Abstract
Four cpt 1 genes (cpt 1α1a, cpt 1α2a, cpt 1α2b, and cpt 1β) were identified in the Nile tilapia genome. Two transmembrane helix domains (TMH) were identified for Cpt 1α1a, Cpt 1α2a, and Cpt 1β, while Cpt 1α2b had only one TMH domain. Evidence was found of conserved gene synteny between cpt 1 genes from Nile tilapia and the cpt 1/CPT 1 genes of zebrafish and human. Phylogenetic analysis showed that Nile tilapia Cpt 1 sequences clustered in distinct clades with their orthologous Cpt 1/CPT 1 from other vertebrates. Nile tilapia cpt 1α1a, cpt 1α2a, cpt 1α2b, and cpt 1β contain 18 coding exons encoding polypeptides of 771, 784, 788, and 786 amino acids in length, respectively. The cpt 1 genes were determined in all the tested tissues with varying tissue distribution patterns. These findings suggest that (1) cpt 1α1a, cpt 1α2a, and cpt 1α2b arose in the Nile tilapia genome as a result of the teleost-specific whole-genome duplication; (2) nonfunctionalization is the most likely cause of the loss of cpt 1α1b in the Nile tilapia genome; (3) the different tissue-specific transcription of cpt 1α2a and cpt 1α2b may be either due to the sub- or the neo-functionalization of transcriptional control side.
Introduction
Fatty acid composition in fish tissues is a complex result of the deposition of dietary lipids, de novo synthesis, and the oxidation of long-chain polyunsaturated fatty acids (LC-PUFAs). In addition to their beneficial effects on human health, LC-PUFAs are the main energy source in many organisms. Based on the metabolic role of the tissue, fatty acids are either degraded to produce phospholipids or triacylglycerols, or they can be oxidized within the mitochondria to produce energy.1,2 β-oxidation is the main fatty acid catabolism mechanism, and more than 90% of polyunsaturated fatty acids (PUFAs) are oxidized through mitochondrial β-oxidation in vertebrates. 3 However, PUFAs cannot cross into the mitochondria matrix by simple diffusion, unlike short or medium-chain fatty acids. Therefore, the carnitine palmitoyltransferase system, which consists of carnitine palmitoyltransferase 1 (Cpt 1) and carnitine palmitoyltransferase 2 (Cpt 2), mediates the entry of the long-chain fatty acyl-CoA into the mitochondrial matrix. 2 Cpt 1, which is found in the mitochondrial outer membrane, catalyzes the conversion of the acyl groups of long-chain fatty acyl-CoAs in carnitine, resulting in the production of acyl carnitine. Cpt 1 (not Cpt 2) is sensitive to malonyl-CoA inhibition and plays a pivotal role in the mitochondrial regulation of β-oxidation.1,2,4,5 The teleost genome has 4 or 5 (mostly) cpt 1 paralogues. However, CPT 1 has 3 isoforms in mammals: CPT 1A (liver isoform), CPT 1B (muscle isoform), and CPT 1C (brain isoform).6,7
The global aquaculture industry has undergone continuous growth over the past several decades and continues to expand more rapidly than other food production sectors. 8 However, the global production of fish oil, which has traditionally been the predominant source of energy in aquaculture diets, has remained constant for the last 2 decades.9,10 Therefore, fish oil in aquaculture diets is being gradually replaced with vegetable oils in order to sustain industry growth in the face of economic (increasing fish oil costs due to higher demand) and ecological (overfishing of natural populations to meet the demand for fish oil) sustainability concerns.
Nile tilapia (Oreochromis niloticus) is one of the world’s most commonly farmed fish species, representing 8% of global aquaculture production in 2016. 8 A reference genome has been developed for Nile tilapia, which provides an unprecedented amount of detail for the study of the evolutionary mechanisms that have shaped the species’ genetic makeup. In this study, therefore, we aimed to examine the genomic architecture of Nile tilapia cpt 1 genes and characterize their tissue-specific gene expression. In addition to characterization of the organization and level of conserved gene synteny of cpt 1 genes within Nile tilapia, we examined the paralogues and orthologous of cpt 1 from other vertebrates, to elucidate the evolution of the cpt 1 genes relative to the teleost specific whole-genome duplication and species divergence.
Material and Methods
Fish and sampling
Nile tilapia (O. niloticus) obtained from the Fisheries Faculty, Çukurova University, Turkey, were transferred to the Biotechnology Laboratory at the Department of Agricultural Biotechnology, Agriculture Faculty, Atatürk University, Turkey. The fish were kept in a 100-liter aquarium for a 2-week acclimatization period at 27 ± 1°C and fed a commercial diet twice daily at satiation. Three female and 3 male fish (mean weight ~155 g) were anesthetized using 0.2% tricaine methanesulfonate (MS-222, Sigma-Aldrich, Germany), and the liver, intestine, muscle, brain, eye, adipose, gill, heart, spleen, kidney, ovary, and testis were immediately dissected.
Identification of Nile tilapia cpt 1 genes
The nucleotide sequences of Nile tilapia cpt 1α1a, cpt 1α2a, cpt 1α2b, and cpt 1β were identified using the amino acid sequences of zebrafish Cpt 1α1a (ENSDARP00000083632), Cpt 1α2a (ENSDARP00000082601), Cpt 1α2b (ENSDARP00000055293), and Cpt 1β (ENSDARP00000077856), respectively, as query for an individual tBLASTn search in the Ensembl genome database (release 99—January 2020; http://useast.ensembl.org/Multi/Tools/Blast?db=core). Another tBLASTn search of the National Center for Biotechnology Information (NCBI) (release 234—October 2019; http://blast.ncbi.nlm.nih.gov) using the Nile tilapia Cpt 1 amino acid sequences from Ensembl was conducted to retrieve Expressed Sequence Tags (ESTs) for each of cpt 1 (Supplementary File 1). No EST for cpt 1α1a, cpt 1α2a, cpt 1α2b, and cpt 1β was found in the NCBI EST database. Therefore, cpt 1α1a, cpt 1α2a, cpt 1α2b, and cpt 1β mRNAs were synthesized as indicated in the next section using reverse transcription PCR (RT-PCR) and sequenced by Genmar Laboratories (İzmir, Turkey). The sequences were aligned with Ensembl-derived cpt 1 genomic sequences to confirm their exon/intron organizations. NCBI Primer-BLAST (http://www.ncbi.nlm.nih.gov/tools/primerblast/) was used to design PCR and quantitative PCR (qPCR) primers (Table 1). Standard RT-PCR conditions were applied to the cpt 1 genes with different annealing temperatures for different cpt 1 transcripts (Table 1).
RT-qPCR primers for Nile tilapia cpt 1 and reference genes (β-actin and ef1α) with optimal annealing temperatures and qPCR efficiencies.

Abbreviations: cpt, carnitine palmitoyltransferase; RT-qPCR, reverse transcriptase-quantitative polymerase chain reaction.
Phylogenetic analysis of Nile tilapia Cpt 1/CPT 1 polypeptides and conserved gene synteny
The orthologous relationships between Nile tilapia cpt 1 genes were determined through individual BLAST search using each putative sequence as query for the OrthoInspector web portal (https://www.lbgi.fr/orthoinspectorv3/). 11 MAFFT multiple alignment program was used to align Cpt 1/CPT 1 polypeptide sequences. 12 The Jones-Taylor-Thornton (JTT) was determined as the best model to Cpt 1/CPT 1 polypeptide sequences using ProtTest tool. 13 The phylogenetic relationship of Nile tilapia Cpt 1s and other vertebrate Cpt 1s/CPT 1s was revealed using a Bayesian phylogenetic tree with an online tool (https://ngphylogeny.fr/) 14 using the JTT model approach with 106 generations, 100 sampling frequency, and a burnin of 10,000. 6 A maximum likelihood phylogenetic tree with JTT substitution model that used the Nile tilapia, Makobe island cichlid, and Zebrafish Cpt 2 proteins as outgroups (Protein IDs: XP_003444294.1, ENSPNYP00000028960, and ENSDARP00000122226, respectively) was constructed with a 1000 bootstrap replicates using MEGA7 software. 15 A maximum likelihood and Bayesian phylogenetic trees were compared to one another via a tangles analysis within the program Dendroscope 3. 16 The amino acid sequences of the Nile tilapia Cpt 1 genes were compared with those of other vertebrates using the BLOSUM62 matrix algorithm to quantify sequence identity and similarity 17 (see Supplementary File 1 for a list of protein accession numbers). In addition, conserved gene synteny of Nile tilapia cpt 1A (cpt 1α1a), cpt 1B (cpt 1β), and cpt1 C (cpt 1α2a and cpt 1α2b) genes with the cpt 1/CPT1 genes of zebrafish and human was identified manually using the Ensembl genome database.
Identification of regulatory elements and secondary structure
The putative TATA boxes, 18 transcription initiation sites, 19 and polyadenylation signals 20 of cpt 1 genes were predicted using online tools (http://gpminer.mbc.nctu.edu.tw/index.php, http://www.fruitfly.org/seq_tools/promoter.html, and http://dnafsminer.bic.nus.edu.sg/PolyA.html, respectively). TMHMM Server v. 2.0 (http://www.cbs.dtu.dk/services/TMHMM/) 21 and SMART conserved domains profile (http://smart.embl-heidelberg.de/) 22 were used to predict hydrophobicity and the protein structure of Nile tilapia Cpt 1 family, respectively.
Reverse transcriptase-quantitative PCR (RT-qPCR) analysis
The tissue-specific distributions of the Nile tilapia cpt 1 transcripts were identified using RT-qPCR. Total RNA from the liver, intestine, muscle, brain, eye, adipose, gill, heart, spleen, kidney, ovary, and testis was extracted using a TRIZOL reagent (Invitrogen, Carlsbad, CA, USA). The integrity and concentration of the total RNA were determined using a NanoDrop Spectrophotometer (ThermoFisher, Multiskan Go). The first-strand complementary DNA (cDNA) was synthesized by Omniscript Reverse Transcription kit (Qiagen, Düsseldorf, Germany) with 2 μg of the total RNA extracted from the tissues. A Rotor-Gene 6000 thermal cycler system with a QuantiTect SYBR Green PCR kit (Qiagen GmbH, Düsseldorf, Germany) was used to perform RT-qPCR analysis to determine the copy number of four cpt 1 transcripts. 23 The PCR conditions were as follows: 15 min at 95°C (initial denaturation), followed by 40 cycles of 20 s at 95°C (denaturation), 30 s at each cpt 1 transcript’s optimum annealing temperature (primer annealing), and 30 s at 72°C (elongation). As a negative control, template cDNA was omitted from the qPCR reaction. 24 The normalized copy number of the Nile tilapia cpt 1 mRNA transcripts in each tissue was calculated by dividing their copy numbers separately by the copy number of the following reference genes: Elongation factor 1 alpha (Gene ID: AB075952.1; ef1α) and ß-actin (Gene ID: XM_003443127; actb), which were selected since they were reported as suitable Nile tilapia reference genes for qPCR analysis.25-27 Then, the mean of the normalized values was calculated.
Statistical analysis
For each of the tissues sampled (liver, intestine, stomach, muscle, brain, heart, eye, gill, kidney, spleen, ovary, and testis) the steady-state levels of cpt 1 gene transcripts were calculated as the mean ± the standard error of six samples (three samples in the case of ovaries and testes). The differences among normalized mRNA transcripts of cpt 1 genes were compared with Duncan’s multiple comparison and one-way analysis of variance (ANOVA) tests were performed, respectively, using the SPSS 10.0 package program. 28 And a significant difference was defined as a p value of less than 0.05.
Results and Discussion
cpt 1 genes in Nile tilapia genome
Four cpt 1 paralogues (cpt 1α1a, cpt 1α2a, cpt 1α2b and cpt 1β) in the Nile tilapia genome were identified using the Ensembl (www.ensembl.org) and National Center for Biotechnology Information (www.ncbi.nlm.nih.gov) genome databases. The open reading frames of cpt 1α1a (XM_003440354.4), cpt 1α2a (XM_019362661.2), cpt 1α2b, (XM_003446465.5) and cpt 1β (XM_005470876.4) consist of 2316, 2355, 2367, and 2361 base pairs encoding 771, 784, 788, and 786 amino acids, respectively. Similar to previous reports that indicated the existence of two conserved membrane spanning domains in the Cpt 1/CPT 1 of vertebrates,7,29 this study identified two transmembrane helix domains for most members of the Nile tilapia Cpt 1 family (50-72 amino acids and 105-127 amino acids in Cpt 1α1a, 49-71 amino acids and 104-126 amino acids in Cpt 1α2a, and 61-83 amino acids and 104-126 amino acids in Cpt 1β), except for Cpt 1α2b, which had only one such domain (106-128 amino acids) (Supplementary File 2). Mutation in the key amino acid site created by teleost specific whole-genome duplication (tsWGD) may be an explanation of one transmembrane helix domain in Cpt 1α2b. 30 This scenario, however, needs future investigation.
Orthology of Nile tilapia cpt 1 genes with cpt 1/CPT 1 genes from other teleosts and tetrapods
Protein sequences of Nile tilapia Cpt 1α1a, Cpt 1α2a, Cpt 1α2b, and Cpt 1β were aligned with Cpt 1/CPT 1 from Makobe island cichlid, midas cichlid, three-spined stickleback, medaka, fugu, zebrafish, grass carp, and humans using CLUSTAL W to determine sequence identity and similarity. 31 The percentage sequence identity varied among the examined Cpt 1s/CPT 1s genes, but each Nile tilapia Cpt 1 shared the highest sequence identity with the Makobe island cichlid (98%, 96%, 97%, and 99% identity respectively) (Supplementary File 3).
Nile tilapia Cpt 1 sequences clustered in distinct clades with their orthologous Cpt 1/CPT 1 from other fishes and vertebrates according to Bayesian phylogenetic inference (Figure 1). While tetrapod genomes have one CPT 1α isoform, teleost genomes have several Cpt 1α duplication isoforms, indicating teleost-specific CPT 1α duplications (Figure 1).29,32 Comparison of maximum likelihood and Bayesian phylogenetic trees supported the topology with the correct grouping of the tetrapod CPT 1 C genes with the cpt 1α2a and cpt 1α2b of teleosts, especially. The expected topology is also observed for the remaining genes (Supplementary File 4). It has been suggested that duplicated proteins in teleost fishes might exhibit higher sequence identity and similarity with their orthologues than with their paralogues due to their derivation from a common ancestral gene,33,34 which is in line with the results of the phylogenetic analysis presented here.

Bayesian phylogenetic tree of Nile tilapia Cpts with Cpts/CPTs from fishes and tetrapods: The tree topology was detected using O. niloticus, Pundamilia nyererei, Astyanax mexicanus, Amphilophus citrinellus, Danio rerio, Ctenopharyngodon idella, Oryzias latipes, Gasterosteus aculeatus, Tetraodon nigroviridis, Takifugu rubripes, Lepisosteus oculatus, Latimeria chalumnae, Callorhinchus milii, Leucoraja erinacea, Gallus gallus, Falco peregrinus, Sus scrofa, Anolis carolinensis, Homo sapiens, and Mus musculus amino acid sequences (Protein ID for each of Cpt/CPT sequences was shown in Supplementary File 1). Numbers at nodes indicate the posterior probability. Abbreviation: CPT indicate carnitine palmitoyltransferase. Scale bar = 0.1 substitutions per site.
Conserved gene synteny of Nile tilapia cpt 1 genes
The level of conserved synteny between Nile tilapia cpt 1α1a and cpt 1β and the following homologs from other species was determined: human (CPT 1A and CPT 1β), and zebrafish (cpt 1α, Cpt 1β, cpt 1α2a, and cpt 1α2b) with zebrafish cpt 1α2a and cpt 1α2b, (Supplementary File 5). The homologs of Nile tilapia cpt 1α1a and cpt 1β within zebrafish were respectively located on chromosomes 25 and 18 of the zebrafish genome, and the human genes CPT 1A and CPT 1β were respectively located on chromosomes 11 and 22 of the human genome. respectively were shown to be conserved with Nile tilapia cpt 1α1a and cpt 1β located on the same scaffold (GL831142.1). Similar conserved gene synteny for Nile tilapia cpt 1α2a located on scaffold GL831136.1 and cpt 1α2b located on scaffold GL831179.1 was determined with cpt 1α2a (chromosome 24) and cpt 1α2b (chromosome 3) from zebrafish. These results suggest that, while cpt 1α1a, cpt 1α2a, and cpt 1α2b genes of Nile tilapia arose because of tsWGD that occurred 230-400 million years ago,33,34 cpt 1β in Nile tilapia genome arose due to duplication of the ancestral CPT 1 gene, which resulted in cpt 1A, cpt 1β and cpt 1 C.29,32,35 Morash et al 6 reported that teleost fish do not exhibit cpt 1α1 isoforms (cpt 1α1a and cpt 1α1b) except zebrafish and rainbow trout. However, cpt 1α1b has been detected in further studies for grass carp and yellow catfish.29,35 In the Nile tilapia genome, cpt 1α1b has been lost, as in the majority of teleosts most likely because of nonfunctionalization, which is the most common case in the evolution of duplicate genes (Table 2).6,36
Carnitine palmitoyltransferase (cpt) genes in some teleost fish.

Abbreviation: cpt, carnitine palmitoyltransferase.
The cpt genes in Nile tilapia genome were identified in this study.
Zebrafish’ cpt genes were obtained from the Ensembl genome database.
Genomic organization of Nile tilapia cpt 1 genes
The nucleotide sequence for each cpt 1 gene was obtained from the Ensembl genome database. Their putative TATA boxes, polyadenylation signals, transcription initiation sites, 5′ and 3′ untranslated regions, and exon-intron locations were determined (Supplementary File 6). All isoforms of Nile tilapia cpt 1 genes contained 18 coding exons separated by 17 introns following the gt-ag rule, 37 expect the 11th intron in cpt 1α2a. Canonical splice nucleotides for this intron was gc-ag, which was species-specific splicing and not resulted in alternative splicing. These results indicated that the genes exhibit a strongly conserved genomic organization. Similar results were reported for grass carp. 29
Tissue-specific transcription of cpt 1 genes
Determination of the steady-state levels of Nile tilapia cpt 1 gene transcripts in various tissues through RT-qPCR provided insights into the evolutionary processes that led to retention of cpt 1 genes. Tissue-specific transcriptional regulation of duplicated genes was shown to have a larger role in determining the overall function of the gene, rather than potential protein structural modifications.31,38 The cpt 1 transcripts were determined in all the tested tissues with varying tissue distribution patterns (Figure 2). While transcript levels for cpt 1α1a and cpt 1α2a were higher in the testis than in the ovary, no significant differences were assayed for steady-state levels of cpt 1α2b and cpt 1β in female and male fish. The mRNA expression level of cpt 1α1a was the highest in the heart with, no statistically significant differences between the intestine, muscle, brain, kidney, and testis (P < 0.05) (Figure 2). Similarly, the highest cpt 1α1a mRNA expressions were determined in the heart of javelin goby, 31 sea bream 39 and grass carp 40 (Table 3). Sister cpt 1 C duplicates (cpt 1α2a and cpt 1α2b) exhibited divergent transcriptional regulation in the intestine, muscle, brain, eye, adipose, heart, spleen, ovary, and testis, suggesting that tsWGD event in Nile tilapia genome following a duplication event in ancestor genome resulted in the divergence of cpt 1 C isoforms 36 and these two isoforms most likely are retained in the Nile tilapia genome because of sub- or neo-functionalization. 41 Nile tilapia cpt 1β had its highest levels of expression in the muscle, kidney, and heart, with very low expression in the spleen, testis, and ovary. This agreed with previously observed cpt 1β expression in rainbow trout, 5 javelin goby, 31 yellow catfish, 35 and sea bream 39 for the muscles and heart, which are central tissues for β-oxidation in fish.

Tissue-specific distribution of (A) cpt 1α1a, (B) cpt 1α2a, (C) cpt 1α2b, and (D) cpt 1β gene in Nile tilapia: liver (l), intestine (i), muscle (m), brain (b), eye (e), adipose (a), gill (g), heart (h), spleen (sp), kidney (k), ovary (o), and testis (t). The line on each bar indicates the standard error of the mean. Letters a-g over each bar show statistically different mRNA expressions among various tissues (P < 0.05). Abbreviation: cpt indicate carnitine palmitoyltransferase.
Tissues reported as having the highest cpt 1 mRNA expression levels within different teleost species.
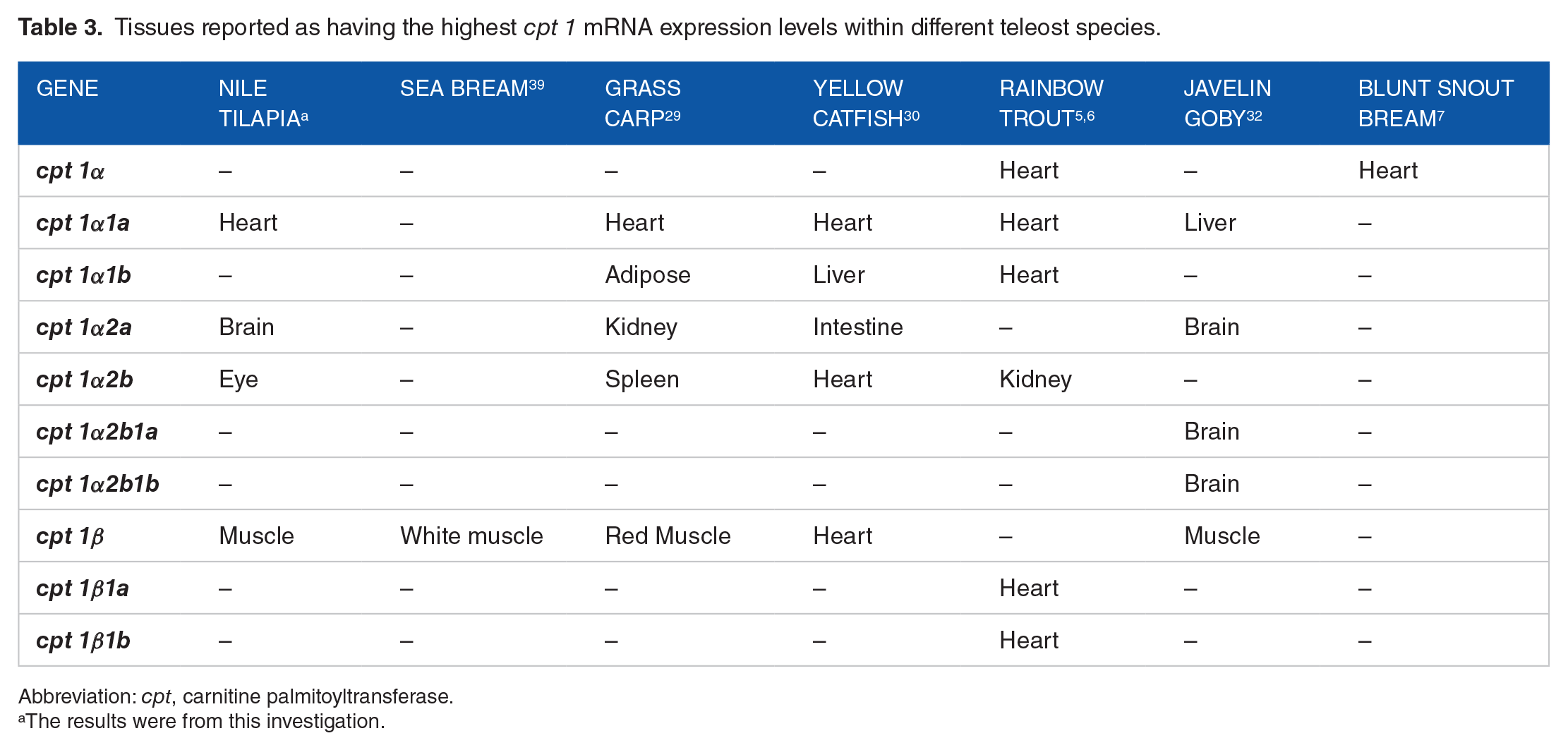
Abbreviation: cpt, carnitine palmitoyltransferase.
The results were from this investigation.
Conclusions
Based on the results of the comparative genomics and gene expression analyses, the following can be concluded: (1) 4 cpt 1 paralogues are retained in Nile tilapia genome; (2) cpt 1α (cpt 1A), cpt 1β (cpt 1B), and cpt 1 C likely arose from a common ancestral gene; (3) cpt 1α1a, cpt 1α2a, and cpt 1α2b, which are homeologs, arose because of the tsWGD; (4) cpt 1α1b has been lost most likely due to nonfunctionalization; (e) the different tissue-specific transcription of cpt 1α2a and cpt 1α2b may be either due to the sub- or the neo-functionalization of transcriptional control side.
Supplemental Material
Supplementary_file_1_xyz338235d34f3b2 – Supplemental material for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus
Supplemental material, Supplementary_file_1_xyz338235d34f3b2 for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus by Mehtap Bayır, Gökhan Arslan and Abdulkadir Bayır in Evolutionary Bioinformatics
Supplemental Material
Supplementary_file_2_xyz338239fe39db7 – Supplemental material for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus
Supplemental material, Supplementary_file_2_xyz338239fe39db7 for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus by Mehtap Bayır, Gökhan Arslan and Abdulkadir Bayır in Evolutionary Bioinformatics
Supplemental Material
Supplementary_file_3_xyz33823c7fbd073 – Supplemental material for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus
Supplemental material, Supplementary_file_3_xyz33823c7fbd073 for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus by Mehtap Bayır, Gökhan Arslan and Abdulkadir Bayır in Evolutionary Bioinformatics
Supplemental Material
Supplementary_file_4_xyz3382348c4bbb0 – Supplemental material for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus
Supplemental material, Supplementary_file_4_xyz3382348c4bbb0 for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus by Mehtap Bayır, Gökhan Arslan and Abdulkadir Bayır in Evolutionary Bioinformatics
Supplemental Material
Supplementary_file_5_xyz33823643304d2 – Supplemental material for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus
Supplemental material, Supplementary_file_5_xyz33823643304d2 for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus by Mehtap Bayır, Gökhan Arslan and Abdulkadir Bayır in Evolutionary Bioinformatics
Supplemental Material
Supplementary_file_6_xyz338236c449136 – Supplemental material for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus
Supplemental material, Supplementary_file_6_xyz338236c449136 for Identification and Characterization of Carnitine Palmitoyltransferase 1 (cpt 1) Genes in Nile Tilapia, Oreochromis niloticus by Mehtap Bayır, Gökhan Arslan and Abdulkadir Bayır in Evolutionary Bioinformatics
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
All authors designed the research. MB performed bioinformatics analyses, the laboratory studies, and wrote the manuscript. AB performed bioinformatics analyses, wrote and revised the manuscript. GA performed the bioinformatics analysis. All authors agree on the final version of manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
