Abstract
Objectives
The present study was undertaken to characterise the viral polypeptide 2 (VP2) gene of parvovirus from domestic cats in India.
Methods
The faecal samples from diarrhoeic/healthy domestic cats were collected from different geographical regions of India for screening by PCR assay followed by sequence analysis of the VP2 gene.
Results
Canine parvovirus (CPV)/feline panleukopenia virus (FPV) infections were found in 12 (11.3%) of 106 faecal samples tested. Two new CPV-2a (297Ala and Asn426) and three FPV strains were identified by VP2 gene analysis. Several unique and existing amino acid mutations were found, suggesting continuous evolution and emergence of newer variants. The phylogenetic analysis of the CPV sequences revealed that the two new CPV-2a strains from Mumbai (MC8) and Puducherry (P15) were clustered together in a single clade but had evolved independently and were ancestrally related to Chinese CPV-2a isolates. The FPV sequences (T-C-6 and T-C-1) from Thrissur, Kerala, formed a different clade (FPV clade) and were closely related to each other and had an ancestral relationship with an FPV isolate from the USA. Another FPV isolate from Goa (GC1) was positioned in the same clade but had evolved independently.
Conclusions and relevance
Detection of CPV in both diarrhoeic/healthy cats and the occurrence of FPV infection in a vaccinated cat provide new insights into parvovirus infections in cats in India.
Introduction
Feline panleukopenia virus (FPV) and canine parvovirus (CPV) infections are highly contagious and serious enteric diseases of cats and dogs, respectively, with high fatality rates. They are members of the genus Protoparvovirus of the family Parvoviridae, which also includes mink enteritis virus. Canine parvovirus caused by CPV-2 is considered a canine-specific variant of the FPV that emerged as a novel pathogen in the late 1970s as a consequence of an interspecies jump from other carnivores to dogs and spread rapidly worldwide. 1 However, by the end of 1980, CPV-2 was completely replaced globally in dogs by a genetic and antigenic variant termed CPV-2a. 2 Subsequently, the VP2 residue 426 changed from Asn to Asp and then from Asp to Glu in the so-called CPV-2b and CPV-2c antigenic variant strains, respectively. However, as the CPV-2b and CPV-2c antigenic strains differ from CPV-2a at only one position (VP2 residue 426), they are now considered to be variants of CPV-2a rather than distinct subtypes, as are all of the CPVs circulating worldwide today. 3 During the early 1990s in German CPV isolates, an additional amino acid difference was observed in CPV-2a and CPV-2b at amino acid position 297 (Ser to Ala) and the mutants were designated ‘new CPV-2a/2b’. 4 CPV and FPV are two closely related viruses, causing disease in their respective hosts, but new variants of CPVs have acquired the feline host range, allowing them to infect both cats and dogs, whereas the original CPV-2 does not replicate in cats. 5
FPV, the prototype parvovirus of carnivores is a highly contagious sickness occurring in cats, characterised by severe leukopenia, gastroenteritis, reproductive disorders and nervous signs. Cats are also susceptible to the new variants of canine parvovirus (CPV-2a, CPV-2b and CPV-2c). As a consequence of the host range shift of CPV-2 from dogs to cats and wild felids, there are several reports of detection of CPV-2a/2b5,6 and CPV-2c infection in cats,7–9 but FPV remains the more prevalent species of parvovirus causing disease in cats. 10 In contrast, in some Asian countries, large numbers of CPV isolates were detected in domestic and wild cats. 7 A recent study frequently isolated CPVs from apparently healthy cats, suggesting the role of CPVs in causing subclinical or very mild disease in this species. 11 Multiple infections could increase the chance of establishing persistent infection in feline hosts emphasising the epidemiological role of cats as reservoirs. Feline hosts could also act as the source of new variants of parvoviruses, emerging in the field, as they are susceptible to both new variants of CPVs and FPV.
Realising the importance of cats as a potential source of genetic diversity for parvoviruses, and as there is absolutely no such study in India on detection and characterisation of parvoviruses in domestic cats, the parvovirus (CPV/FPV) strains detected in diarrhoeic/healthy domestic cats from different states of India were analysed in the present study. The molecular characterisation and sequence diversity and phylogeny were also evaluated on the viral polypeptide 2 (VP2) capsid gene.
Materials and methods
Clinical samples and laboratory processing
A total of 106 faecal samples/rectal swabs were collected between 2011 and 2014 from both CPV/FPV suspected and also apparently healthy cats of different age groups from different states of India, as detailed in Table 1. The faecal samples/rectal swabs collected were emulsified in 1 ml 0.1 M phosphate-buffered saline of pH 7.4, stored at 4ºC and transported to the laboratory, maintaining cold chain during transport. The emulsions were centrifuged at 9300 g for 15 mins at 4ºC and the supernatant was passed through a 0.45 μm membrane filter (Millipore). The filtrate was collected and stored at −40ºC until further use.
Number of samples collected from different states of India (2011–2014) and the results

PCR
The template DNA for the PCR assay was prepared following the procedure described earlier. 12 Total nucleic acid from all the clinical specimens was extracted by the boiling lysis method by boiling at 96ºC for 10 mins followed by chilling immediately in crushed ice. Following high-speed centrifugation, the supernatants were diluted at 1:10 in nuclease-free water (NFW) to reduce residual inhibitors of DNA polymerase activity. 13 The extracted template DNA was stored at −40ºC until further use and was screened for the presence of CPV/FPV using the Hfor/Hrev primer pair listed in Table 2, which amplifies a 630 base pair (bp) fragment of the VP2 gene encoding capsid protein. 14 The DNA prepared from CPV-2 vaccine strain (Strain C154; Intervet India)/CPV-2b vaccine strain (Fort Dodge Animal Health) maintained in the Department of Veterinary Microbiology, Rajiv Gandhi Institute of Veterinary Education and Research, Puducherry, India was used as positive control in the PCR assay.
Oligonucleotide primer sequences used for screening the samples and viral polypeptide 2 gene analysis

The primer positions are based on CPV-2 GenBank accession number M38245
The PCR amplification of the VP2 capsid protein gene was carried out using 100 ng template DNA, 5 µl 10× PCR buffer, 2 mM MgCl2, 1 µl 10 mM deoxynucleotide triphosphates, 10 µM forward and reverse primers, 2 U Taq DNA Polymerase (New England Bio Labs) and the volume was made up to 50 µl with NFW. PCR amplification consisted of initial denaturation at 95ºC for 5 mins followed by 35 cycles of denaturation at 95ºC for 30 s, annealing at 55ºC for 30 s and extension at 70ºC for 1 min, followed by a final extension at 70ºC for 10 mins. The PCR-amplified products were resolved on 1.5% agarose gel in Tris acetate EDTA buffer and visualised under an ultraviolet transilluminator (Syngene).
Virus isolation
Virus isolation was carried out as per the procedure recommended. 17 Eight PCR-positive samples from cats representing the diverse geographical locations of India were randomly selected and subjected to isolation of virus in an A-72 cell line maintained in RPMI-1640 media (Sigma-Aldrich). The clinical samples were filtered and used for virus isolation in the A-72 cell line. The infected monolayers were harvested on day 3 postinoculation (with or without a cytopathic effect [CPE]) by three cycles of alternative freezing, thawing and clarified at 6000 g for 15 mins in a refrigerated centrifuge. The supernatants were stored at −40ºC until further use. The presence of virus in the cell culture fluids at the third passage level was confirmed by PCR assay using the Hfor/Hrev primer pair.
Genotyping of parvovirus samples/isolates
Three randomly selected PCR-positive (Hfor/Hrev) clinical samples and two cell culture isolates representing different geographical areas of India were subjected to another four individual PCR assays using four sets of overlapping sequencing primer pairs listed in Table 2,14,15,16 following the same protocol as detailed earlier. The PCR-amplified products were excised from the gel and extracted using a Qiagen Gel Extraction Kit. The four overlapping PCR products of the VP2 gene (for each sample/isolate) were sequenced for both strands of DNA (5’–3’ and 3’–5’) and aligned to determine their nucleotide/amino acid variations in the VP2 gene. The sequencing was carried out using automated DNA sequencer (Applied Biosystems) at Eurofins Genomics (Bengaluru, India). The sequence chromatogram was visualised in BioEdit version 7.0.5 analysis software (Isis Therapeutics). A nucleotide basic local alignment search tool (BLAST) analysis was performed with the obtained sequences using the NCBI nucleotide database (http://blast.ncbi.nlm.nih.gov/Blast.cgi) to confirm the genus specificity to CPV/FPV. VP2 gene sequences of FPV, CPV-2, CPV-2a, CPV-2b and CPV-2c from different parts of the world were retrieved from the NCBI nucleotide database and multiple sequence alignment was performed using the Clustal W algorithm in MEGA5. 18 The VP2 genes of five complete/partial aligned sequences under this study were submitted to Genbank for allotment of accession numbers.
Phylogenetic analysis
The parvovirus sequences obtained in this study and the 17 parvovirus reference sequences retrieved from GenBank were aligned and evolutionary history was inferred by employing the maximum likelihood method based on the Tamura 3-parameter model implemented in MEGA5.18,19 The branch lengths were measured in terms of the number of substitutions per site. Bootstrap test (1000 replicates) was undertaken to evaluate the confidence level of branching in the phylogenetic tree.
Results
PCR
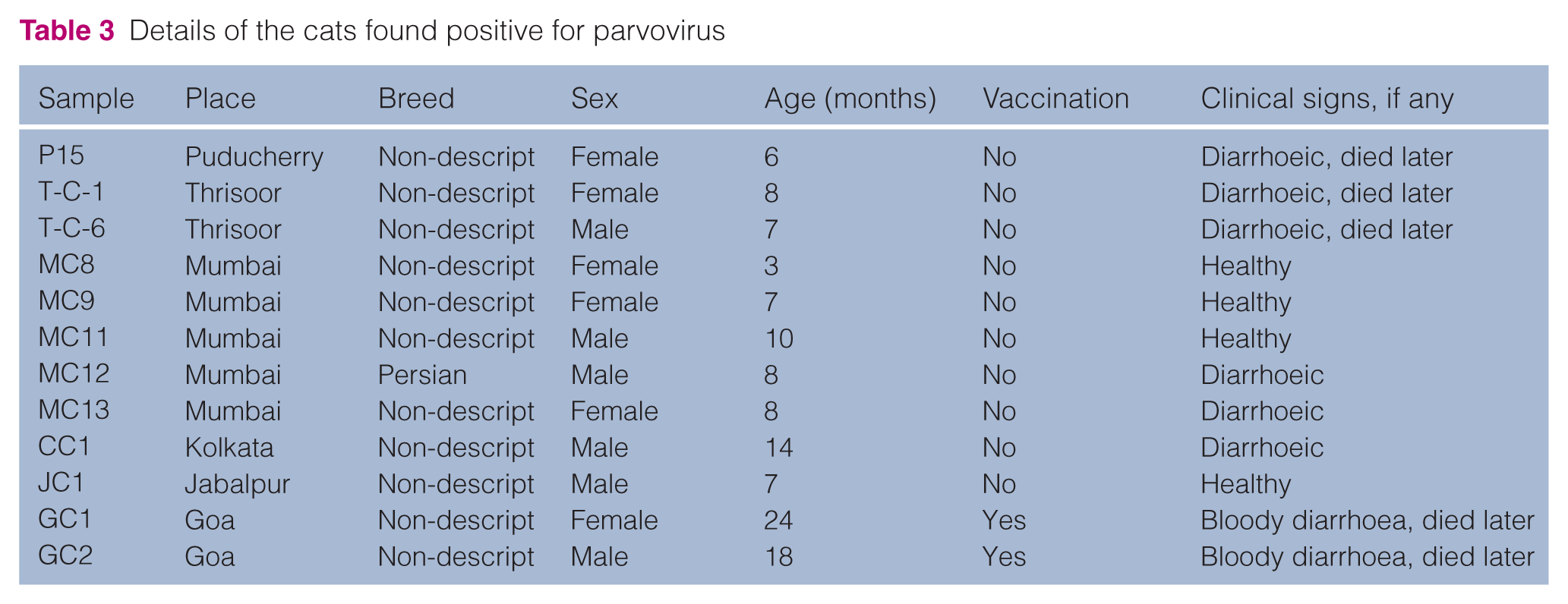
Of the total 106 faecal samples collected from CPV/FPV suspected diarrhoeic/healthy cats of different states and various zones of India from 2011 to 2014, 12 samples (11.3%) were found positive for parvoviruses by PCR assay using Hfor/Hrev primers (Table 3), by yielding a specific product size of 630 bp.
Details of the cats found positive for parvovirus
Virus isolation
In A-72 cell culture a mild cytopathic effect in the form of rounding, increased granularity and detached cells could be seen 3–4 days postinfection for two cat samples at the third passage level. The cryolysates were also confirmed as positive for parvovirus by PCR. The virus in the remaining six samples failed to grow in A-72 cell line.
Sequencing and BLAST analysis
Upon BLAST analysis, the specificity of the sequences of full-length/partial VP2 gene of parvoviruses amplified from faecal samples/cryolysates of five cats representing diverse geographical regions of India was found to be maximally identical (99–100%) with VP2 gene sequences of CPV/FPV strains available in GenBank. The sequences of two cell culture isolates namely MC8 (Mumbai) and P15 (Puducherry) had maximum identity (99%) with the new CPV-2a strain KM457141 (Uruguay) and HQ883267 (China). However, the virus sequence from the GC1 (Goa) sample had maximum identity (99%) with FPV strain EU98686 (Italy), whereas the sequences from the T-C-1 and T-C-6 (Thrissur) samples had maximum identity (99%) with FPV strain D78584 (Japan). Therefore, sequencing results revealed detection of two new CPV-2a strains and three FPV strains.
Nucleotide sequence accession numbers and mutations
GenBank accession numbers JX459572, KF772942, KP090138, KP090139 and KT444622 were obtained (Table 3). Additionally, 16 non-synonymous (Table 4) and 11 synonymous (supplementary material Table S1) mutations were also noted in the CPV/FPV sequences under study.
Non-synonymous mutations at various amino acid residues of the VP2 protein of feline panleukopenia virus (FPV)/canine parvovirus (CPV) obtained from cats

Not sequenced
Phylogenetic analysis
The phylogenetic analysis of the CPV sequences obtained from cats revealed that the two new CPV-2a strains from Mumbai (MC8) and Puducherry (P15) were clustered together in a single clade with Chinese and Uruguayan CPV-2a isolates but had evolved independently. The FPV sequences (T-C-6 and T-C-1) from Thrissur, Kerala, formed a different clade (FPV clade) and were closely related to each other and ancestrally originated from reference FPV strain (M38246) from the USA. Another FPV isolate from Goa (GC1) was positioned in the same clade at a distance along with the Korean FPV strain but appeared to have evolved independently as several non-synonymous and synonymous mutations were noted in it. A commercial FPV vaccine strain was also clustered at a distance within the same FPV clade (Figure 1).

Maximum likelihood tree depicting phylogenetic relationship among parvovirus isolates. Canine parvoviruses (CPV)/feline panleukopenia viruses (FPV) sequenced in this study are shown with solid triangles, vaccine with solid circle and the bootstrap values are shown next to the branches in the phylogenetic tree
Discussion
The present study confirmed the presence of both CPV and FPV infections among the cat population in India. Of the 106 faecal samples screened in this study from various states of India, 12 (11.3%) samples (including four samples from apparently healthy cats) were found positive by PCR assay. Battilani et al analysed faecal samples from 24 cats with a clinical diagnosis of parvovirus infection and found that 22 viral strains were FPV, whereas two strains were intermediate between CPV and FPV. 10 In a recent study, two CPV-2a (Ser297Ala) strains were identified out of 16 samples collected from cats (12.5%) in China. 20 The report of CPV/FPV infection among cats in India is scarce. There is absolutely no report of CPV infection in cats in India and in a recent study in the city of Chennai, India, 60 diarrhoeic cat samples were tested and 17 samples (28.3%) were detected as positive for FPV by PCR assay. 21 Therefore, the present study is the first comprehensive report of detection and characterisation of parvovirus infection among domestic (both diarrhoeic and apparently healthy) cat populations from different geographical regions of India based on complete/partial VP2 gene analysis.
A similar CPE pattern by parvovirus in the A-72 cell line was also noted by several researchers.14,15,22 Both the cryolysates found positive by PCR assay were identified later as CPVs (MC8 and P15) and not FPVs by sequencing. Probably, FPVs require more passages to become adapted to a canine-origin A-72 cell line. FPVs were passaged in the A-72 cell line 15 times, to observe the characteristic CPE. 21
The present finding of the presence of CPV in the faecal samples of healthy (MC8) as well as diarrhoeic (P15) cats indicates the important role the cats play in acting as reservoirs and transmitting the infection to dogs. A similar result was also observed in other research. 11 All three FPV strains were detected from diarrhoeic cats. The cats from Goa alone (GC1 and GC2) were vaccinated with a commercial trivalent FPV vaccine. None of the other cats in the present study were vaccinated with either CPV or FPV vaccines.
The mutation at amino acid change Tyr324Ile was noted in the two CPV-2a strains (MC8 and P15). Similar codon changes at amino acid positions 324 were also reported in the CPVs isolated from dogs in several countries.22–27 Residue 324 was prone to strong positive selection in all carnivorous CPV isolates. 28 This residue was adjacent to residue 323, responsible for canine transferrin receptor binding, and together with residue 93, determined the canine host range. Therefore, mutation at amino acid residue 324 is likely to have an impact on the parvovirus host range. 29 Interestingly, the frequency of CPV strains carrying mutations at the 324Ile residue had increased enormously and had reached a high prevalence among Chinese and Indian CPVs isolated from dogs.22–23,25,26 However, to date, this mutation has never been reported from any CPV strains isolated from cats. In fact, the presence of the mutation at amino acid position 324 (Tyr➔Ile) in CPVs also isolated from cats in this study provides new insights into the theory of involvement of amino acid position 324 as a canine host range mutation.
The next variation was observed in the amino acid residue Thr440Ala in both the sequences (MC8 and P15). Amino acid residue 440 is important because it is located at the top of the three-fold spike (GH loop) of the VP2 protein on the surface of the capsid, the main antigenic site of the virus.30,31 A similar codon change at nucleotide position 4104 (A➔G) was also reported in earlier studies in CPVs isolated from dogs,15,24,25,32,33 and also in CPVs isolated from cats. 8 Therefore, this mutation is likely to change the antigenicity of this CPV strain isolated from the cat.
Interestingly, three unique amino acid changes – Gly10Gln, Gly224Glu and Ala514Thr – were noted in the MC8 sequence. These mutations have not been reported by earlier research and are unique to the CPV strain obtained from cats in the present study. Although the functional impact of these mutations was yet to be ascertained, the present study indicated active evolution of this CPV-2a strain of cat origin.
The FPV sequence (GC1) showed four amino acid changes – Arg81Gly, Ser179Thr, Ala359Pro and Ile401Val. Two of these non-synonymous mutations were detected in the GH loop of the VP2 protein (267–498 residues located between βG and βH strands), exposed on the surface of capsid, and greatest variability was seen in this antigenic region. Six synonymous mutations were also noted in this FPV sequence. This FPV sequence was obtained from a cat from Goa that was vaccinated with a commercial trivalent FPV vaccine. The presence of many non-synonymous and synonymous mutations in this FPV strain would have contributed to antigenic variation, resulting in vaccine failure. Three amino acid mutations – Met551Val, Asn564His and Asn565His – were observed in another FPV sequence (T-C-6), while the other FPV sequence T-C-1 showed only one mutation Thr388Asn. These mutations are all unique to our strains and have not been reported so far from any FPV isolates. The amino acid at position 564 is a differentiating amino acid between FPV (Asn/AAT) and CPV (Ser/AGT) strains. Interestingly, two consecutive unique mutations in one of the FPV strains (T-C-6) at amino acid positions 564 (His/CAT) and 565 (His/CAT) have been reported in this study. This FPV strain (T-C-6) needs to be studied further in relation to its pathogenicity and antigenicity. Therefore, the presence of unique and certain existing non-synonymous mutations in the CPV/FPV strains in India indicate that the CPV/FPV strains are under constant selection pressure and are constantly mutating, leading to the evolution of newer CPV/FPV types/variants among the Indian cat population.
Therefore, as per the present study, mutations and the positive selection of the mutants was found to be the major mechanism of emergence and evolution of CPV/FPVs in India. No cat is vaccinated against CPV in India and very few cats are vaccinated with FPV vaccine. In our study, except for the cats from Goa which were vaccinated with a commercial trivalent FPV vaccine, no cat was vaccinated against CPV/FPV vaccines.
Conclusions
The present study provides new and valuable insight into the evolutionary dynamics of parvovirus infections in cats in India, although further studies involving greater sample size and wider geographical regions are required for complete understanding of the problem. The presence of CPVs in both healthy and diarrhoeic cats emphasises the possible role of cats as a source of infection for dogs. As cats in India were found to be susceptible to both CPV and FPV infections, the role of cats as the source of new variants of parvovirus needs to be further explored. Furthermore, the presence of FPV infection in the domestic cat population raises serious concern regarding the urgent need to educate owners about the importance of FPV vaccination. The cats from Goa succumbing to FPV infection in spite of vaccination also raises serious concerns. Intensifying surveillance of parvovirus infections in cats in India is required.
Footnotes
Acknowledgements
We thank Dean Rajiv Gandhi, Institute of Veterinary Education and Research, Puducherry, India, for providing the necessary funds and facilities to carry out this study.
Supplementary material
Table to show synonymous mutations at various amino acid residues of the VP2 protein of FPV/CPV obtained from cat.
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This project was funded by a grant received from DBT, Government of India (BT/PR14677/ADV/57/111/2010).
