Abstract
Objectives
This aim of this study was to characterize the composition and content of the feline urine metabolome.
Methods
Eight healthy domestic cats were acclimated at least 10 days before starting the study. Urine samples (~2 ml) were collected by ultrasound-guided cystocentesis. Samples were centrifuged at 1000 × g for 8 mins, and the supernatant was analyzed by gas chromatography/time-of-flight mass spectrometery. The urine metabolome was characterized using an untargeted metabolomics approach.
Results
Three hundred and eighteen metabolites were detected in the urine of the eight cats. These molecules are key components of at least 100 metabolic pathways. Feline urine appears to be dominated by carbohydrates, carbohydrate conjugates, organic acid and derivatives, and amino acids and analogs. The five most abundant molecules were phenaceturic acid, hippuric acid, pseudouridine phosphate and 3-(4-hydroxyphenyl) propionic acid.
Conclusions and relevance
This study is the first to characterize the feline urine metabolome. The results of this study revealed the presence of multiple low-molecular-weight substances that were not known to be present in feline urine. As expected, the origin of the metabolites detected in urine was diverse, including endogenous compounds and molecules biosynthesized by microbes. Also, the diet seemed to have had a relevant role on the urine metabolome. Further exploration of the urine metabolic phenotype will open a window for discovering unknown, or poorly understood, metabolic pathways. In turn, this will advance our understanding of feline biology and lead to new insights in feline physiology, nutrition and medicine.
Introduction
All living organisms produce a chemical diversity of low-molecular-weight substances (LMWs; <900 Da), 1 such as amino acids, carbohydrates, fatty acids, organic compounds and peptides. The collection of LMWs in a biological sample is called a metabolome.
Metabolomics is the science of measuring and characterizing LMWs in different matrices, such as plasma, serum, urine, exhaled breath and tissues.2–7 Since its first applications in the 1960s and 1970s,8–11 the use of metabolomics as a research tool has expanded into fields such as physiology, nutrition, infectious diseases, toxicology and pharmacology.2,4,12,13 Metabolomics has enabled the discovery of new therapeutic targets, and it has helped to unveil mechanisms of drug toxicity. 14 It has also become a useful tool in individualized medicine.15,16
Urine is commonly studied in metabolomics because it contains hundreds of LMWs, including metabolic breakdown products from foods, drugs, environmental contaminants, endogenous waste metabolites and bacterial end products.17–21 These substances reflect gene function, non-genetic factors and phenotypic endpoints that indicate the physiological and pathological state of the body. The human urine metabolome has been extensively investigated in the last two decades, leading to a constantly growing, searchable database that contains >3000 metabolites. 22 The urine metabolome has also been mapped out in other species such as dogs, horses and rats.4,23,24 However, the feline urine metabolome remains to be determined.
With the hypothesis that the feline urine metabolome contains hundreds of LMWs, this study endeavored to characterize the composition and content of this metabolome. Using an untargeted metabolomics approach, we found 318 LMWs, many of which were previously not known to be present in feline urine. The LMWs found in the urine metabolome are involved in at least 100 metabolic pathways. These findings significantly expand our knowledge of feline biology.
Materials and methods
Animal population
The Washington State University Institutional Animal Care and Use Committee approved all study procedures prior to its initiation (ASAF#04662).
Study population
Eight female, clinically healthy, intact adult (1–2 years old) cats were obtained from a US Department of Agriculture-licensed commercial breeder (Liberty Research). The primary enrollment criteria were no signs of illness or infection and no history of drug treatment within 30 days prior to initiation of the study. Every cat was given a baseline complete history, physical examination and laboratory evaluation. All had a normal complete blood count, serum chemistry and urinalysis.
Animal management and monitoring
Cats were acclimated at least 10 days before the study. Each was housed separately in a 4’ × 8’ partitioned room. A see-through barrier partitioned the room so the cats could see one another. Bedding, cat trees and additional enrichment, including toys and radio, were provided. The room was temperature- and humidity-controlled with a 12 h light/dark cycle. The cats had free access to drinking water and food (Purina Cat Chow Indoor Formula) throughout the study. The food was withheld 8 h before urine sampling. Cats were examined at least twice daily during the entire study duration to rule out possible health problems.
Sample collection and processing
Each cat was sedated with a combination of intravenous ketamine (15 mg/kg), acepromazine (0.2 mg/kg) and atropine (0.07 mg/kg), and a urine sample (~2 ml) was collected by ultrasound-guided cystocentesis. Immediately after collection, samples were centrifuged at 1000 × g for 8 mins. Supernatants were aliquoted and stored at −80 °C until analysis (all samples were processed within 6 months of collection).
Metabolite determination
Metabolomics analysis was performed at the West Coast Metabolomics Center, a National Institutes of Health Regional resource core. The methods used to determine the urine feline metabolome are fully explained by Fiehn and Kind, 10 and described briefly as follows.
Chemicals and reagents
The following chemicals and reagents were used: acetonitrile and isopropanol (JT Baker), pure water (MilliQ) 18.2 MΩ, pyridine (Acros Organics), methoxyamine hydrochloride (MeOX) and N-methyl-N-(trimethylsilyl)-trifluoroactamide (MSTFA; both Sigma-Aldrich).
Extraction of LMWs
A 30 μl urine sample was added to a 1.0 ml extraction solution of acetonitrile, isopropanol and water (3:3:2 v/v/v). This was mixed using a vortex mixer for ~10 s and then shaken for 5 mins at 4 °C. Samples were centrifuged for 2 mins at 14,000 × g. Next, 450 μl supernatant was evaporated using a cold trap concentrator. The dried aliquot was resuspended with 450 μl 50% acetonitrile and then centrifuged for 2 mins at 14,000 × g. The supernatant was transferred to a new tube and then dried in the cold trap concentrator. Samples were derivatized by adding 10 μl 40 mg/ml MeOX solution to each and then shaken at maximum speed at 30 ºC for 1.5 h. Next, 91 μl MSTFA was added to each sample and shaken at 37ºC at maximum speed. The mixture was transferred to glass vials and immediately capped.
Chromatography
An Agilent 6890 gas chromatograph coupled to a Leco Pegasus IV time-of-flight mass spectrometer and controlled using Leco ChromaTOF software version 2.32 were used for the analyses. Samples were separated using a 30 m long and 0.25 mm internal diameter Rtx-5Sil MS column with 0.25 μm 95% dimethyl/5% diphenyl polysiloxane film and an additional 10 m integrated guard column was used. Pure helium was used at a constant flow of 1 ml/min. The oven temperature was held constant at 50 °C for 1 min, and then ramped at 20 °C/min to 330 °C, and held constant for 5 mins. Mass spectra were acquired at mass resolving power R = 600 from m/z 85-500 at 20 spectra per s and 1550 V detector voltage without turning on the mass defect option. The recording ended after 1200 s.
Metabolite identifications
In accordance with the Metabolomics Standards Initiative, 25 gas chromatography/mass spectrometry peaks were identified using two independent parameters: retention index window ± 2000 U (around ± 2 s retention time deviation) and mass spectral similarity. Hence, spectra that were just matched to library entries without retention time (or retention index) information were considered not identified. All signals were exported by the BinBase database. 26 Database entries were named using the Fiehn library, which currently includes 713 unique metabolites and 1197 unique spectra, for which names, structure graphs, and codes, retention indices and database references are available at FiehnLib 2009 website. 27
LMWs were reported based on BinBase name and external database identifiers for PubChem, 28 and or Kyoto Encyclopedia of Genes and Genomes (KEGG; see supplementary material). 29 Signal intensity is reported as mean value ± SEM.
Metabolic pathway analysis
Metabolic pathway analysis was based on the association between identified LMWs and their related biological processes. Analytes were classified in nine global metabolic pathways. Pathway assignment was made with the help of KEGG, PubChem and the Human Metabolome Database.22,28,29
Results
Urine metabolome
By applying an untargeted metabolomics approach, 318 LMWs were detected in the urine of the eight cats. Of these, 125 have been identified (Table 1), while 193 remain unknown (see supplementary material).
Low-molecular-weight substances identified in feline urine*
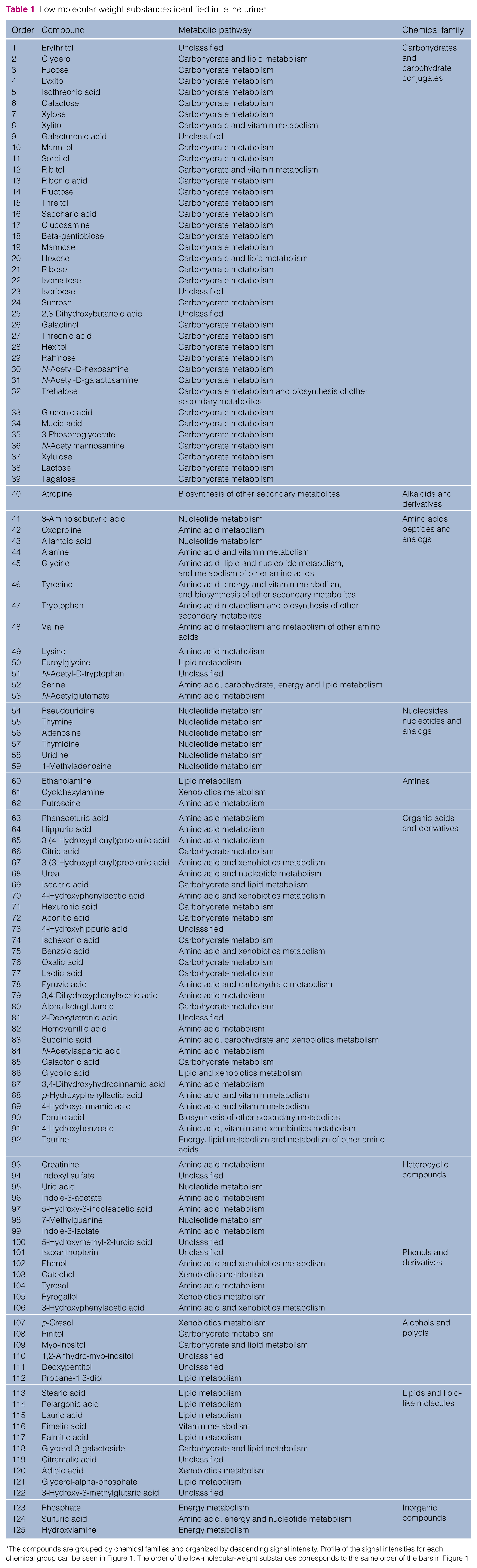
In general, feline urine is dominated by carbohydrates and carbohydrates conjugates, organic acid and derivatives, and amino acids and analogs (Figure 1). These results show that feline urine contains an important number of hydrophilic molecules, as has been reported in human urine. 30 Other chemical families found in feline urine include heterocyclic compounds, phenol and derivatives, nucleosides, nucleotides and analogs, alcohols and polyols, lipids and inorganic compounds. Of the 125 analytes we identified, the five most abundant were phenaceturic acid, hippuric acid, pseudouridine, phosphate and 3-(4-hydroxyphenyl)propionic acid. The supplementary material provides a more detailed description of the identified analytes.

Signal intensities (mean ± SEM) for the known low-molecular-weight substances (LMWs) detected (n = 125) in urine, grouped by their chemical families: (1) alcohols and polyols, (2) amines, (3) amino acids, peptides and analogs, (4) alkaloids and derivatives, (5) carbohydrates and carbohydrate conjugates, (6) inorganic compounds, (7) heterocyclic compounds, (8) lipids and lipid-like molecules, (9) nucleosides, nucleotides and analogs, (10) organic acids and derivatives, and (11) phenols and derivatives. The order of the bars in Figure 1 corresponds to the same order of the LMWs listed in Table 1
Metabolic pathway
The identified LMWs are involved in at least 100 specific metabolic pathways. Most of these compounds participate in metabolic pathways related to the carbohydrate or amino acid metabolism (Table 1). Twenty-eight LMWs participate in 2–4 distinct metabolic pathways: (1) two pathways: alanine, benzoic acid, glycerol, glycerol-3-galactoside, glycolic acid, hexose, 4-hydroxycinnamic acid, 3-hydroxyphenylacetic acid, 4-hydroxyphenylacetic acid, p-hydroxyphenylacetic acid, 3-(3-hydroxyphenyl)propionic acid, isocitric acid, myo-inositol, phenol, pyruvic acid, ribitol, trehalose, tryptophan, urea, valine and xylitol; (2) three pathways: 4-hydroxybenzoate, succinic acid, sulfuric acid and taurine; and (3) four pathways: glycine, serine and tyrosine (Table 1).
Discussion
This study is the first to characterize the composition and content of the feline urine metabolome. Using an untargeted metabolomics approach, we performed a comprehensive analysis of LMWs in feline urine and detected 318 substances. Strikingly, these LMWs are key components of at least 100 chemical pathways.29,31 This study is unique in that it revealed the presence of multiple LMWs that were previously not known to be present in feline urine (aconitic acid, isocitric acid, N-acetylmannosamine; Table 1). Of the 318 LMWs, 125 are considered known metabolites (when the spectra and retention index are matched to library entries). As expected, the origin of the metabolites detected in urine was diverse including endogenous compounds (N-acetyl-D-galactosamine, allantoic acid, galactinol, glycerol-3-galactoside, hexuronic acid, isomaltose, 7-methylguanine, 3-phosphoglycerate, ribonic acid, and tagatose), molecules biosynthesized by microbes (4-hydroxyhippuric acid and indoxyl sulfate), food (erythritol and 5-hydroxymethyl-2-furoic acid), plant (ferulic acid) and exogenous compounds (phenol). Most had been previously reported as components of the human urine metabolome. 30 Interestingly, 19/125 metabolites, are not in the human urine metabolome database: exogenous (beta-gentiobiose, propane-1,3-diol and raffinose), endogenous (N-acetyl-D-galactosamine, allantoic acid, galactinol, glycerol-3-galactoside, hexuronic acid, isomaltose, 7-methylguanine, 3-phosphoglycerate, ribonic acid and tagatose), mixed (N-acetylmannosamine: endogenous and microbial; cyclohexylamine and pelargonic acid: endogenous and food) and unknown origin LMWs (1,2-anhydro-myo-inositol, deoxypentitol and isoribose). It remains to be determined if any of these endogenous LMWs are feline-specific.
Of the 318 LMWs, 193 remain unknown (when spectra or the retention index is not matched to library entries). 32 As with metabolomics studies undertaken in humans and animals, it is the norm to find a large number of unknown metabolites. More than 99% of common metabolites in complex matrices remain unknown because there are no analytical standards at present. The characterization of unknown LMWs in the feline urine metabolome is an exciting, novel area for future research. 33 Identification of each uncharacterized metabolite often requires preparative scale isolation for nuclear magnetic resonance studies or extensive chemical synthesis to enable structural comparisons using tandem mass spectrometry. 34
Even though this study provides a significant amount of novel information, the characterization of the total collection of LMWs in feline urine remains to be determined. Currently, the Human Urine Metabolome Database contains >3000 detectable metabolites, 22 suggesting that the number of the LMWs reported in this study is a fraction of the feline urine metabolome. It is worth noting that the human urine metabolome was not characterized by a single study, but it is the result of multiple studies. 30 This is likely because of the gigantic chemical diversity of LMWs present in biological samples. Despite advances in the methodology and instrumentation, analytical methods able to detect the entire collection of LMWs in a biological sample remains to be developed. 35
In this study we used a relatively small population of female young adult cats that were non-randomly selected and housed under highly controlled experimental conditions, representing a very homogenous population. It is important to note that the experimental conditions and several external factors could influence the urine metabolome. In humans and other species, diet and gastrointestinal microbiota are the main drivers of urinary metabolite variance.36–38 In this study, multiple LMWs detected in urine were also present in the food used to feed the cats (eg, ferulic acid, phosphoric acid, sulfuric acid, taurine, manitol and erythritol). This suggests that the type of diet influences the composition of the urine metabolome. Furthermore, stress, medications, age, exercise, diurnal variation, sex and environment are also important factors that can shape the composition of the urine.39,40 This highlights the need for careful interpretation and comparison of results generated in different studies. Future studies should include a heterogeneous and randomly selected population of cats and different extended analytical platforms of LMWs, such as lipids.
As we expand our knowledge about the feline urine metabolome, it is imperative that we create a harmonized public repository database for metabolome data that is equivalent to a GenBank or Protein Data Bank for mass spectral deposition.41,42 Ideally, this database should be searchable by: (1) elemental composition; (2) parent ion mass; (3) retention time/index; and (4) mass fragment pattern. 38 It would provide researchers and clinicians a convenient, centralized resource from which to learn more about the feline urine metabolome and its unique chemical constituents. In this study, we report the components (parent ion mass; retention time/index and mass fragment pattern) necessary to initiate a complete meaningful feline urine metabolome database.
The study of cellular metabolites in urine provides a powerful minimally invasive mean to assess the status of cellular metabolic processes occurring in the body. The urine metabolome was found to be valuable to study several human systemic diseases such as immuno-mediated conditions, Parkinson’s disease, cancer and liver disease.43–48 Furthermore, studying the dynamics of LMWs in urine helped to understand the pathophysiological mechanisms of acute kidney injury and chronic kidney disease in humans.49–51 These advances in human medicine suggest that the feline urine metabolome could also help to unravel mechanisms associated with kidney disease, a condition that is in great need of scientific progress.
Conclusions
The urine metabolic phenotype is a means to uncover unknown or poorly understood metabolic pathways. The knowledge gained will advance our understanding of feline biology and lead to new insights into feline physiology, nutrition, infectious diseases and medicine.
Supplemental Material
Click here for Supplementary Material
LMWs detected in feline urine, and the parameters used to identify the LMWs: the quantification ion, database identifiers, retention index and the complete mass spectrum encoded as string
Footnotes
Supplementary material
Low-molecular-weight substances (LMWs) detected in feline urine, and the parameters used to identify the LMWs: the quantification ion, database identifiers, retention index and the complete mass spectrum encoded as string.
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by The Autzen Foundation and John and Ruth Ottmar Memorial Endowment intramural funds from the College of Veterinary Medicine at Washington State University. The authors thank Barbara Merchant for excellent editorial help with this manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
