Abstract
Skeletal muscle plays a vital role in mobility, posture, and overall metabolic health, yet it is highly susceptible to injury due to its distribution throughout the body’s upper and lower extremities and torso. When muscle damage occurs, the injured muscle can undergo regeneration to repair or form new myofibers. Muscle regeneration consists of phases starting with degeneration-necrosis and ending with functional recovery (Forcina et al., 2020). Inflammation occurs after the first phase (Forcina et al., 2020) and does not completely resolve until the formation of mature regenerating myofibers (Zhang et al., 2023). Therefore, inflammation plays a critical role in coordinating and supporting skeletal muscle recovery following injury.
Inflammation in response to an acute muscle injury can vary depending on estrogen status. In ovariectomized mice, treatment with 17β-estradiol led to increased chemokine mRNA expression at 24 h after freeze injury to the tibialis anterior muscle compared to placebo-treated mice (Le et al., 2018). Chemokines are the proteins that attract inflammatory cells to the injury. Then one day later, Cd68 and Cd206 (coding for surface markers of the inflammatory cells, macrophages) mRNA expression was lower and greater, respectively, in the 17β-estradiol-treated mice compared to placebo-treated mice (Le et al., 2018). The mRNA of the coded macrophage surface marker, CD163, was higher in the 17β-estradiol-treated than the placebo-treated mice from postinjury days 1–4, although similar CD163 protein levels were detected in these two groups at these time points (Le et al., 2018). Therefore, the presence of estrogen or biological sex may be an important factor to consider in understanding the inflammation related to muscle regeneration and assessing and treating muscle injury.
Myeloid-derived cells are a key immune cell type involved in muscle inflammation. In acute muscle injury models, myeloid-derived cells are common, including neutrophils (Zhang et al., 2023), macrophages (Zhang et al., 2023), and dendritic cells (Saidj et al., 2022). One myeloid cell type detected in the muscle of the idiopathic inflammatory muscle condition, anti-synthetase syndrome, is CD169+ cells (Preuβe et al., 2022). Although the function of muscle CD169+ cells in persons with myositis (Preuβe et al., 2022) or those with long COVID-19 (Aschman et al., 2023) is unclear, these cells can have a repair or regenerative role in other tissues. For example, Batoon et al. (2017) reported that CD169+ macrophages contributed to ossification during bone repair. In an experimental model of liver regeneration, Behnke et al. (2018) observed that CD169+ macrophages stimulated the production of the cytokine, interleukin-6, and the transcription factor, signal transducer and activator of transcription 3 (STAT3), which promoted liver regeneration. Similarly, Ismahil et al. (2025) demonstrated that certain CD169+ macrophages are critical for healing of the heart after myocardial infarction. Specifically, the absence or depletion of these cells in the infarcted heart resulted in prolonged or greater inflammation (Ismahil et al., 2025). These findings suggest that CD169+ cells may have unrecognized roles in injured/regenerating muscle.
Another consideration about CD169 is that this protein is encoded by the gene Siglec1 (The UniProt Consortium, 2025). The Siglec1 gene, CD169 protein, and other interferon-inducible genes are often assessed for type 1 interferon (IFN1) activity because the IFN1s can be difficult to detect in cells and tissue (Boz et al., 2024; Kamperman et al., 2024). Recently, Siglec1 has been identified as a biomarker for IFN1 activity in chronic muscle inflammation or myositis in response to autoimmune conditions, such as dermatomyositis (Boz et al., 2024; Graf et al., 2022; Kamperman et al., 2024), anti-synthetase syndrome (Graf et al., 2022), or systemic lupus erythematous (Boz et al., 2024).
Muscle regeneration involves activation of key signaling pathways, such as the Janus kinase/signal transducer and activator of transcription (JAK/STAT) pathway (Moresi et al., 2019), along with muscle-specific genes including Myh1, Myh2, Myod1, Myh3, Myh8, and Ncam1 (Pinal-Fernandez et al., 2024). While Siglec1 has not been directly associated with the Jak/Stat pathway or these muscle-related genes, interferon (Bolko et al., 2021) and interferon-inducible (Pinal-Fernandez et al., 2024) genes have been associated with these Jak/Stat and muscle-related genes, respectively. The interferon-inducible gene effects include combating viruses indirectly and inducing the production of suppressor cytokine signaling proteins, SOCS, to diminish inflammation (Bolko et al., 2021). Therefore, the possibility exists that Siglec1/CD169 may be associated with both inflammation- and muscle-related genes.
In contrast to bone repair (Batoon et al., 2017), liver regeneration (Behnke et al., 2018), and myocardial infarction (Ismahil et al., 2025) studies, the presence of CD169+ macrophages during muscle regeneration has not been investigated. Moreover, the trajectory of Siglec1 and other interferon-inducible genes after an acute muscle injury has not been examined. Therefore, one aim of this study was to explore the transcriptional expression of Siglec1 and other interferon-inducible genes in injured muscle by comparing microarray gene expression between 17β-estradiol and placebo-treated ovariectomized (OVX) mice at three postinjury time points. Another aim was to assess the presence of CD169+ cells in regenerating muscle areas between 17β-estradiol- and placebo-treated mice. By investigating CD169+ cells and interferon-inducible genes, our understanding of the mechanisms driving muscle inflammation during muscle regeneration will expand and ultimately may lead to identifying interferon-inducible-gene-based therapeutic strategies to promote muscle regeneration.
Methods
Animals
Female C57BL/6NHsd OVX mice, aged 6–10 weeks (N = 77), were purchased from Harlan Laboratories (Indianapolis, IN, USA). The mice were housed individually under a 12-h light/dark cycle with ad libitum access to food and water. They were fed a phytoestrogen-free diet and were acclimated for at least 5 days before experimental procedures began. All experimental protocols were approved by the University of Nevada, Las Vegas Institutional Animal Care and Use Committee (IACUC) and the United States Air Force Animal Use Programs Office of Research Oversight and Compliance. Microarray data from 36 of these mice were originally analyzed using a weighted gene co-expression network procedure in Moore et al. (2019). Here, we performed a secondary analysis of these data. Our analysis is different and does not duplicate the analysis of Moore et al. We selected 86 genes from this dataset and examined the interrelationships. In addition, we added data from muscle samples of other mice to examine the presence of the CD169 protein.
Pellet Implantation
Mice were divided into two groups and received either a 60-day release 17β-estradiol (n = 39) or placebo pellet (n = 38; Innovative Research of America, Sarasota, FL, USA; 0.18 mg). The pellets were inserted subcutaneously approximately 4–9 weeks post-ovariectomy. At the time of pellet implantation, the mean body weight of the 17β-estradiol group was 22.74 g (SD 2.10), while the mean of placebo group was 22.71 g (SD 2.17).
Crush Muscle Injury and Hypobaria Exposure
After 7 days of placebo or 17β-estradiol treatment, a closed crush injury was applied to the right gastrocnemius and quadriceps femoris muscle groups. Mice received subcutaneous buprenorphine (target, 0.05–0.10 mg/kg) prior to anesthesia with inhalation isoflurane (1–5% to effect) and 100% oxygen. The crush injury was induced by applying 45 psi pressure for 30 s, repeated two to four times. A second dose of buprenorphine was administered 9–12 h after the first dose. At 22–24 h postinjury, all mice were exposed to mild hypobaria (565 ± 5 torr) for 8 h to simulate the atmospheric pressure during medical airplane transport.
Mice were either returned to their cages or were euthanized immediately after hypobaria exposure (∼32-h or 1.5-days postinjury). Other mice were euthanized at 96-h (4-days) or 192-h (8-days) postinjury. The first time point represents macrophage infiltration; the second represents the initial appearance of regenerating myofibers, and the third represents regenerating myofiber growth and maturation. After euthanasia, the left and right plantarflexor (including the gastrocnemius muscle) and quadriceps femoris muscle groups were harvested. Upon harvesting the muscles of one mouse, we found a fibula fracture. For this analysis, we focused on the harvested right (injured) lateral gastrocnemius muscle or plantarflexor muscle group.
RNA Isolation/Preparation and Microarray Analysis
Isolation and preparation of the right lateral gastrocnemius samples and generation of the microarray sample files were described in Moore et al. (2019). Samples were processed at the University of Washington Department of Environmental & Occupational Health Sciences microarray facility using the Affymetrix Mouse Gene 1.0 ST array and Sense Target Labeling protocol. Arrays were scanned with a GeneChip 3000, and feature extraction was performed using Affymetrix Command Console software. Raw Affymetrix (CEL) files were processed in R v.4.4.1 (R Core Team, 2022) using Bioconductor packages oligo, affy, AnnotationDbi, and limma. Data were background-corrected, quantile-normalized, and summarized at the probe set level using the robust multiarray average (RMA) method (Bolstad et al., 2003; Irizarry et al., 2003). Probes were annotated with gene symbols, names, and Entrez IDs. Quality control was performed on unique probe IDs; low-expressing probes across all experimental conditions were removed from further analyses. Principal component analysis was used to evaluate the variance in gene expression data across experimental conditions and time points. Differential expression analysis comparing 17β-estradiol- to placebo-treated OVX mice at each postinjury time point (32 h, 96 h, and 192 h) was conducted using limma’s empirical Bayes approach. p values were adjusted using the Benjamini–Hochberg false discovery rate (FDR) method, and genes with adjusted p < 0.05 and |log2FC| > 0.2 were considered statistically significantly differentially expressed. The entire dataset including Probe ID, gene symbols, ENTREZ Gene ID, gene name, transformed expression values, and differential analysis results are included in the NCBI GEO submission (GSE accession number GSE311704). To gain further insights into the differentially expressed genes (DEGs), we uploaded the results to iPathwayGuide (Advaita Bio; Ahsan & Drăghici, 2017) to identify enriched pathways between the 17β-estradiol- and placebo-treated groups at the three time points.
Given that Siglec1 is of particular interest as a biomarker for IFN1 activity, we conducted a gene-wise correlation analysis with Siglec1 using Pearson correlation coefficient. Also, we conducted in-depth analyses of 84 genes that were categorized into four biological themes: interferon-inducible, Jak/Stat pathway, interferon regulatory factor, or muscle regeneration. These genes were selected based on an extensive literature review (Bolko et al., 2021; Boz et al., 2024; Chen et al., 2016; Graf et al., 2022; Kamperman et al., 2024; Pinal-Fernandez et al., 2024; Wang et al., 2024).
Hematoxylin and Eosin Staining
Hematoxylin and eosin (H&E) staining was performed to identify the areas of regeneration and assess the difference in regeneration phases between 17β-estradiol- and placebo-treated OVX mice. Staining was performed on 10-micron-thick cross-sections of the injured plantarflexor muscle group of 17β-estradiol- (n = 21) and placebo-treated (n = 20) mice that recovered at 96-h and 192-h postinjury. The phase of regenerating area was assessed using the classification described by Zhang et al. (2023). Two observers independently assessed these areas. Discrepancies were resolved by a third observer.
Anti-CD169 Immunohistochemistry
CD169 protein is the product of Siglec1. To evaluate the presence of the CD169 protein at an area of regeneration, immunohistochemical staining of 10-micron-thick cross-sections of the plantarflexor muscles was performed at two different times. The primary antibody was a purified rat anti-mouse CD169 antibody (AbD Serotec-Bio-Rad, 1:20 dilution, Hercules, CA; or Biolegend, 1:50 dilution, San Diego, CA). Certain sections were tested with both primary antibodies to ensure consistency. The secondary antibody, biotinylated anti-rat IgG (mouse adsorbed, ∼1:200 dilution), Vectastain® ABC peroxidase kit, and Vector® VIP peroxidase substrate kit were purchased from Vector Laboratories (Newark, CA). Each CD169-stained section had a corresponding negative control in which a separate section underwent all steps except primary antibody incubation and was substituted with phosphate-buffered saline solution incubation.
For the regenerating area observed in the H&E-stained section, we identified the same area in the corresponding anti-CD169-stained and negative control sections. Negative control sections were compared to the anti-CD169-stained sections for false-positive staining using a Nikon Eclipse Ni microscope (Melville, NY) under the 10X and 20X objectives. Anti-CD169-stainted sections that exhibited faint to strong staining intensity and discrete (>2 cells) and/or diffuse staining in this regenerating area at 200X magnification were considered “positive muscles” for CD169+ macrophages. Two observers independently assessed these areas.
Results
Gene Expression
The filtered microarray dataset includes 24,019 unique probe IDs corresponding to 22,283 unique genes. Principal component analysis revealed that gene expression variation was primarily driven by recovery time (Supplemental File Figure 1). Clear separation between treatment groups was observed at 32-h and 192-h postinjury, whereas at 96-h postinjury, 17β-estradiol- and placebo-treated OVX mice showed higher intra-variability. DEGs were further investigated within iPathwayGuide. Supplemental File Table 1 summarizes DEG up- and down-regulation at each time point.
Siglec1 is regulated by IFN1 activity and has been linked to several autoimmune diseases and inflammatory myopathies (Boz et al., 2024; Graf et al., 2022). Its protein product, CD169, has also been implicated in tissue regeneration processes, including bone repair (Batoon et al., 2017), liver regeneration (Behnke et al., 2018), and cardiac healing after myocardial infarction (Ismahil et al., 2025), making it of interest in the context of muscle inflammation and healing after acute injury. Although Siglec1 did not demonstrate differential expression at 32-h or 96-h postinjury, this gene was amongst the most up-regulated DEGs in 17β-estradiol-treated mice compared to placebo-treated mice at 192-h postinjury with a log2FC of 1.595 (adj. p < 1.000e-6).
Gene-wise correlation analysis of Siglec1 expression over time revealed distinct co-expression network dynamics between experimental groups. In placebo-treated mice, Siglec1 exhibited notable positive correlations (Pearson correlation coefficient r ≥ 0.8) with 1605 genes (Figure 1(A)), indicating a broader connectivity across the transcriptome than expected by chance. This extensive co-expression pattern suggests that Siglec1 functions as a hub gene, potentially coordinating immune and inflammatory signaling pathways under baseline conditions. In contrast, in 17β-estradiol-treated mice, Siglec1 correlations followed an approximately normal distribution (Figure 1(B)), consistent with typical gene-gene relationships. Here, Siglec1 exhibited a Pearson correlation coefficient of r ≥ 0.8 with only 78 genes. This shift toward reduced network centrality implies that estrogen modulates Siglec1-driven signaling, diminishing its influence within the broader inflammatory network. In further examination of the top correlated genes in both treatment groups, we observed Pearson correlation coefficients of 0.984 and 0.913 between Siglec1 and Syndecan 3 (Sdc3) in placebo- and 17β-estradiol-treated mice, respectively (Supplemental File Table 2). Figures 1(C) and (D) show the co-expression patterns of Siglec1 and Sdc3 at the three time points. Correlation of gene expression with Siglec1 and temporal expression patterns under placebo- and 17β-estradiol-treatment. (A and B) Distribution of gene-wise Pearson correlation coefficients (r) with Siglec1 within each treatment group across all time points. The placebo group (A) shows a broader, left-skewed distribution indicating a wider range of genes correlated with Siglec1, whereas the 17β-estradiol group (B) exhibits a near-normal distribution centered around zero, suggesting reduced network connectivity. (C and D) Normalized expression profiles of Siglec1 (solid circles, solid line) and Sdc3 (solid triangles, dashed line) measured at 32-h, 96-h, and 192-h postinjury. Under placebo treatment (C), Siglec1 and Sdc3 expression decline over time. Under 17β-estradiol treatment (D), both genes remain relatively stable across time points. Error bars represent standard deviation (SD). h, hours postinjury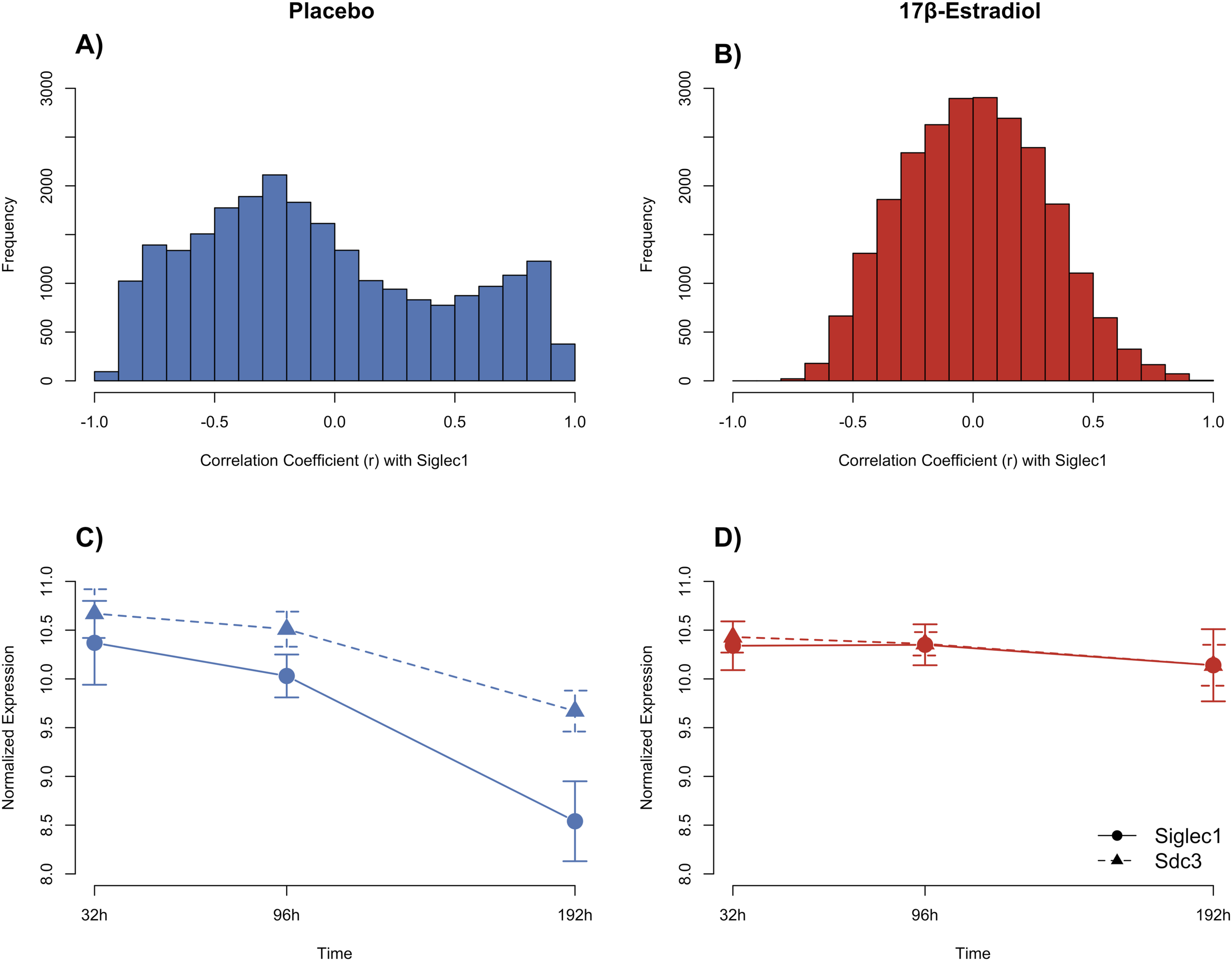
Given our hypothesis regarding Siglec1, the goal was to examine its relationship with genes identified in the literature as being associated with four biological themes: interferon-inducible, interferon regulatory factor, the Jak/Stat pathway, and muscle regeneration.
Siglec1 behaved similarly to the interferon-related genes and over half of the Jak/Stat pathway genes (Supplemental File Figure 2). This similarity in expression was observed over the three time points in the placebo-treated mice (Figure 2). In contrast, these genes did not correlate as strongly with Siglec1 in the 17β-estradiol-treated mice. Differential gene expression patterns associated with Siglec1 and Sdc3 under placebo- and 17β-estradiol-treatment of 86 genes categorized into four biological themes. (A and B) Heat maps showing scaled normalized expression (z-scores) of genes correlated with Siglec1 in the placebo (A) and 17β-estradiol (B) groups across 32-h, 96-h, and 192-h postinjury. Genes are clustered by expression profile and are annotated by biological theme: interferon-inducible (purple), interferon regulatory factor (turquoise), Jak/Stat pathway (yellow), muscle regeneration (blue), and Siglec1 and Sdc3 (red). h, hours postinjury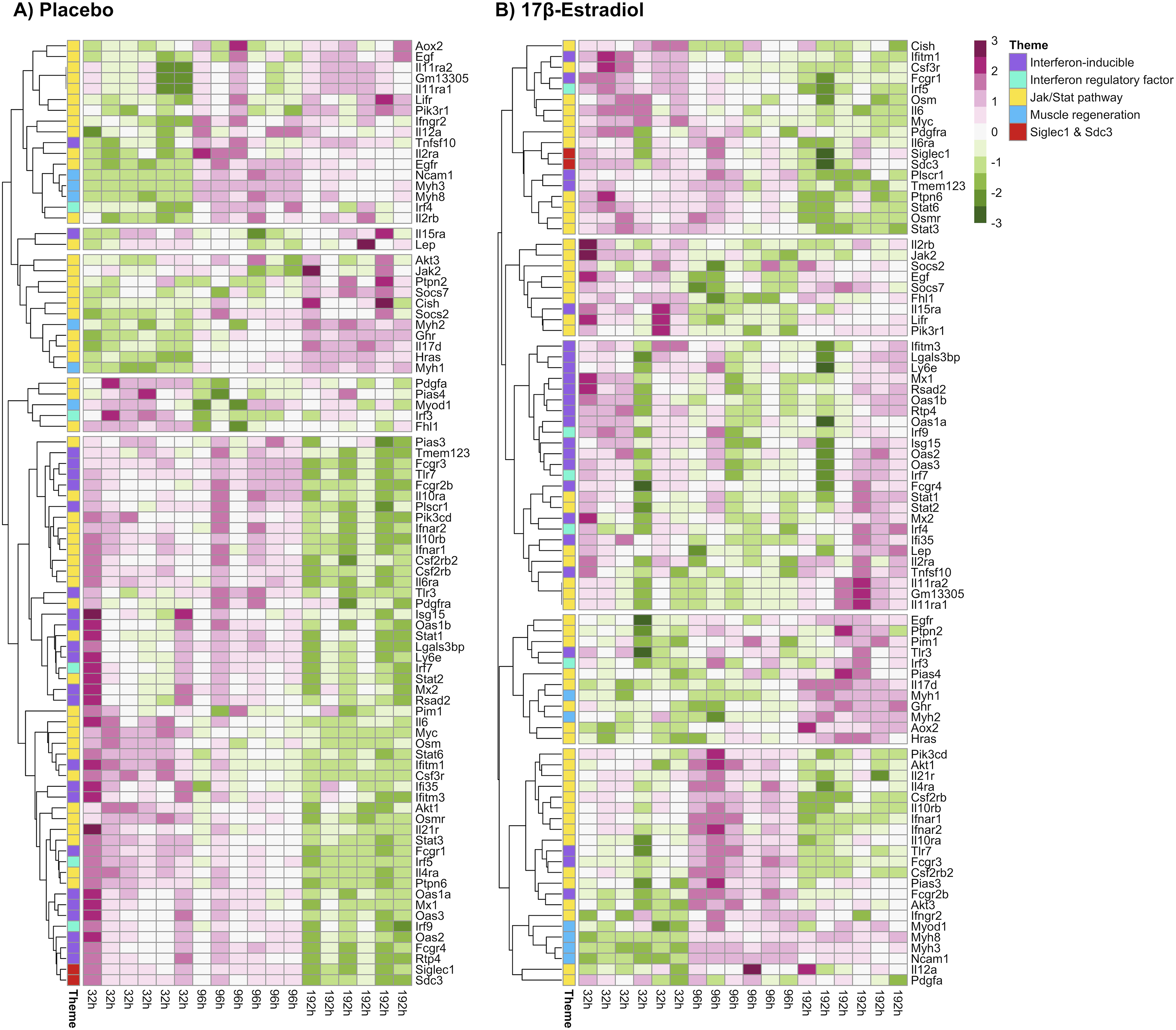
Figure 2 shows heat maps of the z-scored expression values of the biological themes of the two treatment groups. In the placebo-treated group, the expression value of Siglec1, 22 of the 24 interferon-inducible genes, and over half of the Jak/Stat pathway genes declined at 192-h postinjury. The expression value of these genes in the 17β-estradiol-treated group was more stable across all time points. In the 17β-estradiol-treated group, Myod1 clustered with the other three genes associated with early muscle regeneration (Myh3, Myh8, and Ncam1), whereas Myod1 did not cluster with these genes in the placebo-treated group.
Regenerating Phase and CD169+ Macrophages
Number of Crush-Injured Muscles Categorized in Each Regenerating Phase
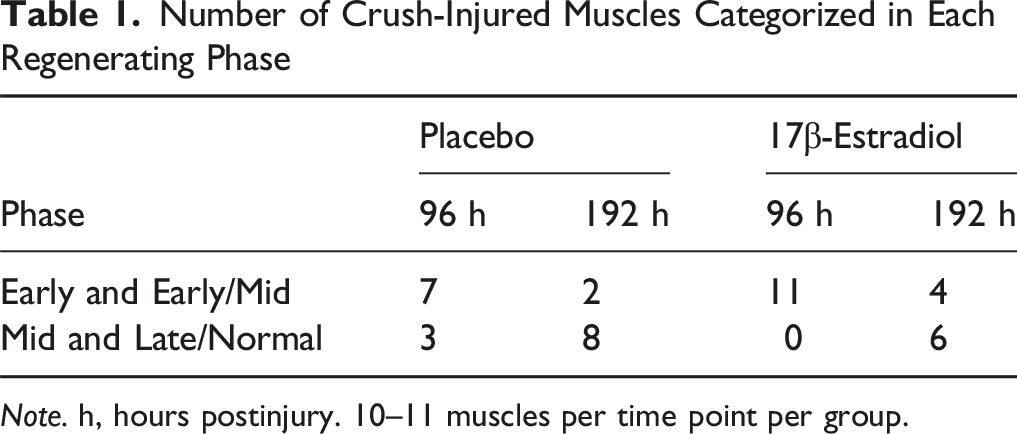
Note. h, hours postinjury. 10–11 muscles per time point per group.
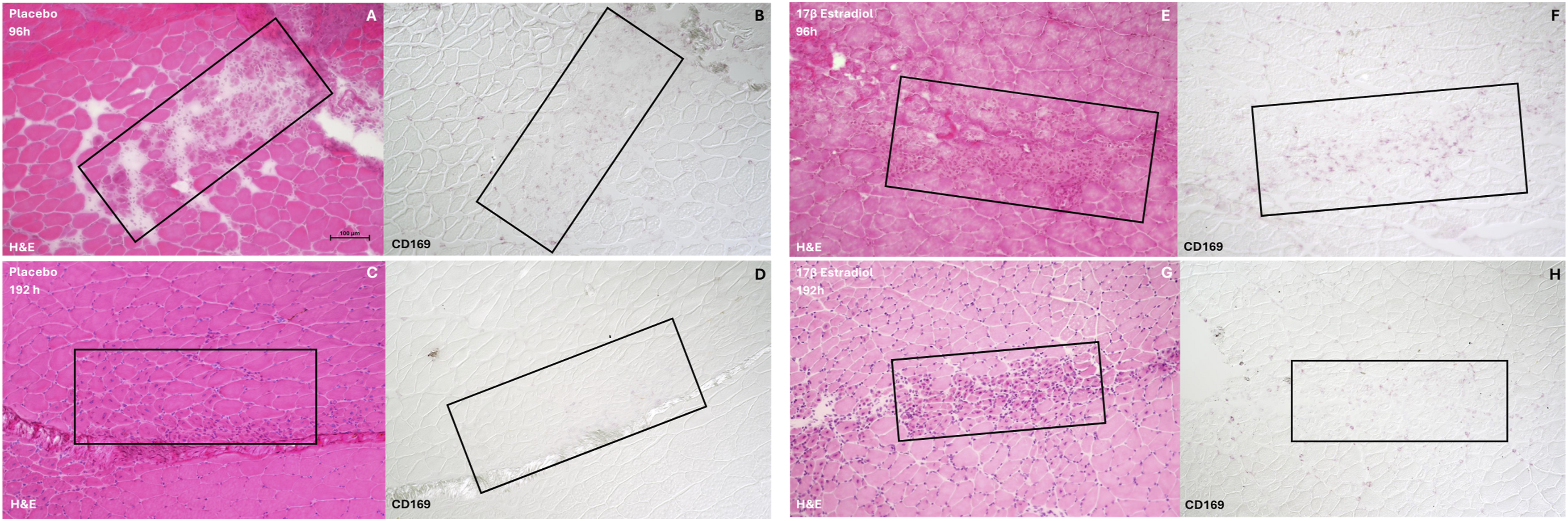
CD169+ macrophages in regenerating areas at 96-h and 192-h postinjury. CD169+ macrophages in the regenerating area of plantarflexor muscle group cross-sections at 96-h (A–D) and 192-h (E–H) postinjury. The regenerating area (black box) in each hematoxylin and eosin (H and E)-stained cross section (A, C, E, G) is matched to an area (black box) in a serial section stained with an anti-CD169 antibody (B, D, F, H) to detect the CD169 protein coded by Siglec1. h, hours postinjury
Quantification of Crush-Injured Muscles Positive for CD169+ Macrophages in a Regenerating Area
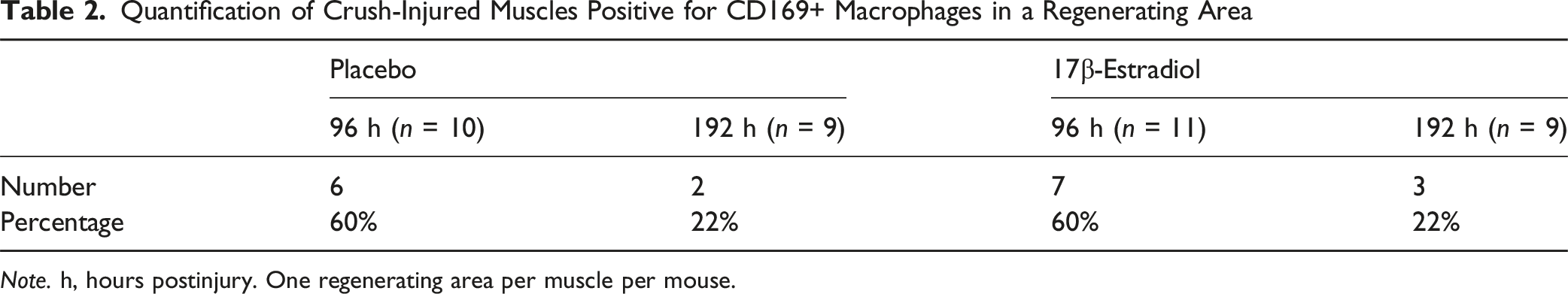

Note. h, hours postinjury. One regenerating area per muscle per mouse.
Discussion
Using between-group contrasts at each time point (17β-estradiol vs placebo), we found the most pronounced transcriptional divergence at 32-h and 192-h postinjury, indicating temporally distinct estrogen effects during recovery (Supplemental Figure 2 and Supplemental Table 1). Siglec1 is a top upregulated gene in 17β-estradiol-treated mice compared to placebo at 192-h postinjury with Siglec1 exhibiting constant mRNA expression levels in the 17β-estradiol-treated mice, while its expression declined in the placebo-treated mice (Figures 1(C) and (D)). Therefore, the differential expression of Siglec1 reflects a change in the muscle of the placebo-treated mice rather than in the muscle of the 17β-estradiol-treated mice. In addition, we observed that more genes correlated positively with Siglec1 in the placebo-treated group than in the 17β-estradiol-treated mice (Figures 1(A) and (B), Supplemental Table 3). This hormonal-related differential expression of Siglec1 and its correlated genes in injured muscle represents a novel finding. Another new finding is that we detected the presence of CD169+ cells in the regenerating area, especially in 17β-estradiol-treated mice at 192-h postinjury (stronger intensity in this group).
The Siglec1 gene and protein expression during the recovery period suggests a role of IFN1 activity during muscle regeneration. Increased Siglec1 expression is an indicator of IFN1 activity in inflammatory myopathies (Graf et al., 2022) and its protein expression has been observed in tissue regeneration (Behnke et al., 2018) or remodeling (Ismahil et al., 2025). We found a correlation between interferon-inducible genes and Siglec1 in the placebo-treated mice, suggesting that IFN1 activity may occur during muscle regeneration. Similarly, Mathes and Lafyatis (2011) detected two interferon-inducible genes, Ifit1 and Mx-2, during muscle regeneration in mice. However, these investigators ruled out IFN1 activity during muscle regeneration because the regenerating muscle of IFN1-receptor-deficient, gonadal-intact mice and wild-type, gonadal-intact mice exhibited similar mRNA expression levels of Myh3 (Mathes & Lafyatis, 2011). Despite an important role of Myh3 in the repair and formation of new muscle fibers, IFN1 activity may affect other targets as in our data set, Myod1, did not cluster with other muscle-regeneration genes (including Myh3) in the placebo-treated mice. Therefore, future muscle regenerating studies examining a wider range of targets may be warranted.
Our finding, the correlation between Siglec1 and Jak/Stat pathway genes in the placebo-treated mice, also may indicate involvement of Siglec1 or IFN1 activity in muscle regeneration. Different genes of the Jak/Stat pathway may play a role in muscle regeneration by promoting myoblast proliferation (Sun et al., 2007) and differentiation (Wang et al., 2008). For example, in murine regenerating muscle, Sun et al. (2007) demonstrated higher JAK1 kinase activity and active STAT1 protein levels at 1-day postinjury compared to 0-day postinjury. In the same experiment, active STAT3 protein levels were elevated at 1-day and 3-days compared to 0-days postinjury (Sun et al., 2007). Both Stat1 and Stat3 correlated with Siglec1 in the injured muscle of the placebo-treated mice in our study. These correlations suggest an association between these Stat genes and Siglec1, and the possibility that Siglec1 or IFN1 activity may be involved in our model of muscle injury/regeneration.
While the expression of Siglec1 was significantly lower in placebo-treated mice compared to 17β-estradiol-treated mice at 192-h postinjury, CD169+ cells were still detected in the regenerating areas of both groups. This finding may be explained by the relationship between interferon-inducible-gene levels and Siglec1 expression. In patients with immuno-rheumatological conditions or acute infections, Boz et al. (2024) reported a positive correlation between the median fluorescence intensity of Siglec1-positive monocytes and a higher interferon score. (Median fluorescence intensity indicates the amount of protein in the cell. An interferon score reflects the positive expression of certain interferon-inducible genes.) In contrast, the percentage of Siglec1-positive cells was more highly correlated with lower interferon scores (Boz et al., 2024). These data suggest that higher interferon-inducible-gene levels may enhance the amount of Siglec1 protein in cells already expressing this protein (Boz et al., 2024). Likewise, in our findings, at 192-h postinjury, the placebo-treated mice exhibited low Siglec1 gene expression and faint, diffuse CD169+ staining in the regenerating areas, while 17β-estradiol-treated mice exhibited stable Siglec1 gene expression and more intense, discrete CD169+ staining in the regenerating areas.
Interestingly, 55% of the muscle cross-sections from the placebo-treated mice appeared to exhibit characteristics of later-stage muscle regeneration compared to 28% from the 17β-estradiol-treated mice. One possible explanation is the occurrence of premature myoblast differentiation in the muscle of the placebo-treated mice. Normally, the phases of myoblast proliferation and differentiation are separate, but in the case of premature differentiation, certain myoblasts may be proliferating whereas other myoblasts advance to differentiating (Sun et al., 2007). The likely consequence may be incomplete regeneration/recovery because the number of myoblasts required for recovery may not be produced (Sun et al., 2007). According to Sun et al. (2007), while the JAK1-STAT1-STAT3 pathway is active in promoting myoblast proliferation, this pathway is concurrently suppressing muscle differentiation genes, such as Mef2, Myod, and myogenin. However, factors that inhibit Jak1 could hasten differentiation gene expression. For example, Sun et al. (2007) transfected the mouse immortalized myoblasts, C2C12, with JAK1-siRNA and examined their proliferation and expression of differentiation genes. Compared to control cells, the C2C12 cells transfected with JAK1-siRNA exhibited reduced proliferation and increased cell cycle inhibitors, p21Cip1 and p27Kip1, which would hinder cells from completing the cell cycle (Sun et al., 2007). In addition, Sun et al. found that these transfected cells expressed the differentiation protein, MyoD, while still in growth medium (as opposed to differentiation medium). While we cannot identify any specific JAK1 inhibitory factors in our study, the drop in Siglec1 expression or other Siglec1 gene correlates in the muscle of the placebo-treated group might be involved either directly by inhibiting the Jak/Stat pathway or via the downstream effects to account for the presence of early differentiation.
The increased connectivity defined by positive correlations of Siglec1 with other genes in the injured muscle of placebo-treated mice, compared to 17β-estradiol-treated mice, suggest that Siglec1 displays more hub-like characteristics under ovarian-hormone-deficient conditions. While Moore et al. (2019) did not identify Siglec1 among the top hub genes in their network analysis of the same dataset, the two approaches offer complementary perspectives. Moore et al. conducted a global analysis to identify major network hubs across the entire transcriptome, whereas the present study focused specifically on Siglec1 and its correlated gene network, providing a more localized view of its potential regulatory interactions.
Interestingly, Moore et al. (2019) identified Socs7, a member of the suppressor of cytokine signaling (SOCS) protein family (Lynch et al., 2024), as a hub gene in the placebo-treated mice, a finding we confirmed. Socs7 showed increased connectivity when considering the absolute correlation values, which is the default approach in weighted gene co-expression analysis (Supplemental File Table 3). Further, Socs7 is strongly negatively correlated with Siglec1 (r = −0.87) and showed increased expression in the placebo-treated mice at 192-h postinjury (Supplemental File Figure 3). The biological effects of the SOCS7 protein include the inhibition of growth hormone, prolactin signaling via STAT5 protein and leptin signaling via STAT3 protein (Martens et al., 2005), and regulation of insulin signaling and glucose levels (Banks et al., 2005). In addition, SOCS7 colocalizes with cytoskeletal proteins, such as vinexin and actin, suggesting that SOCS7 may be involved in cell adhesion signaling and cytoskeletal remodeling (Martens et al., 2004). Further investigations are needed to determine the relation between the Siglec1 and Socs7 genes and whether the known SOCS7 effects relate to the inflammatory or other phases of muscle regeneration.
This study has limitations. The primary aim of the study was to examine the effects of estrogen on skeletal muscle inflammation and regeneration after a combination of acute muscle injury and mild hypobaria exposure. The purpose of including the mild hypobaria exposure was to simulate medical air transport. For this reason, groups of healthy and injured 17β-estradiol-treated OVX mice without mild hypobaria exposure were not included. Without these groups, the effects of 17β-estradiol on muscle regeneration or gene expression cannot be assessed independently of mild hypobaria. Therefore, additional studies without mild hypobaria are needed to determine whether the findings in the current study are unique to this specific pressure exposure. Another limitation is that quantification of CD169+ cells in the regenerating area was not performed. In many cross-sections, the anti-CD169 staining was diffuse, and individual cells could not be distinguished to determine number or concentration; and/or the faint intensity could not yield meaningful values. Furthermore, the system used to classify the regenerating phase of H&E-stained muscle cross-sections was originally based on regenerating quadriceps femoris muscles of male mice (Zhang et al., 2023). While a uniform regeneration process occurs across different muscles, the sex-specific differences in muscle regeneration might have affected the morphological features assigned to create this classification.
Nursing Implications
Recovery from injury, mobility, and the physical activity of persons with and without disease or illness are central to nursing. As such, skeletal muscle health is an important focus of both nurse clinicians and scientists. These preclinical findings may represent a new direction in skeletal muscle health nursing research. Specifically, these findings serve as a foundation for future studies to investigate the role of IFN1 activity/Siglec1 in muscle regeneration using mutant IFN1/Siglec1 mice, inhibiting the Siglec1 gene, and/or overexpressing the Siglec1 gene. In addition, depleting CD169+ macrophages and examining the relation of IFN1 activity to estrogen during muscle regeneration warrants investigation. Although placebo-treated OVX mice are different from postmenopausal women, the data from these mice may lay the groundwork for understanding whether postmenopausal women who sustain an acute muscle injury are at risk for incomplete recovery. If this risk exists and given the importance of physical activity to overall health and well-being, then these women may require adjunct therapy to maintain IFN1 activity to ensure complete muscle regeneration. However, the proposed preclinical studies should be conducted initially before considering and developing adjunct IFN1 therapy for these women.
Conclusion
In this study, we found more stable Siglec1 expression and stronger CD169+ macrophage staining in the injured muscle of 17β-estradiol-treated mice than in the placebo-treated mice. However, Siglec1 was correlated with more interferon-related genes in the muscle of the placebo-treated mice. These findings suggest that Siglec1/IFN1 activity may be part of the inflammatory response during muscle regeneration. Future studies are warranted to investigate this role.
Supplemental Material
Suppplemental Material - Estrogen Stabilizes Interferon-Inducible Genes During Skeletal Muscle Regeneration
Suppplemental Material for Estrogen Stabilizes Interferon-Inducible Genes During Skeletal Muscle Regeneration by Tara Movaghar, Juli Petereit, Barbara St. Pierre Schneider in Biological Research For Nursing
Footnotes
Acknowledgements
We thank Dr. Joachim Voss and the laboratory staff for their RNA and microarray expertise.
Ethical Consideration
All experimental protocols were approved by the University of Nevada, Las Vegas Institutional Animal Care and Use Committee (IACUC) and the United States Air Force Animal Use Programs Office of Research Oversight and Compliance.
Consent for Publication
The review and publication decisions for this manuscript were handled solely by the editor, Carolyn Yucha.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the National Library of Medicine of the National Institutes of Health under Award Number R25LM014215 (Principal Investigator, Gabriela Mustata Wilson, PhD, MSc, FHIMSS, FIAHSI, SNAI), as well as by grants from the National Institute of General Medical Sciences GM103440 (Principal Investigator, Josh Baker, PhD) and 5 U54 GM104944 (Principal Investigator, Francisco Sy, MD, MS, DrPH) from the National Institutes of Health. This work was also supported by a Department of Defense Air Force Grant (FA-7014–10-2–0001) awarded to Dr. Barbara St. Pierre Schneider. Review of this material does not imply Department of the Air Force endorsement of factual accuracy or opinion. The content is solely the responsibility of the authors and does not necessarily represent the official views of the funders.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
NCBI GEO accession number GSE311704.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
