Abstract
Human biobanks are recognised as vital components of translational research infrastructure. With the growth in personalised and precision medicine, and the associated expansion of biomarkers and novel therapeutics under development, it is critical that researchers can access a strong collection of patient biospecimens, annotated with clinical data. Biobanks globally are undertaking transformation of their operating models in response to changing research needs; transition from a ‘classic’ model representing a largely retrospective collection of pre-defined specimens to a more targeted, prospective collection model, although there remains a research need for both models to co-exist. Here we introduce the Health Science Alliance (HSA) Biobank, established in 2012 as a classic biobank, now transitioning to a hybrid operational model. Some of the past and current challenges encountered are discussed including clinical annotation, specimen utilisation and biobank sustainability, along with the measures the HSA Biobank is taking to address these challenges. We describe new directions being explored, going beyond traditional specimen collection into areas involving bioimages, microbiota and live cell culture. The HSA Biobank is working in collaboration with clinicians, pathologists and researchers, piloting a sustainable, robust platform with the potential to integrate future needs.
Translational research in medicine aims to accelerate the introduction of improvements in patient treatment and care. The translational pipeline (Figure 1) describes stages wherein progressive areas of research are referred to as: T1, translation of basic laboratory research into human trials; T2, translation of successful trials into clinical practice; and T3, promoting new policy and practice to achieve system-wide change. 1
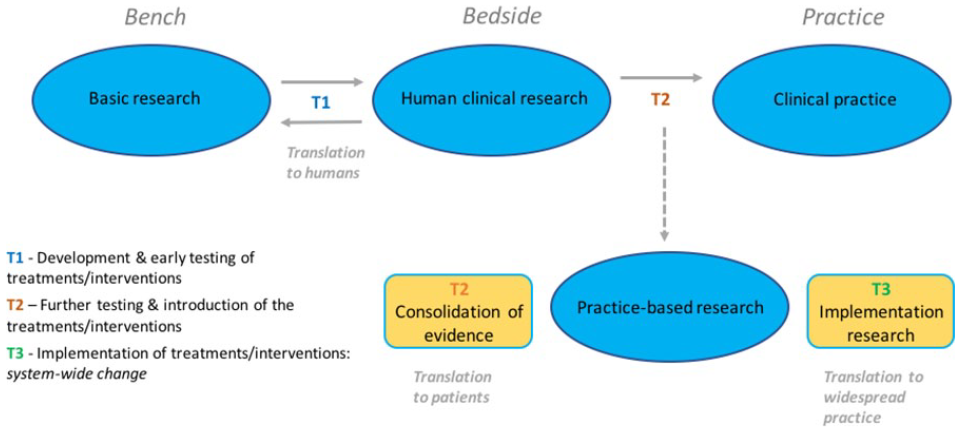
The translational research pipeline (adapted from Westfall et al 1 ).
Human biobanks are recognised as vital components of translational research infrastructure2–4 (Figure 2) and are central in facilitating T1/T2 research in particular. There is high demand for access to human biospecimens to answer research questions; increasingly these are replacing the traditional ‘starting points’ in basic research, that is, cell lines and animal models. 2 Indeed, with the growth in personalised and precision medicine, and the associated expansion of biomarkers and novel therapeutics under development, research efforts to expedite introduction of new advancements in patient care can be strengthened by working with biobanks.5,6 It is critical for the advancement of personalised medicine in particular that researchers can access a strong collection of patient biospecimens, annotated with clinical and pathological data. 7 The considerable burden placed upon individual researchers in organising their own collection of suitable samples, justifies this clear translational need; consequently, over recent decades this has resulted in a global trend for human biospecimen collection and biobanking, on different scales and utilising different models.
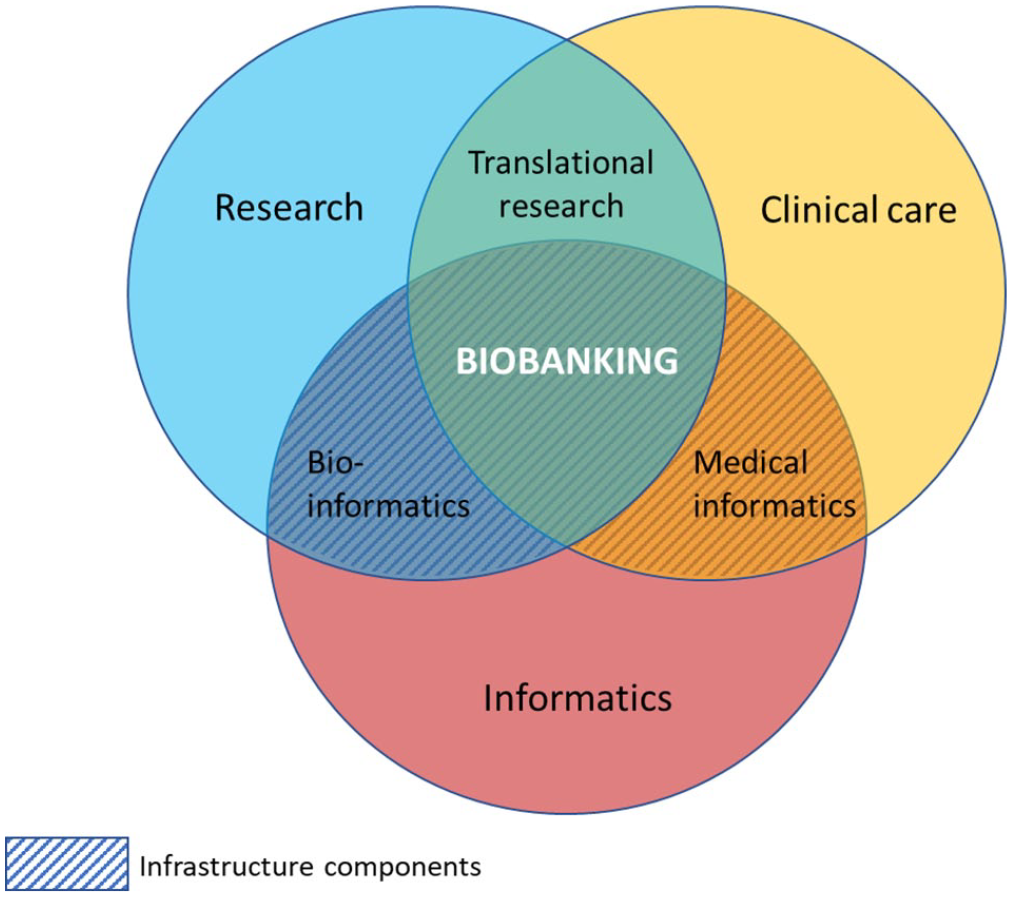
Biobanking is a core component of translational research in medicine.
Cancer researchers are amongst the most prolific users of biobanks. In a Canadian study, of 445 cancer research papers published from 2010 to 2014, 30% of these were found to have used specimens obtained directly from biorepositories 8 ; also, a recent literature review showed an exponential increase in PubMed publications relating to biobanking over time, with the highest number in the area of cancer research. 3
Although the need for cancer biobanks is well established, which models can be sustained, and best suit researcher requirements is a subject of debate. Operating models include the ‘classic’ (specimen-centric) model wherein specimens are collected on the basis of predetermined characteristics, and the ‘prospective’ (investigator-centric, hypothesis driven) model where specimen collection is targeted for specific research questions.
Much discussion has surrounded the advantages and disadvantages of these operational models: for example, classic collections have been criticised for ‘hoarding’ specimens without strategic plans for distribution, which both wastes resources and presents a moral dilemma in that patients have donated specimens intending they be used in research but without the guarantee that this will happen. 9 Prospective collections, though targeted and therefore utilised, have the disadvantages of both the time needed to accumulate specimens as well as a lack of time to allow for follow-up data 4 ; logistics involved in filling bespoke research requests means these collections can also be very resource intensive.
Lessons can be learnt from the many classic biobanks as to how to plan more strategic future collections but also how to maximise distribution of current stocks. 9 A common experience has been the collection of specimens without proper consideration of potential future use; a greater understanding of local research interests drives narrower, better targeted collection of specimens, with a higher chance of utilisation. In practice, it is also easier to facilitate these narrower collections as clinician-researchers local to the biobank are more likely to engage with collection of patient specimens. However, despite the criticisms of classic biobanks, wholesale abandonment of these repositories would presume that the nature of future scientific research is settled, when the emergence of new technologies may confer new importance upon existing specimens. One example is the recently expanded utility of formaldehyde-fixed, paraffin embedded (FFPE) blocks which now have additional research value through the evolution of molecular techniques enabling the use of fixed tissue in genomic studies6,10,11; the maturity of many of these specimens means that accompanying clinical outcome data is available.
Here we present the Health Science Alliance (HSA) Biobank, as a hybrid model. We describe our classic operating model wherein collection of pre-defined specimens has been the default and how we are refining this and now combining it with a prospective approach to collection, recognising the need to be agile in response to changing research needs. New directions are being explored, going beyond traditional specimen collection into areas involving bioimages, microbiota, live cell culture, organoids, 3D tissue models and tissue explants. The HSA Biobank is working in collaboration with clinicians and researchers, piloting and developing a robust platform with the potential to integrate future needs.
The HSA Biobank: Establishment of the biobank
The HSA Biobank was established in 2012 as a collaborative partnership between the University of New South Wales (UNSW Sydney, Australia), the South Eastern Area Laboratory Service, SEALS (now NSW Health Pathology; NSWHP) and the South Eastern Sydney Local Health District (SESLHD). This was established through a Translational Cancer Research Centre (the Translational Cancer Research Network, TCRN), funded by the Cancer Institute NSW (CINSW), a NSW state government cancer control agency. Participating hospitals are those within the Local Health District: The Prince of Wales Hospital (POWH), Prince of Wales Private hospital (POWP), Royal Hospital for Women (RHW), St George Hospital (SGH), St George Private Hospital (SGPH), The Sutherland Hospital (TSH); in addition, there is a regional participating institution, Border Medical Oncology (BMO) located in Albury, NSW.
The HSA Biobank was developed as a cancer biobank, aiming to consent all patients having surgery at participating hospitals for a known or suspected cancer diagnosis; the HSA Biobank is an open collection, any researcher is able to apply to access specimens and/or data on a cost-recovery basis, subject to ethical approval.
The HSA Biobank has a robust governance structure (Figure 3) to ensure that ethical obligations are met and sustainability issues are addressed: The HSA Biobank Management Committee oversees strategic direction of the biobank, and the Research Access Committee (RAC), comprised of scientists and clinicians, assesses each research request submitted. A Material Transfer Agreement (MTA) is required for release of specimens and/or data to researchers external to UNSW Sydney; the MTA has been developed to align with the Australian National Health and Medical Research Council (NHMRC) National Statement on Ethical Conduct in Human Research as well as ISBER Best Practice Guidelines.

HSA Biobank governance structure and process for access to biospecimens and data. Strategic direction is provided by a management committee, with access to ex-officio advisors. When requests for specimens and/or data are received, they are assessed by the Research Access Committee who can consult additional expert clinicians, as necessary. Specimens and/or data can be dispatched on receipt of RAC and ethical approvals; an MTA is required for transfer outside of UNSW Sydney.
The original operating model of the HSA Biobank aimed for a minimum of an FFPE tissue block allocated for banking by an anatomical pathologist, a peripheral blood sample collected with plasma and cell pellet banked, and fresh tissue banked when possible. For known or suspected haematological malignancies, bone marrow biopsy specimens are collected where possible for both cryopreserved cells and cell pellets. Specimens are housed in the UNSW Biorepository and along with associated data are managed through an open-source biobanking software solution, OpenSpecimen 12 ; this ensures that the provenance of all specimens is tracked and interrogation for research requests can be expedited (Figure 4).
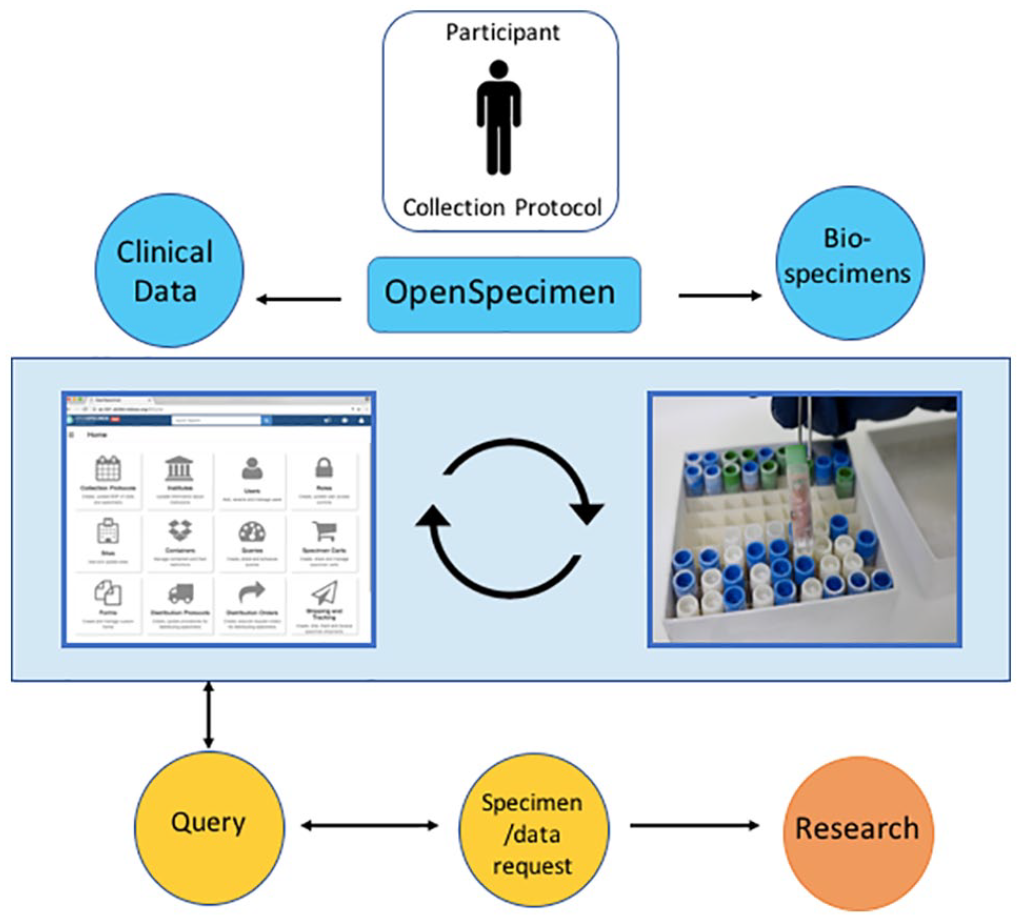
HSA Biobank specimens and associated data are managed using an open-source biobanking software solution, OpenSpecimen. 12 OpenSpecimen permits open access from multiple sites with role-based permissions; it is Collection Protocol driven, with specimens ‘attached’ to each participant.
Patient consent
Patient consent is requested by clinicians (consultants, registrars, nurses) typically during pre-admission appointments, using a SESLHD consent form – the use of an official LHD form is vital to the recognition and acceptance of the biobank consent process within hospital workflows; post-operative consent can also be requested. Consent is sought for collection of biospecimens (tissue/fluid/blood/saliva) and access to clinical hospital information; from 2013 permission was also sought from patients for linkage to the Medicare Benefits Schedule (MBS, ‘Medicare’; government-funded health services other than in-hospital services) and Pharmaceutical Benefits Scheme (PBS; pharmaceutical use) data; all consent is for unspecified future research. Importantly, neither the HSA Biobank participant consent form or the accompanying information brochure refer to ‘cancer’; this enables consent to be sought when diagnosis is not confirmed without causing undue distress, and potentially provides capacity for expansion of the consent model for use in disease streams other than cancer.
Linking the university and hospital IT systems using secure health-grade IT infrastructure
Timely availability of surgical pathology reports is vital for biobanking and the HSA Biobank has developed a system to facilitate this. Pathology reports from the NSWHP laboratory information management system (LIMS), are delivered in real-time into OpenSpecimen via HL7 messaging. The reports are sent only after patient consent has been confirmed and recorded in the NSWHP LIMS. The reports are linked to the consented patients in OpenSpecimen using a machine learning based ensemble algorithm, 13 and they are de-identified and replaced with surrogate information using a deep learning-based algorithm. 14
Annotation of specimens
While data requests from individual researchers will vary, there are common data items requested regularly, for example, the grade, stage and anatomical site of biospecimens. The HSA Biobank has worked collaboratively with other NSW-based biobanks within a ‘NSW Cancer Biobank Stakeholder Network’ (BSN) to establish a minimal data set for annotation of banked specimens and to map the sources for the different data items within each Local Health District. 15 Data for biospecimen annotation is extracted from, or linked to several sources: demographic information is sourced from hospital records, clinical diagnosis and tumour morphology are sourced from pathology reports and electronic medical records (eMR), and as mentioned, the history of health services and prescription use are sourced from MBS and PBS records. Survival data is procured through data linkage with the National Death Index (NDI), administered by the Australian Institute of Health and Welfare (AIHW). De-identified surgical pathology reports are available for all surgical cases. A minimum data set of core data items for all specimens can be seen in Table 1; where not already specified, these data items are available to the HSA Biobank through the eMR.
Data items available* for request by researchers accessing the HSA Biobank.

MBS and PBS data items are also available subject to additional approvals.
The HSA Biobank: Description of the current cohort
Since 2012, the HSA Biobank collection has grown substantially. At the time of writing, the HSA Biobank collection held over 17 000 specimens from more than 3800 consented participants. Consent rates are consistently high. More than two-thirds of participants are female, which reflects the large contribution of gynaecological and breast specimens to the biobank; participant numbers are greater in the older age groups for both male and female (Table 2).
HSA Biobank participant demographics: numbers of consented participants and as % of total consents for each gender (*does not include 20 consented patients for whom a gender or date of birth was not recorded).

Although the HSA Biobank is a universal biobank in that it incorporates biospecimens from any cancer diagnosis, in practice some larger, more comprehensive cohorts are held within the collection in accordance with the relative clinical strengths of the participating hospitals: these include gynaecology, upper and lower gastroenterology, sarcoma and breast. A description of cancer types represented in the HSA Biobank collection is shown in Figure 5a. Importantly, research utilisation of specimens and data has seen significant growth with 35 projects directly supported (27 since 2016) which has resulted in 52 conference presentations and publications (Figure 5b).

(a) Tumour sites of participants consented for HSA Biobank (#pancreatic biliary and liver; *melanoma, skin, genitourinary, endocrine, cervical, CNS, lung, head & neck). (b) The number of projects directly supported by the HSA Biobank over time, and research outcomes generated (including manuscripts under review). (c) Projects supported and research outcomes classified by tumour type; #projects used datasets across several tumour types; *outputs relating to the development of the HSA Biobank. Some projects use specimens/data from more than 1 tumour type. Further details of all research outputs are available in the supplemental material.
Challenges, current and future
Since its inception, the HSA Biobank has become an established resource for cancer research, but challenges remain, including those confronting biobanks more broadly; some of these are discussed here.
Challenge: Ongoing engagement with clinicians and hospital pathology
The creation of strong partnerships spanning the health and research precincts, and involving key stakeholders within the hospitals (surgery, pathology) and infrastructure/research (university), has been essential in establishing a successful biobank. A major strength of the HSA Biobank is the collaborative partnership with hospital anatomical pathology; this is essential for routine assignment of tissue for banking. However, for this to persist, ongoing engagement with the individual pathology departments is key to ensuring ‘buy-in’ from pathologists; wherever possible, individual pathologists (and other clinicians) should be included in research studies facilitated by the biobank and co-authorship offered; indeed, a pathologist is included as a co-author on 90% of HSA biobank publications to date.
The initial approach of embedding both patient consent and allocation of bio-specimens within the routine practice of surgical and pathology departments has endured. However, recurrent engagement from biobank personnel (tracking of eligible cases and regular interaction with incumbent and new personnel) is essential to maintain consent rates; in addition, establishment of biobank technical officer positions within the hospital anatomical departments has proved crucial for the timely allocation of biospecimens.
Challenge: Utilisation of collected bio-specimens
A common complaint of ‘classic’ biobanks is underutilisation of biospecimens, 16 and ways of improving this situation have been discussed.4,9,17,18 Currently, HSA Biobank specimens and data are available for access by any researcher with ethical approval for their project. However, in practice it is the responsibility of the HSA Biobank to promote the collection to researchers and maximise use of current resources: researchers need to be made aware of what the HSA Biobank can offer. Marketing of available biospecimens is important4,17 and requires dedicated resources which can be regarded as peripheral to biobanking and not well supported. However, as well as a commercial need to dispatch biospecimens, there is also an ethical imperative to ensure that as far as possible, all biospecimens donated by participants be used in meaningful research 19 – this is important in instilling participant trust in the biobank and is important for social sustainability. 20 To address this the HSA Biobank is targeting local researchers to inform them of specimens and data available; in particular, if an annotated cohort has been assembled for a specific request, this may be of use as a ‘ready to go’ resource for others. ‘Word of mouth’ advertising is useful in raising awareness of what the HSA Biobank can offer, as well a requirement for acknowledgement on all research outputs generated from banked specimens and/or data. In addition, via the UNSW Biorepository, the biobank is included in the NSW Biobank locator (https://biobank.health.nsw.gov.au/about-the-biobank/nsw-biobank-locator) and has a presence within the Australasian Biospecimen Network Association (ABNA).
Challenge: Biospecimen annotation
The clinical annotation of biospecimens (what data is required and how to access it) is an issue across biobanking, made complicated by the non-conformity of electronic medical record and information systems across hospitals and the sometimes ad-hoc nature of how patient information is recorded. The HSA Biobank has established automated extraction of minimum dataset items from hospital information systems, using a business intelligence application called Crystal Reports, which employs the SQL command language; this data is matched to biospecimen data housed within OpenSpecimen. Although the number of correct matches between the clinical record and OpenSpecimen data using automated extraction is high (>60%), manual adjustments and data quality checking are still required. Improvements to the automated extraction and data matching processes are ongoing. By ensuring that core data items can be sourced reliably, the HSA Biobank is developing the capability to provide ‘complex biospecimens’. 21
Challenge: Universality
Although universality of the HSA Biobank is a strength in that it can accommodate requests from a broad cancer research base; our experience is that depth of acquisition is proportionate to engagement of surgeons who have a vested interest in research – these are most likely to lead to collections which are stronger. Conversely, cancer research groups who have a well-established connection between clinicians and preclinical scientists are more likely to engage with active collection practices which are then subsequently available to other researchers. This is demonstrated in Figure 5: although many tumour types are housed within the biobank, some of the largest collections represent local areas of strength (5a), for example, gynaecological cancers, upper GI cancers and these have been converted into proportionately more research output in these tumour streams (5c). True universality in cancer specimen collection is difficult for biobanks operating within a local area to achieve; identifying local areas of strength to enable efficient, targeted use of biobank resources will aid sustainability; this is the approach being followed by the HSA Biobank.
Challenge: Sustainability
All the challenges discussed so far feed into the overarching challenge of biobank sustainability. Biobanks globally are struggling with this issue (with many having folded 22 ) and it has been discussed widely in the literature.20,21,23,24 It is likely that initiation of a new biobank today would result in something quite different to one established a decade or more earlier. Indeed, initially it was envisaged that HSA Biobank patient consent, and specimen and data acquisition would become embedded as part of routine workflow within the hospital systems and ongoing maintenance would be minimal; in our experience without regular stakeholder engagement by biobank personnel the initiative would not be maintained and consequently, sustainable resourcing is essential.
Watson et al 20 proposed that when considering biobank sustainability, funding is not the only concern. Alongside financial constraints, operational and social aspects should be considered, for example: improvements in efficiency of specimen processing and data acquisition, re-allocation of resources for prospective collections as required, broadening of biospecimen offerings (categorised as operational concerns); consumer engagement and biobank certification (social concerns).
The HSA Biobank is addressing operational issues around sustainability, some of which are described in the Future Directions section below. However, despite employing a cost-recovery model, true cost recovery incorporating all biobank overheads (e.g. salary costs, cold storage and data storage infrastructure costs) would price most researchers out of the biospecimen market. Hence, most biobanks can only continue operating long-term with the support of a large organisation behind them 25 : in the case of the HSA Biobank, this is UNSW Sydney, with the biobank transitioning to a UNSW core facility based on the current operating model.
With regard to social aspects of sustainability, which relate to the acceptability and trust of biobanks, we believe that the HSA Biobank is well-placed. The Biobank is working towards becoming certified through the NSW Health Pathology Biobank Certification Program, (adapted from the Canadian CTRNet Biobank Certification Program; application pending approval); this is important for inspiring researcher and consumer confidence in our operations. In addition, a robust governance structure and consequent adherence to high standards helps further build this trust, which is essential for participant consent and therefore affects the number and variety of specimens available for research. Ideally, consent to take part in biobanking should be ‘patient-driven’, and anecdotally the HSA Biobank finds participants very supportive of their specimens being used in research. The HSA Biobank has a long-term, close collaboration with a group of consumers trained to advise cancer researchers, the TCRN Consumer Advisory Panel (CAP); 1 panel member holds a position on the HSA Management Committee. The consumer perspective has been invaluable in the development of patient-facing aspects of the biobank, and championing of biobanking more broadly can be facilitated by groups such as these.
Transforming the HSA Biobank operational model: Future directions
The HSA Biobank continues to directly support multiple projects in its role as a ‘classic biobank’ 18 demonstrating an ongoing demand for traditional biospecimen wares. Examples include usage of ovarian, endometrial or sarcoma tissue and derivatives in stand-alone projects26–28 as well as contributions to large-scale collaborative cohort studies. 29 The compilation of tumour micro-arrays (TMAs) from cohorts within the HSA Biobank collection is also an efficient and cost-effective way in which researchers can access bio-specimens.
However, the HSA Biobank is aware of the need to be agile 6 : a combination of maintaining relevance in response to changing researcher demands and a re-focus on sustainability has led to exploration of new directions which may augment the classic biobanking model. Areas which build on strengths within HSA Biobank participating institutions are currently being explored, with pilot projects underway in primary cell/organoid culture, microbiome research, bioimage banking and research using participant data alone. Prospective, investigator-driven collection is a pre-requisite for some of these initiatives and importantly, establishing good working relationships with researchers builds ongoing trust in the biobank. 30 The current participant consent model can continue to be used that is, participants consent to use of de-identified biospecimens and data for future research. This hybrid model is summarised in Figure 6.

The HSA Biobank going forward: demonstrating agility by extending the current operational model, adding areas of research demand. Research support: the HSA Biobank will continue to support research by utilising the current collection and building upon this in areas of local research strength. Enhanced specimen collection: introducing prospective collection catering to research needs. Research data: making biospecimen-associated data available for data-only research projects. Bioimage banking: storing DICOM images (radiomics) and histopathology images (pathomics) along with biospecimens and data from the same patient cohorts, enabling imaging biomarker (IB) research. Live cell culture: moving beyond prospective collection to facilitate bespoke collection of biospecimens requiring explicit acquisition and culture conditions. Microbiome research: collection of microbiota specimens from patient cohorts alongside additional matched biospecimens and data, to enable research into the involvement of microbiomes in disease processes and treatment outcomes.
Pilot 1: Live cell culture, organoids, 3D tissue models, tissue explants
The HSA Biobank is piloting a process wherein researchers access tissue directly from surgery for the culture of models involving live tissue/cells. In one example, pancreatic tissue is obtained directly from surgery and processed immediately to create an ex vivo culture model of the pancreatic tumour microenvironment; this culture model has been successfully used elsewhere using prostate tumour. 31 In this case, researchers are studying the stroma within the pancreatic tumour microenvironment which cannot be done effectively without such models. Importantly, this process goes beyond prospective collection – it requires the fresh tissue to be collected under very specific conditions; in this instance time is the critical factor, with tissue required to be in culture within 45 minutes of surgical resection; at time of writing, 35 specimens had been successfully propagated.
This same process of accessing newly-resected tissue could be applied in a range of emerging research areas: organoid culture is one example, wherein stem cells from a variety of tissues can be harvested and cultured to produce self-organised, 3-D tissue architecture which can mimic either complex organ structure, or a restricted selection of cells from within an organ. 32
Another tissue model system, patient-derived xenografts (PDX), relies on use of freshly-resected tissue. These models involve grafting of human tumour tissue into immunosusceptible mice to effectively reproduce the tumour as it exists in vivo; in an example of human renal PDX models, tissue cannot be used beyond 2 hours from resection. 33
In each case, the specificity of the research techniques will determine how soon and under what conditions tissue must be processed after resection, but there is an unequivocal need for the biobank to facilitate rapid release of this tissue to the researcher. This cannot occur without the involvement of collaborating surgeons and anatomical pathologists who need to understand the nature of the research request and be willing to accommodate it when feasible, with diagnostic needs always taking precedence. Relationship building with surgeons, pathologists and researchers is vital for these requests to be satisfied: brokerage provided by biobank personnel enables these researcher needs to be met by providing the ‘umbrella infrastructure’ to assist with for example, ethics and governance requirements, platforms ensuring security of data, and assistance with specimen tracking, transport and processing.
Pilot 2: Microbiome research
Increasingly, the microbiota (the microbes that collectively inhabit the human body), is recognised for its importance in many aspects of human health 34 ; the microbiome is the collection of all the genomes of these microbes. 34 Within the human body, different organs have distinct microbial ecosystems, most notably the gut, skin, vagina and mouth. 35 In particular, the gut microbiota has been implicated in development, treatment and prevention of different cancers.36,37
The current recommended medical treatment for patients with advanced or locally advanced cancer remains cytotoxic drugs, but these drugs frequently have treatment-related morbidity, 38 which additionally impacts on treatment tolerance and response. Studies have shown the gut microbiota to have a major influence on the efficacy and toxicity of chemotherapy agents39–41 and as such, this provides an opportunity for optimising chemotherapy outcomes while minimising toxicity in personalised cancer therapies; this would be particularly beneficial for patients currently deemed unsuitable for chemotherapy.
Links have been proposed between the microbiome and the development of colorectal cancer 42 but to date, there is little evidence for research addressing the impact of an altered microbiota on treatment outcomes. Patients with one of a number of cancers have been shown to exhibit divergent gut microbiota, including breast43,44 and pancreatic cancer. 45
The HSA Biobank has begun collection of stool and oral swab specimens, allowing research into gut and oral microbiota in different cancers and at different stages of treatment; this is done by engaging with individual clinicians who explain and discuss this with their eligible patients. Where possible the same patient is asked to provide specimens at different time points over the course of treatment allowing in depth investigation of the effects of, and upon the microbiota. In keeping with the clinical and research strengths of the HSA Biobank affiliated institutions, focus is given to pancreatic, colorectal and gynaecological cancers. Potential exists within the biobank for the extension of microbiota/microbiome specimen collection for disease cohorts other than cancer.
Pilot 3: Bioimage banking
The advantages of banking bioimages for both research and validation purposes are gaining increasing recognition. The evolution of high-throughput computing has made the prospect of extracting multiple features from digital medical images a reality. The conversion of these digital bioimages into data has become known as ‘radiomics’ (for radiological images) and ‘pathomics’ (for high-resolution histopathology tissue images).46,47 A major driver of expansion in these new fields is the development of Imaging Biomarkers (IBs).
Biomarkers can be defined as characteristics measured as an indicator of normal biological processes, pathogenic processes or responses to an exposure or intervention. 48 This can be applied equally to Imaging Biomarkers (IBs) along with more readily recognisable biospecimen-derived biomarkers. 49
Some IBs are already in clinical use for example, TNM (Tumour, Node, Metastasis) staging in cancer, 46 and BI-RADS (Breast Imaging Reporting and Database System) breast morphology readings in mammography 49 ; many others are used in research settings. The long-term aim in most radiomic and pathomic research is the development and validation of non-invasive, cost-effective measures to assess patient status; this is particularly appealing as almost all cancer patients have radiology and histopathology images generated during the diagnostic process. Within medicine, particularly in oncology, researchers seeking to cross the translational gap from IB discovery to their clinical use cannot do so without sufficient validation 49 – bioimage banks can facilitate this.
An essential requirement for any bioimage bank is annotation of banked bioimages with patient data. However, the power of biomage banking can be additionally enhanced by co-localisation of other ‘omics’ banking from the same patient cohorts, in the same biobank, to promote co-utilisation; this would allow for parallel validation studies to be done, moving from a correlative finding to a causative link between an IB and for example, patient prognosis.
With all this in mind, the HSA Biobank is actively looking to expand operations to include banking of bioimages. A proof of principle study for extraction of hospital radiology report data has been completed, and expansion to include accompanying DICOM (Digital Imaging and Communications in Medicine) images is under development. Unsurprisingly, storage of multiple large image files and ready access to these files is one factor that needs a satisfactory, workable solution.
It is well recognised that differential performance of imaging devices can vary output and while this may not affect use of radiology images for clinical practice, these inconsistencies may affect image use for radiomics. 46 Local collection of images and associated biospecimens and data has the advantage that IBs are generated on the same systems, using the same processes, making interpretation more straightforward. This would allow for repeatability within IB studies. 49
Pilot 4: Research data bank
As discussed, in order for collected biospecimens to have value, annotation with demographic and clinical data is essential; linkage to additional data sources is also in demand. The HSA Biobank accesses several data repositories to annotate biospecimens which posed the question: with such rich data available, can research projects using data alone be facilitated by the biobank? Although data sources for biobanks have small participant numbers compared to cohorts used in mainstream ‘big data’ research, biobank data is rich: data can be examined at a granular level, not possible with large data sets. Importantly, this allows research at a local level, examining health service utilisation and patterns of care within the geographical area in which the biobank operates.
In addition, data held by biobanks can be used in health IT research, areas such as automatic de-identification of patient documents and natural language processing, which have a role in improving future efficiency and cost-effectiveness across the health sector.
The HSA Biobank is unusual in the Australian context in that as well as demographic and clinical data, consent is sought for linkage to patients’ medical appointment and pharmaceutical use data, via the Medical Benefits Schedule (MBS) and Pharmaceutical Benefits Scheme (PBS) respectively; this allows the potential for local level health services research. A pilot project is underway comparing procedures, medications and overall costs of surgical and non-surgical upper gastrointestinal (GI) cancer cases, as well as comparing surgical upper versus lower GI cases 50 ; this can inform about local situations, with further comparisons possible with national and international practices.
In conclusion
In summary, the HSA Biobank is an established, cancer biobank based in South-East Sydney. Although originally developed and operated successfully as a classic (biospecimen-centric) biobank, the HSA Biobank has come full circle is now embracing a hybrid model to include prospective collection for specialised requests (investigator-centric). In addition, expansion into new areas is being explored, centring on areas of local strength. The establishment of a functional operating model such as the HSA Biobank provides a vehicle that is transferable beyond cancer to any surgical specialty and in return increases the viability of a biobank for the long term.
By adapting to a dynamic cancer research environment, the HSA Biobank can continue to operate as an integral translational resource.
Supplemental Material
sj-pdf-1-bmi-10.1177_11772719211005745 – Supplemental material for Moving with the Times: The Health Science Alliance (HSA) Biobank, Pathway to Sustainability
Supplemental material, sj-pdf-1-bmi-10.1177_11772719211005745 for Moving with the Times: The Health Science Alliance (HSA) Biobank, Pathway to Sustainability by Carmel M Quinn, Mamta Porwal, Nicola S Meagher, Anusha Hettiaratchi, Carl Power, Jitendra Jonnaggadala, Sue McCullough, Stephanie Macmillan, Katrina Tang, Winston Liauw, David Goldstein, Nikolajs Zeps and Philip J Crowe in Biomarker Insights
Footnotes
Acknowledgements
We would like to acknowledge the Cancer Institute NSW for funding the Translational Cancer Research Network (TCRN) which has operated the biobank, as well as the contribution of UNSW Sydney for funding aspects of the biobank including infrastructure and salary; additional funding was provided by the Health Science Alliance. We would like to thank the clinicians across the hospital sites within the SESLHD for consenting patients, the NSWHP anatomical pathology staff for facilitating access to specimens, the data custodians who have facilitated access to data for specimen annotation, the researchers with whom we have worked (in particular in the pilot project areas), and the patients themselves for generously agreeing to participate in research through granting access to their specimens and data.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The HSA Biobank received financial support from the Translational Cancer Research Network (a Cancer Institute NSW Translational Cancer Research Centre), UNSW Sydney, and the Health Science Alliance.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
