Abstract
Background
Breast milk is essential for premature infants. It contains a variety of functional and protective nutrients that help to create a suitable microenvironment for intestinal development and maturation. This prospective and observational study was planned to examine the effects of feeding premature infants with fresh or frozen breastmilk on their gut microbiota.
Materials and Methods
The study was carried out with a total of 40 infants, with a gestational age of 28–326 weeks, fed fresh (n = 20) or frozen (n = 20) breastmilk. Stool samples were stored at −80°C until analysis. Infants were included in groups based on the feed type, which accounted for more than 70% of their 10-day feeding. The Mann-Whitney U, Chi-square, and t-tests were used to evaluate the demographic data. Stool samples were analyzed by sequencing the V3–-V4 region of the 16S rRNA gene from the extracted DNA for microbiota analysis.
Results
Streptococcus and Enterobacteriales, the majority of which are considered human pathogens, in infants receiving frozen breastmilk (Streptococcus 69%, Enterobacteriales 79%) was higher than that in infants receiving fresh breastmilk (Streptococcus 16%, Enterobacteriales 49%). Further, the Lactobacillus and Bifidobacterium species were more abundant in infants who received fresh breastmilk (Lactobacillus 17%, Bifidobacterium 12%) than in infants who received frozen breastmilk (Lactobacillus 3%, Bifidobacterium 1%).
Conclusion
Thus, compared to frozen breast milk, fresh breastmilk has an effect on the diversity of preterm infants’ gut microbiota.
Introduction
Approximately 15 million infants are born before 37 weeks of gestation each year (WHO, 2018). Most of these infants stay in neonatal intensive care units (NICU) for days or even months, depending on their individual needs (El-Metwally & Medina, 2020). Nutritional insufficiency is among the most important problems faced by premature infants (Kumar et al., 2017). Breast milk is essential for all newborns and especially for premature infants. It contains a variety of functional and protective nutrients that help to create a suitable microenvironment for intestinal development and maturation (Cabrera-Rubio et al., 2012).
Microbiota refers to all microorganisms living in the human body (Xu & Knight, 2015). The gastrointestinal tract offers the most suitable environment for bacterial colonization because of its large surface area extending from the esophagus to the rectum and rich nutrients (Hillman, et al., 2017). In the first month of life, bacteria from breast milk and the skin flora of the mother predominate the intestines of newborns. Approximately 40% of intestinal bacteria are formed in the first month of breastfeeding (Pannaraj et al., 2017).
Nurses are primarily responsible for feeding the infants treated in the NICU. It is important for neonatal nurses to know the factors that affect the development of microbiota in the newborn, such as nutrition and their role in diseases, from the time of newborn’s admission to the NICU. There are some important care methods that neonatal nurses can apply for the development of an optimal microbiota, and feeding fresh breast milk to infants is one of them (Rodriguez et al., 2017).
Fresh breast milk ensures that the intestine is colonized with beneficial bacteria, especially the probiotic bacteria that use oligosaccharides from breast milk for nutrition. Freezing breast milk has been reported to affect its immunoprotective components and bacterial composition (Sherman et al., 2015). Fresh breast milk is rich in both nutrients and antibodies. However, breast milk is usually frozen and used when direct breastfeeding is not possible, especially for infants monitored in the NICU (Hung et al., 2018). Therefore, this study aimed to determine the effects of feeding fresh or frozen breast milk on the gut microbiota of premature infants.
Methods
Design
This was a prospective observational study. Infants were divided into two groups: (1) those who received fresh breast milk, and (2) those who received frozen breast milk. They were grouped based on the predominant feeding solution (>70% fresh or frozen breast milk) after 10 days of feeding.
Sample Criteria
The research population consisted of premature infants monitored in the third level NICU between December 2018 and February 2020. Inclusion criteria for newborns were as follows: (1) gestational age between 28 and 32 weeks at delivery, (2) treatment with aminoglycoside and cephalosporin group of antibiotics, (3) first day after birth, (4) born by planned cesarean section, and (5) infants fed by means of an orogastric tube. The inclusion criteria for mothers were: (1) 18 years of age and older, and (2) no antibiotic use during pregnancy. The exclusion criteria for newborns were as follows: (1) vaginal birth, (2) cesarean section after the onset of labor, (3) suspected necrotizing enterocolitis, (4) not fed enterally, (5) received a different type of antibiotic treatment other than aminoglycoside and cephalosporin, (6) given any probiotic product that may affect the intestinal flora, (7) receiving kangaroo care, (8) meconium and stools on the 10th day not collected by the researcher, and (9) receiving infant formula for more than 70% of their 10-day feeding. The exclusion criteria for mothers were as follows: (1) taking any probiotic product during pregnancy and lactation, (2) diagnosed with premature rupture of membranes during pregnancy, (3) younger than 18 years of age, (4) inability to provide fresh or frozen breast milk, and (5) regular drug use due to chronic illness during pregnancy and lactation. All infants in the unit were evaluated for eligibility to be included in the study group.
A total of 235 infants, 28–32 weeks old, were admitted to the NICU of the hospital during the specified span, of which 195 infants were excluded from the study based on the exclusion criteria. Due to the limited number of studies on the subject, 20 infants were included in the study, considering the sample numbers of similar studies. The study comprised 20 infants in each group considering equivalence in terms of gender and gestational week (Akagawa et al., 2019; Cong et al., 2017; Ma et al., 2020).
Storage and Preparation of Breast Milk
Three deep freezers (−8°C) and a refrigerator (4°C) were used to store breast milk in the unit. The neonatal nurses informed the mothers about storage and delivering their milk to the unit. Mothers brought their milk to the unit in fresh or frozen form with milk storage bags. The infant’s details were recorded on breast milk bags. Fresh breast milk brought to the unit was stored at 4°C for a maximum of 3 days (Bilgen et al., 2018), while frozen milk was stored in the deep freezer (−18°C). The infant was given fresh milk if the mother brought fresh breast milk. The frozen breast milk was thawed at 4°C for 2–3 hours prior to feeding. It was then placed in warm water before feeding it to the infant. Neonatal nurses ensured that the infants were fed fresh or frozen breast milk.
Data Collection Procedure
In addition to ethical approval from the University Clinical Research Ethics Committee (number: E.55091, date: 30.11.2018), permission to conduct this study was obtained from the hospital. All neonatal nurses and doctors in the unit were informed about the study. During the data collection phase, the parents of the newborns who met the research criteria were interviewed. Information about the research was provided and written and verbal consent was obtained.
A ‘Descriptive Information Form’ and ‘Infant Follow-up Form’ were used to collect data. The descriptive information form was used to gather the demographic characteristics of the newborns and their mothers. It was completed by the mothers and the information for the newborn was filled in by the researcher from the infant’s file. In the Infant Follow-up Form, information was recorded daily (for 10 days) to determine the frequency of feeding the infant, amount of feeding solution taken by the infant, characteristics of the feeding solution (fresh breast milk, frozen breast milk, and infant formula), and time of stool sample collection.
Nutrition
The type and amount of feeding of the infants were recorded by neonatal nurses on the nurse observation form. These records were followed and updated daily in the Infant Follow-up Form. The groups (fresh or frozen breast milk group) in which infants were placed were determined based on the characteristics and amount of 10-day feeding solutions. Infants who received more than 70% of their 10-day feeding with fresh breast milk were included in the fresh breast milk group, while those who received more than 70% of frozen breast milk in their 10-day feeding were included in the frozen breast milk group. Infants whose 10-day feeding was more than 70% of infant formula were excluded from the study (Cong et al., 2017). No intervention was made on the feeding style of any of the infants included in the study and infants were fed routinely. The meconium samples of all infants who were included in the study were collected.
Collection of Sterile Stool Samples
The researchers received online training on collecting, storing, and transporting stool samples with appropriate techniques. The training was given by the bioinformatics specialist in the laboratory where the microbial analysis of stool samples was conducted.
Stool documents were taken by the researcher (first author) using the stool collection protocol. The protocol includes the following content: 1. Hands should be thoroughly washed before collecting stool samples. 2. The diaper should open flat. 3. The stool container with a sterile spoon should be opened carefully and left in a clean area with the inside of the lids on top. 4. There are enough stool samples to fill the cloth frames by sneakily scooping the lids of the sterile stool container. 5. The collected stool sample should be placed in a sterile cabinet and the lids should be closed. 6. Name, date, time, and work hierarchies should be checked and labelled according to sterile stool rules. 7. Stool samples that are not taken immediately are stored in a −80°C freezer (McInnes & Cutting, 2010).
Gut Microbiota Analysis
Novogene Genome Sequencing Company performed gut microbiota analyses. The stool samples were analyzed by sequencing the V3–-V4 region of the 16S rRNA gene from the extracted DNA. QuickGene (tissue DNA extraction kit) extraction device was used during the extraction; a 25 mg stool sample was extracted during the application by following the kit protocol. As a result of the extraction process, genomic DNA was obtained by dilution with 200 μL CDT (elution buffer). Nanodrop measurements were made using the Colibri Titertek Berthold device and Qubit ™ dsDNA HS Kit (ThermoFisher, PicoGreen® Dye) Qubit Fluorometer 2 device to determine the DNA quality. Following the measurement process, the 16S rDNA V4 region in the bacterial DNAs was amplified by PCR, and sequencing was performed on the HiSeq platform (Illumina) by following the 2 × 250 bp pair-end protocol, in which approximately the entire region overlaps. After sequencing, read-pairs with unique molecular barcodes were separated and pair-end reads were combined using FLASH (V1.2.7). Clean taqs with high-quality filters in raw taqs were obtained according to the QIIME (V1.7.0) quality control process. These taqs were compared with the reference database (Gold database) for the detection of chimera sequences using the UCHIME algorithm and the CHIMERIC sequences were extracted. Finally, effective taqs were obtained. In addition, a quality filter was applied to the combined read results and those above the expected p-value of .05 were eliminated. 16S rRNA gene sequences were separated into OTU clusters with 97% similarity as cut-off using the UPARSE (Uparse v7.0.1001) algorithm. OTUs were mapped with the optimized version of the SILVA database containing only 16S V4 regions to decide on taxonomic classes. Intensities were obtained by mapping demultiplexed reads using UPARSE OTUs. MUSCLE (Version 3.8.31), which can compare multiple sequences, was used to obtain the phylogenetic relevance of the representative sequences of all OTUs. Thus, relative species, richness, and distribution of abundance could be analyzed with images such as alpha diversity (observed species, ACE, Chao 1, Shannon and Simpson), beta diversity (Unweighted UniFrac and Weighted UniFrac), and taxa-bar plots. On the other hand, multiple sequence alignments were performed with the OTU clustering results, and phylogenetic trees were constructed. Differences between samples or groups related to the structure of the microbial community were explained by dimension reduction (PCA and NMDS). Statistical methods such as LEfSe, Anosyme, and MRPP were tested with the significance of microbial community structure difference between groups. With the total sum scaling normalization approach, percentages were calculated, and relative abundance values were determined.
Data Analysis
Data were analyzed using IBM SPSS (Statistical Package for Social Sciences Statistics) version 22.0. The conformity of the research data to the normal distribution was evaluated using the Shapiro-Wilk test. The statistical significance was set at p < .05. Percentage, median, standard deviation, and minimum–maximum values were used to determine the distribution of descriptive features. The Mann–Whitney U test was used to compare the continuous variable data that did not show a normal distribution, while the chi-square test was used for the comparison of categorical data. A t test was used for normally distributed independent groups.
Results
Infant Characteristics
Characteristics of the Infants in the Groups.
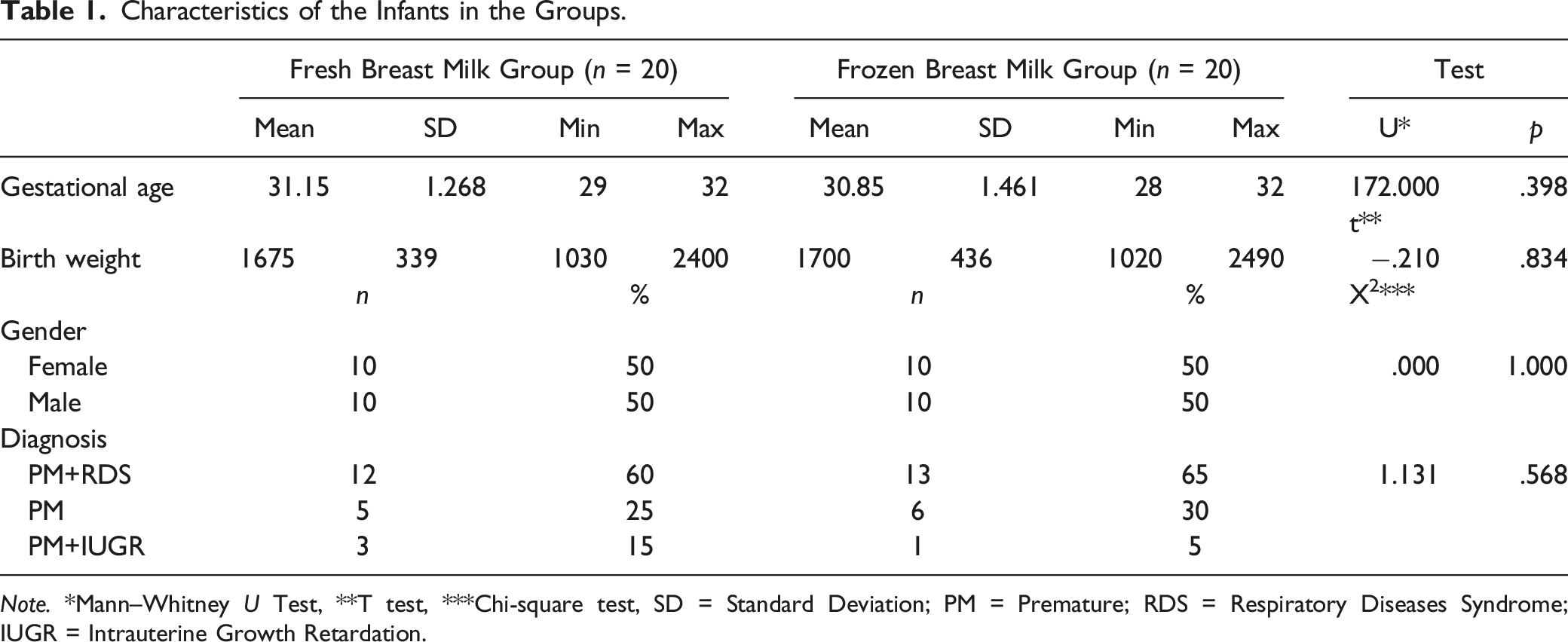
Note. *Mann–Whitney U Test, **T test, ***Chi-square test, SD = Standard Deviation; PM = Premature; RDS = Respiratory Diseases Syndrome; IUGR = Intrauterine Growth Retardation.
Characteristics of the Mothers
Characteristics of the İnfants’ Mothers in the Groups.
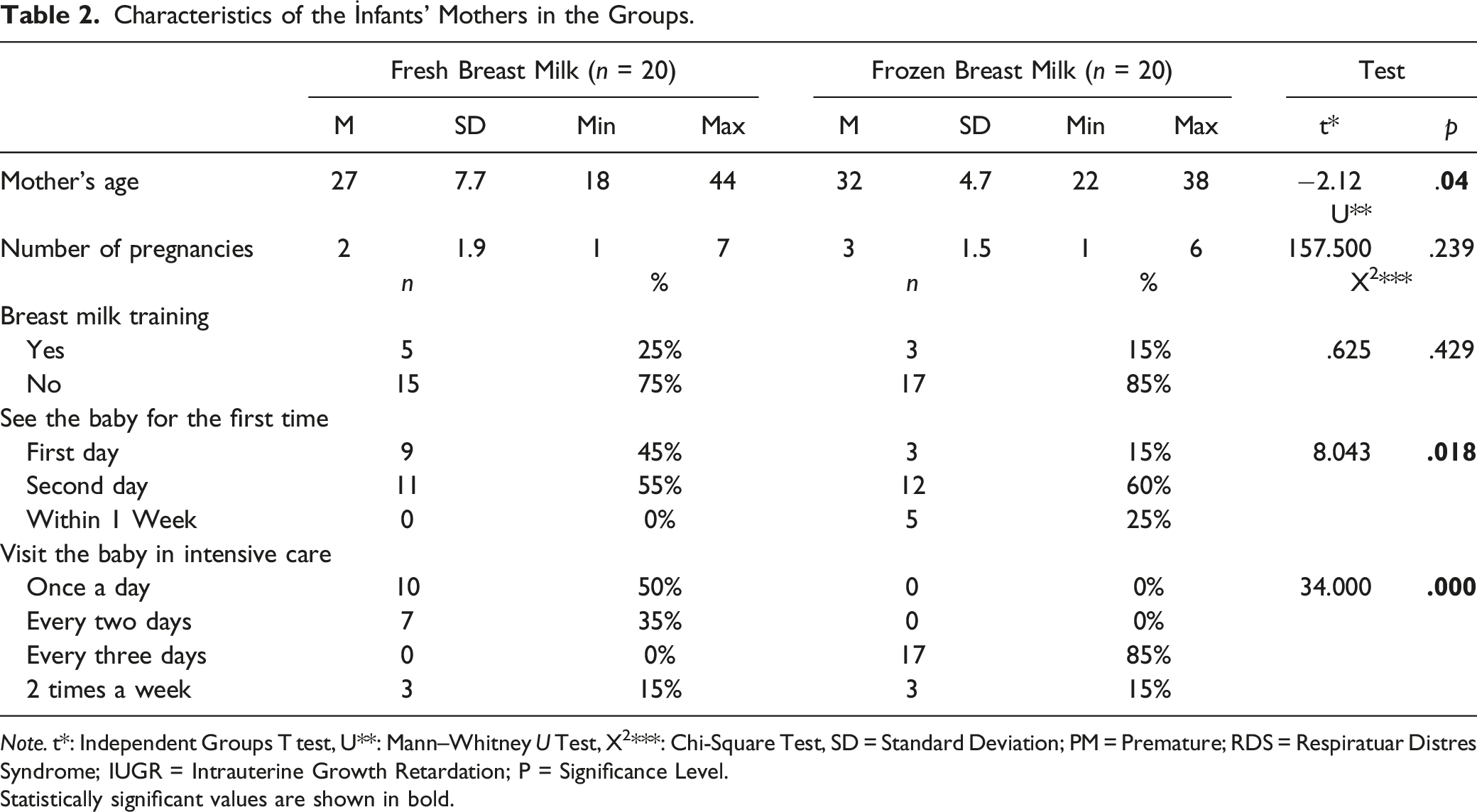
Note. t*: Independent Groups T test, U**: Mann–Whitney U Test, X2***: Chi-Square Test, SD = Standard Deviation; PM = Premature; RDS = Respiratuar Distres Syndrome; IUGR = Intrauterine Growth Retardation; P = Significance Level.Statistically significant values are shown in bold.
Gut Microbiota Analysis of the Infants
Eighty stool samples were collected from 40 premature infants as meconium and stool samples on the 10th day. Microbiota analysis was performed for 52 stool samples (40 stool samples on the 10th day, 12 meconium samples). The statistical significance between the different groups in the microbiota analysis was assessed using Adonis, Anosim, and multiple response permutation procedure (MRPP) analysis (Warton et al., 2012).
Adonis and Multiple Response Permutation Procedure (MRPP) Grouping Analysis.
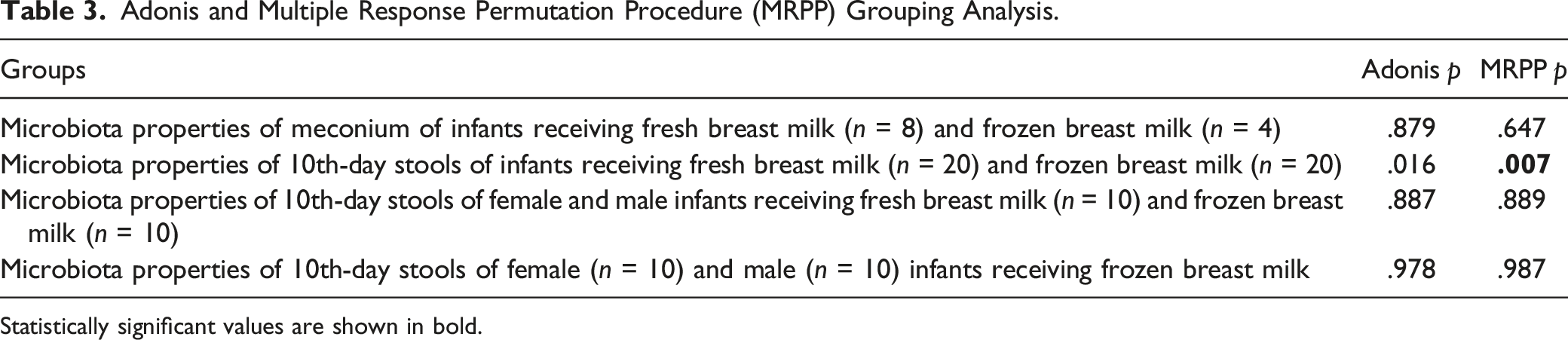

Statistically significant values are shown in bold.
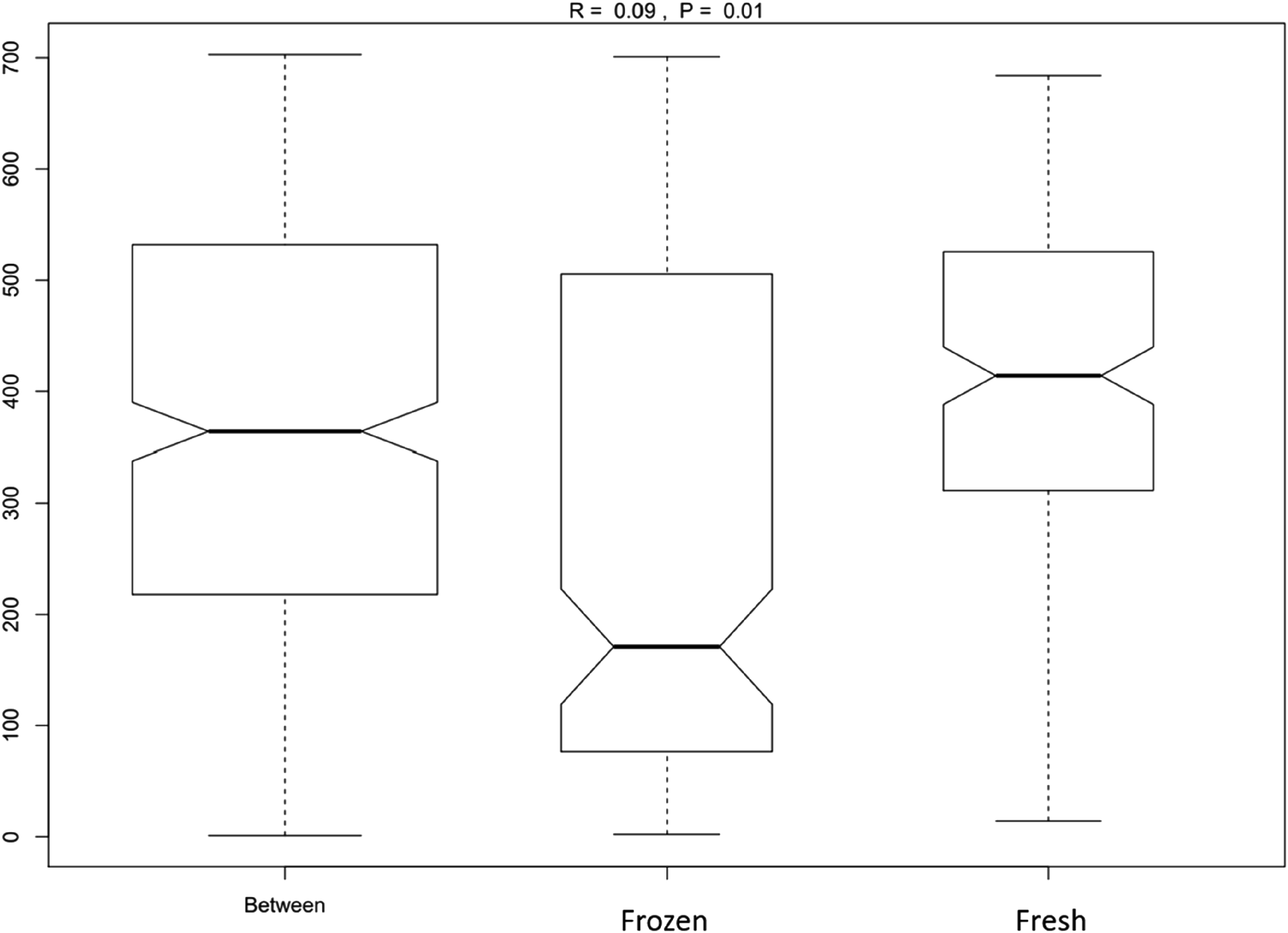
Anosim grouping analysis.
Alpha diversity is the analysis of microbial community diversity, and the richness is assessed in each sample through a series of statistical analysis (Bokulich et al., 2013). The alpha diversity of the gut microbiota from the stool samples of the infants are displayed in box plots. The meconium and 10th-day stool microbiota of all infants who received fresh and frozen breast milk did not show a difference in the alpha diversity (Figure 2, p > .05). Comparison of groups in terms of alpha diversity.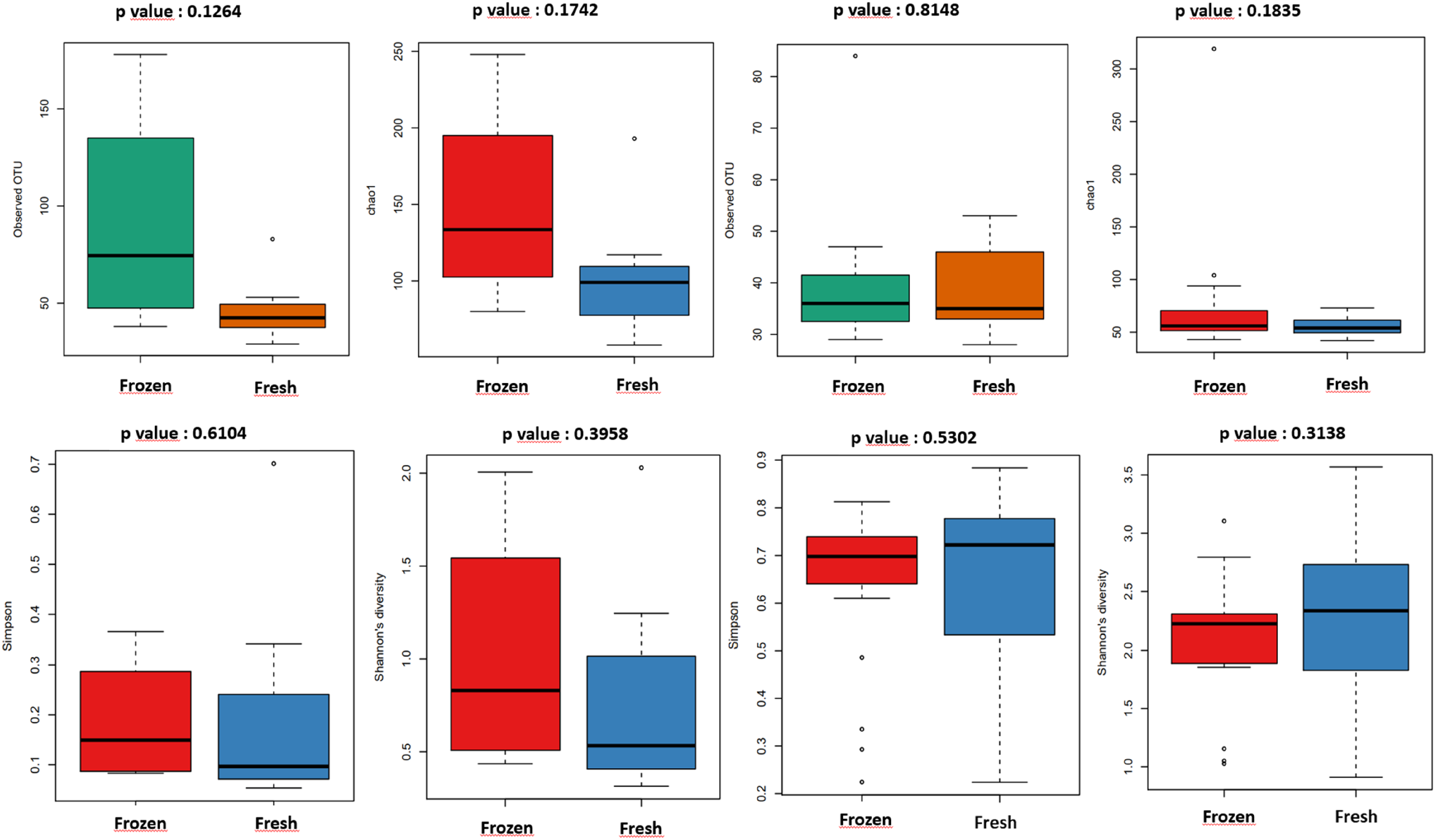
Beta diversity represents a comparison of microbial communities based on composition, where a square ‘distance’ or ‘difference’ matrix is calculated to reflect the differences between samples (Baselga, 2010). The data in this distance matrix is displayed with Principal Component Analysis (PCA). PCA describes how the groups differ in their characteristics. The microbiota content of the meconium of infants in the fresh and frozen breast milk groups was investigated using PCA, performed according to operational taxonomic unit (OTU) values. Although microorganisms with different characteristics were found in both the fresh and frozen breast milk groups, these groups were found to cluster close to each other and were therefore similar (Figure 3a). However, in the microbiota analysis of the 10th-day stools of all infants who received fresh and frozen breast milk, the samples from the infants in frozen breast milk group contained microorganisms with different characteristics, and the groups clustered far from each other. In contrast, the gut microbiota of infants who received fresh breast milk contained microorganisms with similar characteristics, and the groups clustered close to each other (Figure 3b). Comparison of groups in terms of beta diversity.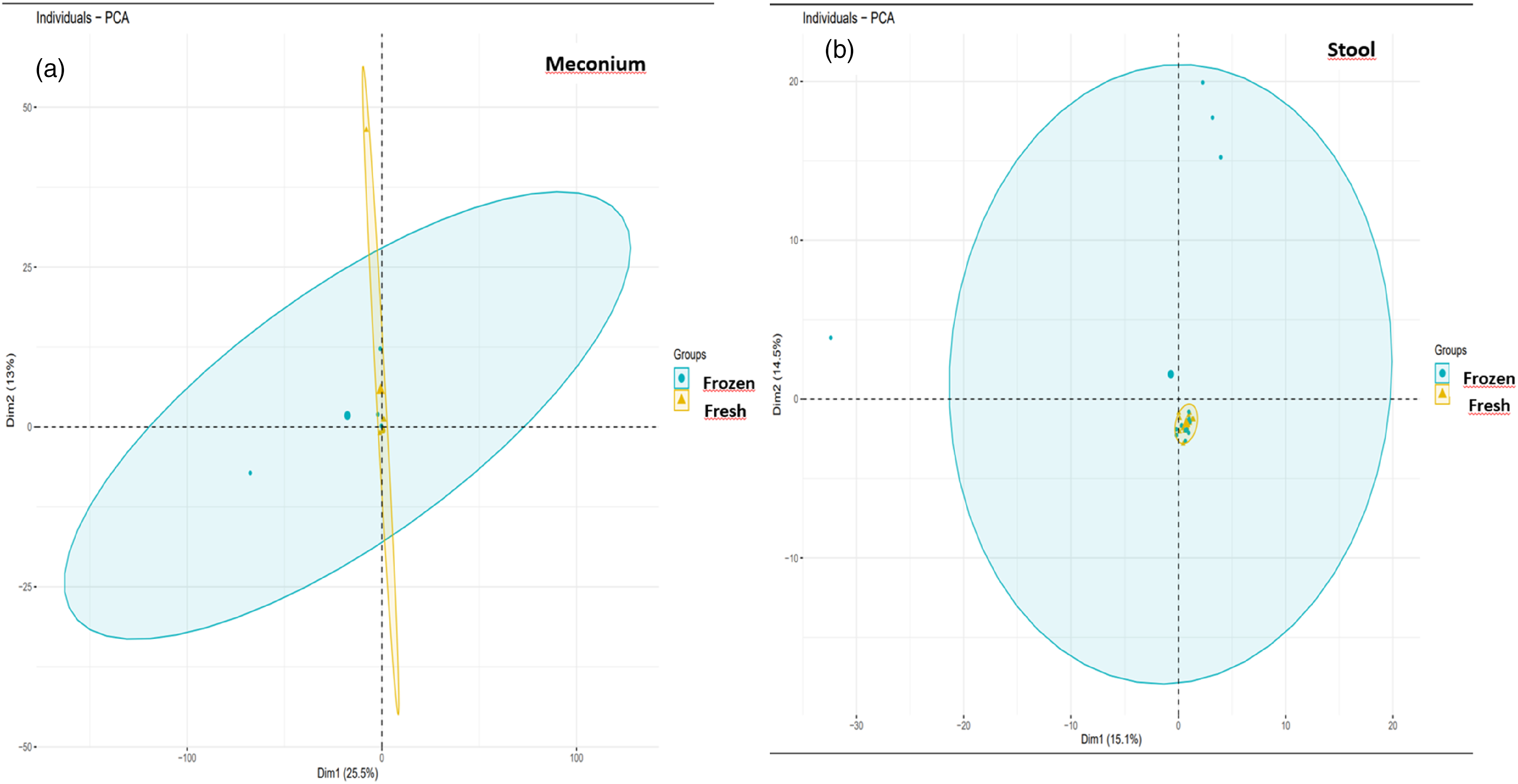
The similarities and differences between the groups in the microbiota analysis for the top 35 genera was represented in the form of a heat map. An increased abundance was observed for Sphingomonadaceae, Rhizobiales, Enterobacteriaceae, and Sphingomonadaceae in the infant meconium samples (Figure 4). The 10th-day stool samples of the frozen breast milk group showed an increase in the abundance of Enterobacteriaceae, which was found to be decreased in the fresh breast milk group. Abundance was observed in Lactobacillus, Bifidobacterium longum, Acinetobacter, Streptococcus, Enterococcaceae, Bacilli, and Clostridium hathewayi genera in the 10th-day stools of infants from the fresh breast milk group, and in the Enterobacteriaceae, Streptococcus, and Gemellales genera in the samples from the frozen breast milk group (Figure 4). Heat map of taxonomic abundances of groups at genus level.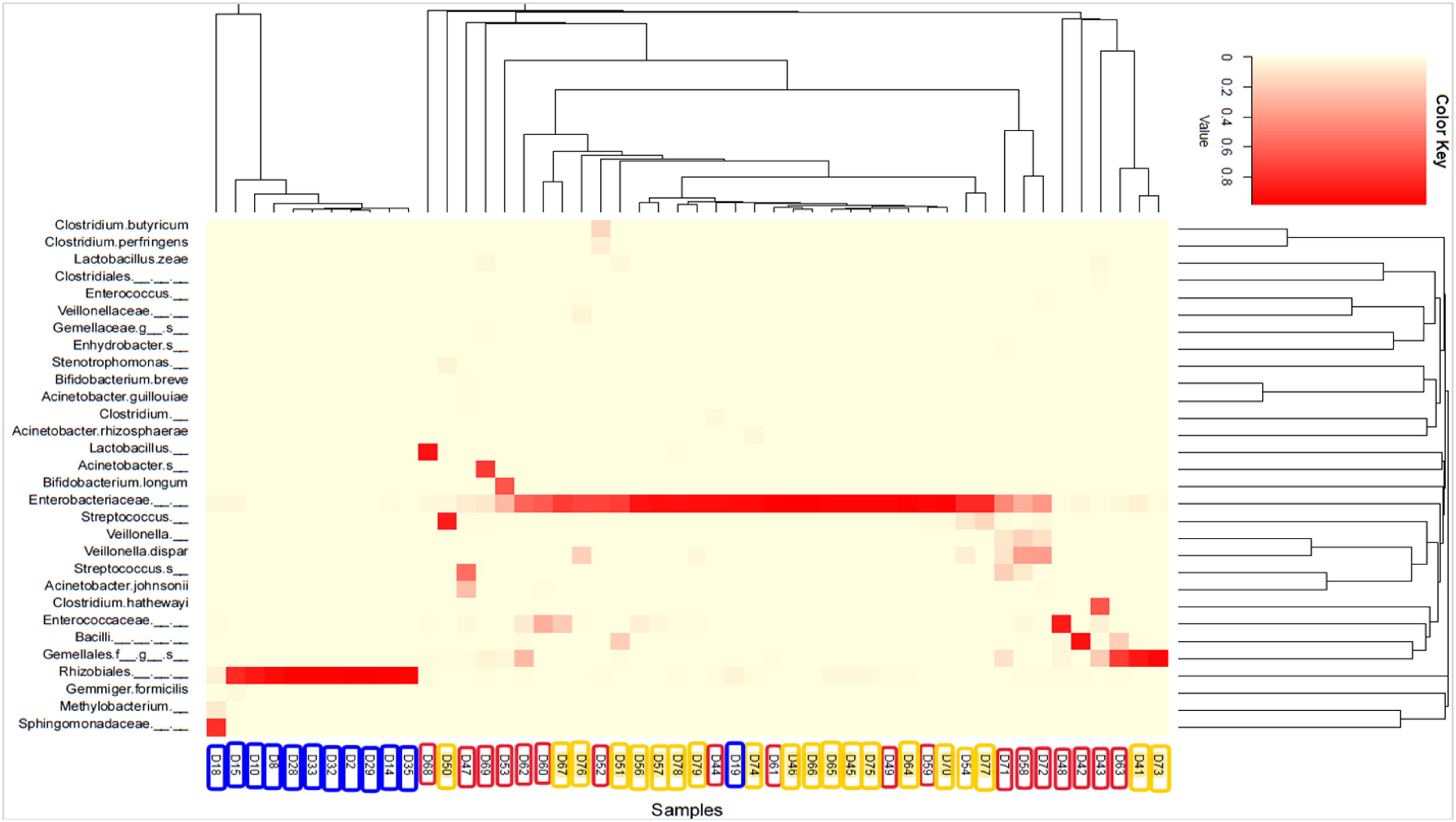
Relative abundance analyses of infant gut microbiota by taxa-bar plots at the phylum and genus levels show abundances of Proteobacteria (fresh: 99%, frozen: 96%) and Firmicutes (fresh: .8%, frozen: 3%) phyla in the meconium of infants in the fresh and frozen breast milk groups were found to be similar (Figure 5a). The meconium of infants in the fresh breast milk group showed a relative abundance of Methylobacterium (58%) genus compared to that of the frozen breast milk group (Figure 5b). Relative changes in the abundance at the phylum and genus level in meconium and 10th-day stool samples of infants receiving fresh and frozen breast milk.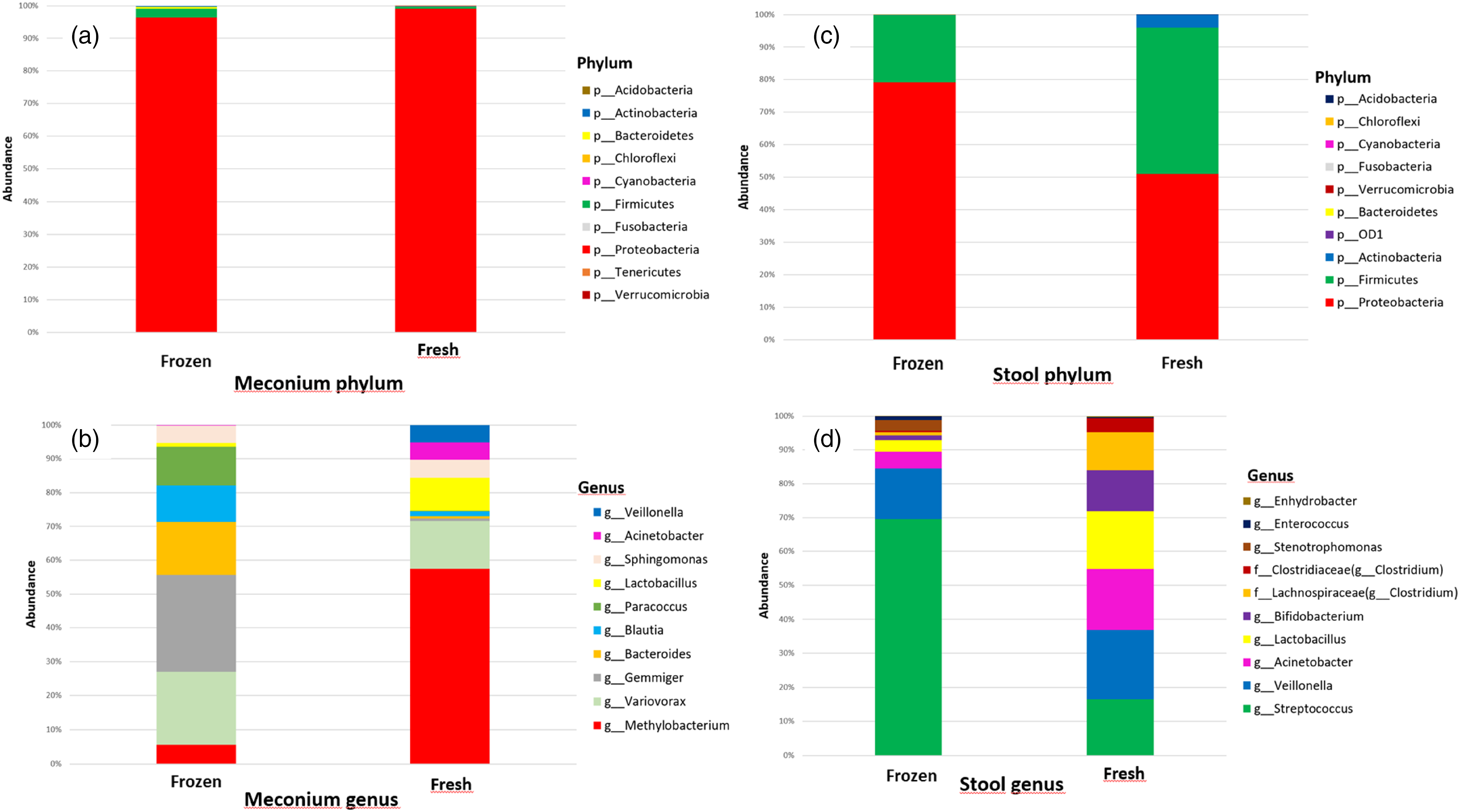
The phylum Proteobacteria ranked first in the 10th-day stool microbiota of infants receiving fresh breast milk (51%) as well as in the frozen breast milk group (79%). This was followed by Firmicutes (45%) and Actinobacteria (4%) in the fresh breast milk group and only Firmicutes (21%) in the frozen breast milk group (Figure 5c). Veillonella (20%) was the predominant genus in the 10th-day stool microbiota of infants receiving fresh breast milk. This was followed by Acinetobacter spp. (18%), Lactobacillus spp. (17%), Streptococcus spp. (16%), and Bifidobacterium spp. (12%). Streptococcus (69%) was the predominant genus in the 10th-day stool microbiota of infants who received frozen breast milk. This was followed by Veillonella (15%), Acinetobacter (5%), Lactobacillus (3%), Stenotrophomonas (3%), Bifidobacterium (1%) and Enterococcus (1%). The relative abundance of the Streptococcus genus in the 10th-day stool microbiota of infants receiving frozen breast milk was higher than that of infants receiving fresh breast milk. In contrast, the relative abundance of Acinetobacter (18%), Lactobacillus (17%), and Bifidobacterium (12%) genera was higher in the 10th-day stool of infants who received fresh breast milk than that of the frozen breast milk group (Figure 5d).
When the 10th-day stool microbiota of the infants who received fresh breast milk were evaluated in terms of gender, the most abundant phyla were Proteobacteria (females: 42%, males: 60%), Firmicutes (females: 50%, males: 40%), and Actinobacteria (female: 8%, male: .3%; Figure 6a). When the 10th-day stool microbiota of the infants who received fresh breast milk were evaluated in terms of gender and genus level, Acinetobacter (female: 1% male: 31%), Streptococcus (female: 5%, male: 25%), Veillonella (female: 21%, male: 20%), and Lactobacillus (female: 37%, male: 2%) were found to be the most abundant genera. Although the microbiota content of the 10th-day stools of the fresh breast milk group was similar in both genders, there were also differences. For example, f_Lachnospiraceae (g_Clostridium) (20%) in male infants and Bifidobacterium (26%) and f_Clostridiaceae (g_Clostridium) (8%) genera in female infants were more abundant (Figure 6b). Relative changes in the abundance at the phylum and genus level in the 10th-day stool samples of infants receiving fresh and frozen breast milk based on gender.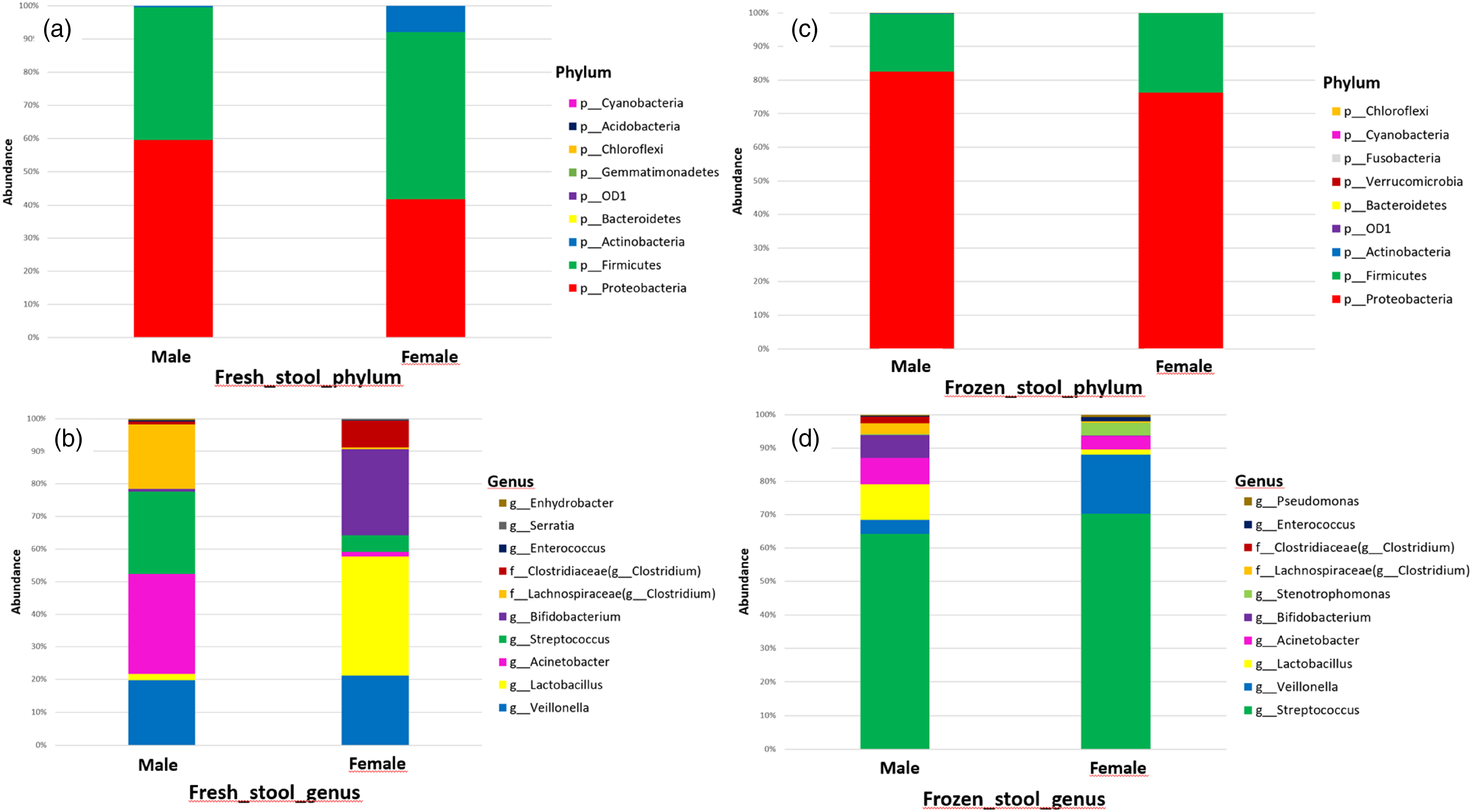
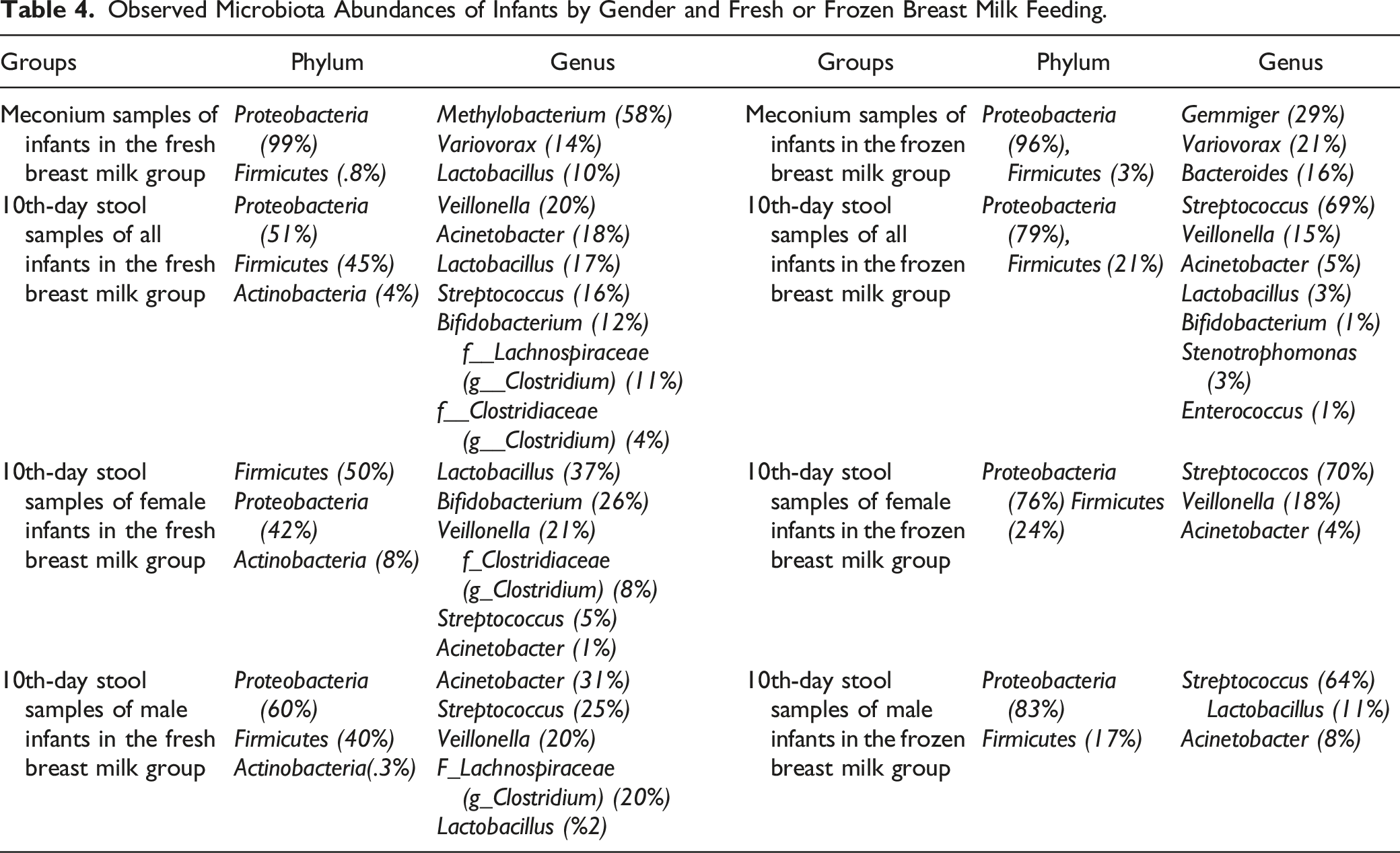
Observed Microbiota Abundances of Infants by Gender and Fresh or Frozen Breast Milk Feeding.
Discussion
In this prospective observational study, feeding fresh or frozen breast milk was found to influence the gut microbiota of premature infants. The groups did not differ in alpha diversity. This is thought to be due to the fact that the study was conducted on newborns who have not yet reached full maturity in terms of microbiota. According to the beta diversity PCA analysis, based on the comparison of the microbial content of the 10th-day stools of infants who received fresh and frozen breast milk, infants' microbial who received fresh breast milk clustered together. In contrast, those who received frozen breast milk did not show clustering because they contained different microorganisms. This can be attributed to the breakdown of bacteria due to the freezing and thawing of breast milk (Shao et al., 2012).
Proteobacteria, the predominant phylum in the meconium microbiota of infants in this study, was highly abundant in both fresh and frozen breast milk groups. This was followed by Firmicutes. The findings obtained from this study aligned with those in the literature. Similar to our study, Stinson et al. (2019) analyzed amniotic fluid samples from 50 mothers who gave birth by non-urgent cesarean section and did not have an intrauterine infection as well as the meconium samples from infants at 34–42 weeks of gestation. They observed that the abundance of Pelomonas puraquae bacteria from the Proteobacteria phylum was high in 43 meconium samples. Dornelles et al. (2020) evaluated the microbiota of the meconium of 84 premature babies younger than 32 weeks of gestation, of which 40 were diagnosed with early onset neonatal sepsis and 44 were not diagnosed. The most abundant phylum in both the groups was Proteobacteria. All the studies suggest Proteobacteria to be the most dominant phylum in the neonatal meconium.
A greater diversity of bacteria was observed in the 10th-day stools of infants who received fresh breast milk (Table 4). Although there were proportional differences in the 10th-day stools of all infants who received fresh and frozen breast milk, Proteobacteria and Firmicutes were observed at the phylum level in both the groups. Considering that the gut microbiota developed from 4 phyla, including Proteobacteria and Firmicutes (Yang et al., 2016) and the infants in our study were fed mainly with fresh or frozen breast milk, it is speculated that the difference may arise from the infants receiving fresh breast milk because the milk had not been exposed to any process such as freezing.
The harmful effects of freezing on bacteria have been attributed to the formation of intracellular ice (Corbett et al., 2018). However, Fonseca et al. (2006) reported that cell damage due to freezing may be a result of intracellular and extracellular osmotic imbalances during warming of the milk. Various studies have shown that freezing breast milk causes changes in milk content and bacterial diversity (Fonseca et al., 2006; García-Lara et al., 2012). García-Lara et al. (2012) found that freezing breast milk at −20°C for 90 days reduced the amount of fat, protein, and calories compared to fresh breast milk. In a study of 18 premature infants younger than 37 weeks of gestation, Hung et al. (2013) provided fresh and thawed breast milk for 2 consecutive feeding hours and then measured their heart rate and oxygen saturation before and after feeding. They found more signs of stress in premature infants fed thawed breast milk than in those fed fresh breast milk.
Commensal bacteria in breast milk play a protective role against gastrointestinal infections in infancy (Carr et al., 2021). In general, a high abundance of beneficial bacteria, like Bifidobacterium and Lactobacillus, is more effective in the formation of healthy microbiota in preterm infants than pathogenic bacteria such as Escherichia coli, Enterobacter, Citrobacter, Proteus, Klebsiella, and Candida. Breast milk oligosaccharides are among the most beneficial prebiotics for premature infants, facilitating the growth of Bifidobacteria and Lactobacillus (Xu et al., 2018). Bifidobacteria and Bacteroidetes have been reported to be dominant in the gut microbiome communities of breastfed infants (Groer et al., 2014). Bacteria from the breast milk stimulate the maturation of cytotoxic Th1 cells and strengthen the immune system against infections (Carr et al., 2021). In particular, Lactobacillus in breast milk increases the number of natural killer cells and CD4+ and CD8+ T cells, which are important for immune functions (Fernández et al., 2013). Commensal bacteria such as Lactobacillus gasseri and Lactobacillus crispatus in breast milk have been reported to have an ability to adhere to intestinal cells, thereby increasing the colonization of beneficial bacteria in the gut of the breastfed infants. Damaceno et al. (2017) also found that Bifidobacterium breve, Lactobacillus gasseri, and Streptococcus salivarius inhibited the attachment of pathogenic bacteria to intestinal epithelial cells. Some differences were observed in our study in terms of bacterial diversity and abundance in the 10th-day stools of babies who received fresh or frozen breast milk. Lactobacillus and Bifidobacterium, which are the beneficial bacteria expected to be present in breast milk, were observed to be more abundant in infants receiving fresh breast milk. The abundance of Bifidobacterium in stool is shown to be increased in infants fed healthy breast milk in the early stages of life (Borewicz et al., 2019). Another study associated high Bifidobacterium abundance in the first week of life with higher levels of IL-13, IL-5, IL-6, and IL-1b in children later in life (Rabe et al., 2020). Consistent with these studies, Bifidobacterium abundance was observed in the 10th-day stools of infants who received fresh breast milk in our study, compared to infants who received frozen breast milk (Table 4).
It is thought that beneficial bacteria, such as Bifidobacterium and Lactobacillus, are less abundant in infants receiving frozen breast milk, which may be related to the freezing of breast milk. Streptococcus (Reglinski & Sriskandan, 2015), mostly found as the skin flora and the majority of which is considered to be a human pathogen and disease agent, showed the highest abundance in the 10th-day stools of the frozen breast milk group. Lactobacillus, which is found in healthy intestinal flora, can adhere to the intestinal epithelial layer. Therefore, they are known to remove pathogenic microorganisms (Kwon et al., 2018). Further, due to the lower Lactobacillus ratio in the group receiving frozen breast milk, pathogenic microorganisms adhere more easily to the infant’s gut, and therefore, the abundance of pathogenic bacteria such as Streptococcus is higher in these infants.
Another order classified as pathogens in humans and animals is Enterobacteriales (Adeolu et al., 2016). In their study of 33 premature infants, Cong et al. (2017) evaluated the gut microbiota according to the feeding patterns and found that breastfed babies had the highest abundance of Clostridiales, Lactobacillales, and Bacillales. However, they observed a high amount of Enterobacteriales in infants fed donor milk or infant formula. Although the results of our study were similar to those of this study, a higher abundance of Enterobacteriales (79%) was observed in the 10th-day stools of babies who received frozen breast milk. This result shows that frozen breast milk can create a microenvironment that allows the development of more pathogenic microorganisms in the infant gut than fresh breast milk.
Various factors, such as nutrition, race, age, antibiotic use, stress, psychological factors, maternal health, mode of delivery, environmental factors, and exercise, can affect the state of the gut microbiota (Osadchiy et al., 2019). Along with these factors, gender is thought to be an important factor in the development of microbiota. Although puberty is thought to affect the composition of the microbiota due to hormonal changes, there are not enough data to support this argument (Martin et al., 2016).
The differences observed in our study can be attributed to both the freezing of breast milk as well as the differences between the genders (Table 4). Haro et al. (2016) found that gut microbiota differs in males and females at the phylum level (Firmicutes/Bacteriodetes ratio), genus level (Bacteroides, Bilophila, Veillonella, and Methanobrevibacter), and species level (B. plebeius, B. caccae, and C. catus). Similarly, a higher diversity of bacteria was observed in our study in the 10th-day stools of both female and male infants who received fresh breast milk than in those of infants who received frozen breast milk.
At the genus level, Lactobacillus was observed in greater abundance in the 10th-day stools of female infants who received fresh breast milk compared to both male infants who received fresh breast milk and all infants who received frozen breast milk (Table 4). Similar to our findings, Martin et al. (2016) found that the total bacterial count in the gut of male infants at birth was higher than that of female infants. They found that Lactobacillus, a dominant genus in the vaginal microbiota and known to be regulated by female estrogens (Romero et al., 2014), colonizes the gut of girls more frequently than that of boys at an early age. Contrary to these findings, Lactobacillus was not observed at the genus level in the 10th-day stools of female infants receiving frozen breast milk, whereas it was found in greater abundance in the 10th-day stools of male infants (Table 4).
Limitations
As this study was conducted on premature infants receiving antibiotic treatment, the results cannot be generalized to healthy term infants where antibiotics were not administered. Forty meconium samples were obtained from 40 infants who received fresh and frozen breast milk, but microbiota analysis could be performed in only 12, while 28 meconium samples could not be analyzed. For this reason, only the analysis results of the 12 meconium samples were considered. As meconium analysis in microbiota research is more difficult than stool analysis, this is one of the limitations of this study.
Conclusion
Dietary patterns of premature infants have a profound effect on the gut microbiota. The study established that feeding fresh breast milk influences the gut microbiota of newborns, and the abundance of beneficial bacteria is higher than that in frozen breast milk. Therefore, feeding fresh breast milk should be widespread in all full-term and premature infants. In this regard, all mothers should be informed about the importance of breastfeeding during pregnancy and after delivery and should be provided with the necessary support. Health professionals should be involved in spreading awareness. Clinical conditions should be improved and NICU facilities should be organized accordingly. Frozen breast milk should be used only if fresh breast milk cannot be provided to infants, and infant formula should be considered as a last resort. Our study is the first to provide substantial evidence on the effects of fresh and frozen breast milk on the gut microbiota of newborns. It highlights the importance of microbial composition and aids in filling the gap in the field.
Footnotes
Acknowledgments
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Gazi University Scientific Research Projects Unit with project code 47/2019-11.
Research ethics
In addition to ethical approval from the University Clinical Research Ethics Committee (number: E.55091, date: 30.11.2018), permission was obtained from the hospital management to conduct the study.
