talk about cancer biomarkers, these promising biomarkers should point to cancer altering properties. In cancer, most molecular targeted therapies are still inhibitors of oncogenes or/and their associated signaling pathways.45,46 In addition, minimally invasive biomarkers are desirable in clinical application and the treatability of the biomarker is another important feature that may be coveted by oncologists. In the face of lung cancer types, in this study we obtained a total of 3008 protein biomarkers for lung cancer, 1189 protein biomarkers for LUAD and 182 protein biomarkers for LUSC. After the applying of the 3 in silico pre-clinical validation steps, these values decreased to 28, 25 and 6 protein biomarkers for lung cancer, LUAD and LUSC respectively (S. Figure 1), clearly demonstrating that several of the reported biomarkers have great potential for clinical testing or use in clinics. Overall, with these 3 sequential in silico pre-clinical validation approaches, molecular cancer biomarkers can be screened according to specific criteria and help researchers to identify robust biomarkers that need to be investigated experimentally. We therefore believe that this approach can make clinical trials easier (i.e., subject to budget, time and equipment requirements) and more affordable for researchers/experts.
In addition to the 3 in silico pre-clinical validation approaches that we have employed in this study to support the translation of biomarkers to the clinic, the following in silico approaches can also be investigated: (i) receiver operating characteristic (ROC) curve analysis, (ii) Kaplan-Meier analysis, and (iii) supervised and unsupervised machine learning approaches. The ROC analyses can be used to understand the diagnostic power of biomarkers by assessing both the sensitivity and specificity of a biomarker. 47 Kaplan-Meier analysis can be used to understand the prognostic power of biomarkers by assessing the specific time to event. 48 For biomarker assessment unsupervised machine learning algorithms that includes clustering (i.e., hierarchical and K-means clustering), association (Apriori algorithm) and dimensionality reduction (principle component analyses) approaches and supervised machine learning algorithms including regression (i.e., linear regression and decision tree) and classification (i.e., logistic regression, support vector machines, and naïve bayes) approaches can be used to assess patients and control groups discrimination capacity as well as to evaluate the disease prediction efficiency.49,50 Therefore, these in silico pre-clinical validation analyses could also reveal the biomarkers that are truly desirable for clinical investment.
Conclusion
While “omics” technologies and/or in silico approaches can accelerate biomarker discovery and increase the possibility of biomarker translation to the clinic, these technologies can also improve existing approaches and pave the way for easier and faster integration of discovered biomarkers into clinical applications. Translating a promising biomarker discovery into clinical practice requires large-scale clinical trials. However, the challenges posed by these large-scale studies can be reduced or overcome by in silico pre-clinical validation approaches which we have proposed in this study. As protein biomarkers are becoming more prominent in the clinic alongside other molecular biomarkers, and lung cancer is the most common cause of cancer-related deaths, in this study we have used the protein biomarkers of lung cancer as an illustrative example of a in silico pre-clinical validation approach that can be applied to any cancer type. We recommend that these in silico pre-clinical validation steps should become a routine procedure for all clinical, biological, epidemiological, biostatistical and health research professionals.
Supplemental Material
sj-jpg-1-bmi-10.1177_11772719241287400 – Supplemental material for Biomarkers From Discovery to Clinical Application: In Silico Pre-Clinical Validation Approach in the Face of Lung Cancer
Supplemental material, sj-jpg-1-bmi-10.1177_11772719241287400 for Biomarkers From Discovery to Clinical Application: In Silico Pre-Clinical Validation Approach in the Face of Lung Cancer by Medi Kori, Esra Gov, Kazim Yalcin Arga and Raghu Sinha in Biomarker Insights
Supplemental Material
sj-xlsx-2-bmi-10.1177_11772719241287400 – Supplemental material for Biomarkers From Discovery to Clinical Application: In Silico Pre-Clinical Validation Approach in the Face of Lung Cancer
Supplemental material, sj-xlsx-2-bmi-10.1177_11772719241287400 for Biomarkers From Discovery to Clinical Application: In Silico Pre-Clinical Validation Approach in the Face of Lung Cancer by Medi Kori, Esra Gov, Kazim Yalcin Arga and Raghu Sinha in Biomarker Insights
Footnotes
Acknowledgements
Not applicable.
Declarations
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
