Abstract
The recombination plays a key role in promoting evolution of RNA viruses and emergence of potentially epidemic variants. Some studies investigated the recombination occurrence among SARS-CoV-2, without exploring its impact on virus-host interaction. In the aim to investigate the burden of recombination in terms of frequency and distribution, the occurrence of recombination was first explored in 44 230 Omicron sequences among BQ subvariants and the under investigation “ML” (Multiple Lineages) denoted sequences, using 3seq software. Second, the recombination impact on interaction between the Spike protein and ACE2 receptor as well as neutralizing antibodies (nAbs), was analyzed using docking tools. Recombination was detected in 56.91% and 82.20% of BQ and ML strains, respectively. It took place mainly in spike and ORF1a genes. For BQ recombinant strains, the docking analysis showed that the spike interacted strongly with ACE2 and weakly with nAbs. The mutations S373P, S375F and T376A constitute a residue network that enhances the RBD interaction with ACE2. Thirteen mutations in RBD (S373P, S375F, T376A, D405N, R408S, K417N, N440K, S477N, P494S, Q498R, N501Y, and Y505H) and NTD (Y240H) seem to be implicated in immune evasion of recombinants by altering spike interaction with nAbs. In conclusion, this “in silico” study demonstrated that the recombination mechanism is frequent among Omicron BQ and ML variants. It highlights new key mutations, that potentially implicated in enhancement of spike binding to ACE2 (F376A) and escape from nAbs (RBD: F376A, D405N, R408S, N440K, S477N, P494S, and Y505H; NTD: Y240H). Our findings present considerable insights for the elaboration of effective prophylaxis and therapeutic strategies against future SARS-CoV-2 waves.
Background
In a short span of time, the Coronavirus-related Severe Acute Respiratory Syndrome-2 (SARS-CoV-2) has drastically affected the worldwide healthcare system and economy.1,2 SARS-CoV-2 is an RNA-positive virus, included in genus Betacoronavirus, subgenus Sarbecovirus, and family Coronaviridae. 3 Its genome is approximately 30 kb in size and consists of untranslated regions (3′UTR and 5′UTR) and 14 Open Reading Frames (ORFs), encoding a total of 31 proteins.4,5 The Spike protein is essential for virus entry and is considered as the most important target for vaccine and neutralizing antibodies. Structurally, two subunits S1 and S2 form the spike. The S1 comprises the N-terminal domain (NTD, 14-317) and the receptor binding domain (RBD, 318-541). 6 The receptor binding motif (RBM, 438-506) is the RBD region that binds directly to the host cell’s receptor ACE2. 7 There are four classes of neutralizing antibodies (nAbs) that recognize different antigenic sites on Spike and are able to abolish the interaction between the RBD and ACE2. 8 Other anti-S neutralizing antibodies have been described and identified to neutralize the SARS-CoV-2 through binding to NTD. 9
Since 2020, variants have been emerged and posed challenges for pandemic control: VOCs (variants of concern), VOIs (variants of interest) and VUMs (Variants under investigation). 10 Five VOCs have been described; the Alpha variant (B.1.1.7 lineage), the Beta variant (B.1.351 lineage), the Gamma variant (P.1 lineage), the Delta variant (B.1.617.2 lineage) and the Omicron variant (B.1.1.529 lineage).11 -13 Since August 2022, the evolutionary process has led to the emergence of new Omicron subvariants BQ.1 (BA.5.3.1.1.1.1.1) and BQ.1.1 (BA.5.3.1.1.1.1.1.1) which dominated in the USA, UK, and several other European countries during the winter 2022 to 2023. 14 Numerous other descendent BQ subvariants (BQ.1.2, BQ.1.12, BQ.1.1.4, BQ.1.3, BQ.1.1.5. . .) have been expanded (Outbtreak.info, accessed in 13th March, 2024), possibly due to improved antibody evasion properties arising from their additional spike mutations and possible recombination.15 -18
RNA recombination driven by RNA polymerase plays an important role in the evolution of SARS-Cov-2. 19 The introduction of new genetic fragments can lead to the generation of new viral strains and confer some advantages in terms of transmissibility of infection and evasion to immune response.20 -23 SARS-CoV-2 recombinants have been registered under the names XA, XB, XC, XD. . . and detected in various countries around the world. XA was the first recombinant lineage detected in the UK between B.1.1.7 and B.1.177. After that, XB and XC variants were detected in the USA and Japan respectively.24,25 Subsequently, other recombinant lineages (XD, XF, XS. . .) have been recognized in different parts of the world.23,26 The majority of them have resulted from recombination between Omicron’s BA.1 and BA.2 lineages particularly in the spike gene. Potential recombinants between Omicron and Delta variants have also been reported in France and Denmark (XD), the UK (XF) and the USA (XS). 27 Numerous sequences submitted to the GISAID database (Global Initiative on Sharing Avian Influenza Data) (https://gisaid.org/) requires further investigation and denoted as “ML,” corresponding to “the presence of markers of multiple lineages from both Delta and Omicron variants.” Such sequences may result from the occurrence of recombination events.
Continuous investigation of SARS-CoV-2 recombinant is crucial to track variant emergence and detect strains with increased potential of transmissibility and virulence that could pose a risk to global health. To the best of our knowledge, few studies have investigated the status of recombinant SARS-CoV-2 lineages, their rates and distribution.28 -31 Turkahia et al 28 focused only on SARS-CoV-2 genomes obtained during May 2021 and revealed the existence of approximately 2.7% of recombinant sequences. The study by Shiraz and Tripathi 29 investigated larger number of sequences collected between November 2019 and July 2022 but reported only the prevalence of recombinant strains. In the other hand, Tamura et al 30 examined the virological characteristics of only one recombinant SARS-CoV-2 variant (XBB) on the basis of lab experimentation. More recently, Pipek et al 31 investigated occurrence of co-infection and revealed also existence of recombination hotspots of Delta–Omicron BA.1 intra-host recombinants. Gaps remain especially on continuous recombinant strains tracking as well as the understanding of the impact of recombination on virus-host interaction, immune evasion, pathogenicity and also vaccine and treatment efficacy.
In this study, we are interested in identifying recombinant strains and exploring their potential impact on virus host-interaction and possible immune evasion, among Omicron sequences from BQ subvariants and the under investigation “ML” sequences. The rates, types and geographic distribution of recombinant strains were investigated. To explore the virus host interaction and potential immune evasion of recombinants, we focused on the interaction between spike protein and its receptor ACE2 as well as interaction with the neutralizing antibodies. Analysis of recombinant mutational profile was also performed.
Materials and Methods
Investigated complete SARS-CoV-2 genome sequences
All available complete SARS-CoV-2 genome sequences (n = 44 230) belonging to the Omicron BQ subvariants (n = 43 749) and ML denoted sequences (n = 484), were extracted from GISAID database (Global Initiative on Sharing All Influenza Data) 32 and the NCBI SARS-CoV-2 Resources (National Center for Biotechnology Information: https://www.ncbi.nlm.nih.gov/sars-cov-2/). They were obtained from samples collected during the period July 2022 and January 2023 and corresponded to all available ML and BQ subvariant sequences in the GISAID database (accessed on January 31, 2023) and the NCBI SARS-CoV-2 database (accessed on January 31, 2023), respectively, presenting less than 10% of ambiguous nucleotides (N).
Recombination analysis
Genomes of first sequenced strain of SARS-CoV-2 Wuhan Hu-1 as well as first detected VOCs sequences (Alpha B.1.1.7, Delta B.1.617.2, Gamma P.1 and Omicron B.1.1.529) were retrieved from NCBI https://www.ncbi.nlm.nih.gov/sars-cov-2/) as potential parents for recombinant strains (Supplemental File 1). Sequences of BA.4, BA.4.1, BA.5, BA.5.1, BA.5.2, and BA.5.2.1, were predominant during the emergence of the investigated ML and BQ subvariant as reported by Nextstrain database, accessed on January 17, 2023. Consequently, they were also retrieved from NCBI https://www.ncbi.nlm.nih.gov/sars-cov-2/) and used as potential parents for recombinant strains (Supplemental File 1). Their first strains detected on each continent (America, Africa, Asia, and Europe) were selected. Genomes with more than 1% of ambiguous nucleotides (N) were avoided.
All sequences of parental lineages as well as the investigated recombinants were aligned using MAFFT version 7 (Multiple Alignment using Fast Fourier Transform).33,34 To detect recombination events, the resulted fasta file of multiple sequence alignment was used as an input for 3seq program, installed in linux system and run as described in the 3seq manual. 35 Recombinant triplets with corrected P-value < .05, according to Dunn-Sidak method, were recorded and considered significant.
Statistical investigation
Statistical analysis of geographic distribution, recombinant type frequency and recombinant region frequency was carried out using dplyr package (https://www.rdocumentation.org/packages/dplyr/versions/0.7.8). The mapping of geographic distribution of recombinants was performed with R package leaflet software (https://cran.r-project.org/web/packages/leaflet/index.html). The code used to generate the plots is shared on the following link: https://github.com/souiai/maps.
Molecular docking
The molecular docking concerned strains with recombination into the spike gene. Given the large number of obtained recombinant sequences only representative sequences were considered for docking.
For each subvariant, the NTD and RBD domain sequences were first extracted using the extractseq tool of the EMBOSS package (http://emboss.open-bio.org/wiki/Appdocs), then aligned and trees were generated based on Neighbor Joining method using ClustalW. 36 Only one representative sequence from the major clusters obtained in the recombinant and non-recombinant trees were taken into account for the molecular docking.
The docking of both domains was achieved against the cellular receptor (ACE2) and 5 different neutralizing antibodies. The Receptor Binding Domain (RBD) was docked against the ACE2 and 4 antibodies (C105, LY-CoV555, S309, and C118) representative of each neutralizing antibodies class. 8 The N-Terminal Domain (NTD) was docked against the unique known neutralizing antibody 4A8. 37
The Spike 3D models were built using the SWISS-MODEL automated server (https://swissmodel.expasy.org/). The ACE2 structure (PDB ID: 6M0J) and neutralizing antibodies (PDB ID: 7C2L, 6XCM, 7KMG, 6WPS, 7RKS) were obtained from RCSB PDB (https://www.rcsb.org/). 3D structures were modified and visualized using PyMOL 0.99rc6 (https://pymol.org/). Molecular docking was performed using Galaxy Tong Dock A server (https://galaxy.seoklab.org/). The generated complexes (RBD-ACE2, RBD-C105, RBD-LYCoV555, RBD-S309, RBD-C118, and NTD-4A8) with the highest docking scores and cluster size were submitted to PRODIGY (PROtein binding enerGY) (https://wenmr.science.uu.nl/prodigy/) to determine the standard binding free energy (ΔG°) and dissociation constant (Kd) at physiological temperature (37°C). The interaction zones between the Spike domains (NTD and RBD) and the cellular receptor as well as the neutralizing antibodies were visualized and determined using PyMOL 0.99rc6 (Interaction zone at an atomic distance of 4 Å).
The validation of docking results was performed by re-docking NTD and RBD against the cell receptor ACE2 and the nAb using PyDock webserver (https://life.bsc.es/pid/pydockweb). Out of the top 10 of docking models, 1 model, fitting the structural conformation of the resulted models of Galaxy Tong Dock A docking, was selected. Both models were aligned using PyMOL 0.99rc6 and the displayed Root-mean square deviation (RMSD) values (Å) were used to evaluate their similarity. Models, expressing RMSD < 3 Å, were validated. 38 Models, expressing RMSD ⩾ 3 Å were discarded from our analysis.
Mutation analysis
Amino acid sequences of NTD and RBD regions of recombinant strains were aligned with Wuhan Hu-1 reference sequence and the non-recombinant sequences using ClustalW. The multiple sequence alignments were visualized on Bioedit program v.7.0.9.0. Only amino acid substitutions, within the interaction zone of RBD and NTD with ACE2 and the neutralizing antibodies, resulted from docking, were considered. Pymol v0.99 program was used to visualize and characterize mutations in the interaction zone.
Results
Geographic distribution of recombinants
The geographic distribution of recombinant subvariants was analyzed during the studied period to identify the hotspot locations and explore the diversity of recombinant lineages in these regions. The geographical map presented in Figure 1 showed a worldwide distribution of recombinant BQ and ML sequences, detected in 34 countries (Figure 1 and Supplemental File 2). America, Europe and Asia constituted hotspots for recombinant strains’ circulation. Recombinants were mostly detected in the USA that showed circulation of 94.98% of the detected recombinant strains while other countries such as the UK and Australia showed lower rates, 3.06% and 0.63% respectively (Supplemental File 2).

Global geographic distribution of recombinants of SARS-CoV-2 BQ and ML subvariants. The pie chart in each country represents the distribution of recombinant lineage sequences collected from that country, with different colors indicating different recombinant lineages.
An important Recombinant subvariant diversity was revealed. In the European countries, twenty three different recombinant lineages were detected and among them fifteen (XAY.3, XAY.2, XBB.1, XBC.1, XBC, XBC.1.1, XAY.1, XB.1, XBC.2, XE, XAW, XAY.1.1, XBB.2.2, XBB.2, and recombinant BQ.1) were predominant (frequency ⩾ 50%). The Asian countries showed circulation of eleven recombinant lineages among them six were predominant: XBC, XBC.1, XBB.1.1, and XBB.1.2 and recombinant BQ.1 and BQ.1.1. In the American countries showed circulation of 7 different recombinant subvariants and the most frequent were XAY.3 (50%) and recombinant BQ.1.1 (40%).
Considering recombinant subvariant diversity per country, 7 to 3 different subvariants, were observed in European and North American countries (Austria, France, Netherlands, Germany, Sweden, Switzerland, USA, and Canada). In Asian countries (India, Philippines, Japan, South Korea), 3 to 4 different subvariants were detected.
Recombinant type frequency and recombinant parental lineages
Using the 3seq program analysis, 25 284 among a total of 44 218 (57.18%) investigated sequences showed possible occurrence of recombination event supported by a statistical P-values (P < .05; corrected by Dunn-Sidak method) (Supplemental File 3). The recombinant sequences included 24896 BQ sequences (56.91% of BQ sequences) and 388 ML sequences (82.20% of ML sequences).
The majority of recombinant BQ.1, BQ.1.11, BQ.1.3, BQ.1.10, BQ.1.12, and BQ.1.1.4 (>93%) showed occurrence of recombination with deltacron XAW or XBC.1 (Figure S1 and Supplemental File 2). More than 80% of recombinant BQ.1.1, BQ.1.2, BQ.1.1.3, and BQ.1.1.5 exhibited double recombination events with Wuhan Hu-1 and XAW as shown in Figure S1 and Supplemental File 2. Other variants (Alpha (B.1.1.7), Delta (B.1.617.2), and Omicron (B.1.1.529, BA.5.2, and BA.5.2.1) were involved in the recombination events with a very low rates (<20%) (Figure S1).
Regarding ML subvariants, strains were double recombinants with XBB.1.5 and XBB.2 (53.61%), XBB.1.5, and XBB.1.2 (56.39%) (Figure S1).
Investigation of recombinant regions
Recombinant regions were identified in the BQ and ML genomes on the basis of breakpoints positions as highlighted by the 3seq program analysis (Supplemental File 3). To determine the hostspot genomic regions of recombination, we calculated the frequency of recombination for each recombinant region in the genome. For BQ subvariants, a total of 49 827 recombination regions were identified, and the recombination hotspot regions are mainly localized in the 3′UTR (25.87%), Spike (23.24%), and Matrice M (18.78%) followed by 5′UTR (13.09%) (Figure 2A). Recombination targets the Spike genomic region for most of the BQ subvariants, notably, BQ.1.1, BQ.1.2, BQ.1.10, BQ.1.12, BQ.1.1.3, and BQ.1.1.5. The highest recombination frequencies into the Spike region are recorded for BQ.1.10 and BQ.1.12 with 97.06% and 95.92%, respectively. Among ML subvariants, recombination events (n = 776) occurred in 3′UTR, E, M, and NSP regions. It affected mainly the ORF1a region: NSP9/NSP10 (26.38%), NSP6/NSP7 (25.74%), NSP8 (21.36%), and NSP7/NSP8 (21.23%) (Figure 2B). Most of ML subvariants show recombination in the NSP6/NSP7 and NSP9/NSP10 regions and the highest recombination frequency is 50%, detected among XE subvariant.

Recombinant region distribution for SARS-CoV-2 BQ and ML subvariants: (A) recombination hotspot identification in genomic regions of BQ subvariants and (B) recombination hotspot identification in genomic regions of ML subvariants. Only the genomic regions presenting 5% of recombination events were displayed among BQ and ML subvariants.
Identification and mapping of the recombination region in the spike
Using the 3seq program, the occurrence of recombination in the spike region among the investigated sequences were only detected in sequences from BQ subvariants (BQ.1.1, BQ.1.2, BQ.1.10, BQ.1.12, BQ.1.1.3, and BQ.1.1.5), with a significant P-value > .05 (Supplemental File 3). BQ.1.1 and BQ.1.1.3 sequences showed the occurrence of one recombination event. The recombination breakpoints were located at the positions 22 354 to 22 856nt (261-428aa) and 22 359 to 22 653nt (262-361aa) respectively. The mapping of recombination regions in the Spike protein showed that they are located in the NTD-RBD junction (Figure 3).

Mapping of recombination regions in Spike protein for major groups of SARS-CoV-2 BQ recombinants BQ.1.1, BQ.1.2, BQ.1.10, BQ.1.12, BQ.1.1.3, and BQ.1.1.5.
For BQ.1.2, BQ.1.10, BQ.1.12, and BQ.1.1.5, sequences harbored two recombination regions located between positions 22 359 and 22 868nt (236-431aa) for the first recombination region and 23 140 to 23 620nt (438-658aa) for the second one (Table S1). The first recombination region takes place in the NTD-RBD junction and the second one is located in the RBD region. We also noted that BQ.1.2 has the largest recombination regions (Figure 3).
Docking analysis of the spike against the cellular receptor ACE2 and the neutralizing antibodies
For all 6 BQ subvariants (BQ.1.1, BQ.1.2, BQ.1.10, BQ.1.12, BQ.1.1.3, and BQ.1.1.5), the docking analysis focus on the recombinant spike region. The binding affinity of the major recombinant group was compared with the main non-recombinant group previously identified by clustering analysis for each of the 6 BQ subvariants (Supplemental File 4).
Docking analysis of the RBD region of the spike protein against the cellular receptor ACE2
Analysis of standard binding free energy and dissociation constant for RBD-ACE2 interaction
The effect of recombination on the binding activity of Spike to the ACE2 receptor was investigated for by comparing the standard binding free energy (ΔG°) as well as the dissociation constant (Kd) of the RBD-ACE2 docking complex for BQ recombinants and non-recombinants.
The docking results showed a decreased ΔG° and Kd values for BQ.1.2 recombinant (Recombinant: ΔG° = −23.0 kcal/mol/Kd = 5.8 × 10−17 M; Non-recombinant: ΔG° = −21.5 kcal/mol/Kd = 6.5 × 10−16 M) indicating a strong interaction of RBD with ACE2 (Figure 4A). For recombinant subvariants BQ.1.1, BQ.1.15 BQ.1.10, BQ.1.12, and BQ.1.13, the recombination has no effect on RBD-ACE2 interaction where a similar or high ΔG° and Kd values were obtained compared to those reported for non-recombinant strains (Figure 4A).

Docking analysis of interaction between the RBD and the cellular receptor ACE2 of SARS-CoV-2 BQ recombinants and non-recombinants: (A) standard binding free energy ΔG° (kcal/mol) and dissociation constant Kd (M) (in parenthesis) of RBD-ACE2 interaction for recombinant (R) and non-recombinant (NR) of BQ subvariants, (B) interaction between RBD of BQ.1.2 recombinant and ACE2 and (C) interaction between RBD of BQ.1.2 non-recombinant and ACE2 (PDB ID: 6M0J). Zoomed-in figures show the interaction zone (IZ).
Investigation of interaction zone between RBD and ACE2 cell receptor
In continuation, this analysis focused only on the BQ.1.2 subvariant that presented increased affinity to ACE2. Structural analysis of the RBD-ACE2 complex showed that the RBD surface of the recombinant is more implicated in the interaction with the ACE2 receptor than the RBD surface of non-recombinant (Figure 4A). The zoomed-in figure of RBD-ACE2 interaction zone revealed a larger region of the non-RBM (318-436aa of RBD) engaged in the RBD attachment to ACE2 (Figure 4B). However, the non-recombinant strain has a reduced engaged non-RBM region (Figure 4B).
Docking analysis of the RBD region of the spike protein against the neutralizing antibodies
Analysis of standard binding free energy and dissociation constant for RBD-neutralizing antibodies interaction
The RBD avidity of recombinant and non-recombinant subvariants was assessed against the neutralizing antibodies (nAbs) C105, LYCoV555, S309, and C118 (Figure 5). The results revealed increased ΔG° and Kd values for BQ.1.12 and BQ.1.1.5 recombinants against two nAbs (C105 and LY-CoV555) reflecting a weak RBD-nAbs interaction. In addition, recombinants BQ.1.10 and BQ.1.1 presented increased ΔG° and Kd values with two antibodies: C105 and C118 for BQ 1.10 and LY-CoV555 and C118 for BQ.1.1. For the other investigated subvariants BQ.1.2 and BQ.1.1.3, similar or low ΔG° and Kd values were obtained for recombinant and non-recombinant strains, showing that there is no effect of recombination on RBD-nAbs interaction (Figure 5).
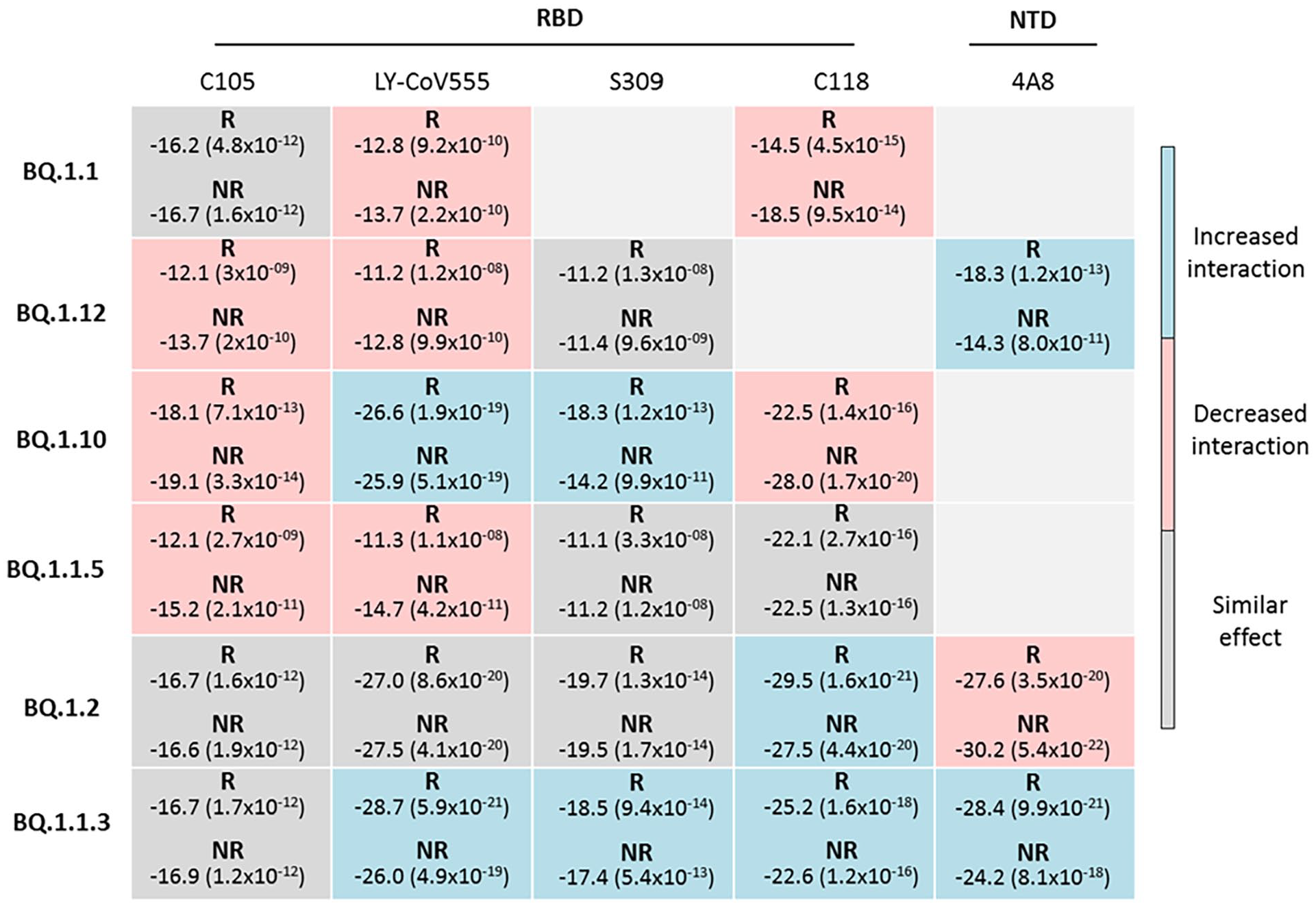
Standard binding free energy ΔG° ( kcal/mol) and dissociation constant Kd (M) (in parenthesis) of RBD and NTD interaction with the neutralizing antibodies C105, LY-CoV555, S309, C118, and 4A8 for recombinant (R) and non-recombinant (NR) of BQ SARS-CoV-2 subvariants. For NTD of BQ.1.1.5, comparison between recombinant and non-recombinant has not been performed as a main phylogenetic cluster of non-recombinants was absent. Docking models of BQ.1.1-S309, BQ.1.12-C118, BQ.1.1-4A8, and BQ.1.10-4A8 were discarded (RSMD ⩾ 3 Å).
Analysis of standard binding free energy and dissociation constant for the NTD region against the neutralizing antibody 4A8
For BQ.1.2 recombinant, an increased in ΔG° and Kd values was observed, resulting in a reduced binding affinity of NTD against the neutralizing antibody 4A8. For the remaining subvariants BQ.1.12, BQ.1.1.3, and BQ.1.1.5, similar or low ΔG° and Kd values against 4A8 were recorded resulting in no changes or enhancement of the NTD-4A8 interaction (Figure 5).
Interaction between RBD and NTD with neutralizing antibodies
With regard to RBD-nAbs (neutralizing antibodies) interaction, docking analysis showed a reduced interaction of BQ.1.1 recombinant with the light chain (LC) of the nAb C118. The zoomed figure reveals a limited involvement of the non-RBM region with the C118 interaction zone (Figure 6). With LY-CoV555, similar configurations were observed between recombinant and non-recombinants strains (Figure S2).

Interaction of RBD with the neutralizing antibodies C105, LY-CoV555, and C118 for recombinant (R) and non-recombinant (NR) strains of SARS-CoV-2 BQ subvariants. Zoomed-in figures present the interaction zone.
For BQ.1.12 recombinant, the RBD region presented an altered interaction with the light chain of the nAb C105. Reduced surfaces of RBM and LC with absence of non-RBM region were noted in the zoomed-in figure of the RBD-C105 interaction zone (Figure 6). Furthermore, the zoomed-in figure of RBD-LYCoV555 complex revealed that the non-RBM region was slightly reduced (Figure 6).
For BQ.1.10 recombinant, Figure 6 shows the dissociation of C118 light chain from the RBD. In the zoomed-in figure, less engagement of the non-RBM region was observed in the zone of C118 interaction (Figure 6). With C105, similar interactions were observed between recombinant and non-recombinants strains (Figure S2).
For BQ.1.1.5 recombinant, a reduced LY-CoV555 light chain interacting with the RBD was noticed in the zoomed-in figure. The results also showed reduction of the non-RBM region. The zoomed-in figures of the interaction zones revealed the absence of the non-RBM region in the interaction zone of RBD-C105 complex (Figure 6).
With regard to the NTD-4A8 interaction, the alteration of LC and HC was clearly noticed among BQ.1.2 recombinants. A reduced interaction with the loop N5 (246-260), known to control with the loop N3 (141-156) the NTD-4A8 interaction, was observed. An alteration in the loop N3 was also detected (Figure 7).

Interaction of NTD with the neutralizing antibody 4A8 for recombinant (R) and non-recombinant (NR) strains of SARS-CoV-2 BQ subvariants. Zoomed figures present the interaction zone.
Mutation analysis
Impact of mutations on interaction of RBD with the cell receptor ACE2
The BQ.1.2 recombinant, that showed an enhanced interaction with ACE2, carried three mutations (S373P, S375F, T376A) in the non-RBM region (Residue 318-437) connecting the RBD to the cell receptor ACE2. As shown by Figure 8, this interaction seems to be consolidated by formation of H-Bonds and direct contact between residues 373P/375F and ACE2. The residue 376A attaches the residue 375F to other residues of RBD, maintaining the interaction of non-RBM region to ACE2. The RBD of non-recombinant strain, that showed weak interaction with ACE2, harbored the same mutations (373P, 375F, 376A), but, they were not involved in the interaction with ACE2 (Figure 8).

Interaction of RBD mutations with the cellular receptor ACE2 for recombinant (R) and non-recombinant (NR) strains of SARS-CoV-2 BQ.1.2 subvariant (6M0J). Yellow: Receptor binding domain (RBD), Gray: ACE2, Pink: Interaction zone in RBD, Cyan: Interaction zone in ACE2. Dotted line: Hydrogen bond. PDB ID: ACE2.
Impact of mutations on interaction of RBD and NTD with the neutralizing antibodies
Mutation analysis revealed that the RBD interaction zone with the neutralizing antibodies (nAb) harbors several mutations, localized in non-RBM (Residue 318-437) and RBM (Residue 438-506) regions, which probably destabilize the interaction of BQ recombinants with the nAb C105, LY-CoV555 and C118 (Figure 9 and Figure S3). For BQ.1.10, BQ.1.12, and BQ.1.1.5 recombinants, the interaction zone in RBD was the same as non-recombinant strains. The RBD of BQ.1.1 recombinant was different from those of non-recombinant BQ.1.1 where 4 substitutions (D405N, R408S, K417N, and N440K) were detected (Figure 9 and Figure S3).

Interaction of RBD mutations with the neutralizing antibodies C105, LY-CoV555, and C118 for recombinant (R) and non-recombinant (NR) strains of SARS-CoV-2 BQ subvariants. Yellow: Receptor binding domain (RBD), Gray: Heavy chain (HC), Purple: Light chain (LC), Pink: Interaction zone in RBD, Cyan: Interaction zone in HC and LC of antibodies. Dotted line: Hydrogen bond. In red: Mutations in RBD connected to residues of antibodies by electrostatic forces or Hydrogen bond. PDB ID: C105 (6XCM), LY-CoV555 (7KMG), and C118 (7RKS).
For BQ.1.1 recombinant, the substitutions (D405N, R408S, K417N, and N440K) and other mutations (S373P, S375F, and T376A) of non-RBM region were lost in the interaction zone RBD-C118. This suggests that their dissociation caused the alteration of RBD-C118 interaction. As observed for non-recombinant model, the strong interaction between the RBD and C118 was maintained by the mutations 373P, 375F, 376A, 408R, and 440N that attach the interaction through electrostatic interactions and H-bonds (Figure 9).
The RBD of BQ.1.10 recombinant harbored six mutations (S373P, S375F, T376A, D405N, D446G, and P494S) that lost Affinity to C118 suggesting their role in the alteration of RBD-C118 interaction. The mutations 375F, 376A, 405N, and 494S are directly connected to C118, thus, they may play an important role in strengthening the interaction RBD-C118 among BQ.1.10 non-recombinant (Figure 9).
The BQ.1.12 recombinant carries about 3 to 7 mutations, within the interaction zone of RBD that lost contact with C105 (Substitutions: D405N, K417N, N460K, S477N, Q498R, N501Y, and Y505H) and LY-CoV555 (Substitutions: D405N, K417N, and N501Y). The altered interaction between the RBD and these nAb seems to be controlled by the residues 405N, 417N, 477N, 498R, 501Y, and 505H which showed their ability to directly attach the heavy and/or the light chains of nAb (Figure 9 and Figure S3).
The RBD of BQ.1.1.5 recombinant contains numerous mutations which lost affinity to the nAb C105 (Substitutions: S373P, S375F, T376A, and D405N) and LY-CoV555 (Substitutions: D405N, K417N, N501Y, and Y505H). The figure showed that the mutations 373P, 375F, 376A, and 405N are strongly engaged in the interaction of RBD with C105 and LY-CoV555 through the electrostatic forces and the H-Bonds formation (Figure 9 and Figure S3), suggesting that these mutations could lead to the destabilization of RBD interaction with the nAb for BQ.1.1.5 recombinants.
The NTD of BQ.1.2 recombinant was different from those of non-recombinant BQ.1.1 by one substitution in position 240 (Y240H) of N5 loop (Figure 10). The mutations D140Y and Y240H, localized in N3 and N5 loops respectively, lost their affinities to 4A8 among the recombinant strain. As revealed by Figure 10, the mutation 240Y directly attaches the NTD to 4A8 among the non-recombinant strain. This suggests that it may play a crucial role in altering the NTD-4A8 interaction.

Interaction of NTD mutations with the neutralizing antibody 4A8 for recombinant (R) and non-recombinant (NR) strains of SARS-CoV-2 BQ.1.2 subvariant. Yellow: N-terminal domain (NTD), Gray: Heavy chain (HC), Purple: Light chain (LC), Pink: Interaction zone in NTD, Cyan: Interaction zone in HC and LC of antibody. Dotted line: Hydrogen bond. In red: Mutations in NTD connected to residues of 4A8 by Hydrogen bond. PDB ID: 4A8 (7C2L).
Discussion
After its first appearance in China in last 2019, the SARS-CoV-2 continues to evolve, and new variants, with enhanced viral fitness and resistance to host immune defenses, are still emerging. Recombination is an important mechanism for molecular evolution and genetic diversity of SARS-CoV-2. 39 In this study, recombination events among Omicron BQ subvariants and “ML” “under-investigation” sequences were explored, with the aim of describing recombinant strains, their burden in terms of frequency and geographic distribution, and characterizing their impact on Spike interaction with the cellular receptor ACE2 and the neutralizing antibodies.
Our analysis revealed a high frequency of recombination events among BQ subvariants and “ML” strains with a worldwide geographic distribution. The USA, the UK, and Australia were recombination hotspots countries. Huge diversity was observed for other countries such as France, Germany, Norway, India, Philippines, and Canada. One hypothesis is that the cosmopolitan features of those countries, their large population and trade exchanges as well as international tourist flows constitute a favorable environment for co-circulation of different SARS-CoV-2 lineages and the genesis of recombinant strains.39,40 From another point of view, the application of supplementary vaccination doses in these countries created an immune pressure that enables the replication of targeted strains and, favor at the same time, the multiplication of mutated and recombinant strains that are have the ability to escape the immune response. 41 Nevertheless, it should be noted that the prevalence of recombinant strains may not indicate the true values, but rather our best estimations, as it may be impacted by the overall rate of available sequences at any given location. In addition, the detection of recombination could be inflated by numerous issues, especially, co-infections in the original sample as well as possible contamination during manipulations, which could be source of detection of many lineages in 1 sample, and thus genesis of chimeric sequences. 31 Furthermore, we cannot deny the possible assembly issues generated by some workflows that lack consistent quality control process 42 and may also produce false recombinant sequences.
Interestingly, most of BQ and ML recombinants inherited the genomic backbone of the Deltacron (XAW) and Omicron recombinant subvariants (XBB*) which spread rapidly in many parts of the world. They exhibited mutations in the Spike protein enabling them to escape from the immune system. 43 The presence of the genomic backbone of XAW and XBB* strains could have an advantageous impact on the viral fitness of the investigated SARS-CoV-2 BQ and ML recombinants.
The majority of BQ subvariants have recombination breakpoints in the Spike. Similar findings were reported by Turakhia et al 28 on the basis of a pandemic-scale phylogenomic analysis. The Spike protein is the primary target of the host immune response, as it is characterized by key multifunctional features related to transmissibility and pathogenicity7 -9,37; the antigenic variability of this protein is therefore necessary for virus adaptation into the infected host environment. The UTRs and the M protein constituted also one of the favorite recombination hotspots for BQ subvariants. Such regions are essential for RNA replication and transcription among SARS-CoV-2. A high genetic diversity has been described for the UTRs, probably related to their importance in conferring resistance against the host miRNA that initially targets the viral replication process.44,45 The M protein is antigenic and appears to play a crucial role in infectivity, viral assembly and morphogenesis via interaction with the spike and the host cell receptors. 46
For ML recombinants, recombination occurred mostly in the ORF1a encoding for Non Structural Proteins (NSPs), which is consistent with the study of Shiraz and Tripathi. 29 NSPs are involved in the transcription and replication processes of SARS-CoV-2. 47
In the Spike protein of BQ subvariants, recombination events occur in the NTD and the RBD regions that control the interaction with the neutralizing antibodies8,9,37 and host cell receptors. 7 Our docking analyses revealed increased affinity to the ACE2 receptor and decreased avidity for anti-RBD/NTD neutralizing antibodies among SARS-CoV-2 subvariants BQ.1.1, BQ1.2, BQ.1.1.5, BQ.1.10, and BQ.1.12. Akerman et al 48 have experimentally demonstrated the emergence of BQ1.1 capable of escaping antibody neutralization in the Australian population.
It seems that the Spike interaction with ACE2 or antibodies is particularly strengthened by the involvement of the non-RBM region. Based on molecular dynamics simulations and experiments, it has been demonstrated that mutations in the non-RBM region are able to change the conformation of RBM and have a significant impact on its hydrogen bonding and hydrophobic interactions; thereby enhancing the binding of ACE2 to the Spike protein. 7 According to our mutation analysis, the mutations 373P, 375F and 376A forms together a network of residues that consolidate the non-RBM interaction with ACE2, allowing a possible enhanced affinity. Zaho et al 49 demonstrated that the mutations S373P and S375F stabilize the up-RBD conformation of the spike, conferring a strong binding affinity with the human cell receptor ACE2. To the best of our knowledge, this is the first report about the contribution of mutation T376A in the binding activity of spike protein. Hence, such new findings certainly need to be confirmed with experimental studies. Several mutations were highlighted in the RBD and NTD regions, potentially related to the observed alteration of spike interaction with the neutralizing antibodies (nAb). Some of them (S373P, S375F, K417N, Q498R, and N501Y) were reported and their impact on immune evasion was confirmed.50,51 For instance, the mutation K417N was associated to immune escape from antibodies and alteration of vaccine efficiency against SARS-CoV-2 infection. 51 The mutations S373P, S375F, Q498R provoke an alteration in spike conformation, allowing immune evasion of SARS-CoV-2 from the 4 classes of nAb. 52 In addition, Lu et al 53 gave experimental evidence about the ability of mutation N501Y to reduce neutralization activity of convalescent sera and monoclonal antibodies. Interestingly, mutations T376A, D405N, R408S, N440K, S477N, P494S, and Y505H seem to be also involved in the destabilization of RBD-nAb interaction. For future investigations, the impact of these mutations on viral fitness as well as immune evasion should be considered. For BQ.1.10, BQ.1.12, BQ.1.1.3 strains, the comparison between the mutational profiles of both recombinants and non-recombinants strains revealed that the interaction zone of RBD harbored the same mutations that interacts differently with the nAbs, changing conformation of RBD-nAb complex binding. Petrie et al 54 demonstrated that the destabilizing mutation can change conformation of proteins, encoded by the same genetic sequence, allowing to viruses to optimize binding to its original receptor.
In addition, our study revealed that the antibody evasion mechanism of BQ subvariants is especially based on the dissociation from the light chain of anti-RBD and anti-NTD neutralizing antibodies. Numerous studies have shown that a certain heavy chain paired with a series of light chains can modify antigens-binding affinity55,56 and even alter the neutralization activity. 57 The light chain could also contribute to the binding and neutralization spectrum by modulating the conformation of the paired heavy chain. 58 As indicated by our docking analysis, interaction with the antibodies LY-555CoV and S309 seems less impacted by the described recombination. In fact, these antibodies have been isolated from recovered individuals, 8 which may explain their observed protective activity against SARS-CoV-2. Moreover, the binding activity of anti-NTD antibody 4A8 was also affected, due to the alteration of the N3/N5 loops in the NTD region. Indeed, these loops constitute the NTD supersite which is the target of all known NTD-specific neutralizing antibodies. 59 Under pressure from the host immune defenses, these exposed regions may constitute a subject of enhanced genetic variability that can affect the N3/N5 loops and promotes escape from neutralizing antibodies. Interestingly, the substitution Y240H, in N5 loop, seems to be a key residue in reducing NTD-4A8 interaction. Complementary studies can be conducted to identify the role of this mutation in consolidating the spike interaction with the neutralizing antibody 4A8.
It’s worth mentioning that our findings are based on “in silico” analysis and complementary experimentations can provide more support. From another point of view, the recombination analysis may also be impacted by possible co-infections, contamination as well as genome assembly artifact that may generate chimeric sequences. One limitation of such studies based on genomic public data is the bias introduced by variations in sampling intensity across geographical regions.
Conclusion
In summary, recombination is an important evolutionary mechanism that mainly affects ORF1a and spike genes of ML and BQ SARS-CoV-2 subvariants. It provides advantages by increasing the affinity with host cell receptors and/or evading immune defenses. Our in silico study gives further lights on the impact of the non-RBM region in the interaction with cell receptors and/or the neutralizing antibodies. It suggests an important role of the collaboration between the non-RBM region and the light chain of neutralizing antibodies that can constitute an object for the engineering of more efficient monoclonal antibodies. It also highlights new key mutations in NTD and RBD regions, contributing in viral fitness related especially to alteration of humoral immune response. For future works, it will be interesting to investigate the contribution of recombination events in the escape of SARS-CoV-2 from T cell-mediated cellular immunity as well.
Supplemental Material
sj-docx-1-evb-10.1177_11769343241272415 – Supplemental material for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies
Supplemental material, sj-docx-1-evb-10.1177_11769343241272415 for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies by Marwa Arbi, Marwa Khedhiri, Kaouther Ayouni, Oussema Souiai, Samar Dhouib, Nidhal Ghanmi, Alia Benkahla, Henda Triki and Sondes Haddad-Boubaker in Evolutionary Bioinformatics
Supplemental Material
sj-docx-2-evb-10.1177_11769343241272415 – Supplemental material for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies
Supplemental material, sj-docx-2-evb-10.1177_11769343241272415 for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies by Marwa Arbi, Marwa Khedhiri, Kaouther Ayouni, Oussema Souiai, Samar Dhouib, Nidhal Ghanmi, Alia Benkahla, Henda Triki and Sondes Haddad-Boubaker in Evolutionary Bioinformatics
Supplemental Material
sj-xlsx-3-evb-10.1177_11769343241272415 – Supplemental material for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies
Supplemental material, sj-xlsx-3-evb-10.1177_11769343241272415 for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies by Marwa Arbi, Marwa Khedhiri, Kaouther Ayouni, Oussema Souiai, Samar Dhouib, Nidhal Ghanmi, Alia Benkahla, Henda Triki and Sondes Haddad-Boubaker in Evolutionary Bioinformatics
Supplemental Material
sj-xlsx-4-evb-10.1177_11769343241272415 – Supplemental material for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies
Supplemental material, sj-xlsx-4-evb-10.1177_11769343241272415 for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies by Marwa Arbi, Marwa Khedhiri, Kaouther Ayouni, Oussema Souiai, Samar Dhouib, Nidhal Ghanmi, Alia Benkahla, Henda Triki and Sondes Haddad-Boubaker in Evolutionary Bioinformatics
Supplemental Material
sj-xlsx-5-evb-10.1177_11769343241272415 – Supplemental material for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies
Supplemental material, sj-xlsx-5-evb-10.1177_11769343241272415 for Recombination Events Among SARS-CoV-2 Omicron Subvariants: Impact on Spike Interaction With ACE2 Receptor and Neutralizing Antibodies by Marwa Arbi, Marwa Khedhiri, Kaouther Ayouni, Oussema Souiai, Samar Dhouib, Nidhal Ghanmi, Alia Benkahla, Henda Triki and Sondes Haddad-Boubaker in Evolutionary Bioinformatics
Footnotes
Acknowledgements
All ML sequences were retrieved from GISAID and we are deeply grateful for the authors and their originating laboratories which contributed in the collection of the specimens, data analysis and sequence submission as well as metadata sharing via the GISAID initiative.
Author Contributions
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
