Abstract
Image-based diagnosis has become a crucial tool in the identification and management of various cancers, particularly lung and colon cancer. This review delves into the latest advancements and ongoing challenges in the field, with a focus on deep learning, machine learning, and image processing techniques applied to X-rays, CT scans, and histopathological images. Significant progress has been made in imaging technologies like computed tomography (CT), magnetic resonance imaging (MRI), and positron emission tomography (PET), which, when combined with machine learning and artificial intelligence (AI) methodologies, have greatly enhanced the accuracy of cancer detection and characterization. These advances have enabled early detection, more precise tumor localization, personalized treatment plans, and overall improved patient outcomes. However, despite these improvements, challenges persist. Variability in image interpretation, the lack of standardized diagnostic protocols, unequal access to advanced imaging technologies, and concerns over data privacy and security within AI-based systems remain major obstacles. Furthermore, integrating imaging data with broader clinical information is crucial to achieving a more comprehensive approach to cancer diagnosis and treatment. This review provides valuable insights into the recent developments and challenges in image-based diagnosis for lung and colon cancers, underscoring both the remarkable progress and the hurdles that still need to be overcome to optimize cancer care.
Introduction
Lung and colon cancers represent significant burdens on global health, in terms of both their prevalence and impact. Lung cancer is one of the most commonly diagnosed cancers in the world. It is often associated with high mortality rates owing to its aggressive nature and limited treatment options, particularly when diagnosed at advanced stages. 1 Smoking, environmental pollutants, and genetic predispositions contribute to its prevalence. 2 Conversely, colon cancer is among the most prevalent cancers globally, with incidence rates varying across regions. 3 Screening efforts have been instrumental in detecting colon cancer at earlier stages, significantly improving the survival rates. However, challenges persist in addressing lifestyle factors such as diet, physical activity, and genetic predispositions that influence its development. Both lung and colon cancers underscore the importance of public health initiatives, early detection strategies, and ongoing research efforts to improve prevention, diagnosis, and treatment outcomes, and mitigate their substantial impact on individuals and healthcare systems worldwide. 4
Early diagnosis plays a pivotal role in improving patient outcomes across various medical conditions, particularly in cancer. Timely detection enables the implementation of prompt and appropriate treatment strategies, often leading to a better prognosis and increased chances of successful treatment. 5 For cancers such as lung and colon cancer, early diagnosis allows for intervention at a stage when the disease may still be localized or minimally invasive, thereby offering more treatment options and potentially curative outcomes. 6 Moreover, early detection can reduce the need for aggressive treatment and decrease associated morbidity and mortality rates. Furthermore, it enables the initiation of supportive care measures and counseling, promoting overall well-being and quality of life for patients and their families. Therefore, prioritizing efforts toward early detection through screening programs, public awareness campaigns, and advancements in diagnostic technologies is crucial for improving patient outcomes and mitigating the burden of disease. 7
Importance of image-based diagnosis
Image-based diagnosis represents a promising approach in modern healthcare, leveraging cutting-edge technologies to enhance diagnostic accuracy and streamline patient care pathways. By harnessing various imaging modalities such as X-rays, CT, MRI, and PET, clinicians can visualize internal structures and detect abnormalities with unprecedented detail and precision.8,9 This noninvasive method allows for the early detection of diseases, including cancers, cardiovascular conditions, and neurological disorders, facilitating prompt intervention and personalized treatment strategies.10,11 Advancements in AI and ML have further revolutionized image interpretation, enabling automated analysis, pattern recognition, predictive modeling, augmenting diagnostic capabilities, and improving clinical decision making. As image-based diagnosis continues to evolve, it has immense potential to revolutionize healthcare delivery by optimizing resource utilization, reducing diagnostic errors, and ultimately, enhancing patient outcomes.
Overview of lung and colon cancer
Each subtype of lung cancer is characterized by a unique set of symptoms and treatment alternatives. The most prevalent subtypes of lung cancer are non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC).12 -14 Hereditary predispositions, tobacco use, and exposure to radon and asbestos are significant risk factors.15,16 The symptoms include weight loss, chest pain, difficulty in inhalation, and persistent wheezing. 17 Precancerous polyps frequently serve as the initial metastatic site for colon cancer, an ailment characterized by aberrant cellular proliferation in the colon or rectum. Risk factors include sedentary lifestyle, obesity, insufficient physical activity, smoking, advanced age, and a prior medical history of inflammatory bowel disease. 18 Indications included irregularities in bowel movements, diarrhea, abdominal discomfort, fatigue, and unanticipated weight loss. Early detection and treatment can improve the prognosis of patients with both the malignancies. Surgical intervention, radiation therapy, chemotherapy, targeted therapy, and immunotherapy are all potential therapeutic components that are contingent on the cancer subtype and stage. This underscores the importance of comprehensive screening initiatives and interdisciplinary treatment strategies.19,20
Scope and objectives of the review
Lung and colon cancers differ significantly in terms of tissue structure, diagnostic procedures, biological origins, and treatment approaches, and there are compelling reasons to include both types of cancer in a unified review. The rationale for this approach is outlined below.
• Both lung and colon cancer share a common pathway by which chronic inflammation contributes to carcinogenesis. Inflammation plays a pivotal role in the development of both cancers, although through different mechanisms and stages. By reviewing both types of cancer, we highlight the broader role of inflammation in cancer development and progression, which can lead to a more comprehensive understanding of this critical factor.
• There are common risk factors that affect both cancers, such as smoking, which is a significant risk factor for lung cancer and has been linked to an increased risk of colorectal cancer. Additionally, certain genetic mutations, such as those involving KRAS, are relevant to both types of cancer.
• Histopathological imaging plays a crucial role in the diagnosis and classification of both lung and colon cancers. Through microscopic examination of tissue samples, pathologists can identify the type and severity of cancer. This involves the analysis of cellular morphology, tissue architecture, and specific staining patterns that are vital for accurate diagnosis.
• Combining information on lung and colon cancer can offer valuable cross-cancer insights. Understanding how similar mechanisms operate in different types of cancer can lead to novel research directions and therapeutic strategies. A holistic review can facilitate the identification of universal principles and strategies that can be adapted to various cancer types.
In summary, although lung and colon cancers have distinct characteristics, the integration of these cancers in a single review provides a valuable opportunity to explore commonalities, shared risk factors, and advancements in diagnostics and treatment. The inclusion of histopathological images further supports the identification and classification of cancer types and severity, thereby enhancing our understanding of cancer mechanisms. This approach not only enriches our understanding of cancer biology but also supports the development of innovative strategies that could benefit research and clinical practice.
Methodology
To conduct a comprehensive review of the advances and challenges in image-based diagnosis for lung and colon cancer, integrating deep learning, machine learning, image processing methodologies, and focusing on X-rays, CT scans, and histopathological images, a systematic approach was meticulously adopted. The search strategy involved querying major academic databases such as PubMed, IEEE Xplore, and Google Scholar using a set of relevant keywords. These keywords included “lung cancer,” “colon cancer,” “deep learning,” “transformer,” “NLP,” “machine learning,” “chest,” “computed tomography,” “image processing,” “medical imaging,” “X-ray,” “CT scan,” and “histopathological images.” Various combinations of these keywords were used, such as “Lung cancer + deep learning + image processing + CT scan + diagnosis,” “Colon cancer + machine learning + histopathological images + chest X-ray + diagnosis,” “Lung cancer + deep learning + machine learning + histopathological images + diagnosis,” among others.
The search was not restricted by publication date to ensure an inclusive representation of the relevant literature across different time periods.
Upon the initial search, a substantial pool of articles was identified that underwent a rigorous screening process focusing on titles, abstracts, and keywords to exclude irrelevant studies. Articles not directly related to the integration of deep learning, machine learning, and image processing techniques in conjunction with X-rays, CT scans, and histopathological images for lung and colon cancer diagnosis were removed.
Subsequently, a final selection process was carried out to curate a diverse collection of articles that addressed the amalgamation of advanced image analysis techniques with X-rays, CT scans, and histopathological images in the domain of lung and colon cancer diagnosis. This meticulous selection procedure ensures comprehensive exploration of the advancements and complexities in utilizing these diagnostic modalities for improved cancer detection and diagnosis.
Brief Overview of the Biology and Pathology of Lung and Colon Cancer
The key clinical features of lung cancer include persistent cough, chest pain, shortness of breath, blood collection, fatigue, and unintentional weight loss. 21 However, lung cancer often presents asymptomatically or with nonspecific symptoms until it reaches an advanced stage, which complicates early detection. Diagnostic challenges include the absence of specific symptoms in the early stages, leading to delayed diagnosis and the overlap of symptoms with other respiratory conditions.22,23 The notable clinical features of colon cancer include changes in bowel habits, rectal bleeding, abdominal discomfort, fatigue, and unintended weight loss. Despite the availability of screening methods such as colonoscopy and fecal occult blood tests, challenges persist in achieving widespread participation in screening programs owing to discomfort, invasiveness, and lack of awareness.24,25 Moreover, the asymptomatic nature of early stage colon cancer can result in missed opportunities for timely diagnosis, highlighting the importance of enhancing screening efforts and public education to address these diagnostic challenges for both lung and colon cancers.
Current diagnostic methods for lung cancer include imaging techniques such as X-rays, CT scans, positron emission tomography (PET) scans, and tissue biopsy for confirmation.24,25 Although these methods are valuable for detecting lung nodules and assessing tumor characteristics, they have limitations such as radiation exposure, false positives, and difficulty distinguishing between benign and malignant nodules.26 -28 Additionally, sputum cytology can be used to identify cancer cells in the respiratory tract; however, its sensitivity is relatively low. Colonoscopy remains the gold standard for colon cancer detection, allowing the direct visualization and removal of precancerous polyps. 29 However, colonoscopy is invasive, uncomfortable, and carries a low risk of complications. Other diagnostic methods include fecal occult blood tests, stool DNA tests, and virtual colonoscopy, each with limitations such as false positives, false negatives, and the inability to detect small polyps. Despite these advances, current diagnostic methods for both lung and colon cancers still face challenges regarding accuracy, invasiveness, and patient acceptance, highlighting the ongoing need for improved screening techniques and technologies.30,31
Overview of Various Imaging Modalities Used in Lung and Colon Cancer Diagnosis
Various imaging modalities play crucial roles in the diagnosis and management of lung and colon cancer. X-ray imaging provides quick and low-cost initial assessments for lung cancer, although it lacks specificity and sensitivity.28,32 CT scans offer detailed cross-sectional images, aiding in detecting and characterizing lung nodules and colon tumors with high resolution and speed. 33 MRI provides excellent soft tissue contrast without ionizing radiation, which is beneficial for assessing the extent of tumor invasion and metastasis in both lung and colon cancer.34,35 PET scans are valuable for evaluating metabolic activity and staging cancer, and are particularly useful for detecting distant metastases. Colonoscopy remains the gold standard for colon cancer screening, offering direct visualization and biopsy capabilities for polyps and tumors.36 -38 Each modality has its advantages and limitations; for instance, while CT and MRI offer high-resolution images, they may require contrast agents and expose patients to radiation (in the case of CT) or longer scan times (in the case of MRI). 39 PET scans provide functional information, but can be costly and have limited spatial resolution. Additionally, image processing and analysis techniques, such as CAD and DL, are increasingly being utilized to enhance diagnostic accuracy and efficiency across these imaging modalities, aiding in the early detection, characterization, and treatment planning for lung and colon cancer patients.40,41 Generally, 2 types of images are used for lung and color cancer diagnosis, that is, radiological and histopathological images. Samples of radiological images of lung and colon cancers are illustrated in Figures 1 and 2, respectively. A sample of the histopathological images is illustrated in Figure 3.
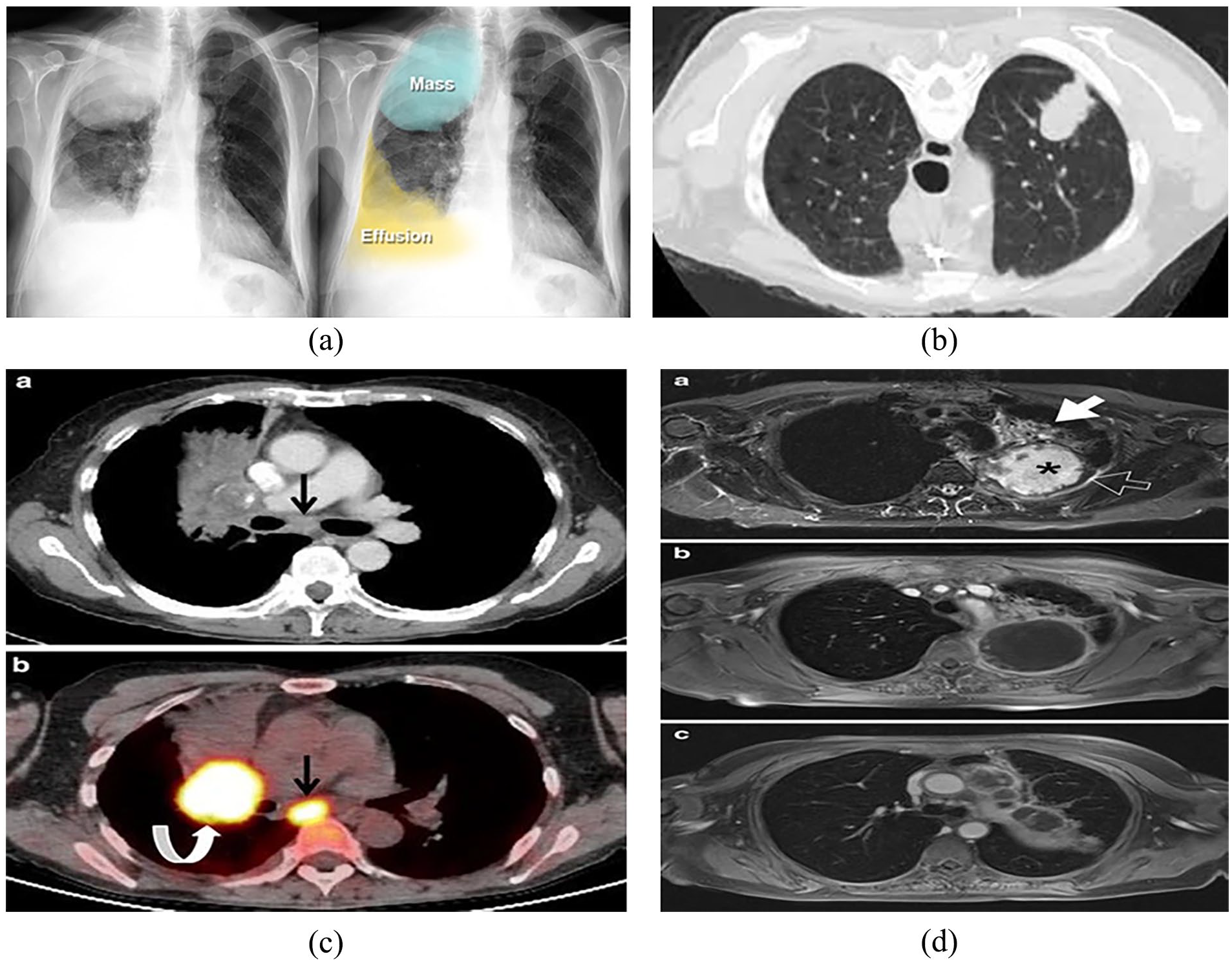
Radiological images of lung cancer: (a) Xray, (b) CT scan, (c) PET, and (d) MRI.
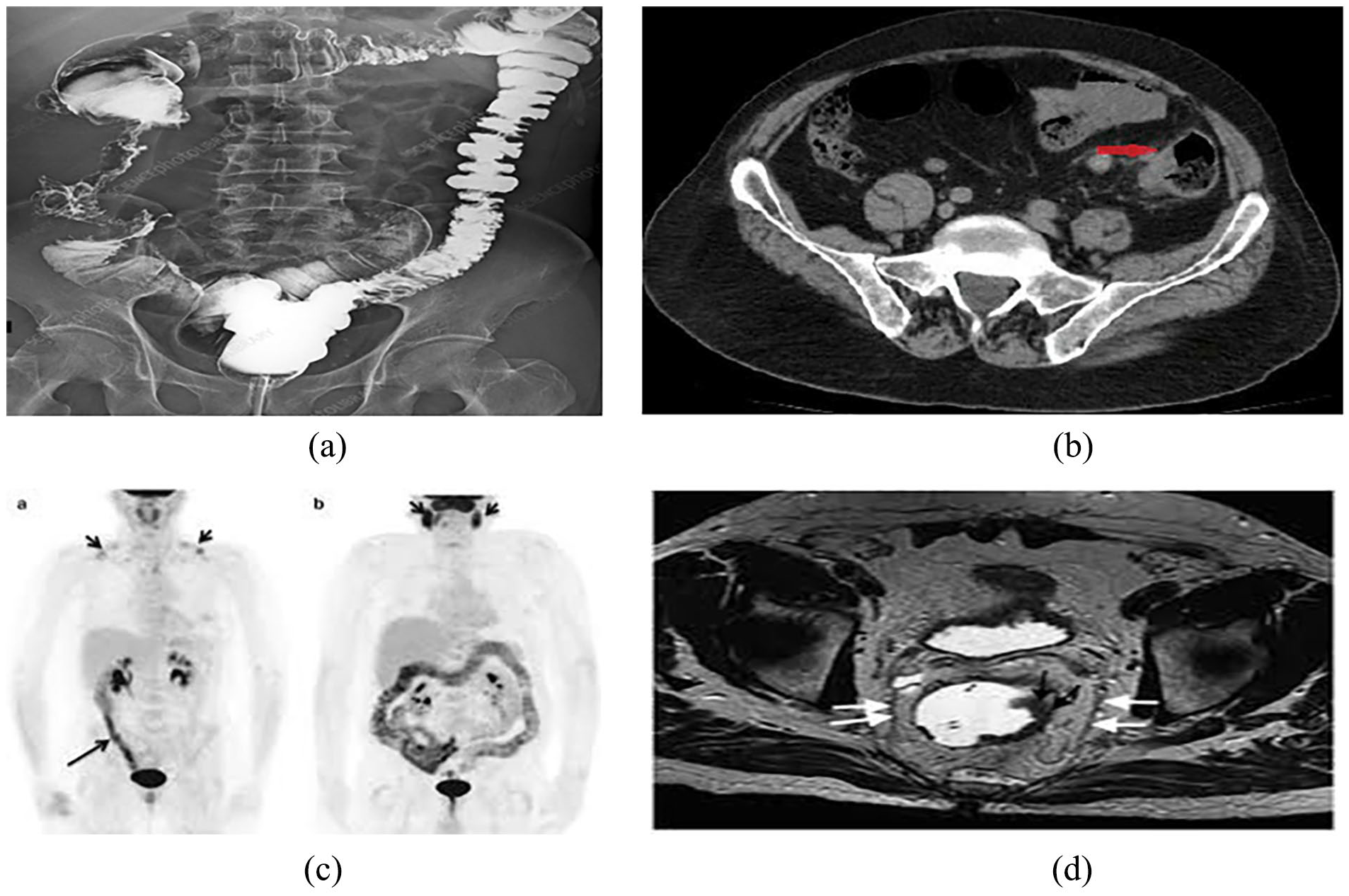
Radiological image of colon cancer: (a) Xray, (b) CT scan, (c) PET, and (d) MRI.

Histopathological images of lung and colon cancer.
Image-Based Diagnostic Approaches for Lung and Colon Cancer
In the past 4 decades, the disciplines of medicine and healthcare have made significant progress. During this timeframe, the true causes of several diseases were revealed, innovative diagnostic techniques were developed, and revolutionary medications were devised. Despite numerous accomplishments, diseases such as cancer persistently affect humans because of their ongoing susceptibility to them. Cancer is the second most prevalent cause of mortality worldwide, with approximately 1 in 6 individuals succumbing to the disease. Among the different types of malignancies, lung and colon cancer are the most prevalent and lethal, respectively. These cancers account for more than a quarter of all cancer cases. Nevertheless, the early detection of the illness greatly enhances the likelihood of survival. Many cancer diagnosis approaches utilize AI, enabling the evaluation of a greater number of cases within a shorter timeframe at a reduced cost. Cancer has a notably high mortality rate owing to its aggressive nature, strong tendency to spread to other parts of the body, and diverse characteristics. Lung and colon cancer are prevalent forms of cancer that can affect individuals of any sex and occur globally. Early and accurate identification of these tumors enhances the survival rate and improves the treatment efficacy. Al-Mamun et al. 42 presented a highly precise and computationally efficient model that is an alternative to existing cancer detection techniques. This model enables the rapid and accurate identification of malignancies in the lung and colon regions. The LC25000 dataset was used for the training, validation, and testing of this project. A cyclic learning rate was implemented to enhance the accuracy and improve the computational efficiency of the proposed technique. This approach is characterized by its simplicity and efficiency, which promotes faster model convergence. Various pre-trained TL models were utilized and compared with the CNN model developed from scratch. The study revealed that the proposed model demonstrated enhanced accuracy by mitigating the influence of inter-class variances between LDC and LSCC. The proposed method achieved a total accuracy of 97% and demonstrated superior computing efficiency compared with existing methods. According to Lakshmanaprabu et al., 43 lung cancer is a highly lethal disease that contributes significantly to cancer-related deaths worldwide. It is impossible to improve patient survival without the early detection of lung cancer. A CT scan is used to accurately localize a lesion and determine the extent of malignancy across the entire body. A novel automated diagnostic classification method for lung CT scan images is introduced in this study. LDA and ODNN were utilized in this study to evaluate CT scans of lung images. The CT lung images were processed to extract significant features, which were then downsized in dimensionality using LDR. The objective of this dimensionality reduction was to determine the malignant or benign nature of lung nodules. The ODNN was initially employed to analyze CT images; this approach was subsequently enhanced through the implementation of MGSA to classify lung cancer. The results of the comparisons demonstrated that the proposed classifier achieved a sensitivity of 96.2%, specificity of 94.2%, and accuracy of 94.56%. According to Lanjewar et al., 44 lung cancer is an exceptionally dangerous disease that poses a significant threat to human life globally. Therefore, the early identification of this condition is critical. Lung cancer was classified into 4 distinct categories using the Kaggle chest CT scan image dataset: adenocarcinoma, large-cell carcinoma, squamous cell carcinoma, and normal cells. An innovative approach utilizing DL was proposed for lung cancer detection by augmenting the DenseNet201 model with additional layers within the initial DenseNet framework. Two distinct methodologies for feature selection were employed to determine the most appropriate features to be extracted from DenseNet201. Next, several classifiers based on ML were implemented using these attributes. The confusion matrix, AUC, MCC, kappa score, 5-fold cross-validation procedure, and p-value were used to assess the performance of the system. After the proposed system adopted a 5-fold approach, it demonstrated exceptional performance, attaining a perfect accuracy rate of 100%, an average accuracy rate of 95%, and a P-value below .001, which was deemed to indicate statistical significance. This study emphasizes the application of ML algorithms to enhance the precision of lung cancer detection using CT images. Owing to its significant influence on patient prognosis, the AJCC employs tumor size as a criterion for classifying various solid tumors into the “T” stage. However, the extent of the tumor was not considered in the staging method for colon cancer. In contrast, emphasis is placed on metastasis (M), nodal status (T), and tumor status (N). As a result, the National Cancer Database was used to ascertain the correlation between survival, tumor size, and TNM stage. A cohort of 300 386 patients was analyzed, with tumor size being the determining factor in the following classifications: S1 (approximately 0-2 cm), S2 (greater than 2-4 cm), S3 (greater than 4-6 cm), and S4 (greater than 6 cm). Statistical analyses were performed to examine the relationships between TNM stage, grade, nodal status, and tumor size. Survival analysis for each “S” stage was conducted using the Kaplan–Meier technique. A total of 13%, 39%, and 18% of the 300, 386 patients were categorized as S1, S2, S3, and S4, respectively. The most prevalent site was the right colon, accounting for 48% of the cases. Positive correlations were observed between the extent of the tumor and its T stage, nodal stage, and grade. Survival was inversely proportional to the magnitude of the tumor. Vital prognostic factors are strongly correlated with the magnitude of malignancy, which is also associated with a decreased likelihood of survival. 44
Gowda et al. 45 aimed to enhance the precision of colon cancer diagnosis by employing DL algorithms instead of the traditional ML techniques. This technique entails preprocessing CT colonography images to eliminate noise and segmenting the colon into separate blocks. A pre-trained CNN was used to categorize these blocks into different types of colons and to identify irregular structures, such as polyps. The findings indicate that the CNN-based method attains superior accuracy (87% for colon segmentation and 88% for polyp detection) compared with the RF and KNN algorithms. This underscores the efficacy of DL in automating the diagnosis of colon cancer. 45 Sharma et al. 46 suggested a 2-step method for early lung cancer diagnosis using high-resolution CT images. The procedure involved extracting a patch from the center of the nodule and accurately segmenting the region of the lung nodule using the Otsu method and morphological procedures. This segmentation was achieved without relying on the complete contour information. In the second stage, a DCNN is employed to enhance the classification accuracy of the segmented patch (distinguishing between malignant and benign images). Experiments conducted on 6306 CT scans from the LIDC-IDRI database demonstrated a test accuracy of 84.13%. The sensitivity and specificity achieved were 91.69% and 73.16%, respectively, surpassing the performance of current cutting-edge algorithms. 46 Kumar et al. (2022) conducted a study focusing on lung and colon cancer classification utilizing 2 approaches: handcrafted feature extraction and deep feature extraction via transfer learning. They employed 6 manually designed strategies based on color, texture, shape, and structure to extract handcrafted features and trained the GB, SVM-RBF, MLP, and RF classifiers. Additionally, they utilized 7 DL frameworks with transfer learning to extract deep features from histopathological images of lung and colon cancers. A significant improvement in classifier performance was observed by comparing handcrafted features with deep features extracted by a deep CNN. The random forest classifier, leveraging DenseNet-121 for deep feature extraction, achieved high accuracy and recall rates of 98.60% and 98.63%, respectively, in identifying lung and colon cancer tissues, with a precision of 98.63% and an F1 score of 0.985. This study highlights the potential of DL techniques for improving cancer tissue classification in histopathological images, aiding in early detection and treatment. Additionally, they utilized visualization techniques such as GradCam and SmoothGrad to enhance classification, and proposed models utilizing MobileNetV2 and InceptionResnetV2, which achieved a precision of up to 99.95% in detecting cancer tissues, offering promise for automated and accurate cancer detection methods. 47 Lung cancer is a highly lethal type of cancer with one of the worst fatality rates globally, as identified by Shandilya et al. 48 “The primary objective of their study was to develop a CAD technique to categorize histopathological images of lung tissues. They used a publicly accessible dataset of 15 000 histopathological images of LDC, LSCC, and benign lung tissue from 3 distinct categories to create and validate a CAD system. Multiscale processing was employed to extract the image features. A comparison study was conducted using 7 pre-trained CNN models: MobileNet, VGG-19, ResNet-101, DenseNet-121, DenseNet-169, InceptionV3, InceptionResNetV2, and MobileNetV2. These models have been used to classify lung cancer. Pretrained models undergo hyperparameter tuning, which involves optimizing characteristics, such as batch size, learning rate, number of epochs, and model correctness. ResNet101 achieved the highest accuracy of all CNN versions, reaching 98.67%. This discovery will assist researchers in developing more effective CNN-based models for lung cancer. 48 ”
Prusty et al. 49 utilized EfficientNetB7 to create a classification model that integrates histopathological images to distinguish between 5 different types of lung and colon tissues, including 2 benign and 3 malignant tissues. Moreover, a lung and colon cancer predictive model was constructed using the histogram images from the Kaggle dataset. Precision, recall, and F1 scores were used to evaluate the model performance. According to the results, the EfficientNetB7 model achieved a notable accuracy of 98%. This model will assist medical professionals in formulating a proficient and suitable methodology for identifying different forms of lung and colon cancers. 49 “In addition, Hadiyoso et al. 50 reported that cancer is a non-communicable ailment and a primary contributor to mortality globally. Lung and colon cancers are the most prevalent types of cancers and are associated with significant fatality rates. An effective strategy to decrease mortality rates is to promptly identify diseases through early detection and appropriate medical treatment. Tissue samples and clinical pathological examinations are considered to be the most reliable and accurate methods for diagnosing cancer. Nevertheless, in certain instances, achieving a high level of precision is crucial when conducting pathological examinations of tissues at the cellular level. This accuracy is contingent upon the contrast of the pathological image, as well as the expertise of the clinician. Hence, it is imperative to employ an image processing methodology integrated with artificial intelligence to achieve automated classification. This study presents a novel technique for automated lung and colon cancer categorization using the DL methodology. The object identified in the image was a histological image depicting normal tissue, benign tissue, and cancer. A CNN using the VGG16 architecture and CLAHE was used to classify 25 000 histopathology images. Simulation results demonstrate that the proposed strategy achieves a maximum classification accuracy of 98.96%. The utilization of CLAHE enhances the performance of the system, resulting in superior detection accuracy compared with the absence of CLAHE. This improvement was stable for all epoch settings. This technique aims to assist clinicians in autonomously identifying cancers using massive datasets. It is expected to have a low cost, high accuracy, and rapid processing capabilities. 50 ” Kauczor et al. 51 mentioned that lung cancer is the most common lethal form of cancer, with a low chance of survival once the disease has progressed. The ability to screen persons at a high risk for lung cancer using annual LDCT demonstrated a survival advantage. ESR and ERS suggest that lung cancer screening should be conducted in well-structured, high-quality, long-term programs as part of a clinical trial or as regular practice at accredited multidisciplinary medical facilities. “The minimum need consists of standardized operating methods for acquiring low-dose images, computer-assisted examination of nodules, managing positive screening results, applying inclusion/exclusion criteria, managing expectations, and implementing smoking cessation programs. It is advisable to make additional improvements to enhance the quality, outcomes, and cost-effectiveness of lung cancer screening. These include incorporating risk models, minimizing radiation exposure, utilizing computer-assisted volumetric measurements, and evaluating comorbidities such as chronic obstructive pulmonary disease and vascular calcification. These parameters must be tailored to the area’s infrastructure and healthcare system. This will allow for precise determination of eligibility by utilizing a risk model, nodule management, and quality assurance strategy. It is highly recommended that a centralized registry, including a biobank and image bank, be created at the European level. 51 ” Coudray et al. 52 utilized a DCNN (specifically, Inceptionv3) to train on whole-slide images obtained from TCGA. This study aimed to automatically categorize lung tumors into 3 categories: LDC, LSCC, and normal lung tissue. The technique entailed instructing the network using diverse datasets such as frozen tissues, formalin-fixed paraffin-embedded tissues, and biopsies. The results demonstrated a performance similar to that of pathologists, with an average AUC of 0.97 for classification. 52 Bychkov et al. 53 utilized DL techniques to predict the outcomes of patients with colorectal cancer without the need for intermediate tissue classification. This approach involves constructing a complex neural network using digitized samples of cancer tissue microarrays (TMAs) stained with hematoxylin and eosin. These samples were obtained from 420 patients and were accompanied by clinicopathological and outcome data. This study showed that using DL to predict outcomes based on small tissue areas yielded better results than visual histological assessments by human experts. Patients were categorized as low- or high-risk at both TMA and whole-slide levels. The hazard ratio was 2.3, and the AUC was 0.69, indicating that DL can extract more prognostic information from colorectal cancer tissue morphology than human observers. 53 Shapcott et al. 54 utilized DL for cell identification algorithm for diagnostic images from TCGA colon cancer repository. The goal was to enhance the performance by employing within-image sampling while maintaining accuracy. The approach involved instructing the DL algorithm using a locally stored dataset, and subsequently utilizing it to analyze TCGA images by dividing them into smaller sections and identifying the cells inside them. The performances of 2 sample approaches, random and systematic random spatial sampling, were examined; the latter demonstrated a 7-fold enhancement in performance while maintaining good accuracy. The findings demonstrated connections between the anticipated cellular characteristics and clinical factors in the TCGA colon cancer data. For instance, a greater number of fibroblasts is associated with metastasis and other invasive factors, underscoring the ability of DL to generate morphological features that reflect cell density and tumor characteristics with clinical significance. 54 Hussain et al. 55 aimed to improve the detection of lung cancer by extracting many types of features such as texture, morphology, entropy, SIFT, and EFD features. This study utilized robust ML classification techniques, including the naïve Bayes, DT, and SVM methods, using Gaussian RBF and polynomial kernels. The findings demonstrate that the NB algorithm, when combined with entropy, SIFT, and texture features, yielded the highest detection accuracy of 100%. In addition, the use of texture features with an SVM polynomial resulted in excellent accuracy. Furthermore, the naïve Bayes classifier achieved the greatest separation, with an AUC value of 1.00 when using entropy, morphological, SIFT, and texture features. The decision tree and SVM polynomial kernel classifiers also achieved high separation using texture features. 55 According to Talukder et al., 56 cancer is a lethal condition that results from a combination of hereditary disorders and several metabolic irregularities. Lung and colon cancers are prominent contributors to mortality and morbidity in the human population. Histological identification of such malignancies typically plays a crucial role in determining the optimal course of therapy. Timely identification of the disease on either side of the body significantly reduces the probability of death. ML and DL algorithms can expedite cancer detection, enabling researchers to analyze many patients within a much-reduced timeframe at a decreased expense. “This study presents a novel hybrid ensemble feature extraction approach for the accurate identification of lung and colon cancer. The system combines advanced techniques such as deep feature extraction, ensemble learning, and high-performance filtering to analyze cancer image datasets. The model was assessed using histopathological (LC25000) datasets of the lungs and colon. The findings indicated that the hybrid model could accurately diagnose lung, colon, and combined lung and colon cancers with accuracy rates of 99.05%, 100%, and 99.30%, respectively. The results of this study demonstrate that the proposed technique significantly outperforms the existing models. Therefore, these models have the potential to be used in clinical settings to assist doctors with cancer detection and diagnosis. 56 ” According to Wahid et al., 57 both lung and colon cancers are highly lethal and can develop concurrently. The primary objective of most studies is to identify the specific ailment that affects a particular body organ. “In this study, a CAD system was developed that utilized a CNN to identify lung and colon cancer tissues in the LC25000 dataset. The LC25000 dataset comprises 25 000 histological color samples of colon and lung tissues, indicating the presence or absence of adenocarcinoma, which is a form of cancer. This study included 3 pre-trained CNN models, ShuffleNet V2, GoogLeNet, and ResNet18, as well as a customized CNN model. The evaluation metric used in this study indicated that ResNet18 achieved the highest accuracy of 98.82% in classifying lung cancer, whereas ShuffleNetV2 had the shortest training time of 1749.5 seconds. The ShuffleNetV2 model achieved the highest accuracy of 99.87% when used for colon classification with a remarkably rapid training time of 1202.3 seconds. The proposed customized CNN model achieved an accuracy of 93.02% in classifying lung cancer and 88.26% in classifying colon cancer. The CNN model suggested in this study had the fastest training time, outperforming both the GoogLeNet and ResNet18 models. 57 ” According to Mengash et al., 58 cancer is a fatal illness that results from a range of biochemical abnormalities and genetic disorders. “Colon and lung cancer have emerged as significant contributors to disability and mortality in humans. Histological identification of these cancers is crucial for selecting the best course of action. Early and prompt diagnosis of illness on either side reduces the likelihood of mortality. DL and ML techniques have been employed to expedite cancer detection, enabling the research community to analyze more patients in a significantly shorter timeframe and at a reduced cost. This paper presents the MPADL-LC3 approach, which utilizes DL to classify lung and colon cancer. The MPADL-LC3 approach was designed to accurately differentiate between various forms of lung and colon cancer based on histopathological images. To achieve this, the MPADL-LC3 approach utilizes CLAHE-based contrast enhancement as a pre-processing step. Furthermore, the MPADL-LC3 method utilizes MobileNet to generate feature vectors. The MPADL-LC3 approach utilizes the MPA as a hyperparameter optimizer. In addition, DBNs can be used to classify the lungs and colors. An analysis was conducted on benchmark datasets to evaluate the simulation values of the MPADL-LC3 approach. The comparative analysis emphasized the improved results of the MPADL-LC3 system in terms of various metrics. 58 ” Ibrahim and Talaat 59 identified lung and colon cancers as the most prevalent and lethal forms of cancer, respectively. It accounts for approximately 25% of all cancer cases. “Detecting the disease at an early stage significantly increased the likelihood of survival. The classification accuracy was improved and cancer detection was automated using Double CLAHE stages and modified neural networks for image enhancement by employing DL methods. This study introduces a novel AI classification system that can accurately identify 5 different types of colon and lung tissue. These included 3 types of malignant tissue and 2 types of benign tissue. This method is based on analyzing histological images, and can classify lung cancer into 3 distinct classes and colon cancer into 2 distinct classes. The findings of this study revealed that the proposed approach has a high level of accuracy and correctly identifies cancerous tissues up to 99.5% of the time. This model will assist medical professionals in creating an automated and dependable system for identifying various types of colon and lung tumors. 59 ” According to Attallah et al., 60 lung and colon cancer are among the primary causes of death and illness in individuals. “They can occur simultaneously in organs and have detrimental effects on human life. If the cancer is not detected during its initial stages, there is a high probability that it may metastasize to many organs. Histological identification of these malignancies is vital for successful treatment. Despite their lengthy and complex nature, DL approaches have significantly accelerated and enhanced the procedure, allowing researchers to efficiently investigate more patients within a shorter timeframe at reduced cost. Prior research has relied on DL models, which require substantial computer power and resources. Most rely on separate DL models to extract high-dimensional data or to conduct diagnoses. Nevertheless, this study introduces a system that relies on several efficient DL models to detect early stage lung and colon tumors. The framework employs multiple transformation techniques to reduce the number of features and enhance data representation. Histopathological scans were inputted into the ShuffleNet, MobileNet, and SqueezeNet models within this framework. The deep features obtained from these models were then reduced using PCA and the FHWT algorithms. Subsequently, the reduced features of the FWHT acquired from the 3 DL models were fused using DWT. Furthermore, the PCA features of the 3 DL models are combined. Ultimately, the reduced features obtained via the PCA and FHWT-DWT reduction and fusion techniques were input into 4 separate ML algorithms, resulting in an impressive accuracy of 99.6%. The findings achieved using the suggested framework, which relies on efficient DL models, demonstrate its ability to differentiate between lung and colon cancer types. Compared with the current approaches, this is accomplished with a reduced set of characteristics and less computational complexity. This study also demonstrated that employing transformation methods to decrease characteristics can provide a more accurate understanding of the data, thereby enhancing the diagnostic procedure. 60 ” According to Hoang et al., 61 cancer treatment poses a significant challenge. Lung cancer, along with colon, stomach, and liver cancer, is one of the most difficult forms of cancer. The early detection and treatment of this condition can significantly extend a patient’s lifespan. DL approaches can be used to perform several tasks across various industries, including medicine. This research suggests modifying the transfer learning of deep neural networks for classifying lung and colon cancers using the GoogLeNet model. The primary concept of the inception module in GoogLeNet involves simultaneously performing numerous operations (such as pooling and convolution) with various filter sizes, thereby eliminating the need for compromise. The second benefit of the inception module is the decrease in dimensionality of the feature maps and the handling of overparameterization. The categorization output was modified to include only 3 or 2 classes to accommodate the specific lung and colon tissue classes. The proposed method achieved accuracies of 99.66% and 100% on lung and colon imaging datasets, respectively. The proposed method outperformed previous algorithms, including VGG16, ResNet50, NASNetMobile, and GoogLeNet. 61 Bychkov et al. 53 utilized a combination of convolutional and recurrent architectures to train a deep network to predict the outcomes of patients with colorectal cancer. “Predictions were made based on images of the tumor tissue samples. They claimed that their approach was unique because they directly forecasted patient prognosis without any intermediary tissue classification. They analyzed a collection of digitized hematoxylin-eosin-stained tumor tissue microarray (TMA) samples from 420 patients with colorectal cancer. Clinicopathological and outcome data were obtained for these patients. The findings indicate that using DL to predict outcomes based on small tissue areas as input is more effective (hazard ratio 2.3; 95% CI 1.79–3.03; AUC 0.69) than visual histological assessments conducted by human experts on both TMA spots (HR 1.67; 95% CI 1.28–2.19; AUC 0.58) and whole-slide levels (HR 1.65; 95% CI 1.30–2.15; AUC 0.57) when categorizing patients into low- and high-risk groups. These findings indicate that advanced DL methods may derive more predictive data from the physical characteristics of CRC cells than a skilled human observer. 53 ” Tharwat et al. 62 argued that addressing colon cancer treatment poses significant social and economic issues, mostly because of the elevated mortality rates associated with the disease. Annually, approximately 500 000 individuals are diagnosed with cancer, including colon cancer, worldwide. Assessment of colon cancer grade mostly relies on examination of the structure of the gland in different tissue regions. This has resulted in the development of multiple screening methods that can be employed to analyze polyp images and detect colorectal cancer. This study provides an extensive examination of the process of colon cancer diagnosis. This comprehensive article encompasses all aspects of colon cancer, including symptoms, grades, and imaging modalities used for analysis, with a special focus on histopathological images. Additionally, it discusses the commonly used diagnostic systems. In addition, this study explored datasets and performance evaluation criteria that are commonly used. They offered an extensive analysis of the existing research on colon cancer, categorized it into DL and ML methods, and highlighted the primary advantages and drawbacks of these methods. These techniques offer substantial assistance in detecting the initial phases of cancer, which enables prompt treatment of the disease and results in a reduced mortality rate compared to the rate observed when symptoms manifest. Furthermore, these techniques can impede the progression of colorectal cancer by eliminating premalignant polyps. This can be accomplished by using screening tests to facilitate early disease detection. The text presents the current issues and prospective research directions that pave the way for further study. 62 According to Hamed et al., 63 colon cancer is one of the primary causes of death and illness in humans. Histopathological diagnosis is a crucial factor in defining cancer type. This article describes the development of a computer-aided design (CAD) system for colon adenocarcinoma. The system utilizes ML techniques to assess digital pathology images. A dataset including 10 000 photos was collected from the LC25000 collection, with an equal distribution of 5000 images for each class. The classification model employed a CNN with a light-gradient boosting machine (CNN-LightGBM) employing multiple threads. The proposed system was compared with other ML algorithms. The claimed diagnostic accuracy for colon cancer has surpassed 90%, surpassing the accuracy of the latest ML algorithms regarding illness classification. Nevertheless, the precision was lower than that of the lung cancer classification achieved using this method. This study demonstrates the ability of ML to enhance the precision and effectiveness of medical diagnoses. It also emphasizes the necessity for additional research to enhance the precision of colon cancer diagnosis. 63 AlGhamdi et al. 64 emphasized the importance of promptly detecting lung and colon cancers to enhance patient outcomes and ensure treatment efficacy. “The HPI has become a reliable and effective diagnostic method for cancer. HPI analysis for LCC diagnosis involves thorough evaluation and analysis of tissue samples obtained from LCC to identify any abnormalities or malignant cells. It plays a crucial role in the staging and diagnosis of this tumor and contributes to the prognosis and treatment planning. However, manually analyzing an image is time consuming and prone to human error. Hence, the detection of LCC using HPIs requires CAD techniques. TL utilizes pre-trained DL algorithms trained on a larger dataset to extract relevant features from HPIs. These features were then used to develop a classifier for tumor diagnosis. This paper presents the invention of the BERTL-HIALCCD approach, which optimizes the Earth’s radius of Al-Biruni using transfer-learning-based histopathological image analysis to detect lung and colon cancer. This study aimed to accurately identify LCC in histopathological images. The BERTL HIALCCD approach employs computer vision and TL principles for precise LCC identification. The BERTL-HIALCCD approach uses an enhanced ShuffleNet model for feature extraction, and its hyperparameters are determined by the BER system. A DCRNN model was used to effectively recognize LCC. The Coati optimization algorithm was used to select the parameters for the DCRNN technique. To evaluate the effectiveness of the BERTL-HIALCCD technique, a series of extensive experiments was performed on a substantial dataset of histopathological images. The experimental results indicate that integrating the AER and COA algorithms achieves superior performance in cancer detection compared to the other tested models. 64 ” According to Xu et al., 65 tumors are dynamic biological systems that are undergoing ongoing evolution. Medical imaging has the distinct advantage of tracking these changes throughout the treatment. Although it may be easy to subjectively follow lesions over space and time, developing automated radiomic systems that combine serial imaging data and are clinically meaningful is much more difficult. This study assessed the performance of DL networks in predicting clinical outcomes by evaluating the time-series CT scans of patients diagnosed with locally advanced NSCLC. Dataset A included 179 patients diagnosed with stage III NSCLC, who underwent definitive chemoradiation. The dataset included pretreatment and post-treatment CT images taken at the 1-, 3-, and 6-months follow-ups, totaling 581 scans. The models were created by applying transfer-learning techniques to combine CNNs with RNNs. These models were trained using a single-point tumor localization approach. Dataset B, which consisted of 89 patients with NSCLC who underwent chemoradiation and surgery, was subjected to pathological response validation. In total, 178 scans were included in the analysis. Time-series scans have been effectively utilized in DL models to accurately predict survival and particular cancer-related outcomes including progression, distant metastases, and locoregional recurrence. The performance of the CNN model improved with each consecutive follow-up scan, as indicated by the increase in the AUC for 2-year overall survival to 0.74, with a significance level of P < .05. The models categorized patients into low- and high-mortality risk groups, which showed a strong correlation with overall survival [hazard ratio (HR) = 6.16; 95% confidence interval (CI), 2.17–17.44; P < .001]. The model accurately predicted the pathological response in dataset B with a substantial level of confidence (P = .016). They proved that DL can include imaging scans taken at various intervals, enhancing the accuracy of predicting clinical outcomes. AI-driven non-invasive radiomic biomarkers can greatly influence clinical practice because of their cost-effectiveness and limited human involvement. Medical imaging is a non-invasive method for monitoring the response and progression of tumors in patients following treatment. Nevertheless, quantitative evaluation using manual measures is laborious, time-consuming, and susceptible to differences across operators, as subjective visual judgment can lack objectivity and be influenced by personal bias. AI can automatically measure and analyze the radiographic features of tumor phenotypes. It can also be used to quantitatively track tumor changes before, during, and after therapy. This study demonstrated the ability of DL networks to accurately predict prognostic outcomes in patients undergoing radiation therapy by utilizing serial CT images that are regularly collected during the follow-up process. They further emphasized their capacity to account for and utilize existing sequential images to extract key time points and image characteristics relevant to predicting survival and response to therapy. This information offers a better understanding of applications such as identifying significant remaining diseases without surgery and other personalized medicine techniques. 65 Agbley et al. 66 powered DL models using extensive datasets. Nevertheless, the availability of medical data is a barrier that affects the reliability of computer-aided diagnostic models. There are multiple reasons for the scarcity of labeled data. One area of expertise is the process of annotating biopsies and scans obtained from the laboratories. Another factor to consider is the delicate and confidential nature of the medical information. “This study aimed to enhance the automatic feature engineering capabilities of DL by utilizing data from other diseases gathered using the same technique, thereby increasing the amount of accessible data. Therefore, this study examined the training of a model that could categorize 2 distinct diseases into their respective subclasses using multiple centers. The data for each disease were stored on individual devices to ensure that the original data remained confidential and exclusive to each device. Each center trains VGG16 locally, and the parameters are then shared and aggregated to create a global model. This experiment employed the LC25000 dataset, which consists of lung and cancer biopsy images. The global model was evaluated using distinct test sets for patients 1 (lungs) and 2 (colon). In addition, they conducted centralized learning (CL) by aggregating the 4 classes used in the decentralized experiment. The methodology achieved exceptional results, surpassing current state-of-the-art methods, while ensuring data confidentiality. 66 ” According to Ren et al., 67 cancer ranks as the second most prevalent cause of mortality globally, with lung cancer exhibiting much higher fatality rates than other cancer forms. “Many innovative computer-aided diagnostic methods utilizing DL have recently been developed to identify early stage lung cancers. However, DL models are prone to overfitting, which leads to poor performance. To address the issue of lung cancer classification, a combined LCGANT approach was used. This system comprises of 2 primary components. The initial component is a DC-GAN designed for lung cancer, which is capable of generating synthetic images. The second component is regularization-enhanced transfer learning, which categorizes lung cancer images into 3 distinct categories. The framework achieved an accuracy of 99.84% ± 0.156%, precision of 99.84% ± 0.153%, sensitivity of 99.84% ± 0.156%, and an F1-score of 99.84% ± 0.156%. The outcome achieved the highest performance in the dataset for classifying lung cancers.” The proposed framework effectively addresses the issue of overfitting in lung cancer classification tasks, surpassing the performance of existing cutting-edge methods. 67
Zheng et al. 68 proposed the Radiology Analysis and Malignancy Evaluation Network (R2MNet) for assessing pulmonary nodule malignancy through radiological characteristics, using radiological features as channel descriptors to emphasize crucial regions for malignancy evaluation. “The study also introduced Channel-Dependent Activation Mapping (CDAM) to visualize features and clarify the decision-making process of deep neural networks (DNNs). Experimental results on the Lung Image Database Consortium Image Collection (LIDC-IDRI) dataset demonstrated that R2MNet achieved an area under the curve (AUC) of 0.92 for nodule radiology analysis and 0.87 for malignancy evaluation. CDAM feature explanations revealed that nodule shape and density are critical factors in malignancy inference, aligning with the diagnostic cognition of experienced radiologists, thereby enhancing the confidence and interpretability of evaluation results. 68 ”
Zhu et al. 69 presented DeepLung, a fully automated lung computed tomography (CT) cancer diagnosis system. “DeepLung consists of 2 components: nodule detection, which identifies candidate nodule locations, and classification, which differentiates benign and malignant nodules. To handle the 3D nature of lung CT data efficiently, 2 deep 3D dual-path networks (DPN) were designed for these tasks. For nodule detection, a 3D faster region with a convolutional neural network (R-CNN) utilizing 3D dual path blocks and a U-net-like encoder-decoder structure is employed to learn nodule features effectively. For nodule classification, a gradient boosting machine (GBM) with 3D dual-path network features was proposed. This classification subnetwork, validated on the LIDC-IDRI public dataset, outperformed state-of-the-art methods and experienced doctors in image-based diagnoses. In DeepLung, candidate nodules are first detected by the nodule detection subnetwork, followed by nodule diagnosis via the classification subnetwork. Extensive experiments showed that DeepLung’s performance is comparable to that of experienced doctors for both nodule- and patient-level diagnoses on the LIDC-IDRI dataset. 69 ”
Pardyl et al. 70 introduces a fully automated pipeline called CompLung for lung cancer screening, utilizing machine learning tools to address challenges in lung segmentation and detection of potentially malignant nodules in CT scans.The proposed tool provides organ segmentation, patient-level cancer probability, and locations of suspicious regions.The authors trained CompLung using the LIDC-IDRI dataset and demonstrate its superior performance and interpretability compared to existing methods for lung cancer diagnosis 70 .
Pardyl et al. 71 addressed the limitations of automated lung cancer classification methods, which often rely on the LIDC-IDRI dataset for training, and typically focus on node-level classification, resulting in poor patient-level diagnostic outcomes. “In their paper, they introduced an end-to-end method that takes a CT scan as input and provides patient-level diagnosis as the output. This study explored 3 approaches under different data regimes to assess how varying levels of supervision impact model performance, aiming to enhance both node-level and patient-level diagnostic accuracy. 71 ”
Souza et al. 72 addressed the challenge of automatically segmenting lung fields in chest X-rays (CXR), especially when dense abnormalities such as opacities from diseases like tuberculosis and pneumonia are present. “These opacities often lead to incomplete segmentation, as they can be misinterpreted as lung boundaries. To address this, the authors proposed a method involving 4 main steps: image acquisition, initial segmentation, reconstruction, and final segmentation, utilizing 2 deep convolutional neural network (CNN) models. Testing 138 CXR images from Montgomery County’s Tuberculosis Control Program showed impressive results, with an average sensitivity of 97.54%, specificity of 96.79%, accuracy of 96.97%, Dice coefficient of 94%, and Jaccard index of 88.07%. The study demonstrated that incorporating a reconstruction step effectively addresses the problem of dense abnormalities, significantly enhancing lung segmentation accuracy. 72 ”
Shen et al. 73 investigate diagnostic lung nodule classification using thoracic Computed Tomography (CT) screening, addressing the challenge of modeling raw nodule patches without predefined nodule morphology. “They proposed a hierarchical learning framework called Multi-scale Convolutional Neural Networks (MCNN) to capture nodule heterogeneity by extracting discriminative features from alternatingly stacked layers. This framework uses multiscale nodule patches to simultaneously learn class-specific features by concatenating the response neuron activations from the last layer of each input scale. Evaluated on CT images from the Lung Image Database Consortium and Image Database Resource Initiative (LIDC-IDRI), this method is effective in classifying malignant and benign nodules without relying on nodule segmentation. 73 ”
Shen et al. 74 address the challenge of insufficient imaging samples with pathologically-proven labels in developing CNN models for predictive lung cancer diagnosis. They proposed a domain-adaptation framework to learn transferable deep features to predict patient-level lung cancer malignancies. “This study utilized CNN-based features from a large discovery set of 2272 lung nodules, labeled with malignancy likelihood based on multiple radiologists’ assessments, which were tested on a smaller diagnosis-definite set of 115 cases with pathologically proven lung cancer labels. Evaluated on the LIDC-IDRI dataset, their approach demonstrated superior predictive performance for patient-level malignancy (Acc = 70.69%, AUC = 0.66), outperforming a nodule-level CNN model (Acc = 65.38%, AUC = 0.63), and comparable to radiologists’ knowledge (Acc = 72.41%, AUC = 0.76).” The model significantly reduces the need for pathologically proven data, offering the potential to enhance cancer diagnosis by leveraging multisource CT imaging datasets. 74
Haghighi et al. 75 explored the untapped potential of rich semantics embedded in medical images for self-supervised learning aimed at enhancing deep semantic representation learning. “They introduced a novel framework that trained deep models to learn semantically enriched visual representations through self-discovery, self-classification, and self-restoration of anatomical structures in medical images, resulting in a pretrained 3D model called Semantic Genesis. This model was evaluated against all publicly available pre-trained models, both self-supervised and fully supervised, across 6 distinct target tasks encompassing classification and segmentation in various medical modalities (CT, MRI, and X-ray). Extensive experiments revealed that Semantic Genesis significantly outperforms its 3D counterparts and the widely used ImageNet-based transfer learning in 2D because of the framework’s ability to leverage consistent anatomical patterns in medical images for compelling semantic representation learning. 75 ”
Asuntha et al. 76 address the significant global mortality caused by lung cancer, with about 5 million deaths annually. Their study aimed to detect cancerous lung nodules on CT scans and classify the severity of lung cancer. “The approach utilizes advanced deep learning methods and effective feature extraction techniques, including the Histogram of Oriented Gradients (HOG), wavelet transform-based features, Local Binary Pattern (LBP), Scale Invariant Feature Transform (SIFT), and Zernike Moment. These techniques extract texture, geometric, volumetric, and intensity features, which are then optimized using the Fuzzy Particle Swarm Optimization (FPSO) algorithm to select the best features. The selected features are classified using a novel FPSOCNN, which reduces the computational complexity of the traditional CNNs. The performance of the method was validated on a real-time dataset from the Arthi Scan Hospital. The experimental results demonstrate that the novel FPSOCNN outperforms other techniques in detecting and classifying lung cancer. 76 ”
Ardila et al. 77 address the high mortality rate of lung cancer in the United States, which caused an estimated 160 000 deaths in 2018. “Lung cancer screening using low-dose computed tomography (CT) has been shown to reduce mortality by 20-43% and is included in the US screening guidelines. However, challenges such as inter-grade variability and high false-positive and false-negative rates persist. To address these issues, the authors propose a deep learning algorithm that predicts lung cancer risk by analyzing a patient’s current and prior CT volumes. Their model achieved state-of-the-art performance with an area under the curve (AUC) of 94.4% on 6716 cases from the National Lung Cancer Screening Trial and showed similar performance on an independent clinical validation set of 1139 cases. In 2 reader studies, the model outperformed all 6 radiologists in scenarios in which prior CT imaging was unavailable, reducing false positives by 11% and false negatives by 5%. When prior imaging was available, the model’s performance was similar to that of radiologists. This demonstrates the potential of deep learning models to enhance the accuracy, consistency, and adoption of lung cancer screening, optimizing the screening process through computer assistance and automation. 77 ”
Lee et al. 78 conducted a study to evaluate the effectiveness of a deep learning algorithm for detecting lung cancer in chest radiographs, supported technically by Lunit but independently designed and analyzed. “This study involved a screening cohort from 2008 to 2012, comprising 10 202 individuals with lung cancer cases determined by experienced radiologists. Validation tests were conducted by board-certified radiologists to compare the performance of the algorithm with that of radiologists. Using 10 285 radiographs, including 10 with visible lung cancers, the algorithm achieved an AUC of 0.99 and a sensitivity of 90% compared to radiologists’ 60% sensitivity, albeit with a higher false-positive rate (FPR) of 3.1% versus 0.3% (P < .001). In a larger screening cohort of 100 525 radiographs, including 47 with visible lung cancers, the algorithm achieved an AUC of 0.97, with 83% sensitivity and 3% FPR. The results indicate that the deep learning algorithm has a lung cancer detection performance comparable to that of radiologists, suggesting its potential to assist in screening populations with a low prevalence of lung cancer. 78 ”
Huang et al. 79 aimed to develop a more accurate lung cancer screening protocol by predicting the 3-year lung cancer risk after 2 CT scans using a deep learning algorithm called DeepLR. “The algorithm was trained on data from the National Lung Screening Trial (NLST) and validated using data from the Pan-Canadian Early Detection of Lung Cancer (PanCan) study. DeepLR performance was compared with that of the Lung CT Screening Reporting and Data System (Lung-RADS) and volume doubling time. The training cohort included 25 097 participants from the NLST and the validation cohort included 2294 individuals from PanCan. DeepLR demonstrated high accuracy, with time-dependent AUC values for cancer diagnosis at 1, 2, and 3 years of 0.968, 0.946, and 0.899, respectively. It identified 94%, 85%, and 71% of lung cancers diagnosed within 1, 2, and 3 years, respectively, in high-risk individuals deemed high risk. Additionally, those with high DeepLR scores had a significantly higher risk of mortality than those with high Lung-RADS scores (hazard ratio 16.07, 95% CI 10.15–25.44; P < .0001).” DeepLR effectively recognizes patterns in nodule and non-nodule features, providing accurate guidance for clinical management following repeat CT screenings. 79
Lu et al. 80 conducted a study to improve the assessment of tumor response in metastatic colorectal cancer (mCRC) patients undergoing anti-vascular endothelial growth factor therapies, where morphological changes in tumors may precede size changes. “Using a deep learning (DL) network, they retrospectively analyzed data from 1028 patients with mCRC in the VELOUR trial (NCT00561470). The DL network outperformed traditional size-based assessments in predicting early on-treatment response, achieving a C-index of 0.649 (95% CI 0.619–0.679) compared to 0.627 (95% CI 0.567–0.638) for size-based methods (P = .009, z-test). Combining DL with size-based methodology further improved the prediction performance to a C-index of 0.694 (95% CI 0.661–0.720), which was significantly superior to either model alone (P < .001, z-test). This study highlights the potential of DL networks to provide a noninvasive, quantitative, and comprehensive assessment of tumor morphological changes, enhancing personalized early on-treatment decision-making in patients with mCRC. 80 ”
Zhang et al. 81 developed and validated a deep learning (DL) method to predict the response of locally advanced rectal cancer to neoadjuvant chemoradiotherapy using diffusion kurtosis and T2-weighted MRI. “In this prospective study, 383 participants with rectal adenocarcinoma (⩾cT3 or N+) were enrolled between October 2015 and December 2017 and were divided into 308 training and 104 test samples. DL models primarily predicted pathologic complete response (pCR) and secondarily assessed tumor regression grade (TRG) and T downstaging. The DL model achieved an area under the receiver operating characteristic curve (AUC) of 0.99 for pCR in the test cohort, significantly outperforming 2 radiologists with AUCs of 0.66 and 0.72 (P < .001). The model’s AUCs for TRG and T downstaging were 0.70 and 0.79, respectively. The AUC of the DL model for pCR also surpassed that of the best-performing diffusion-kurtosis MRI parameter (Dapp value, AUC = 0.76). Radiologists’ subjective evaluations had higher error rates (26.9% and 24.8%) than the DL model (2.2%), but error rates decreased to 12.9% and 14.0% when assisted by the DL model. The study concluded that the DL model based on diffusion kurtosis MRI effectively predicts pCR and assists radiologists in evaluating the response of rectal cancer to neoadjuvant chemoradiotherapy. 81 ”
Zhao et al. 82 developed and validated a deep-learning-based, fully automated lymph node detection and segmentation (auto-LNDS) model for rectal cancer (RC) staging using multiparametric magnetic resonance imaging (mpMRI). The study included 5789 annotated lymph nodes (LNs) from 293 patients with RC. “The auto-LNDS model, based on Mask R-CNN, fused T2-weighted images (T2WI), and diffusion-weighted images (DWI), was validated on both internal (935 LNs) and external (1198 LNs) datasets. For LNs detection, the model achieved a sensitivity of 80.0%, positive predictive value (PPV) of 73.5%, false positive rate per case (FP/vol) of 8.6 in internal testing, sensitivity of 62.6%, PPV of 64.5%, and FP/vol of 8.2 in external testing. These results were significantly better than those obtained by the junior radiologists. The detection and segmentation time was 1.3 seconds per case, compared to 200 seconds per case for radiologists. The Dice similarity coefficient (DSC) for LNs segmentation ranged from 0.81 to 0.82.” The study concluded that the auto-LNDS model significantly improved the efficiency and accuracy of LNs detection and segmentation in RC staging. 82
Shayesteh et al. 83 investigated the feasibility of predicting treatment response in locally advanced rectal cancer (LARC) patients undergoing neoadjuvant chemoradiation therapy (nCRT) using MRI-based pre-, post-, and delta-radiomic features. “The study included 53 patients with LARC, with data split into a training set (36 patients) and an external validation set (17 patients). T2-weighted MRI scans were acquired before and after nCRT, and 96 radiomic features were extracted and harmonized. Various machine learning algorithms, including k-nearest neighbors (KNN), Naïve Bayes (NB), Random Forests (RF), and extreme Gradient Boosting (XGB), have been used for classification. The highest AUC values were achieved with the delta-radiomic-based RF model (0.96 ± 0.01) and NB (0.96 ± 0.04), outperforming pre- and post-treatment features (P-value <.05). The study concluded that delta-radiomic features analyzed with RF classifiers are promising biomarkers for predicting treatment response in patients with LARC. 83 ”
Pizzi et al. 84 developed a machine learning model to predict treatment response in patients with locally advanced rectal cancer (LARC) undergoing neoadjuvant chemoradiotherapy (CRT) followed by total mesorectal excision (TME). “The study utilized pretreatment T2-weighted MRI scans from 72 patients with LARC, with tumors segmented by 2 independent readers. Radiomic features were extracted from both the tumor core (TC) and tumor border (TB). A Partial Least Squares (PLS) regression model was employed, with leave-one-out nested cross-validation used for hyperparameter optimization. The model achieved an AUC of 0.793 (P = 5.6 × 10−5), demonstrating improved predictive performance when combining clinical and radiomic features.” This study highlights the potential of integrating MRI-based clinical and radiomic features for early prediction of treatment response and suggests the need for prospective validation in clinical trials. 84
Zhou et al. 85 developed a deep learning model named CRCNet for the optical diagnosis of colorectal cancer (CRC) using 464 105 images from 12 179 patients. “The model was tested on 2263 patients from 3 independent datasets. At the patient level, CRCNet achieved high performance with an area under the precision-recall curve (AUPRC) ranging from 0.867 to 0.882, outperforming average endoscopists in terms of recall rate and precision in most test sets. At the image level, CRCNet demonstrated exceptional accuracy, with AUPRC values between 0.990 and 0.997.” These results suggest the potential of CRCNet to improve CRC screening, warranting further investigation through prospective clinical trials. 85
Grosu et al. 86 investigated the use of machine learning to differentiate between benign and premalignant colorectal polyps detected using CT colonography. “They used radiomics to extract 1906 features from segmented polyps and applied a random forest classification algorithm. The model was trained on 107 polyps and validated using an external test set of 77 polyps. The random forest model achieved an area under the receiver operating characteristic curve (AUC) of 0.91, with a sensitivity of 82% and a specificity of 85%. A study demonstrated that machine learning can effectively distinguish between benign and premalignant polyps noninvasively, showing promise for improving colorectal cancer screening. 86 ”
Talukder et al. 56 developed a hybrid ensemble feature extraction model for cancer detection, focusing on lung and colon cancers. “Their model integrates deep learning for feature extraction with ensemble learning and high-performance filtering and is tested on LC25000 histopathological datasets for lung and colon cancers. The hybrid model achieved accuracy rates of 99.05% for lung cancer, 100% for colon cancer, and 99.30% for combined lung and colon cancer.” The results of this study indicate that this model significantly outperforms the existing methods and could be a valuable tool in clinical settings to assist in cancer diagnosis. 56
Sirinukunwattana et al. 87 developed an image-based method, imCMS, to predict colorectal cancer (CRC) consensus molecular subtypes (CMS) from standard H&E-stained tissue sections using deep learning. “The method was trained and tested on data from 3 independent datasets: FOCUS trial, GRAMPIAN cohort, and The Cancer Genome Atlas (TCGA). imCMS achieved high classification accuracy with AUCs of 0.84 and 0.85 in unseen datasets. It effectively resolved intratumoral heterogeneity, correlated genomic and epigenetic alterations, and matched the prognostic associations of transcriptomic CMS. This approach offers a cost-effective alternative for biological stratification in routine clinical workflows. 87 ”
Wulczyn et al. 88 developed a deep learning system (DLS) to predict the 5-year disease-specific survival in stage II and III colorectal cancer using 27 300 histopathology slides from 3652 cases. “The DLS achieved AUCs of 0.70 and 0.69 on validation datasets, adding significant predictive value to 9 clinicopathologic features. To enhance interpretability, the study explored human-interpretable features and found that traditional clinicopathological factors (T-category, N-category, grade) explained only 18% of the variance in DLS scores. In contrast, histological features derived from clustering deep learning embeddings explained 73%-80% of the variance. A specific clustering-derived feature, associated with poorly differentiated tumor cell clusters near adipose tissue, was highly prognostic and identifiable by annotators with 87.0%-95.5% accuracy.” This approach provides insight into DLS predictions and identifies potentially novel prognostic features for further validation. 88
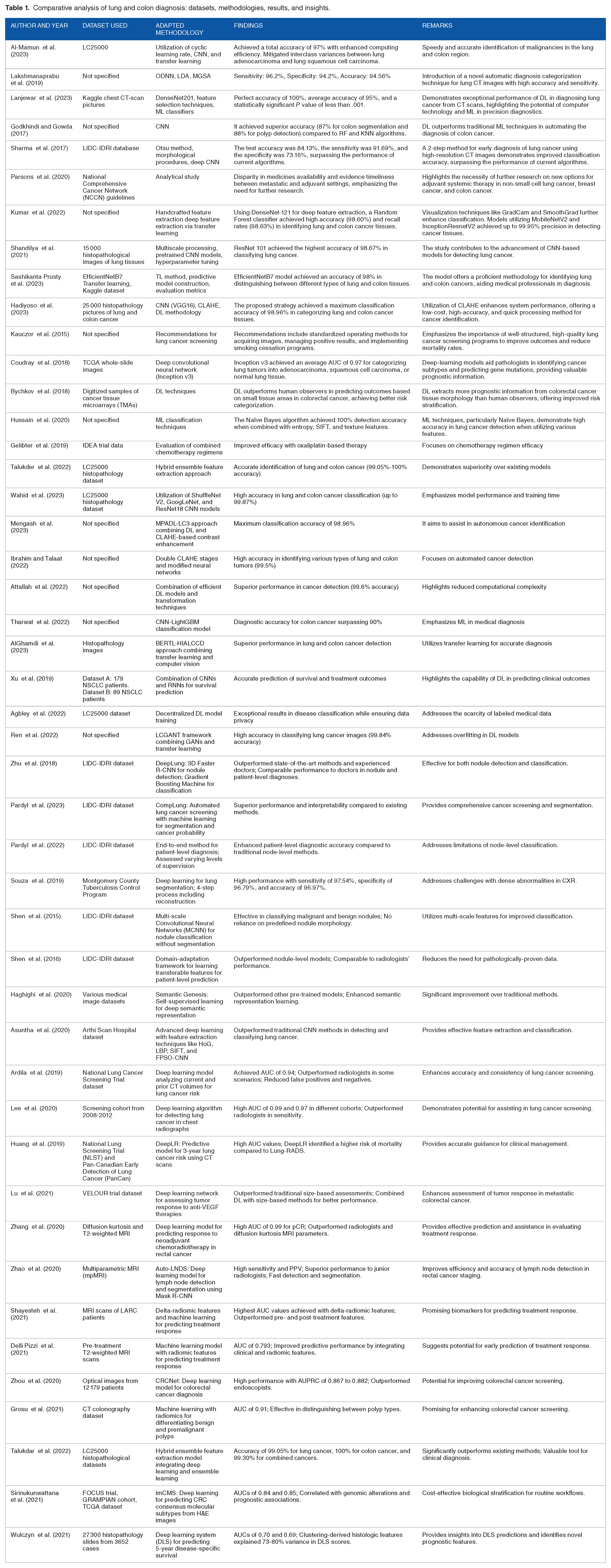
Table 1 encompasses a range of studies focusing on the detection and diagnosis of both lung and colon cancer using ML and DL techniques. These studies have utilized various datasets and methodologies to address the challenges of identifying these 2 types of cancer. The findings demonstrate the efficacy of DL models in accurately diagnosing cancerous tissues, with reported accuracy rates ranging from 88.26% to 100% across different studies. The results underscore the potential of ML algorithms, such as convolutional neural networks (CNNs) and transfer learning, to enhance the precision and efficiency of cancer diagnosis for both lung and colon cancer. However, several limitations were noted, including scarcity of labeled data and risk of overfitting. Despite these challenges, studies have collectively highlighted the significance of leveraging artificial intelligence to improve early detection and ultimately enhance patient outcomes in managing lung and colon cancers.
Comparative analysis of lung and colon diagnosis: datasets, methodologies, results, and insights.
Challenges and Opportunities in Lung and Colon Cancer Imaging Research
Research on lung and colon cancer imaging faces both challenges. Challenges include the complexity of tumor morphology and heterogeneity, making accurate diagnosis and classification difficult. In addition, the availability of labeled data for training ML models remains limited, hindering the development of robust algorithms. Moreover, overfitting and generalizability pose significant obstacles in translating research findings into clinical practice. However, there are several promising opportunities in this field. Advances in imaging technologies such as high-resolution CT scans and hyperspectral imaging offer detailed insights into tumor characteristics and microenvironments. Furthermore, integrating ML and DL techniques has great potential for improving the diagnostic accuracy and personalized treatment strategies. Collaborative efforts to collect and share annotated imaging datasets and the development of innovative algorithms can address some of the current challenges and pave the way for more effective imaging-based approaches for managing lung and colon cancer. The achievements in diagnosing lung and colon cancers using computer intelligence from 2017 to 2023 are listed in Table 2.
Achievement toward lung and colon cancer diagnosis using computer intelligence.

Table 2 illustrates the significant advancements in computer intelligence for the diagnosis of lung and colon cancer. In 2017, while the specific accuracy for diagnosing lung cancer was not provided, there was a notable achievement of 87% accuracy in diagnosing colon cancer. Subsequent years saw varying levels of progress, with 2019 marking a milestone with high accuracies of 94.56% for colon cancer and 95% for both lung and colon cancer. The trend continues to improve, particularly in lung cancer diagnosis, reaching an impressive accuracy of 98.67% in 2021 and a range of 99.05%-100% in 2022. Although the specific accuracy for colon cancer was not provided for 2023, a substantial achievement of 97% accuracy was reported for lung cancer, with a remarkable 99.87% accuracy for both lung and colon cancers. Overall, the consistent enhancement in accuracy underscores the growing potential of computer intelligence in facilitating the early and precise detection of lung and colon cancers, offering promising prospects for improved patient outcomes and healthcare management.
Specific studies and cases provide context and make the information more tangible on challenges and opportunities in lung and colon cancer imaging research.
Examples of the major challenges are as follows:
A study by Gerlinger et al. 89 highlighted the significant intratumour heterogeneity in lung cancer, which complicates accurate diagnosis and classification. Researchers have found that different regions of the same tumor can have distinct genetic mutations, making it challenging to develop uniform diagnostic criteria. 89
Litjens et al. 90 emphasized the scarcity of large annotated datasets for training machine-learning models in medical imaging. This limitation hinders the development of robust algorithms and their subsequent validation across diverse patient populations. 90
Yamashita et al. 91 demonstrated that deep learning models trained on limited datasets often exhibit overfitting, reducing their ability to generalize to new, unseen data. This issue is particularly evident in lung cancer imaging, where the variability in tumor appearance can lead to model performance degradation in clinical settings. 91
Specific examples of opportunities.
• Bejnordi et al. 92 explored the use of high-resolution CT and hyperspectral imaging for lung cancer detection. Advanced imaging techniques provide detailed insights into tumor characteristics and microenvironments, facilitating more accurate diagnosis and treatment planning. 92
• Esteva et al. 93 demonstrated the potential of deep learning algorithms in improving diagnostic accuracy for various cancers, including colon cancer. Their study showed that deep learning models could achieve performance comparable to that of dermatologists in diagnosing skin cancer, suggesting similar potential for lung and colon cancers. 93
• The Cancer Imaging Archive (TCIA) is instrumental in providing publicly available annotated imaging datasets. Clark et al. 94 highlighted the impact of TCIA in facilitating collaborative research and the development of innovative algorithms for cancer diagnosis, including lung and colon cancers. 94
• Kather et al. 95 demonstrated the use of deep learning algorithms to predict microsatellite instability in colorectal cancer using histopathological images. This innovation could lead to personalized treatment strategies and better patient outcomes. 95
Summary of Key Findings and Insights from the Review
Future research on image-based lung and colon cancer diagnosis holds immense promise in advancing early detection, treatment planning, and patient outcomes. One direction is the integration of multimodal imaging techniques, combining data from various imaging modalities such as CT, MRI, PET, and histopathology. This integration can provide comprehensive information regarding tumor morphology, metabolism, and microenvironment, thereby enhancing diagnostic accuracy and treatment decision-making. The development of artificial intelligence (AI) algorithms and DL models tailored for lung and colon cancer diagnosis will also remain a focal point. These models can analyze large-scale imaging datasets with high accuracy, assisting radiologists and pathologists in interpreting complex images and identifying subtle features indicative of malignancy. Furthermore, there is a growing interest in leveraging radiomic and radiogenomic approaches to extract quantitative imaging biomarkers associated with tumor phenotypes, genotypes, and treatment responses. Integrating these biomarkers into clinical practice can enable personalized medical strategies and facilitate prognostication. Moreover, as imaging technology evolves, advancements in image acquisition techniques such as higher spatial and temporal resolutions and novel contrast agents will further enhance the sensitivity and specificity of cancer detection. Finally, efforts to standardize imaging protocols, establish robust quality assurance measures, and promote data sharing and collaboration across institutions will ensure the reproducibility and generalizability of the research findings, ultimately translating into improved patient care and outcomes.
In conclusion, image-based diagnosis powered by cutting-edge technologies, such as artificial intelligence and multimodal imaging, has the potential to revolutionize cancer care and significantly improve patient outcomes. By enabling the earlier detection of lung and colon cancers, these advanced imaging techniques can facilitate timely interventions, leading to better treatment responses and increased survival rates. Moreover, the integration of quantitative imaging biomarkers and radiomic/radiogenomic approaches is promising for use in the era of personalized medicine, where treatment strategies can be tailored to individual patient characteristics and therapeutic efficacy can be optimized while minimizing adverse effects. Furthermore, the standardization of imaging protocols and establishment of quality assurance measures will ensure consistency and reliability across diagnostic procedures, enhancing the accuracy and reproducibility of the results. Overall, the continued advancement and adoption of image-based diagnosis holds great promise for transforming the landscape of cancer care, offering hope for improved patient care and outcomes in the future.
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was not supported by any funding agencies.
Declaration Of Conflicting Interests:
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: On behalf of all authors, the corresponding author states that there are no conflicts of interest.
Abbreviations
CT Computer tomography
MRI Magnetic resonance imaging
PET Positron emission tomography
ML Machine learning
DL DL
AI Artificial intelligence
NSCLC Non-small cell lung cancer
SCLC Small cell lung cancer
CAD Computer aid diagnosis
LDC Lung adenocarcinoma
LSCC Lung squamous cell carcinoma
DNN Deep neural network
ODNN Optimized deep neural network
MGSA Modified gradient search algorithm
MCC Mathew’s correlation coefficient
KS Kappa score
AJCC American joint committee on cancer
RF Random forest
KNN K-nearest neighbors
SVM Support vector machine
CNN Convolutional neural network
NCCN National comprehensive cancer network
GB Gradient boosting
MLP Multilayer perceptron
LDCT Low- dose computed tomography
ESR European society of Radiology
ERS European respiratory society
AUC Area under the curve
TCGA The cancer genome cancer
SIFT Scale invariant Fourier transform
EFDs Ellipse Fourier descriptors
RBF Radial base function
DT Decision tree
IDEA International duration evaluation of adjuvant treatment
PCA Principal component analysis
FHWT Fast Walsh Hadamard transform
DWT Discrete wavelet transforms
HPI Histopathological imaging
TL Transfer learning
DL DL
DCRNN Deep convolutional recurrent neural network
GAN Generative adversarial network
LDA linear discriminant analysis
CLAHE contrast limited adaptive histogram equalization
DBNs deep belief networks
LCC lymphocyte-to-carcinoma cell
RNN Recurrent neural network
DC-GAN Deep convolutional Generative adversarial network
Data Availability
No data were associated with this study.
Ethical Approval
Ethical approval was not required for this study.
Consent for Participation
Consent for participation was not needed.
Code Availability
Not Required
Consent for Publication
Not applicable.
