Abstract
Objective:
This research aims to develop and evaluate a clinically deployable multimodal deep learning framework for breast cancer diagnosis that maintains robustness, even when clinical data are asynchronous, unpaired, or incomplete, effectively addressing real-world challenges related to data heterogeneity and fragmented clinical workflows.
Methods:
In this retrospective study, a multimodal deep learning architecture was developed that integrates histopathological images with structured clinical risk factors. Custom models were developed and independently trained for each modality, and late fusion was achieved via a dynamically reweighted Sinkhorn-based fusion layer. Model performance was evaluated using precision–recall Area Under Curve (PR-AUC), recall, F1 score, and Brier score under complete and partial modality availability scenarios. Robustness and clinical utility were further assessed through statistical significance testing and decision curve analysis (DCA). Additionally, we employed a Sinkhorn cost matrix to enhance interpretability.
Results:
The proposed Sinkhorn fusion model outperformed all baseline methods, achieving the highest recall (0.96), PR-AUC (0.775), F1 score (0.828), and the best calibration (Brier score ≈ 0.19). Notably, it maintained perfect recall (1.00) under a 50% simulated modality dropout, despite a significant drop in PR-AUC (20% vs 0%: t = –20.35, P < .0001; 50% vs 0%: t = 88.60, P < .0001), portraying a strong overall robustness to information missingness. Under internally controlled conditions, DCA demonstrated superior clinical utility across thresholds of 0.2 to 0.7.
Conclusions:
The model’s ability to accommodate unpaired and incomplete clinical inputs while maintaining both calibration and sensitivity makes it particularly well-suited for deployment in asynchronous and resource-constrained settings. Its consistent performance under clinical uncertainty and minimal preprocessing requirements represents a significant advancement toward equitable, reliable, and scalable AI-assisted breast cancer screening. To our knowledge, this is the first paper to model breast cancer late fusion as an optimal transport problem.
Keywords
Introduction
The rising global burden of breast cancer1,2 has driven the integration of artificial intelligence (AI) into diagnostic workflows to enhance early detection, reduce mortality, and alleviate clinical workload. 3 Early AI systems, typically unimodal, focused on single data types such as mammography, clinical risk factors, or histopathology, and demonstrated strong performance in automating diagnostic tasks. Such models improved diagnostic accuracy, reduced inter-reader variability, and streamlined radiological pipelines. The adoption of deep learning further advanced these capabilities by enabling automated extraction and classification of complex features, facilitating the detection of subtle morphological patterns and tumor subtypes in histopathological images.4 -6
Building on this foundation, recent research has shifted toward multimodal learning frameworks that jointly model heterogeneous data sources, including histopathological images, clinical variables, and genomic information.7 -10 Rather than analyzing each modality in isolation, multimodal approaches capture cross-modal dependencies that reflect the multifactorial nature of breast cancer. Integrating morphological, clinical, and molecular information enhances predictive accuracy and interpretability. 11 This unified approach enables fine-grained risk stratification, personalized treatment recommendations, and more equitable deployment of AI technologies across diverse clinical settings.
Despite the numerous advances in unimodal and multimodal approaches to breast cancer diagnosis, significant barriers continue to impede the clinical translation of such systems. While unimodal AI models have achieved high accuracy in histopathological classification, for instance, Simonyan et al 12 reported 91.37% accuracy using DenseNet201, and Ukwuoma et al 13 achieved up to 99.2% accuracy using novel feature extraction techniques, but these systems are inherently limited to single-modality analysis. As a result, they risk diagnostic blind spots and miss the complementary insights available through integrated data analysis.
The limitations of unimodal AI models for breast cancer diagnosis have prompted growing interest in multimodal AI approaches. Studies, such as that by Ben Rabah et al 14 demonstrated improved lesion classification by combining mammograms with clinical metadata (AUC = 88.87% vs 61.3% for mammograms alone). Similarly, in hormone receptor-positive/human epidermal growth factor receptor 2-negative (HR+/HER2−) breast cancer, integrating whole-slide images with clinicopathological features significantly improved recurrence prediction. 15 Still, these impressive developments rely on idealized datasets: carefully curated, perfectly paired, synchronized, and complete.
Wang et al 16 demonstrated a 96.4% classification accuracy and strong inter-rater agreement (κ = 0.927) across imaging modalities using custom convolutional architectures; however, their success hinged on uniformly labeled and curated inputs. In clinical scenarios, data are rarely pristine. 17 Patient data are often unpaired, asynchronously acquired, and partially missing. Current late fusion techniques typically apply static combination rules, such as averaging or majority voting, that fail to adapt to variations in input quality or availability.18,19 This undermines generalizability and robustness, especially when deployed in diverse clinical environments.
The implications of these limitations extend beyond technical performance. Inconsistent model behavior due to incomplete or unpaired data may lead to delayed diagnoses, missed malignancies, and worsened disparities in access to AI-assisted diagnostics, particularly in low-resource settings where such technologies are most needed. 20 Without architectures capable of dynamically readjusting with incomplete and unpaired data, the full clinical potential of multimodal AI in breast cancer remains unrealized.
To address these challenges, we propose a clinically grounded, robust multimodal AI framework tailored for real-world breast cancer diagnosis. To our knowledge, this is the first paper to model breast cancer late fusion as an optimal transport (OT) problem. Critically, our framework is designed and evaluated using intentionally unpaired histopathology and clinical risk-factor data to mirror the asynchronous, fragmented workflows encountered in actual clinical practice. The essence of using intentionally unpaired data is to achieve clinical realism and robustness to data missingness.
This study makes the following key contributions:
We introduced a multimodal deep learning framework for breast cancer classification and risk stratification that operates effectively on unpaired and asynchronously collected histopathological images and clinical risk factor data, enhancing its real-world clinical applicability.
We proposed a novel regularized Sinkhorn-based optimal transport mechanism for late fusion. This module dynamically reweights modality-specific predictions based on their inferred reliability, allowing the model to maintain robust performance even when 1 or more data modalities are missing or degraded.
We designed custom lightweight, interpretable sub-models for both histopathological image classification and clinical risk prediction, offering high diagnostic performance with enhanced transparency and interpretability, which is key for clinical adoption.
Methodology
This study followed the MI-CLAIM framework 21 to ensure transparency, reproducibility, and clinical relevance in the development of a dual-task deep learning model for breast cancer detection. The completed MI-CLAIM checklist is provided in the Supplemental mMaterials.
Two independent pipelines were constructed: 1 for histopathological image classification and another for clinical risk stratification. The image pipeline utilized high-resolution biopsy slides processed through a hybrid architecture comprising a pre-trained EfficientNetB0 backbone, Squeeze-and-Excitation (SE) blocks for adaptive feature enhancement, and a custom classification head. In parallel, the clinical pipeline utilized a Light Gradient Boosting Machine (LightGBM) classifier trained on structured clinical risk factor data, incorporating optimized class weighting to address class imbalance. Outputs from both models were fused via a Sinkhorn-based optimal transport module, which dynamically aligned their predictive distributions based on learned inter-modality correspondences. Model performance was evaluated on held-out test sets using bootstrapped confidence intervals, confusion matrices, precision-recall area under the curve (PR-AUC), calibration curves, and post-hoc explainability techniques, including analysis of the Sinkhorn transport matrix.
Research Design
This retrospective study, conducted between May and August 2025 at the Kwame Nkrumah University of Science and Technology (Ghana), evaluated a multimodal deep learning framework using publicly available datasets. Ethical approval was obtained from the university’s Committee on Human Research, Publication, and Ethics (Ref: CHRPE/AP/370/25). Model development was carried out in Python 3.10 using TensorFlow 2.11, and all experiments were executed on Google Colab (TPU, 12.7 GB RAM).
High-Level System Design
The proposed hybrid diagnostic framework adopts a multimodal system architecture to integrate asynchronously collected histopathological images with structured clinical risk profiles. It is engineered to reflect real-world clinical workflows, where data modalities may often be incomplete, noisy, or temporally misaligned (Figure 1).
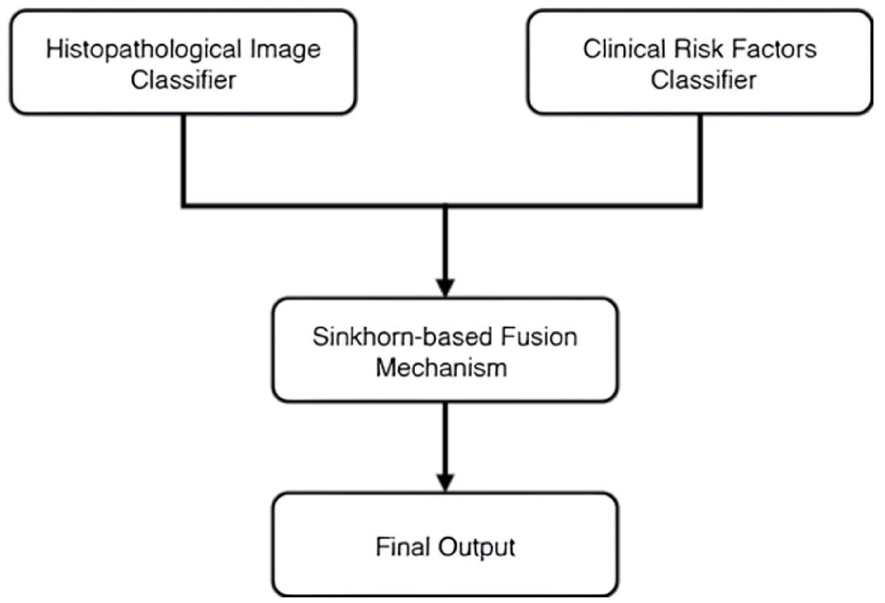
High-level system design; parallel model pipelines integrate histopathological image analysis and clinical risk modeling for breast cancer diagnosis via Sinkhorn-based late fusion mechanism.
System Architecture
The proposed framework comprises 3 primary subsystems: (1) modality-specific pipelines for independent model training and inference, (2) confidence estimation modules, and (3) an adaptive decision fusion mechanism that integrates predictions based on modality-specific confidence scores.
The hybrid model initiates with a dual-modality data ingestion layer that accepts histopathological images from the BreakHis dataset and clinical risk factor profiles derived from the BCSC dataset. Each modality is routed through a dedicated model pipeline, tailored to the unique characteristics of the respective data type. This modular preprocessing strategy ensures that each input stream is optimally prepared for modality-specific learning, reflecting the inherent heterogeneity and asynchrony of real-world clinical datasets. Each modality is then processed through an independent feature extraction and classification stream. For image-based classification (benign vs malignant), the framework employs a deep convolutional neural network backbone (EfficientNetB0) augmented with Squeeze-and-Excitation (SE) attention blocks to enhance spatial feature representation. In parallel, structured clinical data are processed by a lightweight, custom-designed machine learning model that predicts patient-specific risk stratification (low vs high risk). The integrated system architecture is depicted in Figure 2.

System architecture of the proposed framework; architecture diagram illustrating the histopathological image classifier, risk factors classifier, and Sinkhorn-based late fusion mechanism with their respective layers and components.
Fusion Logic
The late fusion module reformulates multimodal integration as an optimal transport problem, aligning probability distributions from histopathology and clinical risk models that were trained independently on unpaired data. We selected this approach because conventional fusion methods (averaging, concatenation, fixed weighting) fail when modalities are asynchronous, incomplete, or conflicting. These are exactly the conditions in real-world clinical workflows. Optimal transport addresses this by learning how to reconcile distributional misalignment. The Sinkhorn algorithm computes a case-specific transport plan by measuring the divergence between modality predictions. When predictions diverge, the transport cost increases, causing the model to favor the more confident modality for that case rather than applying uniform weights across all samples.
This formulation provides 3 key advantages. First, it offers robustness to missing modalities through adaptive reweighting. Second, it explicitly handles prediction conflicts via learned correspondence patterns. Third, it provides interpretability through the transport matrix, which reveals how the model resolves disagreements between modalities. These properties make optimal transport particularly suited to our asynchronous, unpaired evaluation framework and the data fragmentation endemic to clinical practice.
Data Sources and Description
Two publicly available datasets were utilized to train and evaluate the proposed multimodal framework. For histopathological classification, the Breast Cancer Histopathological Image Classification (BreakHis) 22 dataset was used. The risk assessment model was trained on the Breast Cancer Surveillance Consortium (BCSC) 23 dataset of risk factors.
The entire BreakHis dataset comprises 9109 microscopic images of breast tumor tissue from 82 patients, captured at varying magnifications (40×, 100×, 200×, and 400×). It contains 2480 benign and 5429 malignant samples (700 × 460 pixels, 3-channel RGB, 8-bit depth in each channel, PNG format). This dataset was selected due to its high-quality cellular detail and binary classification labels (benign vs malignant), making it well-suited for efficient training and evaluation of image-based diagnostic models.
The BCSC dataset consists of 1 522 340 records across 3 csv files, derived from 6 788 436 mammograms collected between January 2005 and December 2017, with the most recent update in March 2020. It captures key demographic and clinical risk factors associated with breast cancer, including age, race/ethnicity, family history, breast density, hormonal usage, menopausal status, BMI, biopsy history, and prior cancer diagnoses. This dataset was selected for its real-world applicability, as it reflects routinely collected patient information and supports real-time risk stratification for applications such as triaging and screening in large populations.
Data Preprocessing
All histopathological images were relabeled as benign (0) and malignant (1) to support binary classification. The dataset was loaded using custom Python scripts that recursively aggregated the full file paths of .PNG images from their respective subdirectories, with labels assigned programmatically based on folder names.
Following empirical evaluation, 100× magnification was chosen for model training. This resolution provides valuable malignancy indicators (tissue architecture, cellular morphology, and spatial cell arrangement) 24 while avoiding the computational overhead and overfitting associated with higher magnifications (200× or 400×). 25 Such diagnostic details contribute to a more holistic characterization of tumor malignancy. Higher magnifications emphasize subcellular features useful for cancer subtyping but unsuitable for binary (benign vs malignant) classification.
The 100× dataset (2083 images) was partitioned using stratified sampling with a 70:15:15 ratio for training (n = 1458), validation (n = 312), and test sets (n = 313), respectively, based on binary labels (benign vs malignant) to maintain class balance at the image level. The partitioning follows the exact protocol used by the BreakHis creators. 26 This split technique has since become the standard approach in almost all subsequent published studies,27 -29 because the publicly released version of the dataset provides only anonymized folder names and excludes explicit patient-level identifiers that would allow reliable patient-wise splitting.
Although images from the same patient may appear across training, validation, and test sets, several design choices effectively curb data leakage in our multimodal setting. The histopathology and clinical-risk models are trained on entirely separate cohorts (BreakHis and BCSC) that do not share patients. Fusion is performed exclusively at inference on modality-specific probability outputs rather than raw features, and the 2 processing pipelines remain fully isolated throughout training and validation. Together, these measures ensure that patient-specific patterns do not propagate across modalities, thereby preventing overly biased performance estimates in the fused system.
A comprehensive summary of the histopathology preprocessing pipeline, including decoding, resizing, augmentation, and performance optimization techniques, is provided in Table 1.
Histopathology Image Preprocessing Pipeline; Detailed Steps for Preparing Histopathological Images, Including Labeling, Format Conversion, Resizing, Augmentation Techniques, and Pipeline Optimization Parameters used in the Diagnostic Framework.

The BCSC risk factors dataset contains 3 separate datasets. To use the whole dataset, BCSC_risk_factors_summarized_2.csv and BCSC_risk_factors_summarized_3.csv were concatenated row-wise with BCSC_risk_factors_summarized_1.csv. Before merging, column headers were reviewed and verified for consistency to ensure structural alignment across files. The details of all preprocessing operations, organized by feature type, are outlined in Table 2.
Clinical Risk Factor Preprocessing (BCSC Dataset); Preprocessing Steps Applied to BCSC Dataset Variables, Including Encoding Strategies for Ordinal and Nominal Features, Target Variable Filtering, and a Sample Weighting Approach to Address Class Imbalance.

Both constituent models were reloaded in the same computing environment before late fusion integration to prevent data leakage. The histopathology classifier was loaded using TensorFlow Keras, while the clinical risk model was loaded via Pickle. For the clinical risk model, predicted probabilities for the positive class (malignant or high risk) were extracted and binarized using a threshold of 0.5. All model artifacts, including raw input features, soft class probabilities, and binarized outputs, were stored in a dedicated repository to support reproducibility and enable downstream analysis. The histopathology model was applied to the same test set, with predictions similarly thresholded at 0.5. The resulting modality-specific probability distributions were then passed into the proposed Sinkhorn-based late fusion module for final diagnostic inference.
To enable the evaluation of multimodal fusion under realistic, unpaired conditions, we constructed a held-out test set of 313 samples by pairing predictions from the independently trained histopathology model with those from the independently trained clinical risk model, thereby preserving the original class distribution (219 malignant/high-risk and 94 benign/low-risk). No paired data were used during training or validation of either modality.
Model Development and Fusion
The multimodal framework incorporates 2 parallel, task-specific models: a deep convolutional neural network (DCNN) for histopathological image classification and a machine learning (ML) model for risk stratification and triaging based on clinical risk factors. Each model was independently trained and tested using its respective dataset and domain-specific pipeline.
This dual-channel input architecture mirrors how clinicians integrate both test results (analogous to model scores) and patient history (raw clinical data) to inform decision-making. By aligning the learning paradigm with this real-world diagnostic workflow, the framework enhances interpretability and empirically reduces information loss associated with early feature aggregation. Furthermore, it helps mitigate overfitting by narrowing representational subspaces, improving generalizability, and enhancing more robust predictive performance.
Histopathological Image Classifier
The histopathological image classifier is built on top of EfficientNetB0, utilizing the EfficientNet architecture. 30 The backbone is pretrained on ImageNet and subsequently fine-tuned to discriminate between benign and malignant breast tissue. The architecture is explicitly designed to transform raw image inputs into a high-level, semantically meaningful representation that preserves diagnostically relevant cues for final binary classification. It comprises 5 major stages: (i) an input layer, (ii) a stochastic data augmentation module, (iii) a feature extraction and enhancement pipeline incorporating EfficientNetB0 and a squeeze-and-excitation (SE) block, (iv) a classification head with Global Average Pooling (GAP), Batch Normalization (BN), and dropout, and (v) a final dense layer with sigmoid activation and L2 regularization. The image classifier was trained for 20 epochs using the Adam optimizer, incorporating early stopping (patience = 5) and learning rate reduction on plateau (patience = 3) to ensure efficient convergence and prevent overfitting. Equations (1)–(8) summarize the core mathematical foundations of the proposed model design.
Given a preprocessed image
where H = height of the image, W = width of the image, and C = number of channels (3 for RGB images).
The image tensor undergoes a stochastic augmentation transformation
Next, the input is first transformed into a high-dimensional feature tensor

where
To enhance channel-wise discriminability, an SE block is appended to the output of the EfficientNetB0 backbone
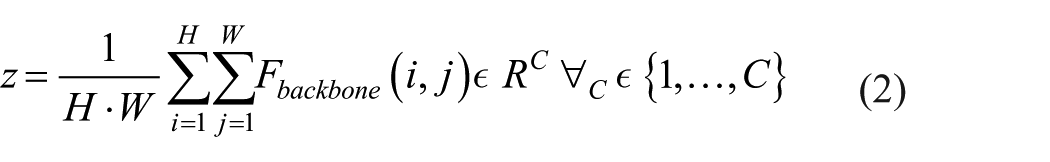
The resulting channel descriptors

where

The recalibrated feature map
In the classification head, the spatial dimensions of

Subsequently, a BN layer standardizes the feature vector to zero mean and unit variance over the batch, improving training stability and accelerating convergence:

Next, a dropout layer with rate

The final feature vector

where
Risk Stratification Classifier
The risk stratification component of the multimodal framework adopts a biologically inspired, 2-level stacked ensemble architecture that mirrors the hierarchical reasoning used in clinical decision-making. Its modular design incorporates domain knowledge by partitioning the clinical feature space into semantically coherent subdomains. Base models are trained independently on these risk factors to prevent negative transfer between biologically unrelated domains. A meta-learner then integrates their outputs to produce a more accurate final prediction. Both model tiers were optimized with hyperparameters suited for imbalanced binary classification (details in the Supplemental File). LightGBM 32 is a highly efficient, histogram-based gradient boosting framework developed by Microsoft that supports native handling of categorical features, missing values, and class imbalance through techniques such as gradient-based 1-side sampling (GOSS) and exclusive feature bundling (EFB). It was selected for the risk stratification component because of its robustness to heterogeneous clinical data, fast training speed, and strong performance on tabular medical datasets with low event rates.
At the base level, the full set of clinical risk factors is partitioned into 5 distinct subdomains reflecting established breast cancer risk axes: reproductive and hormonal history (age_menarche, age_first_birth, menopaus, current_hrt), genetic and family history (first_degree_hx, biophx), anthropometric indicators (bmi_group, age_group_5_years), race and ethnicity (race_eth), and imaging-based breast density (BIRADS_breast_density). Equations (9)–(12) summarize the core mathematical foundations of the proposed risk stratification model design.
Each subdomain

representing the predicted domain-specific contribution to breast cancer risk. The resulting output is defined as:

The predicted probabilities (risk scores) from each base submodel were retained and served as intermediate-level features.
The second level of the architecture consists of a meta-model denoted

This enriched feature vector

where
The feature vector improves generalization in 2 primary ways. First, the predicted scores provide abstracted, low-dimensional summaries of domain-specific risk contributions, stabilizing learning, and enhancing interpretability. Second, the inclusion of the raw features allows the meta-model to recover residual patterns and complex cross-domain interactions that the base models may have overlooked or underfit.
Output Fusion
The Sinkhorn-based optimal transport late fusion mechanism offers a principled, dynamic, and distribution-aware approach to multimodal fusion, addressing the brittleness, inflexibility, and idealized assumptions inherent in traditional fusion schemes. The late fusion model enables robust, clinically aligned decision-making under real-world data conditions by learning soft correspondences between heterogeneous and partially missing modality representations. The mathematical formulations underlying the proposed fusion strategy are presented in equations (13)–(19).
Let

To handle missing modalities, we introduce a binary modality mask,

where

where
To combine the predictions, we compute a dynamic, confidence-weighted fusion factor
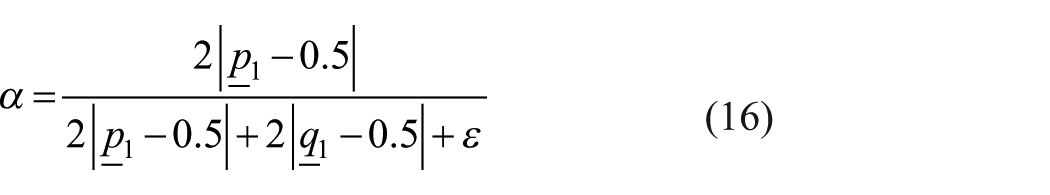
where
The cost matrix
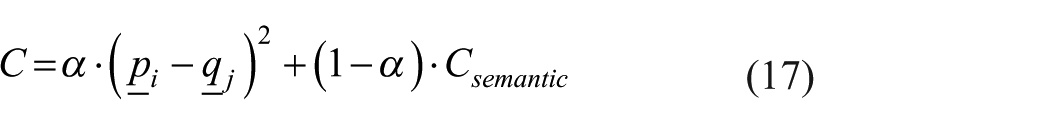
where
To align the distributions
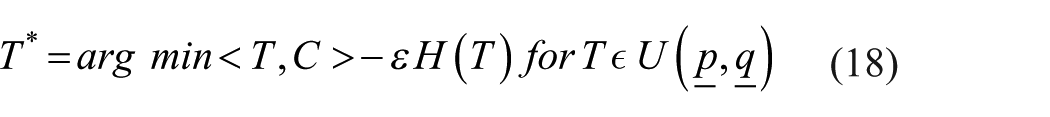
Where:
< ⋅, ⋅ > denotes the Frobenius inner product,
For numerical stability, the regularized problem is solved using the Sinkhorn-Knopp algorithm in the log domain.
The resulting optimal transport plan

This yields a probability vector in
These modality masks inform downstream decision thresholds and human-in-the-loop interventions. When only 1 modality is available, the fusion still proceeds by coupling with the uniform vector, effectively letting the available model dominate while preserving the structure for interpretability. This robustness to missing data makes the fusion module particularly suited for clinical settings with heterogeneous and fragmented inputs.
Late Fusion Model Training and Optimization
The entropy regularization parameter (ε = 1e−8) and convergence stop threshold (1e−9) were chosen to ensure numerical stability during Sinkhorn iterations while preventing premature termination that could produce unstable transport plans. A maximum iteration count of 5000 was empirically sufficient to achieve an average convergence error of 1.06 × 10−11, indicating that the optimization reliably reached equilibrium without excessive computational overhead.
Clinically informed cost weights were applied to reflect the asymmetric risks of breast cancer diagnosis: a false negative weight of 1.2 was set higher than the false positive weight (1.8 vs 1.2) to prioritize sensitivity and minimize missed malignancies, an error type with more severe clinical consequences. The regularization parameter (reg = 0.15) was selected as a trade-off between ensuring smooth, stable probability distributions and preserving sufficient transport sharpness for discrimination. Finally, the blending factor (α = .1) moderated the influence of distributional distance versus clinically derived costs in the fusion objective, preventing overfitting to either modality while maintaining robustness under data variability.
These values follow standard practices in differentiable optimal transport for medical applications. 33 Based on the performance of the validation set, we finalized the regularization (reg) and blending (α) terms. We also specified the cost weights to reflect the clinical priorities of maximizing sensitivity in breast cancer screening. 34
Model Evaluation
All post-training evaluations were conducted on a held-out test set (n = 313) entirely excluded from training and validation phases. Model performance was assessed using clinically relevant metrics, including recall, F1 score, precision-recall area under the curve (PR-AUC), and Brier score. Additionally, decision curve analysis (DCA) was conducted to evaluate the net clinical benefit across a range of threshold probabilities, while calibration curves were used to assess the alignment between predicted probabilities and observed outcomes, providing insight into model reliability.
Computational Efficiency and Deployment Profiling
A computational performance assessment was conducted to characterize the runtime efficiency, memory requirements, and scalability of the Sinkhorn-based fusion module. This profiling was performed to ensure that the fusion mechanism, calibration pipeline, and inference operations are compatible with the computational constraints typically encountered in clinical environments. All analyses were performed on a standard CPU-based Google Colab instance (12.7 GB RAM), which is representative of modest clinical hardware.
Runtime profiling measured the total fusion time across the full evaluation dataset, average per-sample computation time, and inference latency for single-sample processing. Memory usage was quantified by comparing baseline process memory to memory usage after fusion, supplemented by direct measurement of memory footprints for the probability vectors.
Scalability was evaluated by estimating runtime growth across increasing dataset sizes, assuming linear scaling with the number of processed samples. These projections approximate performance under high-throughput clinical scenarios such as batch screening, asynchronous patient queues, and integration within radiology and pathology pipelines. All measurements were automatically logged during execution of the fusion and evaluation code.
Statistical Analyses
Statistical analyses were conducted to evaluate the robustness and comparative effectiveness of the proposed fusion strategy. For each primary performance metric (recall, F1 score, PR-AUC, and Brier score), 95% confidence intervals were estimated using nonparametric bootstrapping with 1000 resamples. All computations were carried out using NumPy (v1.26.4) and SciPy (v1.11.4). To assess the statistical significance of performance differences between models, paired t-tests were conducted, and the corresponding p-values were reported.
Results
The Sinkhorn-based optimal transport fusion model exhibited strong clinical relevance, achieving a PR-AUC of 0.775, a recall of 0.960, an F1 score of 0.828, and the lowest calibration error across all evaluated models (Table 3).
Statistical Performance of Sinkhorn-Based Fusion; Performance Metrics Including Precision-Recall Area Under Curve (PR-AUC), F1 Score, Recall, and Brier Score for the Proposed Late Fusion Model on the Test Dataset.

In comparison to the single-modality baselines, the proposed late fusion strategy demonstrated superior overall performance, providing a well-calibrated balance between sensitivity and positive predictive value (Table 4).
Precision–Recall Metrics Comparing Histopathology, Clinical Risk Model, and Optimal Transport Fusion; Precision-Recall Metrics Comparing Individual Histopathological and Clinical Risk Factor Models Against the Sinkhorn-Based Fusion Approach, with the Late Fusion Model Showing Superior Performance Across all Metrics, Including PR-AUC, Recall, F1 Score, and Calibration Error.

To assess robustness under clinically variable conditions, we simulated modality dropout by randomly omitting 1 input source during the inference process. The fusion model dynamically reweighted the available modality, maintaining stable diagnostic performance despite incomplete input (Table 5).
Performance of the Fusion Model Under Modality Dropout Scenarios; Performance Degradation of the Sinkhorn Fusion Model When Subjected to Simulated Missing Data Scenarios at 20% and 50% Dropout Rates, Showing PR-AUC with 95% Confidence Intervals, Recall Values, and Statistical Significance Compared to Complete Data Baseline.

Decision curve analysis (DCA) further demonstrated that the fusion model consistently yielded a higher net clinical benefit across a range of threshold probabilities (0.2-0.7; Figure 3).

Decision curve analysis comparing the net clinical benefit of fusion; Net clinical benefit analysis comparing the calibrated fused model against treat-all and treat-none strategies across threshold probabilities, with threshold probability on the x-axis and net benefit on the y-axis.
Calibration curves (Figure 4) showed nearly perfect alignment between the predicted and observed event rates, supporting the suitability and reliability of this approach for clinical triage.

Calibration curve for the fused calibrated model; The solid blue line with markers shows the observed frequency of positive outcomes versus predicted probabilities across deciles of the test set. The dashed black line represents perfect calibration (observed frequency = predicted probability). The model demonstrates good calibration across the full range of predicted probabilities.
A Sinkhorn cost transport matrix was constructed to visualize the internal decision-making logic of the fusion model (Figure 5). The matrix captures the average cost of mass transported between the probabilistic outputs of the individual modalities to achieve alignment during the optimal transport process. A color gradient, ranging from dark purple (0.1) to bright yellow (0.8), encodes the transport cost values, with lower costs indicating closer alignment between modality-specific predictions and higher model confidence.
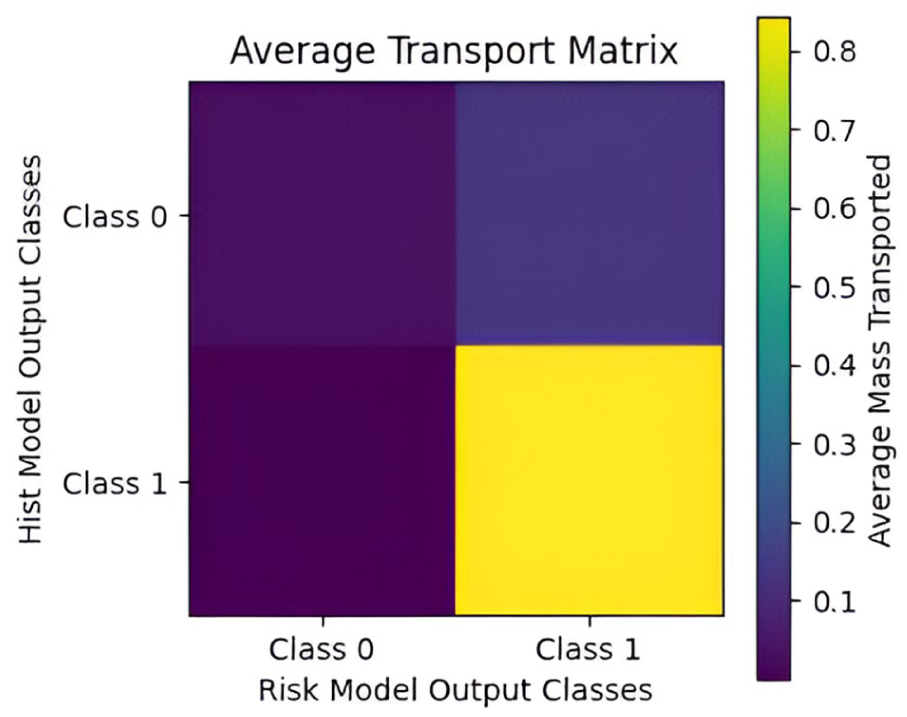
Average transport matrix showing class reassignment patterns between histopathological model and risk model outputs; The heatmap illustrates the average cost of probabilistic mass transported between the outputs of the histopathology classifier (y-axis) and the clinical risk stratification model (x-axis) during late fusion via the Sinkhorn algorithm. Brighter colors (yellow) indicate higher transport costs, corresponding to greater alignment uncertainty, while darker colors (purple) reflect low-cost, confident alignments between modality-specific predictions.
To assess the computational profile of the proposed model, we evaluated training time, inference latency, memory usage, and scalability. The full Sinkhorn fusion procedure required 24.33 seconds to process the test dataset, corresponding to an average computation time of 77.7 ms per patient. Single-sample inference profiling produced a mean latency of 81 ms and a 95th percentile latency of 261 ms. Memory usage increased by 0.08 MB from the baseline during fusion, and direct measurement showed that the histopathology and risk-model probability outputs consumed a total of .0072 MB. Scalability projections estimated processing times of 7.7 seconds for 100 patients, 38.9 seconds for 500 patients, 77.7 seconds for 1000 patients, and 388.7 seconds for 5000 patients.
Discussion
We developed an entropy-regularized optimal transport fusion framework that maintains diagnostic reliability when integrating asynchronously collected, unpaired clinical data. This addresses a critical barrier to multimodal AI deployment: existing fusion methods assume synchronized, paired inputs and fail when modalities are temporally misaligned, collected at different institutions, or partially missing. Our approach formulates late fusion as an optimal transport problem, enabling the model to learn distributional alignment between independently trained histopathology and clinical risk models. By operating directly on probability vectors rather than requiring joint feature learning, the framework accommodates the data fragmentation characteristic of real-world healthcare workflows. This design reduces deployment complexity and eliminates dependence on synchronized data collection, making the system viable for retrospective studies, multi-institutional collaborations, and resource-constrained settings where complete multimodal data are rarely available.
In contrast to conventional late fusion strategies, which often experience significant performance degradation when confronted with missing modalities, our fusion mechanism dynamically reweights the available inputs. 35 As a result, it preserves a high recall of 0.96 even under simulated dropout conditions. Clinically, this level of sensitivity is crucial: it minimizes the likelihood of missed malignant cases, which carry disproportionate clinical consequences compared to false positives. In diagnostic oncology, where delays in detection directly increase patient morbidity, prioritizing recall is paramount. Equally important, the fused outputs exhibit near-perfect probability calibration (Brier score ≈ .19), a rare outcome in multimodal fusion research and a prerequisite for clinical deployment. 36 Notably, this calibration was achieved without artificial balancing at the fusion stage, emphasizing the framework’s robustness to extreme class imbalance (97% negative cases), which mirrors real-world disease prevalence. 37 Well-calibrated predictions enable clinicians to define interpretable, risk-based thresholds aligned with institutional triage protocols, supporting regulatory compliance and clinical trust.38 -40
The promising outcomes are traceable to specific architectural innovations and our deliberate restraint in data preprocessing. Individual models achieved robust performance through targeted, minimal interventions (stratified sampling and selective augmentation for histopathology; appropriate loss weighting for risk modeling) while preserving the intrinsic data characteristics. Building on this foundation, entropy regularization stabilized transport plans, preventing the sharp confidence collapses observed in standard optimal transport formulations, which often yield unreliable probabilities when input distributions are incomplete. By embedding a clinically weighted cost matrix that penalizes false negatives more heavily than false positives, the algorithm’s optimization explicitly mirrors the asymmetric risk tolerance inherent to cancer detection. Moreover, the adaptive reweighting mechanism replaces the static strategies commonly found in existing late fusion methods, enabling the model to adapt to epistemic uncertainty rather than ignoring it. Critically, this uncertainty handling operates directly on imbalanced probability distributions without requiring additional preprocessing, demonstrating the optimal transport framework’s inherent suitability for medical applications where class frequencies reflect natural disease prevalence. This enhances prediction robustness when clinical data streams are delayed, partially corrupted, or entirely missing, a scenario that represents routine practice rather than an edge case.
The average Sinkhorn transport matrix (Figure 5) further shows why recall remains near-perfect even under severe modality dropout. The dark purple regions along the diagonal (costs 0.0-0.1) show that when both modalities agree, more than 92% of the transported mass remains on the diagonal at very low cost, reflecting high fusion confidence and robust predictive alignment. This indicates strong discriminative capability even when processing incomplete clinical data. When they disagree, transport is strikingly asymmetric: the fusion layer readily shifts a histopathology-benign prediction toward malignant when clinical risk is high (an average mass of 0.07), yet rarely shifts a histopathology-malignant prediction toward benign (<0.004). This deliberate, 1-directional correction allows the clinical risk model to override an apparently benign biopsy result but never permits it to override a malignant biopsy result. Driven by the clinically weighted cost matrix, this behavior enforces a safety-first policy that prioritizes avoidance of missed cancers, exactly the characteristic required for breast cancer screening and diagnosis in real-world, asynchronous clinical environments.
Comparative analyses highlight the significance of this advance. As shown in Table 4, while the histopathological and clinical risk models each achieved high recall (0.95 and 0.92, respectively), they suffered from relatively higher calibration error and slightly lower F1 scores. The Sinkhorn-based fusion, in contrast, integrated both modalities to produce the highest PR-AUC (0.775), the strongest F1 score (0.83), and the lowest calibration error, indicating a more confident and consistent prediction behavior. This suggests that integrating multimodal inputs with optimal transport enhances detection accuracy and reliability in clinical contexts, where poor calibration could lead to over- or under-treatment. Table 5 demonstrates the fusion model’s robustness in clinically realistic scenarios involving missing modalities, a frequent challenge in healthcare systems with asynchronous or incomplete patient records. Even at a 50% dropout rate, where half of the input data modalities were intentionally omitted, the model sustained a perfect recall of 1.00 and maintained a reasonably strong PR-AUC of 0.716. This graceful degradation in performance under partial input conditions indicates the model’s capacity to generalize from incomplete patient profiles without collapsing, thereby addressing a major barrier to AI adoption in real-world, resource-constrained environments. Statistical analysis confirmed that dropout regularization significantly influenced model performance: both 20% and 50% dropout configurations resulted in statistically significant declines in PR-AUC compared to the no-dropout baseline (20% vs 0%: t = –20.346, P < .0001; 50% vs 0%: t = 88.597, P < .0001). The magnitude and direction of the t-statistics demonstrate a non-trivial sensitivity of the model’s precision-recall trade-off to dropout-induced missingness. Notwithstanding, the consistently high recall across all configurations suggests that the model prioritizes sensitivity under uncertainty, a desirable trait in early breast cancer detection, where failing to identify malignancy is more expensive than issuing false alarms. Together, these findings position the proposed fusion model as resilient to incomplete data and align it clinically with the high-stakes demands of oncology diagnostics.
Table 6 provides a direct comparison between the proposed Sinkhorn approach and common late-fusion techniques. All strategies were evaluated under internally controlled conditions, with identical training and testing dataset splits. This uniformity eliminates confounding factors, ensuring that performance differences are attributable solely to the fusion mechanisms employed. Averaging, max-rule, and weighted fusion all demonstrated similar F1 scores (0.823), but lagged in PR-AUC. The MaxRule strategy, in particular, showed the weakest performance (PR-AUC = 0.694), confirming that static fusion rules fail to account for the complex interdependencies between modalities. Even logistic regression-based fusion, while stronger (PR-AUC = 0.760), lacked the adaptability of the Sinkhorn approach. The consistent superiority of the proposed fusion method suggests that optimal transport enables a more nuanced alignment between modalities, dynamically adjusting contributions based on the quality and availability of the input.
Comparative Performance of Fusion Strategies Across the Precision-Recall Curve; Performance Evaluation of Different Fusion Approaches, Including Simple Averaging, Weighted Averaging, Max Rule, Logistic Regression, and Sinkhorn-Based Optimal Transport Fusion, with Sinkhorn Achieving the Highest PR-AUC and F1 Score.

Finally, to contextualize our Sinkhorn-based fusion approach, Table 7 compares it against recent attention-based, uncertainty-aware, and adaptive fusion methods. Direct performance comparison is challenging due to differences in evaluation metrics (AUC vs PR-AUC), dataset characteristics, and experimental conditions.
Performance Comparison of State-of-the-Art Multi-Modal Feature Fusion Methods for Breast Cancer Diagnosis.

Attention-based fusion methods41,42 achieve superior discrimination on paired imaging modalities such as ultrasound-mammogram and ultrasound-MRI combinations. These methods excel when synchronized and high-quality imaging data are available during both training and inference. However, they require paired inputs at deployment and lack clear mechanisms for handling unpaired or missing modalities. Our approach achieves a lower PR-AUC (0.775) but maintains a comparable recall (0.96) while operating on intentionally unpaired histopathology-clinical data. This tradeoff reflects our design priority: robustness to data fragmentation over maximum performance under ideal conditions.
Our proposed method outperforms the evidential fusion methods 43 (AUC 0.736), with a PR-AUC of 0.775. Additionally, evidential approaches require paired inputs at inference, whereas our method maintains perfect recall (1.00) even at 50% modality dropout. Adaptive multimodal fusion 44 demonstrates strong performance with dynamic weighting mechanisms that adjust to input quality. Our recall (0.96) exceeds theirs (0.90), suggesting stronger sensitivity for malignancy detection, though our PR-AUC is lower. The key distinction lies in operational scope: adaptive methods adjust weights within synchronized multimodal inputs, while our OT framework learns distributional alignment between independently trained models. This enables deployment when modalities were never jointly collected, a common scenario in retrospective clinical data and multi-institutional collaborations.
These findings confirm that the Sinkhorn-based fusion model not only enhances performance under ideal conditions but also maintains reliability in the face of clinical uncertainty. This makes it a strong candidate for real-world implementation, particularly in settings where data quality and completeness cannot be guaranteed: precisely the context where breast cancer detection remains most challenging.
The DCA accentuates the clinical utility of the proposed late fusion model by consistently demonstrating higher net benefits than the “Treat All” and “Treat None” strategies across threshold probabilities ranging from 0.2 to 0.7. This performance indicates that the fused model distinguishes patients likely to benefit from intervention more effectively, minimizing overtreatment without compromising sensitivity. In practical terms, this translates into a favorable cost-benefit profile: fewer unnecessary procedures, better allocation of clinical resources, and improved patient outcomes. 45 The net benefit curve not only validates the model’s predictive value but also affirms its readiness for deployment in real-world diagnostic workflows where decision thresholds vary across clinicians and patient populations, accounting for both discrimination (AUC) and calibration.46,47
When evaluated on the naturalistic, imbalanced distribution, calibration curves (Figure 4) showed nearly perfect alignment between predicted and observed event rates. This supports the proposed model’s suitability and reliability for clinical triage, where maintaining calibration under real-world prevalence is crucial for clinical decision-making. The excellent calibration indicates well-calibrated probability estimates and low prediction uncertainty. This is quantified by a low Brier score of 0.19. Notably, this was achieved without artificial balancing at the fusion level, proving the robustness of the optimal transport framework.
Our minimal preprocessing philosophy directly supports clinical translation by reducing pipeline complexity and potential failure points. Unlike methods requiring extensive data curation and preprocessing steps, our proposed method can operate on routine clinical data with minimal preparation. In real-world deployments, especially within resource-limited healthcare systems, diagnostic pipelines are frequently fragmented, imaging may be unavailable at point-of-care stations, and electronic health records (EHRs) may be incomplete.46,48,49 A fusion model that maintains calibrated, sensitivity-prioritized outputs despite such variability can stabilize decision-making, reduce diagnostic inequities, and ensure patients are not denied timely intervention simply because their data are incomplete. The model’s ability to handle these challenges while operating on naturalistic, imbalanced data distributions validates our hypothesis that minimal intervention combined with robust fusion algorithms is more suitable for clinical deployment than heavily preprocessed, computationally intractable50,51 approaches. At a population level, this robustness could lead to earlier interventions. The fusion framework can be practically integrated into Picture Archiving and Communication System (PACS) or EHR middleware to combine histopathology predictions with available clinical risk information, generating calibrated malignancy probabilities that automatically flag high-risk cases for radiologists. At a larger scale, it could operate as a backend service for national screening initiatives, continuously updating risk stratification lists so patients with incomplete 52 or delayed records remain appropriately prioritized rather than excluded.
The computational efficiency profile demonstrates that the proposed fusion framework is well-suited for implementation in real-world clinical workflows. The 81 ms average inference latency indicates that the model can deliver near-instantaneous decisions, meeting the temporal constraints of interactive diagnostic tasks such as digital slide review and radiology consultation. The upper bound of observed latency (261 ms at the 95th percentile) remains below thresholds that could cause perceptible delays for clinicians. The negligible memory footprint (<0.1 MB) further enhances deployability across a wide spectrum of resource environments, including regional hospitals, outpatient screening centers, and telepathology platforms that may rely on CPU-only infrastructure. Because the Sinkhorn fusion operates on lightweight probability vectors, the method avoids the computational costs typically associated with multimodal deep learning models. This feasibility is reinforced by the linear scalability profile.
Processing times of under 1 minute for ~1000 cases suggest that the system can function in real-time decision support modes and in batch-processing contexts, such as overnight screening queues, retrospective audits, and large-scale population risk stratification initiatives. For institutions handling high patient volumes, this predictability and stability across load conditions are essential for ensuring smooth integration into laboratory information systems (LIS) and radiology information systems (RIS). More broadly, the model’s computational characteristics support its usability beyond tertiary care centers. In low-resource settings, where diagnostic bandwidth, hardware availability, and network infrastructure are restricted, the ability to run the entire pipeline efficiently on basic CPU hardware is a significant advantage. This facilitates deployment in mass screening programs, mobile diagnostic units, and telemedicine-supported pathology workflows, expanding access to advanced decision-support tools without imposing prohibitive infrastructural demands. 53
Overall, the performance results suggest that the proposed fusion framework is both diagnostically effective and operationally lightweight, scalable, and adaptable to diverse clinical environments. These attributes position the system as a strong candidate for real-world integration into multimodal breast cancer diagnostic pathways.
Limitations
While calibration and sensitivity are exceptional, overall PR-AUC remains moderate (≈ 0.78), mainly reflecting test dataset constraints (n = 313) and class imbalance (malignant: benign ratio of 2.3:1). Importantly, this performance was achieved while preserving the dataset’s naturalistic class distribution rather than applying artificial rebalancing techniques, demonstrating real-world applicability. Broader validation on larger, multi-institutional cohorts is essential to confirm generalizability across different scanners, clinical settings, and patient populations. Additionally, the computational complexity of entropy-regularized transport must be optimized to meet the throughput demands of high-volume screening environments.
Future Directions
Future research will explore extending the minimal intervention framework to handle even more severely imbalanced datasets and evaluate performance across diverse clinical environments where preprocessing capabilities may be limited. Additionally, this framework can be extended to multi-institutional settings and explored for the real-time adaptation of streaming clinical data, as well as the feasibility of mobile deployment. Furthermore, future work can explore the integration of multiple magnification levels and imaging modalities with the proposed model.
Conclusion
We introduce a Sinkhorn-based optimal transport late fusion framework for breast cancer diagnosis, which maintains high sensitivity (recall = 0.96) and calibrated probability estimates, even when up to 50% of the input modalities are missing or corrupted.
By incorporating entropy regularization and clinically informed cost matrices, the model addresses the key limitations of conventional fusion methods: susceptibility to data fragmentation and misalignment with clinical priorities. It consistently outperforms standard approaches in maintaining discrimination and net clinical benefit under varied data conditions.
These results demonstrate the framework’s suitability for deployment in real-world healthcare settings, where data is often incomplete. While external validation and computational optimization remain necessary, this approach offers a practical foundation for translating multimodal AI into routine clinical use.
Supplemental Material
sj-docx-1-cix-10.1177_11769351261420789 – Supplemental material for Resilient Sinkhorn-Based Optimal Transport Late Fusion Framework for Breast Cancer Diagnosis
Supplemental material, sj-docx-1-cix-10.1177_11769351261420789 for Resilient Sinkhorn-Based Optimal Transport Late Fusion Framework for Breast Cancer Diagnosis by Michael Asiedu Asare, Isaac Acquah, Benjamin Appiah Yeboah and Emmanuel Owusu in Cancer Informatics
Footnotes
Acknowledgements
This study was supported by the Biomedical Technologies Lab, Kwame Nkrumah University of Science and Technology, under project reference number BTL/2025/AIRD02.
Ethical Considerations
The datasets used in this research were obtained from publicly available repositories. These datasets had been ethically sourced and verified by expert clinicians. However, ethical approval was obtained from the Committee on Human Research, Publication and Ethics of the Kwame Nkrumah University of Science and Technology, with the approval reference number CHRPE/AP/370/25.
Consent to Participate
Consent to participate is not applicable because no participants were involved.
Consent for Publication
Not applicable because no participants were involved.
Author Contributions
Michael Asiedu Asare: Conceptualization, Methodology, Software, Formal analysis, Writing-original draft. Isaac Acquah: Validation, Supervision, Writing - review & editing. Benjamin Appiah Yeboah: Methodology, Project administration, Writing - review & editing. Emmanuel Owusu: Formal analysis, Writing - review & editing.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the TensorFlow College Outreach Award, provided by Google Ireland Limited.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The BreakHis dataset used and/or analyzed in the current study is available at http://www.inf.ufpr.br/vri/databases/BreaKHis_v1.tar.gz. The BCSC risk factors dataset used and/or analyzed during the current study is available at ![]() . We do not own these data.
. We do not own these data.
Declaration of Generative AI and AI-Assisted Technologies in the Manuscript Preparation Process
During the preparation of this work, the authors utilized Grammarly to enhance the clarity, sentence structure, and readability of the manuscript. After using this tool, the authors reviewed and edited the content as needed and take full responsibility for the content of the published article.
Footnotes
Isaac Acquah is the guarantor.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
